Abstract
The primary aim of the present study was to reveal the major differences between benzene-degrading bacterial communities evolve under aerobic versus microaerobic conditions and to reveal the diversity of those bacteria, which can relatively quickly degrade benzene even under microaerobic conditions. For this, parallel aerobic and microaerobic microcosms were set up by using groundwater sediment of a BTEX-contaminated site and 13C labelled benzene. The evolved total bacterial communities were first investigated by 16S rRNA gene Illumina amplicon sequencing, followed by a density gradient fractionation of DNA and a separate investigation of “heavy” and “light” DNA fractions. Results shed light on the fact that the availability of oxygen strongly determined the structure of the degrading bacterial communities. While members of the genus Pseudomonas were overwhelmingly dominant under clear aerobic conditions, they were almost completely replaced by members of genera Malikia and Azovibrio in the microaerobic microcosms. Investigation of the density resolved DNA fractions further confirmed the key role of these two latter genera in the microaerobic degradation of benzene. Moreover, analysis of a previously acquired metagenome-assembled Azovibrio genome suggested that benzene was degraded through the meta-cleavage pathway by this bacterium, with the help of a subfamily I.2.I-type catechol 2,3-dioxygenase. Overall, results of the present study implicate that under limited oxygen availability, some potentially microaerophilic bacteria play crucial role in the aerobic degradation of aromatic hydrocarbons.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Benzene is a known carcinogenic organic compound, classified as a priority pollutant by the US Environmental Protection Agency. Its occurrence in gasoline contaminated subsurface environments is general, and due to its relatively high solubility in water (1.79 g/L), considerably threatens subsurface water reservoirs. Thus, the removal of benzene from polluted sites is crucial. Since a plethora of microorganisms can use benzene as a sole source of carbon and energy, bioremediation is one of the most frequently applied processes of benzene removal. Still, bioremediation of benzene contaminated subsurface ecosystems is challenging due to the fact that the vast majority of the degraders use aerobic metabolic pathways for the activation and cleavage of the benzene ring. Under aerobic conditions these key steps are catalysed by mono- and dioxygenases, which require molecular oxygen as a co-substrate. However, availability of oxygen in subsurface environments, even in pristine ones is usually limited. In highly contaminated environments, the aerobic degraders relatively quickly deplete oxygen, giving chance to anaerobic degraders (nitrate-, iron-, or sulphate-reducers, methanogens) to cope with the contamination Meckenstock et al. (2016). Nevertheless, benzene is very slowly degraded without molecular oxygen and this fact makes air sparging technique a preferred and successful bioremediation process in case of large-scale subsurface BTEX-contaminations Kabelitz et al. (2009). As cost effectiveness is becoming an increasingly important factor in the case of bioremediation processes as well, it is a valid aim to look for bacteria, which can aerobically degrade BTEX-compounds even in the presence of tiny amount of oxygen. It has been known for more than two decades, that aerobic benzene degradation is feasible even under microaerophilic conditions (~ 0.2 mg/L dissolved oxygen) and that phenol is the major intermediate metabolite during the degradation of benzene at low concentrations of dissolved oxygen Yerushalmi et al. (2002). Recently, it has been shown that toluene and xylene can be degraded under microaerobic conditions and mainly a well-defined group of Betaproteobacteria in the families Comamonadaceae, Zoogloeaceae and Rhodocyclaceae were reported as the key toluene- or xylene-degraders under such conditions (Táncsics et al. 2018, 2023). Moreover, it has been shown that meta ring-cleaving catechol 2,3-dioxygenases belonging to I.2.C subfamily of extradiol dioxygenases (EDOs) are functional under hypoxic conditions and play prominent role in the degradation of methyl-substituted aromatics under microaerobic conditions (Kukor and Olsen 1996). Nevertheless, benzene can be degraded through the ortho-cleavage pathway as well, and an ortho ring-cleaving catechol 1,2-dioygenase, adapted to low oxygen availability has already been reported earlier Balcke et al. (2007). Still, it is clear that our knowledge on the diversity of bacteria and ring-cleavage enzymes playing a key role in the degradation of benzene under microaerobic conditions is still in its infancy (Bedics et al. 2022). Therefore, the aims of the present study were to reveal differences between microbial communities degrading benzene under aerobic versus microaerobic conditions and to clearly identify microaerobic benzene-degraders by a microcosm experiment combined with DNA-stable isotope probing.
Materials and methods
Sampling site and sample acquisition
Groundwater sediments were taken from the deeply studied Siklós BTEX-contaminated site of Hungary (Táncsics et al. 2012, 2013, 2023). Previously, sediment sample originated from the same groundwater well (designated as ST2) was used successfully in a SIP study, which aimed to reveal diversity of bacteria capable of degrading toluene under microaerobic conditions (Táncsics et al. 2018). Sediment rich groundwater was pumped out from the bottom of the monitoring well and collected into sterile, 1 L brown coloured glass bottles. Samples were stored at 4 °C and transported to the laboratory immediately.
Incubation of sediments
Triplicates of 5 g (wet weight) homogenously mixed sediment material were transferred into sterile 100-mL serum bottles containing 50 mL of mineral salts (MS) medium. The MS medium was prepared with 100 ml each of solutions A and B, 1 ml of solution C and 800 ml of water. Solution A: 5 g MgSO4·7H2O and 1 g CaCl2·H2O l−1 (filtered). Solution B: 11.1 g Na2HPO4, 2.5 g KH2PO4 and 6.7 g NH4Cl l−1 (autoclaved). Solution C: 10 g FeSO4·7H2O, 0.64 g Na2EDTA·3H2O, 0.1 g ZnCl2, 0.015 g H3BO3, 0.175 g CoCl2·6H2O, 0.15 g Na2MoO4·2H2O, 0.02 g MnCl2·4 H2O and 0.01 g NiCl2·6 H2O l−1 (filtered). Additionally, the medium was supplemented with 1 mg/L thiamine, 15 μg/L biotin, and 20 μg/L vitamin B12. Microaerobic microcosms were sparged aseptically with N2/CO2 (80:20, v/v) for 10 min, after which the desired volume of sterile (0.2-µm-pore-size-filtered) air was injected into the bottles through butyl rubber stoppers. Dissolved oxygen concentration in the microaerobic microcosms was set to 0.5 mg/L and kept between 0.5 and 0 mg/L throughout the experiment. In the aerobic microcosms dissolved oxygen concentration was kept between 6–8 mg/L. Oxygen was replenished once every 24 h. A 5 µL of fully labeled (13C6) benzene (Sigma-Aldrich) was injected into the microcosms. Abiotic control bottles (autoclaved three times) amended with unlabelled toluene were also prepared to exclude abiotic benzene loss. The bottles were incubated at 16 °C in a rotary shaker at 145 rpm.
Process measurements
Dissolved oxygen concentration in the liquid phase of the microcosms was measured by using planar oxygen sensor spots and a Fibox 3 Oxygen Meter (PreSens, Germany). At each sampling spot, dissolved oxygen concentrations were registered every second for 1 min, and the results were displayed by using the OxyView-PST3 software (V7.01, PreSens). Benzene concentrations were determined by headspace analysis on an ISQ Single Quadrupole GC–MS (ThermoFischer Scientific) via a SLB-5 ms fused silica capillary column (Supelco Analytical). The oven temperature was set to 40 °C for 3 min, then ramped at a rate of 20 °C/min to 190 °C, and held for 1 min. The mass spectrometer (MS) was operated at 250 °C in full scan mode.
Nucleic acid extraction and ultracentrifugation
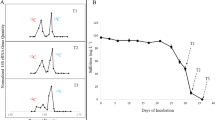
Sediments were collected from sacrificed microcosms after 4 d (aerobic microcosms) or 7 d (microaerobic microcosms) of incubation by centrifugation at 2360 g at 4 °C for 10 min using a Rotanta 460 R (Hettich). DNA was isolated from the sediment pellets by using the NucleoSpin Soil Kit (Macherey–Nagel) according to the instructions of the manufacturer. DNA samples were stored frozen at −80 °C until downstream analyses. For isopycnic density gradient ultracentrifugation, equal volumes (30 µL) of eluted DNA from the triplicate aerobic and microaerobic microcosms were pooled, respectively. This was necessary to get enough amount of DNA for the ultracentrifugation. Subsequently, approximately 1 µg of NanodropOne quantified (Thermo Fisher Scientific) DNA extract was loaded onto 18.5 mL gradient medium of CsCl (average density 1.72 g/mL, Fisher Chemical) in gradient buffer (0.1 M Tris–HCl at pH 8, 0.1 M KCl, 1 mM EDTA) and centrifuged (39,000 rpm, ~ 122,000 RCFav, 72 h) by using a Sorvall Combi ultracentrifuge (DuPont). Centrifugation was performed by using 18.5 mL centrifuge tubes (DuPont) and vertical rotor (Sorvall TV-865B). Following centrifugation, eighteen gradient fractions (1 mL in volume) were collected into sterile 1.5 mL Eppendorf tubes from “heavy” to “light” fractions. Bouyant density (BD) of the fractions was calculated based on the mass of 0.5 mL of each of the gradient fractions. Subsequently, recovery of the DNA from the gradient fractions was performed by using 30% (m/V) polyethylene glycol 6000 (PEG 6000) (Fluka AG) as described by Griffiths et al. (2000).
16S rRNA gene targeted qPCR, Illumina amplicon sequencing and genome-resolved metagenomics
Twelve DNA fractions (from 3rd to 14th) of each gradient were selected for bacterial 16S rRNA gene-targeted qPCR, which was performed as described earlier (Kunapuli et al. 2007; Pilloni et al. 2011).
Illumina 16S rRNA gene amplicon sequencing was carried out both in case of non-density resolved, unique DNA samples of triplicate aerobic and microaerobic microcosms, and selected “light”, “medium” and “heavy” DNA fractions resulted after the isopycnic density gradient centrifugation. The variable V3 and V4 regions of the 16S rRNA gene were amplified by using primers recommended by Klindworth et al. (2013). PCR amplification was carried out using the KAPA HiFi HotStart Ready Mix (KAPA Biosystems) according to the 16S metagenomics sequencing library preparation guide of Illumina. Paired-end fragment reads were generated on an Illumina MiSeq sequencer using a MiSeq Reagent Kit v2 (500 cycles) by Eurofins BIOMI Kft. (Gödöllő, Hungary). Primary data analysis (base-calling) was conducted with Bbcl2fastq^ software (v2.17.1.14, Illumina). Finally, the sequences were processed using Mothur v1.41.1 software Schloss et al. (2009) following the recommendations of the MiSeq SOP manual (http://www.mothur.org/wiki/MiSeq_SOP) Kozich et al. (2013). Based on the sequence alignment as determined by the SILVA 132 SSURef NR99 database Quast et al. (2013), sequences were then assigned. The detection of chimeras was performed with Mothur’s uchime command and the ‘split.abund’ command was also used to remove singleton reads (Edgar et al. 2011; Kunin et al. 2010). A 97% similarity threshold level was used to assign sequences into Operational Taxonomic Units (OTUs) as suggested by Tindall et al. (2010) for prokaryotic species delineation. Amplicon sequence data were deposited at NCBI under the BioProject number PRJNA704528 (BioSamples: SAMN39829928 and SAMN39829929).
The process of metagenome sequencing and reconstruction of bacterial genomes (genome binning) has been described earlier Táncsics et al. (2023). Briefly, paired-end fragment reads (2 × 250 nucleotides) were sequenced using the MiSeq Reagent Kit v2 (500-cycles) on an Illumina MiSeq sequencer by Seqomics Kft. (Mórahalom, Hungary). Quality-controlled (BBDuk and sickle v1.33) reads were assembled using the MetaSPAdes pipeline, version 3.15.0. Nurk et al. (2017). Subsequently, Abawaca Brown et al. (2015) was run twice, first to include reads with a minimal length of 3000 base pairs (bps) and sequences split after 5000 bps and second to set the minimal scaffold length of 5000 bps and sequences split after 10,000 bps. MaxBin2 Wu et al. (2016) was also run on both marker sets. All bins created in the four runs were optimized and filtered using the DAS Tool Sieber et al. (2018) and hand curated with uBin Bornemann et al. (2023). Phylogenetic affiliation of the bins was ascertained using the pipeline of the MiGA–Microbial Genomes Atlas Rodriguez-R et al. (2018). The Azovibrio-related bin genome analyzed in the present study can be found at NCBI under the BioProject number PRJNA818156.
Phylogenetic analyses
Reference genomes for comparison purposes were retrieved from the GenBank database (https://www.ncbi.nlm.nih.gov/genbank). The phylogenomic tree shown in the study was constructed using a concatenated alignment of 92 core genes with UBCGs software Na et al. (2018). The catechol 2,3-dioxygenase-based phylogenetic tree was created with MEGA version 7 Kumar et al. (2016) by using the neighbor-joining algorithm Saitou and Nei (1987). Orthologous average nucleotide identity (OrthoANI) value reported in the study was calculated by using the OAT software Lee et al. (2016), while average amino-acid identity value was determined by the MiGA pipeline Rodriguez-R et al. (2018).
Results and discussion
Benzene degradation in the aerobic vs microaerobic microcosms
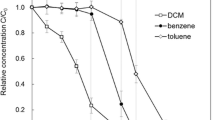
Both aerobic and microaerobic degradation of benzene proved to be a rather rapid process. Under clear aerobic conditions, benzene was depleted after 4 days of incubation (96 h), while in the microaerobic microcosms the degradation was a bit slower process and additional 3 days were needed (a total of 7 days) for the complete degradation of benzene (Fig. S1). This is at least because anoxic periods occurred in the microaerobic microcosms, which slowed down the degradation process. A similar observation was made in our earlier SIP study, in which a seven days-long period was needed for the complete microaerobic degradation of toluene (1 mM) Táncsics et al. (2018). In earlier degradation experiments, which we performed with pure strains and where microaerobic conditions prevailed throughout the whole experiment (due to a much smaller amount of benzene and bacterial biomass) no significant difference in terms of degradation time was observed between aerobic and microaerobic degradation of benzene (Banerjee et al. 2022; Bedics et al. 2024).
Bacterial community composition of aerobic and microaerobic benzene-degrading microcosms
At first, microbial communities were investigated by using the non-density resolved DNA samples obtained from each of the enrichments. In the aerobic enrichments members of the genus Pseudomonas were overwhelmingly dominant (with relative abundance values between 40–51%), Rhizobium (10–15%), Thauera (12–15%) (Fig. 1). The major Pseudomonas OTUs were related to P. aromaticivorans (35–45%) and members of the P. stutzeri complex (4–5%) (Table 1). Both lineages contain prominent aromatic hydrocarbon degrading microorganisms and P. aromaticivorans was described by us previously as a benzene-degrading bacterium (Banerjee et al. 2022; Li et al. 2022). The genus Thauera was represented by T. aminoaromatica, which was described as an aromatic compound-degrading bacterium (Mechichi et al. 2002). The genus Rhizobium was mainly represented by an OTU most closely related to a lineage containing R. alamii and R. mesosinicum. The relatively high abundance of Rhizobium in a benzene-degrading enrichment was unexpected, however, recently Rhizobium cremeum was described as an aromatic compound-degrading bacterium, capable of degrading benzene as well (Yang et al. 2022). Notable minor community members detectable in all of the aerobic enrichments were Zoogloea (2–3%), Rugosibacter (~ 2%) and Malikia (~ 1.5%). In the case of the microaerobic enrichments, members of genera Malikia (26–40%) and Azovibrio (20–28%) dominated the communities, which later lineage was completely missing from the aerobic enrichments. The genus Malikia was represented by an OTU most closely related to M. spinosa. Although the type strain of this bacterium was isolated from a pristine freshwater, it has been shown by us earlier that this bacterium can acquire the ability to degrade aromatic hydrocarbons and can play major role in aerobic benzene degradation Révész et al. (2020). The Azovibrio genus was represented by an OTU most closely related A. restrictus, but only at 95.7% 16S rRNA gene sequence similarity. The same Azovibrio-like bacterium was enriched by us earlier in microaerobic xylene-degrading enrichment cultures Táncsics et al. (2023). Nevertheless, the role of Azovibrio in the degradation of BTEX has never been reported earlier. Moreover, the ecological role of this bacterium often remains hidden in relevant environmental microbiological studies Abbas et al. (2019). It was also observed that the relative abundance of genus Pseudomonas considerably decreased and became a minor player of the microaerobic communities. This decrease was due to the fact that the abundance of the P. aromaticivorans OTU was only 0.5–1% in the microaerobic enrichment communities, while the abundance of the other Pseudomonas OTU remained between 4–6%. Another minor community players of the microaerobic enrichments were Geobacter (4–6%), Extensimonas (2–3%) and Rhizobium with slightly decreased abundance (3–8%). Overall, non-density resolved DNA samples indicated that highly different benzene-degrading microbial communities evolved during the experiment period.
Analysis of density resolved DNA samples
In order to clearly identify the bacterial lineages which were degrading the isotopically labelled benzene (13C6-benzene), community DNA samples were density resolved by isopycnic gradient ultracentrifugation. As a result, “heavy” and “light” DNA fractions were differentiated (Fig. S2) and analysed further by Illumina 16S rRNA gene amplicon sequencing. In case of the aerobic enrichments, genera Pseudomonas and Rhizobium were clearly identified as benzene-degraders, due to the notable increase in the abundance of their 16S rDNA fragments in the heavy DNA fraction compared to that of the light fraction (Fig. 2). In case of the P. aromaticivorans OTU, its 36.8% abundance observed in the light fraction increased to 43% in the heavy DNA fraction. However, in case of the P. stutzeri OTU, an opposite trend was observable, as its abundance was higher in the light fraction (4.0% vs 2.7%). Accordingly, P. aromaticivorans was identified as a prominent benzene degrader under clear aerobic conditions. On the other hand, this OTU was hardly detectable in the heavy DNA fraction of the microaerobic enrichments, reaching only 0.15% abundance. Nevertheless, this bacterium was described as a microorganism capable of degrading benzene even under microaerobic conditions Banerjee et al. (2022) in pure culture. Consequently, the fact that a bacterium can degrade aromatic compounds under microaerobic conditions in pure culture does not mean that it will be a competitive microaerobic degrader as a member of a community. Interestingly, the abundance of Thauera was lower in the heavy DNA fraction than in the light DNA fraction (12.0% vs 16.6%). On the other hand in the medium fractions its abundance was considerably higher, e.g. in fraction A8 (BD = 1.715) it was ~ 23% (data not shown). Similar observation was made by us earlier in case of a microarobic, 13C7-toluene–degrading enrichment community, where the DNA of an aromatic compound-degrading Rhodoferax bacterium was primarily found in medium DNA fractions Táncsics et al. (2018). As mentioned above, Rhizobium was identified as benzene-degrader, since its abundance was considerably higher in the heavy DNA fraction than in the light one (23.0% vs 12.5%). In case of the microaerobic enrichments the two major players, Malikia and Azovibrio were clearly identified as benzene-degraders. DNA of both lineages showed considerably higher abundance in the heavy fraction than in the light fraction (42% vs 33% and 33% vs 26%, respectively). Regarding minor community members of the microaerobic enrichments, Sulfuricurvum kujiense-related DNA was detected only in the light DNA fraction. This result is not surprising, as this bacterium was described as facultatively anaerobic, chemolithoautotrophic, sulfur-oxidizing bacterium, which was first isolated from an underground crude-oil storage (Kodama and Watanabe 2004). Similarly, Geobacter-related DNA was detected only in the light DNA fraction. Although some of these strictly anaerobic bacteria can degrade benzene or toluene anaerobically (Kunapuli et al. 2010; Zhang et al. 2013), they have no role in the aerobic degradation of aromatic compounds. Consequently, this lineage was not labelled during the experiment. The abundance of Rhizobium-related DNA was slightly enriched in the heavy DNA fraction, while Extensimonas-related DNA was almost equal in the heavy and the light DNA fractions. This means that the genus Rhizobium-related bacteria could have some minor role in the degradation under microaerobic conditions, while the role of Extensimonas remained unknown. This later genus belongs to the family Comamonadaceae, which contains numerous genera involved in aromatic hydrocarbon degradation (e.g. Acidovorax, Comamonas, Hydrogenophaga, Polaromonas, Variovorax etc.). On the other hand, only handful of studies link the genus Extensimonas to hydrocarbon degradation Wang et al. (2021).
Investigation of a relevant metagenome-assembled Azovibrio genome
As mentioned above, an Azovibrio bacterium was highly abundant in the microaerobic benzene-degrading microcosms. Besides, in one of our previous enrichment experiments of the Siklós sediment, which aimed to investigate xylene degradation under microaerobic conditions, the same Azovibrio lineage was dominant Táncsics et al. (2023). Although the whole genome of this most probably aromatic hydrocarbon-degrading bacterium was assembled earlier by using metagenomic data, it has not been analysed yet. The results of the present study directed our attention to this metagenome-assembled genome, designated as Azovibrio sp. XYLBin2 (WGS accession number JANLJF000000000). Thus, its phylogenomic and functional analysis was performed. Accordingly, it was revealed by the MiGA pipeline, that this Azovibrio-related genome is 2926 790 bp, while its completeness is 99.1%, with low level of contamination (3.8%). The UBCG phylogenomic tree indicated that Azovibrio sp. XYLBin2 clustered most closely to Azovibrio restrictus DSM 23866 T (Fig. 3). On the other hand, the low OrthoANI and AAI values (82.2% and 72.6%, respectively) between the two genomes indicated that the XYLBin2 genome represents a yet uncultured member of the genus Azovibrio.
Functional analysis of the metagenome-assembled genome revealed a gene cluster encoding all necessary elements of multicomponent phenol hydroxylase (mPH) system. Activation of the aromatic ring is often performed by benzene/toluene monooxygenases or dioxygenases. However, if these are not available in the genome, the mPH system performs this task in the same way. Although its primary substrate is phenol, it can also catalyze the transformation of benzene into catechol Cafaro et al. (2004). The activity of the enzyme requires the coordinated operation of the dmpKLMNOP genes, which are responsible for the expression of the P0, P1, P2, P3, P4 and P5 components of the enzyme system. As mentioned above, all of these genes were found in the corresponding gene cluster, assuming that benzene was converted to catechol through the activity of the mPH system in the case of Azovibrio sp. XYLBin2. The next step in the aerobic degradation of benzene is the cleavage of the aromatic ring, either through ortho- or meta-cleavage. In the XYLBin2 genome a catechol 2,3-dioxygenase (C23O) gene was found, thus benzene is degraded through the meta-cleavage pathway by this Azovibrio-related bacterium. Phylogenetic analysis of the C23O gene revealed that it encodes an extradiol dioxygenase (EDO), which belongs to the recently described EDO subfamily I.2.I (Fig. 4) Táncsics et al. (2023). It showed the highest similarity to a C23O gene of Dechloromonas denitrificans strain 110 (85% similarity at nucleotide level), followed by C23O genes of Thauera and Azoarcus strains, e.g. Azoarcus olearius DQS-4 T, which was isolated from oil-contaminated soil, but primarily described as a nitrogen-fixing plant growth-promoting bacterium (Chen et al. 2013; Faoro et al. 2016). Besides the C23O gene, genes encoding the oxalocrotonate branch of the lower meta-cleavage pathway enzymes (e.g. 2-hydroxymuconate semialdehyde dehydrogenase; 4-oxalocrotonate tautomerase, 4-oxalocrotonate decarboxylase etc.) were also found in the XYLBin2 genome, suggesting that benzene was degraded through the meta-pathway.
Neighbor-joining phylogenetic tree of the deduced amino acid sequences of catechol 2,3-dioxygenase (C23O) genes encoding class I EDO enzymes. The Azovibrio sp. XYLBin2-related catechol 2,3-dioxygenase revealed by the present study is highlighted with boldface type. Class I EDO subfamilies are indicated at the main branches of the tree. Only bootstrap values > 50 are indicated. Scale bar, 0.2 substitutions per amino acid position
It has to be noted here, that the genus Azovibrio contains only one single species, A. restrictus, which was reported to have limited metabolic abilities (Reinhold-Hurek and Hurek 2000). Members of the genus were shown to grow on a limited number of carbon sources (e.g. L-malate, DL-lactate, succinate, fumarate, acetate, propionate and ethanol, and amino acids such as L-glutamate), and they cannot grow on mono- or disaccharides. Their most widely known characteristic is that they can fix atmospheric nitrogen. Besides, some strains show strict microaerophilic lifestyle (Reinhold-Hurek and Hurek 2000) Overall, increasing number of evidence suggests that these potentially microaerophilic bacteria of the Rhodocyclales, aquiring subfamily I.2.I-type EDO genes have another ecological role, namely the degradation of monoaromatic hydrocarbons in petroleum hydrocarbon contaminated subsurface ecosystems.
Conclusion for future biology
The main conclusion of the present study, which potentially affects future bioremediation procedures of petroleum hydrocarbon contaminated subsurface ecosystems, is that distinctly different bacterial communities took part in the degradation of benzene under aerobic versus microaerobic conditions. While under clear aerobic conditions members of the genera Pseudomonas, Thauera and Rhizobium were the key degraders, their role under microaerobic conditions was rather marginal. On the other hand, under microaerobic conditions, members of the genera Malikia and Azovibrio mostly replaced these bacteria. Consequently, results confirmed the previous observation, that Malikia species can be prominent benzene degraders in contaminated aquifers. Additionally, the present study provided further evidence on the assumption that members of the genus Azovibrio, which are potentially microaerophilic bacteria, can play a significant role in the degradation of aromatic hydrocarbons under limited oxygen availability. Whether the subfamily I.2.I-type extradiol dioxygenase provides further competitive advantage to these bacteria over other aerobic hydrocarbon degraders (e.g. over Pseudomonas spp.), remains a question and needs to be further elucidated.
References
Abbas SZ, Rafatullah M, Khan MA, Siddiqui MR (2019) Bioremediation and electricity generation by using open and closed sediment microbial fuel cells. Front Microbiol 9:3348. https://doi.org/10.3389/fmicb.2018.03348
Balcke GU, Wegener S, Kiesel B, Benndorf D, Schlömann M, Vogt C (2007) Kinetics of chlorobenzene biodegradation under reduced oxygen levels. Biodegradation 19:507–518. https://doi.org/10.1007/s10532-007-9156-0
Banerjee S, Bedics A, Tóth E, Kriszt B, Soares AR, Bóka K, Táncsics A (2022) Isolation of Pseudomonas aromaticivorans sp nov from a hydrocarbon-contaminated groundwater capable of degrading benzene- toluene- m- and p-xylene under microaerobic conditions. Front Microbiol 13:929128. https://doi.org/10.3389/fmicb.2022.929128
Bedics A, Táncsics A, Tóth E, Banerjee S, Harkai P, Kovács B, Bóka K, Kriszt B (2022) Microaerobic enrichment of benzene-degrading bacteria and description of Ideonella benzenivorans sp. nov., capable of degrading benzene, toluene and ethylbenzene under microaerobic conditions. Antonie van Leeuwenhoek 115:1113–1128. https://doi.org/10.1007/s10482-022-01759-z
Bedics A, Táncsics A, Banerjee S, Tóth E, Harkai P, Gottschall GG, Bóka K, Kriszt B (2024) Acidovorax benzenivorans sp nov a novel aromatic hydrocarbon-degrading bacterium isolated from a xylene-degrading enrichment culture. Int J Syst Evol Microbiol. https://doi.org/10.1099/ijsem.0.006219
Bornemann TLV, Esse SP, Stach TL, Burg T, Probst AJ (2023) uBin: A manual refining tool for genomes from metagenomes. Environ Microbiol 25:1077–1083. https://doi.org/10.1111/1462-2920.16351
Brown CT, Hug LA, Thomas BC, Sharon I, Castelle CJ, Singh A, Wilkins MJ, Wrighton KC, Williams KH, Banfield JF (2015) Unusual biology across a group comprising more than 15% of domain Bacteria. Nature 523:208–211. https://doi.org/10.1038/nature14486
Cafaro V, Izzo V, Scognamiglio R, Notomista E, Capasso P, Casbarra A, Di Donato A (2004) Phenol hydroxylase and toluene/o-xylene monooxygenase from Pseudomonas stutzeri OX1: interplay between two enzymes. Appl Environ Microbiol 70:2211–2219. https://doi.org/10.1128/AEM.70.4.2211-2219.2004
Chen MH, Sheu SY, James EK, Young CC, Chen WM (2013) Azoarcus olearius sp. nov., a nitrogen-fixing bacterium isolated from oil-contaminated soil. Int J Syst Evol Microbiol 63:3755–3761. https://doi.org/10.1099/ijs.0.050609-0
Edgar RC, Haas BJ, Clemente JC, Quince C, Knight R (2011) UCHIME improves sensitivity and speed of chimera detection. Bioinformatics 27:2194–2200. https://doi.org/10.1093/bioinformatics/btr381
Faoro H, Menegazzo RR, Battistoni F, Gyaneshwar P, do Amaral FP, et al (2016) The oil-contaminated soil diazotroph Azoarcus olearius DQS-4T is genetically and phenotypically similar to the model grass endophyte Azoarcus sp. BH72. Env Microbiol Rep 9:223–238. https://doi.org/10.1111/1758-2229.12502
Griffiths RI, Whiteley AS, O’Donnell AG, Bailey MJ (2000) Rapid method for coextraction of DNA and RNA from natural environments for analysis of ribosomal DNA-and rRNA-based microbial community composition. Appl Environ Microbiol 66:5488–5491. https://doi.org/10.1128/aem.66.12.5488-5491.2000
Kabelitz N, Machackova JImfeld G, Brennerova M, Pieper DH, Heipieper HJ, Junca H, (2009) Enhancement of the microbial community biomas and diversity during air sparging bioremediation of a soil highly contaminated with kerosene and BTEX. Appl Microbiol Biotechnol 82:565–577. https://doi.org/10.1007/s00253-009-1868-0
Klindworth A, Pruesse E, Schweer T, Peplies J, Quast C, Horn M, Glöckner FO (2013) Evaluation of general 16S ribosomal RNA gene PCR primers for classical and next-generation sequencing-based diversity studies. Nucleic Acids Res 41:e1. https://doi.org/10.1093/nar/gks808
Kodama Y, Watanabe K (2004) Sulfuricurvum kujiense gen. nov., sp. nov., a facultatively anaerobic, chemolithoautotrophic, sulfur-oxidizing bacterium isolated from an underground crude-oil storage cavity. Int J Syst Evol Microbiol 54:2297–2300. https://doi.org/10.1099/ijs.0.63243-0
Kozich JJ, Westcott SL, Baxter NT, Highlander SK, Schloss PD (2013) Development of a dual-index sequencing strategy and curation pipeline for analyzing amplicon sequence data on the MiSeq Illumina sequencing platform. Appl Environ Microbiol 79:5112–5120. https://doi.org/10.1128/AEM.01043-13
Kukor JJ, Olsen RH (1996) Catechol 2,3-dioxygenases functional in oxygen-limited (hypoxic) environments. Appl Environ Microbiol 62:1728–1740. https://doi.org/10.1128/aem.62.5.1728-1740.1996
Kumar S, Stecher G, Tamura K (2016) MEGA7: Molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33:1870–1874. https://doi.org/10.1093/molbev/msw054
Kunapuli U, Lueders T, Meckenstock RU (2007) The use of stable isotope probing to identify key iron-reducing microorganisms involved in anaerobic benzene degradation. ISME J 1:643–653. https://doi.org/10.1038/ismej.2007.73
Kunapuli U, Jahn MK, Lueders T, Geyer R, Heipieper HJ, Meckenstock RU (2010) Desulfitobacterium aromaticivorans sp. nov. and Geobacter toluenoxydans sp. nov., iron-reducing bacteria capable of anaerobic degradation of monoaromatic hydrocarbons. Int J Syst Evol Microbiol 60:686–695. https://doi.org/10.1099/ijs.0.003525-0
Kunin V, Engelbrektson A, Ochman H, Hugenholtz P (2010) Wrinkles in the rare biosphere: Pyrosequencing errors can lead to artificial inflation of diversity estimates. Environ Microbiol 12:118–123. https://doi.org/10.1111/j.1462-2920.2009.02051.x
Lee I, Kim OY, Park S-C, Chun J (2016) OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103. https://doi.org/10.1099/ijsem.0.000760
Li X, Yang Z, Wang Z, Li W, Zhang G, Yan H (2022) Comparative genomics of Pseudomonas stutzeri complex: taxonomic assignments and genetic diversity. Front Microbiol 12:755874. https://doi.org/10.3389/fmicb.2021.755874
Mechichi T, Stackebrandt E, Gad’on N, Fuchs G, (2002) Phylogenetic and metabolic diversity of bacteria degrading aromatic compounds under denitrifying conditions, and description of Thauera phenylacetica sp. nov., Thauera aminoaromatica sp. nov., and Azoarcus buckelii sp. nov. Arch Microbiol 178:26–35. https://doi.org/10.1007/s00203-002-0422-6
Meckenstock RU, Boll M, Mouttaki H, Koelschbach JS, Tarouco PC, Weyrauch P, Dong X, Himmelberg AM (2016) Anaerobic degradation of benzene and polycyclic aromatic hydrocarbons. J Mol Microbiol Biotechol 26:92–118. https://doi.org/10.1159/000441358
Na SI, Kim YO, Yoon SH, Ha SM, Baek I, Chun J (2018) UBCG: up-to-date bacterial core gene set and pipeline for phylogenomic tree reconstruction. J Microbiol 56:281–285. https://doi.org/10.1007/s12275-018-8014-6
Nurk S, Meleshko D, Korobeynikov A, Pevzner PA (2017) MetaSPAdes: A new versatile metagenomic assembler. Genome Res 27:824–834. https://doi.org/10.1101/gr.213959.116
Pilloni G, von Netzer F, Engel M, Lueders T (2011) Electron acceptor-dependent identification of key anaerobic toluene degraders at a tar-oil-contaminated aquifer by Pyro-SIP. FEMS Microbiol Ecol 78:165–175. https://doi.org/10.1111/j.1574-6941.2011.01083.x
Quast C, Pruesse E, Yilmaz P, Gerken J, Schweer T, Yarza P, Peplies J, Glöckner FO (2013) The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res 41:D590-596. https://doi.org/10.1093/nar/gks1219
Reinhold-Hurek B, Hurek T (2000) Reassessment of the taxonomic structure of the diazotrophic genus Azoarcus sensu lato and description of three new genera and new species, Azovibrio restrictus gen. nov., sp. nov., Azospira oryzae gen. nov., sp. nov. and Azonexus fungiphilus gen. nov., sp. nov. Int J Syst Evol Microbiol 50:649–659. https://doi.org/10.1099/00207713-50-2-649
Révész F, Farkas M, Kriszt B, Szoboszlay S, Benedek T, Táncsics A (2020) Effect of oxygen limitation on the enrichment of bacteria degrading either benzene or toluene and the identification of Malikia spinosa (Comamonadaceae) as prominent aerobic benzene-, toluene-, and ethylbenzene-degrading bacterium: enrichment, isolation and whole-genome analysis. Environ Sci Pollut Res 27:31130–31142. https://doi.org/10.1007/s11356-020-09277-z
Rodriguez-R LM, Gunturu S, Harvey WT, Rosselló-Mora R, Tiedje JM, Cole JR, Konstantinidis KT (2018) The Microbial Genomes Atlas (MiGA) webserver: taxonomic and gene diversity analysis of Archaea and Bacteria at the whole genome level. Nucleic Acids Res 46:282–288. https://doi.org/10.1093/nar/gky467
Saitou N, Nei M (1987) The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425. https://doi.org/10.1093/oxfordjournals.molbev.a040454
Schloss PD, Westcott SL, Ryabin T, Hall JR, Hartmann M, Hollister EB, Lesniewski RA, Oakley BB, Parks DH, Robinson CJ, Sahl JW, Stres B, Thallinger GG, Van Horn DJ, Weber CF (2009) Introducing mothur: open-source, platform-independent, community-supported software for describing and comparing microbial communities. Appl Environ Microbiol 75:7537–7541. https://doi.org/10.1128/AEM.01541-09
Sieber CMK, Probst AJ, Sharrar A, Thomas BC, Hess M, Tringe SG, Banfield JF (2018) Recovery of genomes from metagenomes via a dereplication, aggregation and scoring strategy. Nat Microbiol 3:836–843. https://doi.org/10.1038/s41564-018-0171-1
Táncsics A, Szoboszlay S, Szabó I, Farkas M, Kovács B, Kukolya B, Mayer Z, Kriszt B (2012) Quantification of subfamily I.2.C catechol 2,3-Dioxygenase mRNA transcripts in groundwater samples of an oxygen-limited BTEX-contaminated site. Environ Sci Technol 46:232–240
Táncsics A, Farkas M, Szoboszlay S, Szabó I, Kukolya J, Vajna B, Kovács B, Benedek T, Kriszt B (2013) One-year monitoring of meta-cleavage dioxygenase gene expression and microbial community dynamics reveals the relevance of subfamily I.2.C extradiol dioxygenases in hypoxic BTEX-Contaminated Groundwater. Syst Appl Microbiol 36:339–350. https://doi.org/10.1016/j.syapm.2013.03.008
Táncsics A, Banerjee S, Soares A, Bedics A, Kriszt B (2023) Combined omics approach reveals key differences between aerobic and microaerobic xylene-degrading enrichment bacterial communities: Rhodoferax - a hitherto unknown player emerges from the microbial dark matter. Environ Sci Technol 57:2846–2855. https://doi.org/10.1021/acs.est.2c09283
Táncsics A, Szalay RA, Farkas M, Benedek T, Szoboszlay S, Szabó I, Lueders T (2018) Stable isotope probing of hypoxic toluene degradation at the Siklós aquifer reveals prominent role of Rhodocyclaceae. FEMS Microbiol Ecology. https://doi.org/10.1093/femsec/fiy088
Tindall BJ, Rosselló-Móra R, Busse HJ, Ludwig W, Kämpfer P (2010) Notes on the characterization of prokaryote strains for taxonomic purposes. Int J Syst Evol Microbiol 60:249–266. https://doi.org/10.1099/ijs.0.016949-0
Wang M, Garrido-Sanz D, Sansegundo-Lobato P, Redondo-Nieto M, Conlon R, Martin M, Mali R, Liu X, Dowling DN, Rivilla R, Germaine KJ (2021) Soil microbiome structure and function in ecopiles used to remediate petroleum-contaminated soil. Front Microbiol 9:624070. https://doi.org/10.3389/fenvs.2021.624070
Wu YW, Simmons BA, Singer SW (2016) MaxBin 2.0: An automated binning algorithm to recover genomes from multiple metagenomic datasets. Bioinformatics 32:605–607. https://doi.org/10.1093/bioinformatics/btv638
Yang E, Liu J, Chen D, Wang S, Xu L, Ma K, Zhang X, Sun L, Wang W (2022) Rhizobium cremeum sp nov isolated from sewage and capable of acquisition of heavy metal and aromatic compounds resistance genes. Syst Appl Microbiol 45:126322. https://doi.org/10.1016/j.syapm.2022.126322
Yerushalmi L, Lascourreges JF, Guiot SR (2002) Kinetics of benzene biotransformation under microaerophilic and oxygen-limited conditions. Biotechnol Bioeng 79:347–355. https://doi.org/10.1002/bit.10320
Zhang T, Tremblay PL, Chaurasia AK, Smith JA, Bain TS, Lovley DR (2013) Anaerobic benzene oxidation via phenol in Geobacter metallireducens. Appl Environ Microbiol 79:7800–7806. https://doi.org/10.1128/AEM.03134-13
Acknowledgements
The authors acknowledge the valuable help of János Kiss in the isopycnic gradient ultracentrifugation.
Funding
Open access funding provided by Hungarian University of Agriculture and Life Sciences. This research has been supported by the National Research Development and Innovation Office of Hungary (NKFIH) through the OTKA Grants No. FK 134439 and K 146358. André Soares was supported by the European Union’s Horizon 2020 research and innovation programme under project PROSPECTOMICS (grant agreement n 899667).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interests
The authors declare that there are no conflicts of interest.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Táncsics, A., Bedics, A., Banerjee, S. et al. Stable-isotope probing combined with amplicon sequencing and metagenomics identifies key bacterial benzene degraders under microaerobic conditions. BIOLOGIA FUTURA (2024). https://doi.org/10.1007/s42977-024-00232-4
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s42977-024-00232-4