Abstract
Physiological dormancy is a characteristic of the seeds of plants that are exposed to adverse harsh environmental conditions. Many tree seeds from the northern hemisphere have acquired deep dormancy as an adaptation to the winter period. Such kinds of dormancy can be removed by cold stratification. This physiological process is regulated through abscisic and gibberellic acids signal transduction, in which ABI5, 14-3-3 and RGL2 have a negative response to cold stratification and cause dormancy breaking. Our study is, to our knowledge, the first to report tissue localisation of ABI5 and RGL2 in deeply physiologically dormant seeds. Localisation of these proteins differs in time (weeks of stratification) and space (anatomy of the embryo root). Studies showed that changes occurred on three levels: (1) tissue, as the fluorescence signal throughout the weeks of stratification was localised in different regions of the embryo axes, and these changes were associated with changes in development regulation of the individual regions; (2) cell, either in nucleus or in cytoplasm, involving regulation of gene expression, and synthesis and inactivation in cytoplasm; and (3) organelle, specifically in nuclei/nucleoli, indicating transcription regulation of the specific genes. At the end of stratification, when dormancy is broken, ABI5 and RGL2 were not noticeable in the cells of the apical meristem. ABI5 likely blocked germination through inhibition of meristem activity, whereas RGL2 through blocking of procambial cell differentiation. We speculate that similar molecular and cellular mechanisms exist among other seeds characterised by physiological dormancy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dormancy is a key ecological adaptation which protects seeds against germination during unfavourable conditions, such as during the winter period (Black et al. 2008). In temperate zones, seeds of some plant species are characterised by deep physiological dormancy, located in embryo tissues (Baskin and Baskin 2004). The mechanism of dormancy acquisition and breaking is complex and still far from completely understood, especially regarding deep physiological embryo dormancy. According to Luckwill’s (1952) theory, transition from dormancy to germination is under the antagonistic influence of inhibitors, such as abscisic acid (ABA), and promotors, such as gibberellins (GA). Transduction pathways conduct a signal from the environment through receptors and hormonal pathways to the nucleus. Investigating the regulation of the signal cascade involving ABA and GA at different levels could bring us closer to environmental and hormonal control of dormancy.
ABA is a plant hormone with roles in developmental processes, stomatal movements, stress response and regulations of dormancy (Pawłowski 2007, 2009; Hauser et al. 2011; Pawłowski and Staszak 2016). The signal is transmitted by the PYR/PYL/RCAR receptor. The hormone and the receptor form a complex, which impedes the activity of 2C protein phosphatases (PP2Cs) (Ma et al. 2009; Park et al. 2009). Deactivation of PP2Cs leads to enabled autophosphorylation of SnRK2 kinases, which take part in transcription regulation through the triggering of bZIP transcription factors AREB/ABF/ABI5 (Nakashima et al. 2009; Ma et al. 2009; Park et al. 2009; Fujii et al. 2009). ABA activates the expression of many genes via ABA-responsive elements (ABREs) in their promoter regions (Saavedra et al. 2010). The ABI5 protein is a transcriptional activator for genes controlled by ABREs (Hattori et al. 2002) is present mainly in seeds and plays a role in seed development, germination and seedling development (Lopez-Molina et al. 2001). In dry Arabidopsis seeds treated with ABA, ABI5 accumulated, but after germination or elimination of ABA, ABI5 degraded quickly (Finkelstein and Lynch 2000; Lopez-Molina et al. 2001; Piskurewicz et al. 2008). Mechanism of ABI5 stabilisation by ABI five binding protein (AFP) is involved in post-embryonic growth cessation (Lopez-Molina et al. 2003). ABI5 cooperates with ABI3, which enhances transcription (Lopez-Molina et al. 2002), and is regulated by itself and ABI4, suggesting a complex regulatory interaction (Xu et al. 2014). ABI5 is involved in stress response and development adjustments (Fujita et al. 2011). The role of ABI5 in dormancy induction is discussed (Finkelstein et al. 2008) mostly because of the lack of such research concerning Arabidopsis seeds (Dekkers et al. 2016). Research with orthologues, however, has confirmed that ABI5 homologues are more highly expressed in dormant sorghum cultivars (Rodríguez et al. 2009). Despite this, Finkelstein et al. (2008) concluded that further studies were required to confirm the role of ABI5 in seed dormancy.
The 14-3-3 proteins act as a modulator of the activity of target proteins by attaching to their phosphorylated motifs (Ferl 1996). In plants, they take part in the regulation of a wide range of physiological functions, such as development and response to stress conditions (Denison et al. 2011; de Boer et al. 2013). They can affect signalling pathways, which transduce information from the environment, and also have influence on effector proteins that induce the physiological response (Denison et al. 2011). 14-3-3 proteins have been linked to ABA signalling, with its role in seed germination (Schoonheim et al. 2007). They interplay specifically with members of the ABF/AREB/ABI5 family (e.g. ABI5) (Jakoby et al. 2002), leading to alterations in the transcription of ABA-responsive genes. 14-3-3s enable contact between ABI3/VP1 (co-activator of transcription) and the ABF/AREB/ABI5 family of transcription factors (Himmelbach et al. 2003; Schoonheim et al. 2007). Additionally, gibberellins influence 14-3-3 protein accumulation (Schoonheim et al. 2009), which implies that 14-3-3 proteins take part in cross-talk of the hormonal pathways of ABA and GA (Schoonheim et al. 2009).
Gibberellins influence plant growth and development via DELLA proteins, which are a member of the GRAS family (Sun et al. 2012). DELLAs are repressors of GA-responsive genes (de Lucas et al. 2008). They act as integrators and regulators of environmental signals (Achard et al. 2006; Sun et al. 2010). Aromatic or hydrophobic domains of DELLA proteins directly interplay with the GA-tied receptor GID1 and that connection plays a crucial role in transducing GA signals during the control of plant development (Murase et al. 2008; Sun et al. 2010). Creation of GA-GID1-DELLA complex leads to DELLA decay by the 26S proteasome through addition of a polyubiquitin chain to DELLAs by the GA-dependent SCFGID2/SLY1 E3 ubiquitin ligase (Heckmann et al. 2006). Mutants overexpressing the DELLAs show insensitivity to GA and dwarfism, besides mutants underexpressing the DELLAs have constitutive GA-response phenotypes (de Lucas et al. 2008; Feng et al. 2008). Disintegration or deactivation of DELLAs modulates plant growth by GA, thereby de-repressing GA-regulated genes. DELLAs could act as transcriptional co-activators of phytochrome interacting factors (PIFs), belonging to the basic helix-loop-helix (bHLH) transcription factor family (Itoh et al. 2005). In the absence of GA, DELLAs prevent PIF3 and PIF4 from binding to their target promoters. Disintegration of DELLAs by GA releases PIF3 and PIF4, enabling them to perform their function and regulate gene expression. DELLAs regulate seed germination by interaction with PIF1 (Oh et al. 2009).
RGL2 is a DELLA protein associated with germination repression (Lee et al. 2002; Cao et al. 2005), and it has been recognised as a factor that is associated with dormancy acquisition by up-regulation of several genes associated with dormancy (Stamm et al. 2012). RGL2 links to the GA and ABA signalling pathway by influencing ABA biosynthesis and activation of ABI5 and ABI3 (Piskurewicz et al. 2008; Piskurewicz et al. 2009). RGL2 controls cell growth by influencing cell wall-affecting genes, such as EXPA3 and EXPA8, by direct inhibition of transcription of these genes (Stamm et al. 2012).
The objects of dormancy research are usually seeds with coat-imposed dormancy, which is defined as a non-deep physiological dormancy (Finch-Savage and Leubner-Metzger 2006), for example seeds of Arabidopsis thaliana. Tree species such as Norway maple (Acer platanoides L.) (Pawłowski 2009; Staszak and Pawłowski 2014), sycamore (Acer pseudoplatanus L.) (Pawłowski and Staszak 2016) and beech (Fagus sylvatica L.) (Reyes et al. 2006; Pawłowski 2007; Saavedra et al. 2010) have been the objects of research concerning embryo dormancy, defined also as deep physiological dormancy (Finch-Savage and Leubner-Metzger 2006). Norway maple produces seeds that are deeply physiologically dormant (Pawłowski 2010). This species is categorised as orthodox according to seed storage behaviour and requires cold stratification lasting approximately 3 months for germination.
Despite all of the above findings, there is a lack of knowledge of the details concerning the involvement of abscisic and gibberellic acid pathways in regulation of dormancy breaking during cold stratification. New findings in signal transduction cascades could allow for better understanding of ABA- and GA-mediated dormancy. The current research concentrated on three main regulators of ABA (ABI5 and 14-3-3) and GA (RGL2) signals. The main goal of this study was to describe the relationship between cold treatment dormancy breakage and ABA and GA signal regulators in deeply physiologically dormant seeds. We investigated the accumulation and immunolocalisation of ABI5, 14-3-3 and RGL2 proteins. This investigation showed the role of ABA and GA signalling regulators in dormancy and germination regulation in tree seeds.
Materials and methods
Materials
Norway maple (Acer platanoides L.) seeds were picked up in the Kórnik Arboretum, Poland (N52°14′ and E17°05′) during the autumn of 2009. The seeds were dried at ambient temperature and humidity until they reached a moisture content of 9%. They were then stored in plastic containers at − 3 °C. Prior to the experiments, the seeds were allowed to imbibe water for 48 h at room temperature. The seeds were then subjected to cold stratification at 3 °C (a dormancy-breaking temperature) for up to 17 weeks in closed plastic boxes without medium and in the dark. Germination tests (four replicates of 50 seeds each) were carried out at 3 °C in accordance with the recommendations of the International Seed Testing Association (1999).
Western blotting
Embryo axes were taken weekly from seed samples during the period of stratification at 3 °C. Samples ranged from dry dormant seeds to germinated seeds, which were defined by the protrusion (∼ 1 mm) of the radicle. Embryos were removed and immediately frozen in liquid nitrogen and subsequently stored at − 80 °C until undergoing further processing. Samples were ground in liquid nitrogen and incubated in acetone at −20 °C. After several washes in acetone and centrifugation (20,800×g for 5 min at 4 °C), the resulting pellets were dried in vacuum, and then, extraction buffer was added (0.5 M Tris pH 6.8; 10% SDS; 7% glycerol; 0.015% DTT 15). Samples were incubated for 10 min at 100 °C and centrifuged again (20,800×g for 10 min at 4 °C). Supernatants were taken, and protein content was determined via Bradford assay (Bradford 1976).
Proteins were separated using 12% SDS-PAGE on Mini-PROTEAN® 3 (Bio-Rad, Hercules, Ca, USA) with protein weight markers SeedBlue MagicMarkt (Life Technologies, Carlsbad, CA, USA) after being transferred onto PVDF membrane (Bio-Rad, Hercules, Ca, USA) at 260 mA for 75 min in transfer buffer [25 mM Tris, 192 mM glycine, 20% (v/v) methanol, pH 8.7]. After blocking the membranes with TBS-T and 5% BSA, incubation with primary rabbit polyclonal antibody was carried out. Primary antibodies were used against ABI5, 14-3-3 and RGL2 (Agrisera, Vännäs, Sweden), dilution 1:7500. Bound antibodies were detected using an antirabbit horseradish peroxidase-conjugated secondary antibody (diluted 1:50,000 for ABI5 and 1:30,000 for 14-3-3 and RGL2) and visualised with enhanced chemiluminescent reagent Clarity (Bio-Rad, Hercules, Ca, USA). Signal was detected on X-ray film (Kodak), and then, an X-ray film for each antibody was scanned using optical scanner Image Scanner III in LabScan programme (GE Healthcare, Little Chalfont, UK). Western blot images were analysed using 2D Image Master 7 Platinum (GE Healthcare, Little Chalfont, UK) software. Patterns from three independent biological samples were aligned, and normalised spot volumes of the bands were determined. Statistical analyses were carried out for the obtained results using ANOVA and the Tukey–Kramer HSD test. Results were considered significant for P < 0.05. The data are presented in relative units calculated as the intensity and the area of the band.
Immunolocalisation
Embryo axes of Norway maple were isolated from intact seeds during stratification (from dry to germinated seeds). Plant material was fixed in an ethanol solution of paraformaldehyde (4%) and glutaraldehyde (0.25%) for 4 h (for 1.5 h in a vacuum) at room temperature. In the next step, material was washed in PBS three times for 15 min, and then, material was dehydrated in a series of PBS with ethanol (5, 10, 20, 30, 50, 70, 80, 90, 95, 100, 100%) for 15 min each. Embryo axes with 100% ethanol were supplemented with a proportional amount of Steedman wax (Electron Microscopy Sciences, Hatfield, PA, USA) at 37 °C. Material was cut into 10-µm-thick slides and affixed on microscope glasses with protein glycerol (Roth, Karlsruhe, Germany). Slides were stained after rehydration in a series of PBS with ethanol (100, 100, 95, 90, 80, 70, 50, 30, 10%) for 15 min each. Slides were treated with ammonium chloride (0.1 M) for 5 min and then 5% BSA for 30 min, at room temperature. Slides were incubated overnight at 4 °C with primary antibodies in a dilution of 1:140 for ABI5 and RGL2 (Agrisera, Vännäs, Sweden). Afterwards, slides were washed with PBS. Slides were incubated with secondary antibodies conjugated with fluorescein isothiocyanate isomer I (FITC; Sigma-Aldrich, Poznan, Poland) at a dilution of 1:50 for 2 h at room temperature. Staining was done with propidium iodide for 20 min; then, each slide was sealed with the antifadent solution AF1 (Citifluor, London, UK). Confocal imaging of stained tissues was performed using a Leica SP5 confocal microscope (Leica Microsystem GmbH, Wetzlar, Germany). Imaging used 488 nm and emission wavelengths were collected at 525 nm.
Results
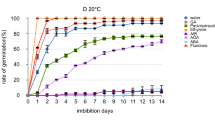
An analysis of Norway maple seed germination (Figs. 1, 2, 3), after dormancy breaking caused by cold stratification, showed that seeds started to germinate during week 7 of stratification, and by week 12, seeds reached 43% of germination. Stratification ended in week 17 when seeds reached 94% of germination.
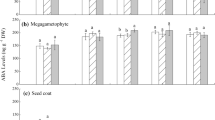
Changes in accumulation of ABI5 protein in Norway maple (Acer platanoides L.) seeds during dormancy breaking. Western blot analyses were performed during stratification (dry—germinated seeds). Statistically significant differences were indicated with different letters. Data are mean ± SD of three independent replications, n = 3
Changes in accumulation of 14-3-3 protein in Norway maple (Acer platanoides L.) seeds during dormancy breaking. Western blot analyses were performed during stratification (dry—germinated seeds). Statistically significant differences were indicated with different letters. Data are mean ± SD of three independent replications, n = 3
Changes in accumulation of RGL2 protein in Norway maple (Acer platanoides L.) seeds during dormancy breaking. Western blot analyses were performed during stratification (dry—germinated seeds). Statistically significant differences were indicated with different letters. Data are mean ± SD of three independent replications, n = 3
Immunodetection of ABI5, 14-3-3 and RGL2 proteins
Western blot analysis used to estimate the accumulation of transcription factor ABI5 indicated the presence of this protein in dormant dry Norway maple seeds at a relatively high level. From the beginning of stratification, ABI5 increased in abundance until week 2 and then diminished gradually until week 8 (Fig. 1), reaching its minimum value in germinated seed. ABI5 accumulation increased in week 10 of stratification, however, when germination started to rise. A decrease in ABI5 accumulation was associated with beginning of seed germination.
Analysis of 14-3-3 protein accumulation during dormancy release of Norway maple seeds showed the presence of this protein in dormant dry seeds at a relatively high level. Generally, the level of protein increased from the beginning of stratification until week 5 and then decreased to levels similar to those found in dry seeds (Fig. 2). Similar to ABI5, a decrease in 14-3-3 accumulation was observed together with the start of germination.
Analysis of RGL2 protein accumulation in Norway maple seeds showed the presence of this protein in dormant dry seeds at a relatively high level, and it decreased after 48 h of imbibition. In week 4 of stratification, accumulation of RGL2 increased to level similar to this found in dry seeds. During week 12, when seeds reached 43% of germination, RGL2 reached its maximum level. Germinated seeds of Norway maple showed a similar pattern of RGL2 accumulation as that found in dry seeds.
Immunolocalisation of ABI5 and RGL2
Analysis of ABI5 immunolocalisation showed the fluorescence signal in each part of the embryo axes of dormant dry Norway maple seeds (Fig. 4). Fluorescence was indicated in the nuclei of cells of the root apical meristem. High intensity of fluorescence was noted also in nuclei of epidermal cells and the outer layers of the cortex. In the fourth week of stratification, we observed changes in the localisation of the fluorescent signal. Along with still being observed in all regions of the embryonic root, strong fluorescence was observed in nucleoli, usually two in the nucleus of some cortex cells. In the eighth week of stratification, fluorescence intensity was very poor in the apical meristem cells and the upper layers of the root cap columella. Fluorescence also decreased in the cells of the primary cortex and the axial cylinder, whereas it remained strong in the nuclei of root cap cells. After germination, strong fluorescence was still observed in the nuclei of the root cap, epidermal and the outer layers of the cortex cells. In the other regions, no ABI5 signal was observed.
Localisation of ABI5 fluorescence signal in embryo root of Norway maple isolated from dry seeds (Dry), following weeks of seed stratification (4, 8) and germinated seed (Germ.); longitudinal section, CLSM; Ep epidermis. Dry A = part of root apex with root apical meristem (RAM; frame); fluorescence of nucleus is visible in all parts. Dry B = part of embryo root, fluorescence signal of nucleus in root cap cells, epidermis cells, cortex and vascular cylinder. 4 = intense fluorescence of large nucleoli in nuclei of some cells of the cortex. 8 = root apex with fluorescence in nuclei of cap cells, epidermis; in RAM (frame), weak fluorescence of nuclei is visible in only some cells. Germ. = fluorescence of nuclei in root cap cells, epidermis and cortex cells; in some cells of epidermis, fluorescence signals are visible in two nucleoli in the nucleus
The RGL2 visualisation experiment showed a very weak fluorescence signal in the cell nuclei of the epidermis and outer layers of the cortex in dormant dry seeds (Fig. 5). Fluorescence was observed in all cells of the root cap. Within the root cap, a particularly intense signal was indicated in a specific region consisting of a couple of cells. Strong fluorescence was noted in the endoderm cells and also in the cells forming into vascular bundles. In the other regions of the root, fluorescence was weak and present only in single cells. In the fourth week, a strong increase in the intensity of the fluorescent signal was observed. In addition to the previously mentioned regions, fluorescence was also found in all the cortex cells and the axial cylinder. In the nuclei of primary cortical cells, the presence of nucleoli with a particularly strong fluorescence was indicated. Weaker, unevenly located signal was also present in the rest part of the nuclei. This RGL2 position was maintained in the sixth week, wherein two large nucleoli were indicated in the nuclei of primary cortex cells. In the eighth week, fluorescence was weaker and nearly invisible within the apical meristem. A strong signal was observed only in the apical part of the root cap and the area of vascular bundles. Similar images were also recorded in the tenth week. The change in RGL2 localisation occurred in the twelfth week. At that time, fluorescence of nuclei in the cortex cells was not observed, but strong point signals were observed in the cytoplasm. In germinated seeds, there was no fluorescence in the region of the root apical meristem. Instead, the signal was intense in the nuclei of epidermal cells and primary cortex cells. Fluorescence was also reported, but weaker, in a part of axial cylinder cells. In the nuclei, we observed lack of nucleoli and the fluorescence signal did not cover the whole surface of the nucleus; signal was instead diffused.
Localisation of RGL2 fluorescence signal in embryo root of Norway maple isolated from dry seeds (Dry), following weeks of seed stratification (4, 6, 8, 10, 12) and germinated seed (Germ.); longitudinal section, CLSM; Ep epidermis, Nu nucleoli, RC root cap. Dry A = root cap cells with fluorescence signal in nuclei. Dry B = fluorescence in RAM is not visible; very weak signal in epidermis and cortex cells. Dry C = intensive signal in procambium; weak in epidermis cells. 4 A = fluorescence signal is visible in nuclei of epidermis, cortex cells, procambium (signal especially intense) and vascular cylinder cells. 4 B = in nuclei of cortex there are large nucleoli with an intense fluorescent signal; unevenly distributed, weak signal is also visible in chromatin area. 6 = in cortex, fluorescence was observed only in large nucleoli (often two in nucleus). 8 = a root cap with fluorescence in nuclei. 8 B = in RAM signal is poorly visible. 10 = intense signal in epidermis, cortex cells and procambium; fluorescence in nuclei of vascular cylinder is weak. 12 = fluorescence observed as a spot in cytoplasm of cortex cells. Germ. A = in RAM fluorescence is not visible. Germ. B = fluorescence signal visible in nuclei of all parts of root. Germ. C = fragment of Germ. B; in nucleus, signal is unevenly distributed as a spot. Germ. D = different fragments of Germ. B; intense signal in nuclei of epidermis cells
Discussion
It was hypothesised that seed dormancy status is controlled according to antagonistic influences of GA and ABA (Luckwill 1952). This hypothesis has been confirmed in previous studies carried out on seeds characterised mostly by non-deep dormancy (Piskurewicz et al. 2008). In this report, we present the results of an investigation of the association of ABA and GA signalling regulators with deep dormancy breaking caused by cold stratification. Norway maple seeds were chosen as the model for investigation because of their deep physiological embryo dormancy (Staszak and Pawłowski 2012).
Quantitative changes in ABI5 and 14-3-3 protein accumulation
ABI5 plays a key role in ABA signal transduction, but its association with regulation of seed germination and seedling growth is not yet well elucidated (Dekkers et al. 2016). ABI5 functions as a transcriptional regulator by binding to an ABA-responsive element, ABRE, which is found in the ABA-induced gene promoters (Hattori et al. 2002). The high level of ABA in dry seeds promotes SnRK2-dependent phosphorylation and stabilisation of ABI5 proteins (Nakashima et al. 2009). ABA stimulates ABI5 gene expression, resulting in germination suppression (Piskurewicz et al. 2009). During stratification, when dormancy is breaking, the level of ABA decreases (Piskurewicz et al. 2008). The reduction in ABA content in Norway maple seeds associated with loss of dormancy was observed by Pinfield et al. (1990). In the present results, accumulation of ABI5 in Norway maple seeds also decreased with loss of dormancy, which probably blocked ABA-dependent gene expression and caused dormancy breaking and, consequently, seed germination. High level of ABI5 in dry Norway maple seeds is responsible for deep dormancy maintenance. The increase in ABI5 accumulation observed after imbibition and during the first few weeks of stratification suggests the preventing role of this ABA signalling regulator in germination control. Cold stratification diminished this influence, as ABI5 levels decreased and seeds germinated. Germinated Norway maple seeds have a relatively small amount of ABI5 protein, which was also observed for Arabidopsis (Lopez-Molina et al. 2002). The presence of ABI5 in germinated seeds may be connected with preparation for post-germination growth, which is regulated via the ABI5 protein (Lopez-Molina et al. 2001; Dai et al. 2013). Increase in ABI5 accumulation observed when germination started to rise was associated probably with activation of some, finally unsuccessful, processes inhibiting germination. We can conclude that accumulation of ABI5 is associated with level of dormancy in Norway maple seeds.
The 14-3-3 proteins are the modulators of effector proteins that bind phosphorylated motifs (Ferl 1996). These proteins interact with ABI5 in ABA signal pathway (Schoonheim et al. 2009). RNAi silencing of 14-3-3 inhibits GA induction of the alpha-amylase gene (Schoonheim et al. 2009). The 14-3-3 protein interplays with ABA binding factors (ABFs), which also interplay with the ABA-responsive kinase PKABA1, a mediator kinase between the GA and ABA pathways. 14-3-3s and ABFs play a regulatory function in the GA signalling, and PKABA1 and ABFss are cross-talk intermediates in ABA and GA pathways (Schoonheim et al. 2009). Analysis of 14-3-3 accumulation during dormancy breaking of Norway maple seeds showed similar patterns to ABI5 accumulation. The increase in accumulation of these proteins in the first few weeks of stratification was likely associated with dormancy maintenance. Start of germination was associated with decrease in accumulation of these proteins. Overall, this suggests an association between the ABA signal pathway and dormancy level in Norway maple seeds.
Localisation of ABI5
Little is known about tissue localisation of the ABI5 protein in seeds. This protein has shown nuclear localisation in epidermal cells (of onion and Nicotiana benthamiana), stamen and corn (of gladiolus) (Wu et al. 2015). In dry Norway maple seeds, ABI5 was noticeable in cell nuclei in each part of the embryo axes. During stratification, the localisation changed and ABI5 was observed mainly in the cortex. Localisation of ABI5 was noticed in the nucleoli of some cortex cells, which suggests that de novo transcription of ABA-dependent genes associated with dormancy breaking was activated. At the end of stratification, when dormancy is broken, ABI5 was not noticeable in the cells of the apical meristem. In germinated seeds, ABI5 was present in the nuclei of epidermal cells and the outer layers of the cortex. It can be concluded that ABI5 localisation changed during dormancy release, from apical meristem to cortex. The presence of ABI5 in germinated seeds suggests that it has a part in regulation of processes during post-germinative growth.
Nuclear localisation of ABI5 was characteristic for Norway maple, as well as for Arabidopsis seeds, which is associated with the fact that gene activation and deactivation occurred in the nucleus (Lopez-Molina et al. 2002). The KEG E3 protein maintains low level of ABI5 by targeting it for degradation in the cytoplasm (Liu and Stone 2013). Thus, in the absence of ABA, nuclear accumulation of ABI5 is reduced (Liu and Stone 2013). In the present study, we noticed reduction in the ABI5 fluorescence signal in nuclei of meristem cells. KEG E3 is likely involved in this degradation.
Quantitative changes in RGL2 concentration
GA fosters germination by increasing the disintegration of DELLA repressors by ubiquitin/proteasome mechanism (Piskurewicz et al. 2009). Members of DELLA protein family play a role in different processes of plant life, such as development of chloroplasts, fruits, leaves and senescence (Chen et al. 2014; Gallego-Giraldo et al. 2014). RGL2 takes part in the dormancy maintenance as a key negative controller of seed germination (Lee et al. 2002), but it is also involved in floral development, in cooperation with RGA and RGL1 (Cheng et al. 2004; Tyler et al. 2004). DELLAs control gene expression through interactions with bHLH transcription factors, such as PIF3, PIF4, PIL5, PIL2 and SPT (de Lucas et al. 2008; Gallego-Bartolomé et al. 2010), as well as through interaction with the GID1 GA receptor (Ueguchi-Tanaka et al. 2005; Murase et al. 2008). GA promotes expression of expansin genes by removing the DELLA repression effect during seed germination (Yan et al. 2014).
In seeds of Norway maple, the level of RGL2 accumulation was relatively high in dry seeds and diminished with imbibition, but increased again during stratification. Our results differ from the results of Wang et al. (2013), who observed that the expression of PpRGL2 was highest in dormant Pyrus pyrifolia seeds, but decreased significantly in the imbibed and germinating embryos. This decrease was suppressed by exogenous ABA (Wang et al. 2013). RGL2 inhibited germination of Arabidopsis seeds by stimulation of ABA synthesis and activity of ABI5 and ABI3 (Piskurewicz et al. 2008, 2009). RGL2 also regulated ABA release in the endosperm to control embryo growth (Lee et al. 2010). These previous studies suggest that there is a link between ABA and GA signalling pathways in germination control, but in the present study this was not confirmed by the results. Similar results were noticed by Tyler et al. (2004) in imbibed Arabidopsis seeds, where a decrease in RGL2 accumulation was associated with high amounts of GA. Gallego-Bartolomé et al. (2010) showed that the level of RGL2 expression increased after even 190 h of imbibition in light exposition. A similar effect was noticed in the present study, but it appeared to be caused by cold stratification. Germinated seeds had relatively low level of RGL2 accumulation which suggests that its inhibitory effect was removed. The high RGL2 in after-ripened seeds of Arabidopsis sly1 mutant also did not stop seed germination, which may indicate that repressive activity of RGL2 can be lost over time (Ariizumi and Steber 2007).
Localisation of RGL2
During dormancy release of Norway maple seeds, changes were observed in localisation of the RGL2 protein. In dry dormant seeds, RGL2 was indicated in nuclei of the root cup. The nuclear localisation of RGL2 transcripts was also observed by Gallego-Bartolomé et al. (2010) in Arabidopsis 7-day seedlings. During the first few weeks of stratification of Norway maple seeds, RGL2 was noticed mainly in the nuclei of cortex cells and procambium. In the following weeks, it was visible in primary cortical cell nuclei, mostly in nucleoli, which indicates its involvement in the transcription process. Chen et al. (2015) reported the presence of nucleoli in plumular apical meristem and the first true leaves in Acer morrisonense seeds stratified for 12 weeks. At the end of stratification, when Norway maple seeds started to germinate, RGL2 was almost invisible in the apical meristem but appeared in the area of vascular bundles. RGL2 regulates several aspects of seed germination, such as cell elongation growth via inhibition of EXPA3 and EXPA8 gene activity by binding to the promoters (Stamm et al. 2012). The role of RGL2 in vascular bundle formation could perhaps be explained by inhibition of its growth.
In the germinated Norway maple seeds, RGL2 was highly noticeable in the nuclei of cells of the epidermis and primary cortex cells. Nuclear localisation of the RGL2 protein in Arabidopsis seedlings seven days after germination also demonstrated that the protein was prevalent in the outer layers of the root (Gallego-Bartolomé et al. 2010). RGL2 gene expression was also limited to the elongation region of the root after seeds imbibed water (Lee et al. 2002). The presence of RGL2 in germinated seeds shows that it is likely an important element of the GA pathway even after germination. This confirms our results, as well as the results of Gallego-Bartolomé et al. (2010), wherein the expression of RGL2 in seeds of Arabidopsis was up-regulated in germinated seeds.
These changes in RGL2 presence in tissues of Norway maple embryos can be explained by proteasome degradation via SCFSLY1/GID2 and de novo synthesis of RGL2 (Achard and Genschik 2009). During dormancy breaking, strong point signals were observed in the cytoplasm, suggesting degradation or synthesis of this GA signal regulator.
Author contribution statement
TAP conceived and designed the research. AMS, MG and TAP conducted experiments. AMS, TAP and MG analysed data. AMS and TAP wrote the manuscript. MG revised it critically. All authors read and approved the manuscript.
References
Achard P, Genschik P (2009) Releasing the brakes of plant growth: how GAs shutdown DELLA proteins. J Exp Bot 60:1085–1092. doi:10.1093/jxb/ern301
Achard P, Cheng H, De Grauwe L et al (2006) Integration of plant responses to environmentally activated phytohormonal signals. Science 311:91–94. doi:10.1126/science.1118642
Ariizumi T, Steber CM (2007) Seed germination of GA-insensitive sleepy1 mutants does not require RGL2 protein disappearance in Arabidopsis. Plant Cell 19:791–804. doi:10.1105/tpc.106.048009
Baskin J, Baskin C (2004) A classification system for seed dormancy. Seed Sci Res 14:1–16. doi:10.1079/SSR2003150
Black M, Bewley JD, Halmer P (2008) The encyclopedia of seeds. Science, technology and uses. CABI, Wallingford
Bradford MM (1976) A rapid and sensitive method for the quantitation of microgram quantities of protein utilizing the principle of protein-dye binding. Anal Biochem 72:248–254
Cao D, Hussain A, Cheng H, Peng J (2005) Loss of function of four DELLA genes leads to light- and gibberellin-independent seed germination in Arabidopsis. Planta 223:105–113. doi:10.1007/s00425-005-0057-3
Chen M, Maodzeka A, Zhou L et al (2014) Removal of DELLA repression promotes leaf senescence in Arabidopsis. Plant Sci 219–220:26–34. doi:10.1016/j.plantsci.2013.11.016
Chen S-Y, Chou S-H, Tsai C-C et al (2015) Effects of moist cold stratification on germination, plant growth regulators, metabolites and embryo ultrastructure in seeds of Acer morrisonense (Sapindaceae). Plant Physiol Biochem 94:165–173. doi:10.1016/j.plaphy.2015.06.004
Cheng H, Qin L, Lee S et al (2004) Gibberellin regulates Arabidopsis floral development via suppression of DELLA protein function. Development 131:1055–1064. doi:10.1242/dev.00992
Dai M, Xue Q, Mccray T et al (2013) The PP6 phosphatase regulates ABI5 phosphorylation and abscisic acid signaling in Arabidopsis. Plant Cell 25:517–534. doi:10.1105/tpc.112.105767
de Boer AH, van Kleeff PJM, Gao J (2013) Plant 14-3-3 proteins as spiders in a web of phosphorylation. Protoplasma 250:425–440. doi:10.1007/s00709-012-0437-z
de Lucas M, Daviere J-M, Rodriguez-Falcon M et al (2008) A molecular framework for light and gibberellin control of cell elongation. Nature 451:480-U11. doi:10.1038/nature06520
Dekkers BJW, He H, Hanson J et al (2016) The Arabidopsis DELAY OF GERMINATION 1 gene affects ABSCISIC ACID INSENSITIVE 5 (ABI5) expression and genetically interacts with ABI3 during Arabidopsis seed development. Plant J 85:451–465. doi:10.1111/tpj.13118
Denison FC, Paul A-L, Zupanska AK, Ferl RJ (2011) 14-3-3 proteins in plant physiology. Semin Cell Dev Biol 22:720–727. doi:10.1016/j.semcdb.2011.08.006
Feng S, Martinez C, Gusmaroli G et al (2008) Coordinated regulation of Arabidopsis thaliana development by light and gibberellins. Nature 451:475-U9. doi:10.1038/nature06448
Ferl RJ (1996) 14-3-3 proteins and signal transduction. Annu Rev Plant Physiol Plant Mol Biol 47:49–73. doi:10.1146/annurev.arplant.47.1.49
Finch-Savage W, Leubner-Metzger G (2006) Seed dormancy and the control of germination. New Phytol 171:501–523. doi:10.1111/j.1469-8137.2006.01787.x
Finkelstein RR, Lynch TJ (2000) The Arabidopsis abscisic acid response gene ABI5 encodes a basic leucine zipper transcription factor. Plant Cell 12:599–609. doi:10.1105/tpc.12.4.599
Finkelstein R, Reeves W, Ariizumi T, Steber C (2008) Molecular aspects of seed dormancy. Annu Rev Plant Biol 59:387–415. doi:10.1146/annurev.arplant.59.032607.092740
Fujii H, Chinnusamy V, Rodrigues A et al (2009) In vitro reconstitution of an abscisic acid signalling pathway. Nature 462:660–664. doi:10.1038/nature08599
Fujita Y, Fujita M, Shinozaki K, Yamaguchi-Shinozaki K (2011) ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res 124:509–525. doi:10.1007/s10265-011-0412-3
Gallego-Bartolomé J, Minguet EG, Marín JA et al (2010) Transcriptional diversification and functional conservation between DELLA proteins in Arabidopsis. Mol Biol Evol 27:1247–1256. doi:10.1093/molbev/msq012
Gallego-Giraldo C, Hu J, Urbez C et al (2014) Role of the gibberellin receptors GID1 during fruit-set in Arabidopsis. Plant J 79:1020–1032. doi:10.1111/tpj.12603
Hattori T, Totsuka M, Hobo T et al (2002) Experimentally determined sequence requirement of ACGT-containing abscisic acid response element. Plant Cell Physiol 43:136–140
Hauser F, Waadt R, Schroeder JI (2011) Evolution of abscisic acid synthesis and signaling mechanisms. Curr Biol 21:R346–R355. doi:10.1016/j.cub.2011.03.015
Heckmann AB, Lombardo F, Miwa H et al (2006) Lotus japonicus nodulation requires two GRAS domain regulators, one of which is functionally conserved in a non-legume. Plant Physiol 142:1739–1750. doi:10.1104/pp.106.089508
Himmelbach A, Yang Y, Grill E (2003) Relay and control of abscisic acid signaling. Curr Opin Plant Biol 6:470–479
International Seed Testing Association (1999) International rules for seed testing. Seed Sci Technol 27:1–133
Itoh H, Sasaki A, Ueguchi-Tanaka M et al (2005) Dissection of the phosphorylation of rice DELLA protein, SLENDER RICE1. Plant Cell Physiol 46:1392–1399. doi:10.1093/pcp/pci152
Jakoby M, Weisshaar B, Dröge-Laser W et al (2002) bZIP transcription factors in Arabidopsis. Trends Plant Sci 7:106–111
Lee S, Cheng H, King KE et al (2002) Gibberellin regulates Arabidopsis seed germination via RGL2, a GAI/RGA-like gene whose expression is up-regulated following imbibition. Genes Dev 16:646–658. doi:10.1101/gad.969002
Lee KP, Piskurewicz U, Turecková V et al (2010) A seed coat bedding assay shows that RGL2-dependent release of abscisic acid by the endosperm controls embryo growth in Arabidopsis dormant seeds. Proc Natl Acad Sci USA 107:19108–19113. doi:10.1073/pnas.1012896107
Liu H, Stone SL (2013) Cytoplasmic degradation of the Arabidopsis transcription factor ABSCISIC ACID Insensitive 5 is mediated by the RING-type E3 ligase KEEP ON GOING. J Biol Chem 288:20267–20279. doi:10.1074/jbc.M113.465369
Lopez-Molina L, Mongrand S, Chua N-H (2001) A postgermination developmental arrest checkpoint is mediated by abscisic acid and requires the ABI5 transcription factor in Arabidopsis. Proc Natl Acad Sci USA 98:4782–4787. doi:10.1073/pnas.081594298
Lopez-Molina L, Mongrand S, McLachlin DT et al (2002) ABI5 acts downstream of ABI3 to execute an ABA-dependent growth arrest during germination. Plant J 32:317–328. doi:10.1046/j.1365-313X.2002.01430.x
Lopez-Molina L, Mongrand S, Kinoshita N, Chua N-H (2003) AFP is a novel negative regulator of ABA signaling that promotes ABI5 protein degradation. Genes Dev 17:410–418. doi:10.1101/gad.1055803
Luckwill L (1952) Growth-inhibiting and growth-promoting substances in relation to the dormancy and after-ripening of apple seeds. J Hortic Sci Biotechnol 27:53–277
Ma Y, Szostkiewicz I, Korte A et al (2009) Regulators of PP2C phosphatase activity function as abscisic acid sensors. Science 324:1064–1068. doi:10.1126/science.1172408
Murase K, Hirano Y, Sun T, Hakoshima T (2008) Gibberellin-induced DELLA recognition by the gibberellin receptor GID1. Nature 456:459-U15. doi:10.1038/nature07519
Nakashima K, Fujita Y, Kanamori N et al (2009) Three Arabidopsis SnRK2 protein kinases, SRK2D/SnRK2.2, SRK2E/SnRK2.6/OST1 and SRK2I/SnRK2.3, involved in ABA signaling are essential for the control of seed development and dormancy. Plant Cell Physiol 50:1345–1363. doi:10.1093/pcp/pcp083
Oh E, Kang H, Yamaguchi S et al (2009) Genome-wide analysis of genes targeted by PHYTOCHROME INTERACTING FACTOR 3-LIKE5 during seed germination in Arabidopsis. Plant Cell 21:403–419. doi:10.1105/tpc.108.064691
Park S-Y, Fung P, Nishimura N et al (2009) Abscisic acid inhibits type 2C protein phosphatases via the PYR/PYL family of START proteins. Science 324:1068–1071. doi:10.1126/science.1173041
Pawłowski TA (2007) Proteomics of European beech (Fagus sylvatica L.) seed dormancy breaking: influence of abscisic and gibberellic acids. Proteomics 7:2246–2257
Pawłowski TA (2009) Proteome analysis of Norway maple (Acer platanoides L.) seeds dormancy breaking and germination: influence of abscisic and gibberellic acids. BMC Plant Biol 9:48. doi:10.1186/1471-2229-9-48
Pawłowski TA (2010) Proteomic approach to analyze dormancy breaking of tree seeds. Plant Mol Biol 73:15–25. doi:10.1007/s11103-010-9623-6
Pawłowski TA, Staszak AM (2016) Analysis of the embryo proteome of sycamore (Acer pseudoplatanus L.) seeds reveals a distinct class of proteins regulating dormancy release. J Plant Physiol 195:9–22. doi:10.1016/j.jplph.2016.02.017
Pinfield NJ, Stutchbury PA, Bazaid SA, Gwarazimba VE (1990) Abscisic acid and the regulation of embryo dormancy in the genus Acer. Tree Physiol 6:79–85
Piskurewicz U, Jikumaru Y, Kinoshita N et al (2008) The gibberellic acid signaling repressor RGL2 inhibits Arabidopsis seed germination by stimulating abscisic acid synthesis and ABI5 activity. Plant Cell 20:2729–2745. doi:10.1105/tpc.108.061515
Piskurewicz U, Turečková V, Lacombe E, Lopez-Molina L (2009) Far-red light inhibits germination through DELLA-dependent stimulation of ABA synthesis and ABI3 activity. EMBO J. doi:10.1038/emboj.2009.170
Reyes D, Rodríguez D, González-García MP et al (2006) Overexpression of a protein phosphatase 2C from beech seeds in Arabidopsis shows phenotypes related to abscisic acid responses and gibberellin biosynthesis. Plant Physiol 141:1414–1424. doi:10.1104/pp.106.084681
Rodríguez MV, Mendiondo GM, Maskin L et al (2009) Expression of ABA signalling genes and ABI5 protein levels in imbibed Sorghum bicolor caryopses with contrasting dormancy and at different developmental stages. Ann Bot 104:975–985. doi:10.1093/aob/mcp184
Saavedra X, Modrego A, Rodriguez D et al (2010) The nuclear interactor PYL8/RCAR3 of Fagus sylvatica FsPP2C1 is a positive regulator of abscisic acid signaling in seeds and stress. Plant Physiol 152:133–150. doi:10.1104/pp.109.146381
Schoonheim PJ, Sinnige MP, Casaretto JA et al (2007) 14-3-3 adaptor proteins are intermediates in ABA signal transduction during barley seed germination. Plant J Cell Mol Biol 49:289–301. doi:10.1111/j.1365-313X.2006.02955.x
Schoonheim PJ, Costa Pereira DDA, De Boer AH (2009) Dual role for 14-3-3 proteins and ABF transcription factors in gibberellic acid and abscisic acid signalling in barley (Hordeum vulgare) aleurone cells. Plant Cell Environ 32:439–447. doi:10.1111/j.1365-3040.2009.01932.x
Stamm P, Ravindran P, Mohanty B et al (2012) Insights into the molecular mechanism of RGL2-mediated inhibition of seed germination in Arabidopsis thaliana. BMC Plant Biol 12:179. doi:10.1186/1471-2229-12-179
Staszak AM, Pawłowski T (2012) Forest tree research in post genomic era. Introduction to systems biology of broadleaves. Dendrobiology 68:113–123
Staszak AM, Pawłowski TA (2014) Proteomic analysis of embryogenesis and the acquisition of seed dormancy in Norway maple (Acer platanoides L.). Int J Mol Sci 15:10868–10891. doi:10.3390/ijms150610868
Sun X, Jones WT, Harvey D et al (2010) N-terminal domains of DELLA proteins are intrinsically unstructured in the absence of interaction with GID1/gibberellic acid receptors. J Biol Chem 285:11557–11571. doi:10.1074/jbc.M109.027011
Sun X, Jones WT, Rikkerink EHA (2012) GRAS proteins: the versatile roles of intrinsically disordered proteins in plant signalling. Biochem J 442:1–12. doi:10.1042/BJ20111766
Tyler L, Thomas SG, Hu J et al (2004) DELLA proteins and gibberellin-regulated seed germination and floral development in Arabidopsis. Plant Physiol 135:1008–1019. doi:10.1104/pp.104.039578
Ueguchi-Tanaka M, Ashikari M, Nakajima M et al (2005) GIBBERELLIN INSENSITIVE DWARF1 encodes a soluble receptor for gibberellin. Nature 437:693–698. doi:10.1038/nature04028
Wang Y, Dai M, Zhang S, Shi Z (2013) Exploring the hormonal and molecular regulation of sand pear (Pyrus pyrifolia) seed dormancy. Seed Sci Res 23:15–25. doi:10.1017/S096025851200027X
Wu J, Seng S, Sui J et al (2015) Gladiolus hybridus ABSCISIC ACID INSENSITIVE 5 (GhABI5) is an important transcription factor in ABA signaling that can enhance Gladiolus corm dormancy and Arabidopsis seed dormancy. Front Plant Sci. doi:10.3389/fpls.2015.00960
Xu D, Li J, Gangappa SN et al (2014) Convergence of light and ABA signaling on the ABI5 promoter. PLoS Genet 10:e1004197. doi:10.1371/journal.pgen.1004197
Yan A, Wu M, Yan L et al (2014) AtEXP2 is involved in seed germination and abiotic stress response in Arabidopsis. PLoS One 9:e85208. doi:10.1371/journal.pone.0085208
Acknowledgements
The authors wish to thank Sylwia Masłowska for technical support. The work was supported by a Grant from the National Science Centre, Poland (Grant number 2011/01/B/NZ9/02868), to TAP, and a Grant from Jung Scientist, Ministry of Science and Higher Education, Poland, to AMS.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by H. Li.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Staszak, A.M., Guzicka, M. & Pawłowski, T.A. Signalling regulators of abscisic and gibberellic acid pathways are involved in dormancy breaking of Norway maple (Acer platanoides L.) seeds. Acta Physiol Plant 39, 251 (2017). https://doi.org/10.1007/s11738-017-2544-0
Received:
Revised:
Accepted:
Published:
DOI: https://doi.org/10.1007/s11738-017-2544-0