Abstract
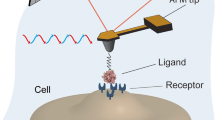
The invention of atomic force microscopy (AFM) has provided new technology for measuring specific molecular interaction forces. Using AFM single-molecule force spectroscopy (SMFS) techniques, CD20-Rituximab rupture forces were measured on purified CD20 proteins, Raji cells, and lymphoma patient B cells. Rituximab molecules were linked onto AFM tips using AFM probe functionalization technology, and purified CD20 proteins were attached to mica using substrate functionalization technology. Raji cells (a lymphoma cell line) or lymphoma patient cells were immobilized on a glass substrate via electrostatic adsorption and chemical fixation. The topography of the purified CD20 proteins, Raji cells, and patient lymphoma cells was visualized using AFM imaging and the differences in the rupture forces were analyzed and measured. The results showed that the rupture forces between the CD20 proteins on Raji cells and Rituximab were markedly smaller than those for purified CD20 proteins and CD20 proteins on lymphoma patient B cells. These findings provide an effective experimental method for investigating the mechanisms underlying the variable efficacy of Rituximab.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Binnig G, Quate C F, Gerber C. Atomic force microscope. Phys Rev Lett, 1986, 56: 930–933
Dufrene Y F. Atomic force microscopy and chemical force microscopy of microbial cells. Nat Protoc, 2008, 3: 1132–1138
Dufrene Y F. Using nanotechniques to explore microbial surfaces. Nat Rev Microbiol, 2004, 2: 451–460
Muller D J, Dufrene Y F. Atomic force microscopy as a multifunctional molecular toolbox in nanobiotechnology. Nat Nanotechnol, 2008, 3: 261–269
Chen P P, Dong H T, Chen L, et al. Application of atomic force microscopy to living samples from cells to fresh tissues. Chinese Sci Bull, 2009, 54: 2410–2415
Li M, Liu L Q, Xi N, et al. Imaging and mechanical property measurement of the lymphoma cells by atomic force microscopy (in Chinese). Chinese Sci Bull (Chinese Ver.), 2010, 55: 2188–2196
Dufrene Y F, Evans E, Engel A, et al. Five challenges to bringing single-molecule force spectroscopy into living cells. Nat Methods, 2011, 8: 123–127
Muller D J, Helenius J, Alsteens D, et al. Force probing surfaces of living cells to molecular resolution. Nat Chem Biol, 2009, 5: 383–390
Florin E L, Moy V T, Gaub H E. Adhesion forces between individual ligand-receptor pairs. Science, 1994, 264: 415–417
Hinterdorfer P, Baumgartner W, Gruber H J, et al. Detection and localization of individual antibody-antigen recognition events by atomic force microscopy. Proc Natl Acad Sci USA, 1996, 93: 3477–3481
Puntheeranurak T, Wildling L, Gruber H J, et al. Ligands on the string:single-molecule AFM studies on the interaction of antibodies and substrates with the Na+-glucose co-transporter SGLT1 in living cells. J Cell Sci, 2006, 119: 2960–2967
Shi X, Xu L, Yu J, et al. Study of inhibition effect of Herceptin on interaction between Heregulin and ErbB receptors HER3/HER2 by single-molecule force spectroscopy. Exp Cell Res, 2009, 315: 2847–2855
Beers S A, French R R, Chan H T C, et al. Antigenic modulation limits the efficacy of anti-CD20 antibodies: Implications for antibody selection. Blood, 2010, 115: 5191–5201
Glennie M J, French R R, Cragg M S, et al. Mechanisms of killing by anti-CD20 monoclonal antibodies. Mol Immunol, 2007, 44: 3823–3837
Cartron G, Watier H, Golay J, et al. From the bench to the bedside: Ways to improve Rituximab efficacy. Blood, 2004, 104: 2635–2642
Lim S H, Beers S A, French R R, et al. Anti-CD20 monoclonal antibodies: Historical and future perspectives. Haematologica, 2010, 95: 135–143
Li B, Zhao L, Guo H, et al. Characterization of a Rituximab variant with potent antitumor activity against Rituximab-resistant B-cell lymphoma. Blood, 2009, 114: 5007–5015
Wang H, Bash R, Yodh J G, et al. Glutaraldehyde modified mica: A new surface for atomic force microscopy of chromatin. Biophys J, 2002, 83: 3619–3625
Stroh C, Wang H, Bash R, et al. Single-molecule recognition imaging microscopy. Proc Natl Acad Sci USA, 2004, 101: 12503–12507
Ebner A, Wildling L, Kamruzzahan A S M, et al. A new, simple method for linking of antibodies to atomic force microscopy tips. Bioconjugate Chem, 2007, 18: 1176–1184
Hinterdorfer P, Dufrene Y F. Detection and localization of single molecular recognition events using atomic force microscopy. Nat Methods, 2006, 3: 347–355
Stevens F, Lo Y S, Harris J M, et al. Computer modeling of atomic force microscopy force measurements: Comparisons of Poisson, histogram, and continuum methods. Langmuir, 1999, 15: 207–213
Kada G, Kienberger F, Hinterdorfer P. Atomic force microscopy in bionanotechnology. Nano Today, 2008, 3: 12–19
Muller D J, Engel A, Amrein M. Preparation techniques for the observation of native biological systems with the atomic force microscope. Biosens Bioelectron, 1997, 12: 867–877
Henderson R M, Schneider S, Li Q, et al. Imaging ROMK1 inwardly rectifying ATP-sensitive K+ channel protein using atomic force microscopy. Proc Natl Acad Sci USA, 1996, 93: 8756–8760
Kirat K E, Burton I, Dupres V, et al. Sample preparation procedures for biological atomic force microscopy. J Microsc, 2005, 218: 199–207
Kada G, Blayney L, Jeyakumar L H, et al. Recognition force microscopy/spectroscopy of ion channels: Applications to the skeletal muscle Ca2+ release channel (RYR1). Ultramicroscopy, 2001, 86: 129–137
Wang H, Kutner L O, Lin M, et al. Imaging glycosylation. J Am Chem Soc, 2008, 130: 8154–8155
Muller D J, Dufrene Y F. Force nanoscopy of living cells. Curr Biol, 2011, 21: R212–R216
Yamada K M, Cukierman E. Modeling tissue morphogenesis and cancer in 3D. Cell, 2007, 130: 601–610
Werten P J L, Remigy H W, Groot B L, et al. Progress in the analysis of membrane protein structure and function. FEBS Lett, 2002, 529: 65–72
Wang Q, Lu Y, Li S, et al. Distribution and force spectroscopy of CD20 antigen-antibody binding on the B cell surface (in Chinese). Chin J Biotech, 2011, 27: 131–136
Gu X, Jia X, Feng J, et al. Molecular modeling and affinity determination of scFv antibody: Proper linker peptide enhances its activity. Ann Biomed Eng, 2010, 38: 537–549
Author information
Authors and Affiliations
Corresponding authors
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Li, M., Liu, L., Xi, N. et al. Detecting CD20-Rituximab interaction forces using AFM single-molecule force spectroscopy. Chin. Sci. Bull. 56, 3829–3835 (2011). https://doi.org/10.1007/s11434-011-4789-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-011-4789-0