Abstract
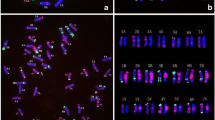
Haynaldia villosa (L.) is a wild relative species of common wheat that possesses many beneficial genes that can be used for wheat improvement. The accurate detection of H. villosa chromosomes in the genetic background of wheat is critical for transferring its beneficial genes to common wheat by chromosome engineering. The aim of the present study was to investigate the distribution patterns of two repeated DNA sequences, pSc119.2 and pAs1, as well as two rDNA multigene family sequences, 45S rDNA and 5S rDNA, in the individual chromosomes of H. villosa for the future precise identification of alien chromatin in germplasm development and breeding programs. A set of common wheat-H. villosa disomic addition 1V-7V lines was used to determine these specific signals on individual chromosomes of H. villosa. The results showed that two rDNA probes, pTa71 (45S rDNA) and pTa794 (5S rDNA), were located on 1VS and 5VS, respectively, and the signal could be discriminated exclusively in the common wheat background as effective markers of 1VS and 5VS. Furthermore, all seven chromosomes of H. villosa could be distinguished clearly by fluorescence in situ hybridization using pSc119.2 and pAs1 as probes in combination. The utilization of these cytogenetic markers of repetitive sequences, combined with other molecular markers sometimes, will make it possible for a precise identification of alien chromosomes with high efficiency.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Hyde B B. Addition of individual Haynaldia villosa chromosomes to hexaploid wheat. Am J Bot, 1953, 40: 174–182
Chen P D, Liu D J. Cytogenetic studies of hybrid progenies between Triticum aestivum and Haynaldia villosa (in Chinese). J Nanjing Agric Coll, 1982, 5: 1–16
Chen Q, Conner R L, Li H, et al. Expression of resistance to stripe rust, powdery mildew and the wheat curl mite in Triticum aestivum-Haynaldia villosa lines. Can J Plant Sci, 2002, 82: 451–456
Murray T D, De la Pena R C, Yildirim A, et al. A new source of resistance to Pseudocercosporella herpotrichoides cause of eyespot disease of wheat located on chromosome 4V of Dasypyrum villosum. Plant Breed, 1994, 113: 281–286
Zhang Q P, Li Q, Wang X E, et al. Development and characterization of a Triticum aestivum-Haynaldia villosa translocation line T4VS-4DL conferring resistance to wheat spindle streak mosaic virus. Euphytica, 2005, 145: 317–320
Blanco A, Resta P, Simeone R, et al. Chromosomal location of seed storage protein genes in the genome of Dasypyrum villosum (L.) Candargy. Theor Appl Genet, 1991, 82: 358–362
De Pace C, Snidaro D, Ciaffi M, et al. Introgression of Dasypyrum villosum chromatin into common wheat improves grain protein quality. Euphytica, 2001, 117: 67–75
Chen P D, Liu D J. Identification of Haynaldia villosa chromosomes in alien wheat addition. In: Li Z S, Swaminathan M S, eds. Proceedings of the International Symposium on Chromosome Engineering in Plants. Xi’an, China, 1986. 31–33
Liu D J, Chen P D, Wu P L, et al. Triticum durum-Haynaldia villosa amphidiploid (in Chinese). Acta Agron Sin, 1986, 12: 155–162
Chen P D, Qi L L, Zhou B, et al. Development and molecular cytogenetic analysis of wheat-Haynaldia villosa 6VS/6AL translocation lines specifying resistance to powdery mildew. Theor Appl Genet, 1995, 91: 1125–1128
Friebe B, Cermeno M C, Zeller F I. C-banding polymorphism and the analysis of nuclear activity in Dasypyrum villosum (L.) Candargy, its added chromosomes to hexaploid wheat and the amphiploid Triticum dicoccum-D. villosum. Theor Appl Genet, 1987, 73: 337–342
Dong F G, Chen P D, Liu D J. An improved C-banding technique for Haynaldia villosa chromosomes (in Chinese). Acta Genet Sin, 1991, 18: 525–528
Liu D J, Chen P D. N-banding in Haynaldia villosa and Triticum durum-H. villosa Amphidiploid (in Chinese). Acta Genet Sin, 1984, 11: 106–108
Gupta P K, Fedak G, Molnar S J, et al. Distribution of a Secale cereale sequence among 25 Hordeum species. Genome, 1989, 32: 383–388
Rayburn A L, Gill B S. Use of biotin-labeled probes to map specific DNA sequences on wheat chromosomes. J Hered, 1985, 76: 78–81
Vershinin A V, Svitashev S K, Gummesson P O, et al. Characterization of a family of tandemly repeated DNA sequences in Triticeae. Theor Appl Genet, 1994, 89: 217–225
Gill B S, Friebe B, Endo T R. Standard karyotype and nomenclature system for description of chromosome bands and structural aberrations in wheat (Triticum aestivum). Genome, 1991, 34: 830–839
Zhang P, Li W L, Friebe B, et al. Simultaneous painting of three genomes in hexploid wheat by BAC-FISH. Genome, 2004, 47: 979–987
Sharp P J, Kreis M, Shewry P. Location of π-amylase sequence in wheat and its relatives. Theor Appl Genet, 1988, 75: 289–290
Yuan W Y, Tomita M, Sun S C, et al. Multicolor fluorescence in situ hybridization of the rRNA genes in wheat relatives, Dasypyrum villosum and Thinopyrum intermedium. Bull Fac of Agric, Tottori Univ, 2000, 53: 7–12
Bie T D, Cao Y P, Chen P D. Mass production of intergeneric chromosomal translocatons through pollen irradiation of Triticum durum-Haynaldia villosa amphiploid. J Integr Plant Biol, 2007, 49: 1619–1626
Cao Y P, Bie T D, Wang X E, et al. Induction and transmission of wheat-Haynaldia villosa chromosomal translocations. J Genet Genomics, 2009, 36: 313–320
Flavell R B. Repetitive DNA and chromosome evolution in plants. Philos Trans R Soc Lond B Biol Sci, 1986, 312: 227–242
Jiang J, Nasuda S, Dong F, et al. A conserved repetitive DNA element located in the centromeres of cereal chromosomes. Proc Natl Acad Sci USA, 1996, 93: 14210–14213
Rayburn A L, Gill B S. Molecular identification of the D-genome chromosomes of wheat. J Hered, 1986b, 77: 253–255
Mukai Y, Friebe B, Gill B S. Comparison of C-banding patterns and in situ hybridization sites using highly repetitive and total genomic rye DNA probes of ‘Imperial’ rye chromosomes added to ‘Chinese Spring’ wheat. Jpn Jpn J Genet, 1992, 67: 71–83
Anamthawat-Jonsson K, Heslop-Harrison J S. Isolation and characterization of genome-specific DNA sequences in Triticeae species. Mol Gen Genet, 1993, 240: 151–158
Zhang P, Li W L, Fellers J, et al. BAC-FISH in wheat identifies chromosome landmarks consisting of different types of transposable elements. Chromosoma, 2004, 112: 288–299
Badaeva E D, Zoshchuk S A, Paux E, et al. Fat element-a new marker for chromosome and genome analysis in the Triticeae. Chromosome Res, 2010, 18: 697–709
Yuan W X, Tomita M. Centromeric distribution of 350-family in Dasypyrum villosum and its application to identifying Dasypyrum chromatin in the wheat genome. Hereditas, 2009, 146: 58–66
Qi Z J, Du P, Qian B L, et al. Characterization of a wheat-Thinopyrum bessarabicum (T2JS-2BS·2BL) translocation line. Theor Appl Genet, 2010, 121: 589–597
Jia J Q, Yang Z J, Li G R, et al. Isolation and chromosomal distribution of a novel Ty1-copia-like sequence from Secale, which enables identification of wheat-Secale africanum introgression lines. J Appl Genet, 2009, 50: 25–28
Schneider A, Linc G, Molnár-Láng M. F Fluorescence in situ hybridization polymorphism using two repetitive DNA clones in different cultivars of wheat. P Plant Breed, 2003, 122: 396–400
Cuadrado A, Jouve N. Distribution of highly repeated DNA sequences in species of the genus Secale. Genome, 1997, 40: 309–317
Zhang R Q, Cao Y P, Wang X E, et al. Development and characterization of a Triticum aestivum-Haynaldia villosa T5VS·5DL translocation line with soft grain texture. J Cereal Sci, 2010, 51: 220–225
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Zhang, W., Zhang, R., Feng, Y. et al. Distribution of highly repeated DNA sequences in Haynaldia villosa and its application in the identification of alien chromatin. Chin. Sci. Bull. 58, 890–897 (2013). https://doi.org/10.1007/s11434-012-5598-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5598-9