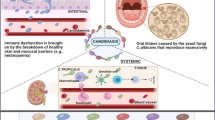
Abstract
Fungal infections (mycoses) represent a major health issue in humans. They have emerged as a global concern for medical professionals by causing high morbidity and mortality. Fungal infections approximately impact one billion individuals per annum and account for 1.6 million deaths. The diagnosis of Candida infections is a challenging task. Laboratory-based Candida species identification techniques (molecular, commercial, and conventional) have been reviewed and summarized. This review aims to discuss the mycoses history, taxonomy, pathogenicity, and virulence characteristics.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
1 Introduction
Mycoses (fungal infections) are characterized by high morbidity and mortality, which affect a huge global population every year [1,2,3,4,5]. Fungal infections have been a constant threat to humans [6]. The occurrence of fungal diseases is on the rise and hospital-residing immunocompromised patients are particularly the primary victims [7]. The main fungal pathogens (50–90% mycoses isolates) involved in the infections include Trichosporon spp., Aspergillus spp., Scedosporium spp., Zygomyces spp., Paracoccidioides spp., Cryptococcus spp., Rhodotorula spp., Candida spp., Histoplasma capsulatum, Geotrichum spp., Coccidioide simmitis, and Fusarium spp. [8,9,10]. The yeasts belonging to the Candida genus have been frequently isolated from fungal infections [11]. There are almost 200 known Candida species whereas almost twenty species are associated with human infections [12]. Candida-related human infections are considered a major issue, especially in hospitalized patients suffering from severe underlying diseases and immunodeficient ICU patients [13, 14].
The parasitic fungus, Candida species, is a eukaryotic yeast that accounts for about 8% of global nosocomial infections [5, 15]. Taxonomically, Candida species belong to the Kingdom Fungi (Mycota), phylum Ascomycota, subphylum Saccharomycotina, class Saccharomycetes, and order Saccharomycetales [16]. The emergence of Candida albicans as a common pathogenic yeast has been reported [5, 17,18,19]. Candida clinical isolates that could colonize the human tissues to exert infections include Candida krusei, Candida dubliniensis, Candida lusitaniae, Candida parapsilosis, Candida utilis, Candida tropicalis, Candida famata, Candida glabrata, Candida rugosa, Candida kefyr (pseudotropicalis), Candida guilliermondii, Candida lipolytica, and Candida haemulonii [8, 20,21,22,23]. The significant role of non-Candida albicans Candida species (NCAC) in invasive candidiasis has also been established [24]. The occurrence of candidiasis is quite higher than the overgrowth of Candida. This review discusses mycoses-related topics and describes virulence factors related to candidiasis along with their participation in pathogenicity and future approaches for better candidiasis diagnosis.
2 The virulence and pathogenicity of Candida spp.
The survivability of Candida species under harsh conditions makes them highly detrimental pathogens, which could threaten the lives of immunocompromised patients [25, 26]. Previously, yeast microorganisms were supposed to infect only immunocompromised patients, but virulence factors have revealed their pathogenicity to other patients as well. Multiple aggression mechanisms of these microorganisms participate in the pathophysiology of the disease [27, 28]. Several virulence factors of Candida enhance their pathogenicity, which includes invasion and adhesion to inanimate surfaces and body tissues, metabolic adaptation, dimorphism, phenotypic switching, secretion of hydrolytic enzymes, and formation of biofilms [29].
Candida albicans, a polymorphic yeast, exhibits different cellular morphologies such as hyphae, yeast cells (white phase), GUT (commensalism-related), pseudohyphae, chlamydospores, and opaque (mating cell types). Different polymorphic forms affect the pathogenicity of C. albicans [30, 31]. The parameters such as nutrients, pH, temperature (37–40 °C), Co2 concentrations (5.5%), and amino acids facilitate their morphological transition, which is crucial for pathogenicity. The yeast forms could conveniently spread inside the host tissues whereas filamentous shapes possessing higher adhesion capability help in the invasion of the host tissues [32].
Candida spp initially attach to the host cell through adhesion proteins present on the fungal cell surface (pga1, als1-7, hwp1, als9, and eap1), and immobilized ligands (cadherins, integrins, or other microorganisms). Fungal cells invade the tissue after the adhesion. The invasion and damage of the epithelium are considered pathogenic [8, 33,34,35]. It could occur via two mechanisms (active penetration or endocytosis) depending upon the type of host cell. For example, the invasion of oral cells by C. albicans occurs through active penetration and endocytosis, whereas only active penetration is possible in intestinal invasion [36, 37].
Candida is highly adaptable to various environmental conditions (low oxygen, limited nutrition, pH fluctuations, and nitrosative, cationic, temperature, osmotic pressure, and oxidative stresses) [8, 31, 38, 39]. This adaptability is of key importance for the C. albicans pathogenicity [8]. The formation of mycotic biofilm is a complicated process, which generates a highly organized structure (Fig. 1). Multiple studies have been performed to investigate the biofilm formation in Candida species [40,41,42,43]. The National Institute of Health has reported that more than 80% of the total microbial infections in the United States are caused by fungal biofilms [5]. Biofilm consists of adhered, attached, and accumulated microorganisms that form extracellular polymeric substances (EPSs) to provide a structural matrix [44]. Planktonic C. albicans cell adhesion to the surfaces is the initial step that induces an organized strong extracellular matrix (ECM) structure [44]. There are four consecutive C. albicans biofilm phases including the Adhesion phase, Initiation phase (early phase), Maturation phase (intermediate phase), and Dispersal phase (dispersion phase) (Fig. 1).
The consecutive phases of biofilm formation in C. albicans (1). Adhesion phase, yeast cells adhere promptly to the surface. (2). Initiation phase (early phase), spherical yeast cells replicate and start to secrete extracellular matrix (ECM) and develop Pseudohyphae. (3). Maturation phase (intermediate phase), the mature biofilm develops with hyphal filaments extending far from a basal layer (yeast forms). (4). Dispersal (dispersion phase), yeast cells disperse from the biofilm and diffuse, expanding the infection and starting the cycle again
Extracellular hydrolytic enzymes (phospholipases and proteases) are necessary for the pathogenicity-causing Candida yeast virulence factors. These enzymes facilitate the C. albicans invasion through host protein degradation (hemoglobin and keratin) and cell membrane structure alteration. These steps help in the targeting and invasion of the host’s immunity cells by avoiding antimicrobial agents. Different Candida species follow this process including C. albicans, C. tropicalis, C. parapsilosis, and C. dubliniensis [8].
3 Roles of virulence-associated genes in pathogenicity
Yeasts can induce infection and overwhelm the host defense systems due to the existence of various proteins and genes associated with their pathogenicity, known as virulence factors [45]. C. albicans genes encode several pathogenic virulence factors. These genes and their products contribute to mycological pathogenicity and are called virulence factors. Numerous genes associated with C. albicans pathogenicity have been described [8, 46, 47]. These factors include the ability of C. albicans to transition from yeast form to hyphal form, adhesins, biofilm formation, and hydrolytic enzymes secreted (aspartyl proteases, and phospholipases) [8, 48].
Candida albicans can grow in the form of yeast and mold. The transition between yeast and hyphal forms is termed (dimorphism) [49]. The dimorphism of C. albicans is a unique characteristic of yeast pathogenicity. Both morphologies have their role to support their virulence [8]. It has been reported that the form of hyphal is more invasive than the form of yeast [50].
The dimorphism of C. albicans plays an important role in the pathogenicity of both systemic and superficial infections. It should be noted that both yeast and filamentous forms of C. albicans were detected in infected tissues [51]. The capability of C. albicans to switch from yeast to filamentous form contributes to the various nature of its infection phases, such as adherence to epithelial and endothelial cells, invasion, iron acquisition from host sources, biofilm formation, escape from phagocytes, and immune evasion [51].
Adhesins are the yeast surface molecules that intermediate the binding of C. albicans to the surface of human or microorganism cells, inert polymers, or proteins [52]. ALA1, ALS1, Hwp1, INT1, MMT1, PMT1, PMT6, and, Als1p are candidate genes considered as encoding adhesins [52,53,54,55]. Other putative adhesins are mannan, chitin, factor 6 oligomannosaccharide, 66-kDa fimbrial protein, fibronectin-binding protein, iC3b binding protein, fucose binding protein, GIcNAc or glucosamine, and secreted aspartyl proteinase (SAP) [52, 53]. The Sapproteins of C. albicans were encoded by a family of 10 SAP genes i.e. SAP1, SAP2, SAP3, SAP4, SAP5, SAP6, SAP7, SAP8, SAP9, and SAP10. The major functions of the C. albicans Saps are nutrition for the yeast cells, assisting penetration and invasion, and avoiding host immune responses [56].
Phospholipases hydrolyze glycerophospholipids, which are the main components of mammalian cell membranes. It destabilizes the membranes by cleaving fatty acids from phospholipids [57]. There are seven phospholipase genes have been identified i.e. PLA, PLB1, PLB2, PLC1, PLC2, PLC3, and, PLD1. However, the role of the enzymes encoded by these genes are not yet clear [58]. In a comparative study conducted by Ibrahim et al.[59], evidence was obtained that phospholipase acts as a virulence factor, and a series of C. albicans obtained from candidemia patients were compared with the isolates obtained from the oral cavities of healthy people. In candidemia cases, higher phospholipase activity was found, reflecting the virulence of these isolates.
4 Antimycotic-resistance of Candida
Candida resistance to various antimycotic agents poses a serious public health concern. The Candida incidence in the bloodstream has increased from 2.2 to 3.2 cases/100,000 population/annum in Europe [60, 61]. Antifungal resistance (azoles and echinocandins) in Candida could hinder their treatment. C. parapsilosis and C. glabrata are commonly found clinical strains, which cause invasive candidiasis by modifying prevalence at various locations [62]. Multidrug-resistant (MDR) cases featuring non-albicans Candida (NAC) and C. albicans strains have raised serious concerns [63]. Antifungal drug preservation has increased because of Candida auris based global nosocomial outbreaks featuring higher morbidity and mortality. Centers for disease control and prevention (CDC) added these strains to priority antibiotic resistance threats in 2019 [62]. The world health organization (WHO) convened the first meeting in 2020 for establishing a health-related pathogens (mycoses) priority list. They also defined the research and development (R&D) priorities for encouraging the development of new drugs, diagnosis methods, and strategies. Global antimicrobial-resistance surveillance system (GLASS) of antimycotic resistance (AMR) has developed a protocol to counter Candida spp. based bloodstream infections (BSIs). Antimycotic susceptibility data related to blood Candida isolates especially from patients in high-risk hospital units (ICUs) is available through GLASS reports [64, 65].
5 Candidiasis
Candidiasis is a global Candida yeast-based major human fungal disease. Candidiasis refers to disseminated, visceral, and mucosal-cutaneous infections of the genus Candida [66]. These infections could occur at any age and are easily identifiable infection risk factors [67]. Candidiasis infections are complex and of different types. The types are distinguished through different morphology, and relationships between mucosal and immune systems [68]. The epidemiological records have established the association of five species with candidiasis including C. albicans (65.3%), C. glabrata (11.3%), C. tropicalis (7.2%), C. parapsilosis (6.0%), and C. krusei (2.4%) (Table 1 and Fig. 2) [69].
The infections such as oral candidiasis, vaginitis, candidemia, systemic infections, and cutaneous candidiasis are linked to Candida. Different candidiasis types are presented in Fig. 3 [70,71,72]. Oropharyngeal candidiasis (OPC), also known as “thrush” (different oral mucosal sites and tongue infection) is characterized by the overgrowth of mycoses and superficial tissues’ invasion [73,74,75]. Urogenital or Vulvovaginal candidiasis (VVC) is a common mycosis worldwide that infects female genital tracts [76]. This infection is quite frequent in diabetic individuals, pregnant women, and patients under antibiotic and corticosteroid treatments [77, 78].
Invasive candidiasis (IC) can substantially implicate any organ and refers to deep-seated infections such as osteomyelitis (bone infection), peritonitis (tissue covering the inner abdominal wall and organs), and intra-abdominal abscess [71, 79,80,81,82]. Candidemia (BSI) refers to the Candida species infection in the blood of patients suffering from fever [83]. Candida parapsilosis is the most frequent agent of bloodstream infection (BSIs) among non-C. albicans Candida species [84]. BSIs-associated Candida ranks fourth in nosocomial-associated infections in the USA whereas it is at the sixth number in Europe [67, 85]. The occurrence of mucosal candidiasis is common and they are more invasive than fungal candidiasis. Candidiasis-infected mucosal surfaces could be oral mucosa, pharynx, esophagus, urogenital, intestine, and urinary system [86, 87]. Systemic candidiasis is also referred to as acute organ invasive or systemic hematogenous candidiasis. During systemic candidiasis, the Candida cells spread to the whole body and rapidly form abscesses in vital organs. The infections of systemic candidiasis lead to visceral lesions (deep candidal focus) or Candida septicemia (candidemia) [15, 88]. Nail and cutaneous candidiasis is a secondary sub-acute or chronic infection of predisposed patients, which could be local or spread to the nails and skin. There are various types of cutaneous candidiasis such as otomycosis, intertrigo candidiasis, diaper rash, Candida folliculitis, paronychia, and onychia [71].
Candidiasis could also infect the surfaces of different medical devices including urinary catheters, central venous catheters, and cardiovascular devices. These infections are commonly associated with morbidity and deaths of hospitalized patients [89]. Candida-related infection of medical devices is among the major pathogenesis-related factors [89,90,91,92].
6 Microbiological and clinical tests for the diagnosis of Candida
The diagnosis of candida infections is complicated. This section summarizes the clinical laboratory tests (molecular, conventional, and commercial) for the isolation and identification of Candida species.
6.1 Phenotypic methods
6.1.1 Conventional methods
Non-albicans and C. albicans identification was initially carried out through phenotypic traits (biotyping, morphotyping, chemical resistance, and serotyping). However, the reproducibility and differentiation levels of these approaches are very low, which limits their reliable diagnosis and epidemiological analysis [73, 93].
6.1.2 Germ-tube test (GTT)
The germ-tube test (GTT) method refers to the formation of tube-like structures by Candida's reaction and it primarily identifies and differentiates C. dubliniensis and C. albicans from other species. This is a simple, rapid, and economical identification technique with 98% sensitivity. Candida is incubated in serum at 37 °C for 2–4 h and then observed for the presence of tube structures. This is an effective method that is widely applied for C. albicans identification [94, 95].
6.1.3 Chlamydospore formation
The chlamydospore formation test is based on the appearance of chlamydospores during the last stage. This test differentiates C. albicans from C. dubliniensis. During the test, C. albicans is subjected to dormant growth under a controlled environment using agar media (cornmeal and rice extract), which leads to chlamydospore formation [96].
6.1.4 Carbon assimilation
This is an economical, simple but time-consuming test that specifically identifies Candida species. The method involves Candida growth on carbon substrates followed by incubation and growth examination [97].
6.1.5 Carbohydrates fermentation
Fermentation tests rely on the acid and carbon dioxide formation in the liquid media. Carbohydrates facilitate the fermentation process during the test. However, this Candida identification test is complex, time-consuming, and less sensitive as compared to the carbohydrate assimilation test. Traditionally, carbohydrate fermentation test was performed for Candida species differentiation, but now this method is not often used [98].
6.2 Rapid identification systems
The commercial and rapid identification methods are based on conventional methods (sugar assimilation, germ tube, and chlamydospore formation tests). However, these methods are comparatively easy, rapid, and cheaper [99].
6.2.1 API 20C Aux system
API 20C Aux (bioMerieux Vitek, Hazelwood, MO, USA) system was developed from API 20 system, which is a carbohydrates assimilation-based process. It involves the assimilation tests of 19 carbohydrates in cupules that are incubated (24–72 h) at 30 °C followed by turbidity analysis using computer programs [99]. This method accurately differentiates between C albicans, C. dubliniensis, and other Candida species [100]. However, certain limitations of this method have also been reported during C. krusei identification [101].
6.2.2 API Candida system
API Candida system containing ten tubes could detect five carbohydrates through various acidification and enzymatic tests (seven). The testing procedures are based on sugars acidification and enzyme activity [102, 103]. This is a highly accurate and simple method that completes without the involvement of computers. It produces observable color changes, and morphological characteristics to identify Candida species [103].
6.3 Chromogenic media-based commercial systems
6.3.1 CHROMagar technique
CHROMagar is a rapid, precise, and straightforward Candida identification technique. It involves the media with multiple chromogenic substrates, which react with Candida species enzymes leading to the growth of colonies in different colors [104, 105].
6.3.2 Candida ID system
Candida ID system (bioMerieux, Marcy l’Etoile, France) is a rapid and more developed CHRMagar system. A chromogenic indolyl glucosaminide substrate in the growth medium reacts with Candida species to produce different insoluble colors. Turquoise/ blue color is produced by C. albicans, C. guilliermondii, and C. lusitaniae whereas C. tropicalis produces pink color [106].
6.3.3 Fluorogenic membrane filtration method
This is another developed and accurate CHROMagar method. The enzymatic reaction is detected by passing fluorogenic substrates through a nylon membrane filter. Different Candida species (C. albicans, C. krusei, C. tropicalis, and C. glabrata) could be differentiated through this method [107].
6.3.4 Fungichrom I and fungi-fast I twin systems
The fungichrom I twin system (International Microbio, Parcd’activites-allee D’athenes, France) consists of sixteen whereas the fungi-fast twin system consists of ten test cupules. The samples are incubated at 30 °C for 24–48 h and observed for color changes. Candida identification rate is high in the fungichrom twin system and it is a more rapid and simple method as compared to the fungi-fast I twin system [108].
6.3.5 Biggy agar system
Biggy agar (Oxoid Company, Wade Road, Basingstoke, Hampshire, UK) is a bismuth sulfite-containing chromogenic medium. Candida species convert it to bismuth sulfide. The reaction generates specific colors depending upon the growth of Candida species. The color of C. albicans becomes light brown whereas the color of C. tropicalis changes to dark brown. However, this method could not differentiate among certain species such as C. krusei and C. parapsilosis, which produce similar colors in their colonies [109].
6.4 Automated methods
Recent automated methods are fast, reliable, and broad, which could facilitate the development of new patient management and therapeutic techniques.
6.4.1 Vitek YBC system
The Vitek YBC system (bioMerieux Vitek, Inc., Hazelwood, MO, USA) is an auto-microbial system that is widely applied in research centers and laboratories. This method could simultaneously perform twenty-six biochemical tests from the same inoculum. Therefore, it could identify several Candida species including C. parapsilosis, C. albicans, C. glabrata, and C. tropicalis. Furthermore, it contains a computerized assessment system for more reliable information on Candida species. The handling of this method is easy and does not require an experienced person [110].
6.4.2 VITEK® 2 ID-YST system
VITEK® 2 Yeast identification (YST) is a developmental system. This is a high-speed, rapid, and simple system that could simultaneously conduct multiple reactions. This system can perform 47 carbohydrate assimilation-based fluorescent biochemical reactions including deamination and oxidation reactions with various arylamidases and oxidases of Candida species. This system is more suitable for differentiating C. dubliniensis and C. glabrata in addition to the identification of yeast microorganisms [111].
7 Serological methods
The serological commercial tests are used for the identification of Candida identification. There are several highly sensitive reagents for fungal identification. (1-3)-Beta-D-Glucan is a novel reagent to identify fungal infection. Similarly, a new fungal surrogate marker is highly sensitive and specific for the Candida infection diagnosis. This test also helps in the diagnosis of Candidemia [112]. A polysaccharide (Galactomannan) of the fungal cell wall could diagnose fungal infection and invasive fungi processes including Candidemia. This laboratory test can track the fungi infection and the patient's state to assess the treatment efficacy [113].
8 Molecular methods
Non-DNA and DNA-based molecular identification methods are gaining popularity. These are highly specific, accurate, and sensitive techniques for the identification and differentiation of C. albicans from other Candida species. These features encourage their widespread applications [93, 114].
8.1 Non-DNA-based techniques
8.1.1 Multi-locus enzyme electrophoresis (MLEE)
MLEE could estimate enzymatic protein and its polymorphism through gel-based analysis of enzyme migration. The charge and size of the protein molecule determine the length of enzymatic migration. MLEE is used in epidemiology and genetic studies as it could accurately differentiate between unknown and novel strains. However, indirect genome evaluation by this technique is the main disadvantage, which results in slow rates of evaluating variations accumulated in the species and incorrect outcomes. MLEE is also does not detect whole nucleotide variations (Table 2) [115, 116].
8.2 DNA-based methods
DNA-based tests could detect and differentiate among microorganisms’ DNA. This category consists of DNA-based conventional methods and Exact DNA-based methods [104].
8.2.1 Conventional DNA-based methods
These are DNA components-based oldest microorganism identification methods. DNA of eukaryotic organisms is extracted through the cell membrane hydrolysis by hydrolyzer enzymes [104].
8.2.2 Pulsed-field gel electrophoresis (PFGE)
Several studies have discussed the application of DNA genome components for microorganism identification, which is derived after the hydrolysis of the cell membrane [117, 118]. Therefore, different electrophoresis methods were developed including pulsed-field gel electrophoresis (PFGE), orthogonal-field alternative gel electrophoresis (OFAGE), contour-clamped homogeneous electric field (CHEF), transverse alternate gel electrophoresis (TAFE), and field inversion gel electrophoresis (FIGE). This is an ideal technique for separating of chromosome-sized DNA molecules due to the size of C. albicans chromosomes are about (1- 4 Mb) (Table 2) [117, 118].
8.2.3 Restriction enzyme analysis (REA)
REA technique was initially applied to conduct an epidemiological investigation of C. albicans infections. This is a complex method that may require a computer-assisted software. REA involves the purification of the total genomic DNA followed by cleavage through multiple endonuclease enzymes (e.g., EcoRI, MspI, BglII, HinF1, or HindIII) that produces small fragments resulting in a sequence-dependent restriction fragment length polymorphism (RFLP). The produced fragments were separated using the agarose gel electrophoresis technique. Then visualized on the gel after dyeing with ethidium bromide. In this technique, the interpretation and differentiation of strains are very difficult due to the production of complex patterns with enormous bands of unequal intensities (Table 2) [120].
8.2.4 Random amplified polymorphic DNA (RAPD)
RAPD is a genomic DNA amplification-based technique. The amplified products are isolated through agarose gel electrophoresis according to the size of the amplified fragments. However, short genomic DNA could generate a complex pattern and segments that are different from the source. RAPD technique is widely performed for the identification and differentiation of Candida species, especially C. albicans (Table 2) [121, 122].
8.2.5 Amplified fragment length polymorphism (AFLP)
This method involves the hydrolysis enzymes-based digestion of genomic DNA, which is then amplified to obtain different DNA fragments. The fragments are further isolated using a high-performance instrument such as fluorescent dye-labeled primers. The amplification is carried out under highly specific conditions as compared to the RAPD method. Therefore, AFLP is more reliable, accurate, and reproducible than RAPD. However, the application of AFLP for Candida spices, especially C. albicans is limited as it is comparatively costly and requires an experienced person to perform its multiple complex steps (Table 2) [123, 124].
8.3 Exact DNA-based methods
8.3.1 Polymerase chain reaction (PCR) based-Candida detection methods
PCR is a landmark DNA molecular microbiology technique, which facilitates microorganism identification in human cells. PCR can detect Candida species during the invasive fungal infection phase such as Candidemia. PCR could also diagnose various genetic disorders. PCR is a rapid method, which can amplify several DNA fragments within minutes to detect millions of scarce DNA copies [125]. Fungal identification through PCR could be conducted in two ways: (i) Nested PCR technique amplifies DNA molecules in two steps to reduce the amplification errors. Therefore, it is considered a highly accurate method (ii) Real-time PCR could quantify amplified DNA in real-time at each PCR cycle. Real-time PCR is carried out using two types of fluorescence such as labeling probes and recently developed double-stranded dyes. Several studies have recommended the application of real-time PCR as it is more accurate and rapid than nested PCR [125]. PCR amplification (nested PCR or real-time PCR) is followed by the analysis of amplicons and conclusions. Different methods are used for the amplicon analysis, but direct sequencing is the most accurate technique as compared to single-strand conformational polymorphism (SSCP) and polyacrylamide gel electrophoresis (Table 2) [127].
8.3.2 Nucleic acid sequence-based amplification (NASBA)
This technique is based on RNA amplification and does not need a thermal cycling instrument for the specific detection of microorganisms. In contrast to DNA, RNA is quickly degradable outside the microorganism cells. NASBA method uses three expensive enzymes (RNase H polymerase, reverse transcriptase, and T7 RNA polymerase). Therefore, it is widely used for the amplification of Candida species. The results are generated within a few hours and it can differentiate up to six different Candida species (Table 2) [128].
8.3.3 Peptide nucleic acid-fluorescent in situ hybridization (PNA-FISH)
FISH is a classical yeast detection method that does not require purification and isolation steps. It only uses fluorescein-labeled oligonucleotide probes. The accuracy of this method further increases in combination with peptide nucleic acid (PAN) probes. These probes lead to hybridized microorganism cells in the samples, which are examined through advanced fluorescence microscopy. The studies have elaborated that the results of the PAN-FISH technique could be compared to PCR, but it generates the results faster than the PCR technique. Furthermore, the PAN-FISH approach contains highly specific probes for the Candida species. Therefore, it could specifically differentiate between Candida species (Table 2 and Fig. 4) [129].
8.3.4 Microsatellite length polymorphism (MLP) typing
This technique has been used in a Candida species-related epidemiology study. This is a simple, rapid, and highly reproducible method. MLP technique is based on the amplification of microsatellite stretch in the cell nucleotides. The sensitivity of this method depends on the type of microsatellite marker, which detects primer pair flanking in a specific microsatellite area. High polymorphism of amplified microsatellite fragments favors the use of the MLP typing technique in genetic analysis to determine the type of alleles (heterozygous or homozygous). Finally, high-resolution gel electrophoresis measures the allele length. The numerical results could easily be compared for the identification of various microorganisms (Table 2) [130].
8.3.5 Multi-locus sequence typing (MLST)
MLST is based on the amplification of internal fragments of nucleotide sequence polymorphisms in the independent loci genes. These are housekeeping genes, which are selected due to their stability. This technique could amplify the DNA fragments up to the size of 400–500 bp. The difference in alleles corresponding to multi-locus sequence type distinct alleles characterize each housekeeping locus. The ability to analyze only seven sequences in 300–400 bp loci genes is the main limitation of this method. Furthermore, the analysis of diploid microorganisms could generate two identical products in this method whereas they vary in heterozygous bases on the polymorphic databases (Table 2) [132].
8.3.6 DNA-microarrays
Microarrays are microscopic high-density oligonucleotide probes, which hybridize the nucleic acid samples followed by immobilization on a solid surface. Microarray-based systems depend upon strain typing. These are highly sensitive, specific, and automatic standards that do not require a prior database. Microarrays hybridization-based bound sequence is detected using a highly efficient fluorescent scanning instrument and advanced computer software (Table 2 and Fig. 5) [133].
9 Modern methods
9.1 Matrix-assisted laser desorption ionization-time of flight mass spectrometry (MALDI-TOF MS)
MALDI-TOF MS is a modern approach that is now widely available in clinical microbiology laboratories. MALDI-TOF MS is a rapid, cost-effective, reliable, and powerful identification method. MALDI-TOF MS generates protein fingerprints of each microorganism in the sample, which could be easily compared at the reference library. Briefly, the fungi degradation is carried out at the temperature of the curie pyrolysis point followed by the production of volatile fragments from the cleavage of tiny molecules. Finally, the mass spectrums of volatile fragments are analyzed using a mass spectrometer, which represents the fingerprint image of each microorganism. The lack of spectra characterization for comparison is a major limitation of the current analysis (Table 3 and Fig. 6) [126, 135, 136].
9.2 Candida species genomes: genome sequence and comparative identification
A significant variation in composition and size has been reported between the sequenced genomes of Candida (Table 3) [137, 138]. Table 3 presents a high continuity range between scaffolds (9–27). Scaffold size and number closely relate to each candida microorganism. The field gel electrophoresis could assess the genomes in each Candida microorganism where telomeric arrays finally link to the scaffolds. 10.6–15.5 Mb difference could occur in the genome size of approximately 50% of Candida species whereas the difference in GC content (guanine and cytosine) could range between 33 and 45% (Table 3). The transportation and repetitive capability of these elements could differ between assemblies in numbers and type [139]. Candida species are primarily different in genomic size and phenotype, however, they are quite similar in protein-coding gene numbers (5.733–6.318) as presented in Table 3. The genome of the smallest Candida species (C. guilliermondii) contains more genes than the genome of the largest Candida species (L. elongisporus). Therefore, genome size and gene numbers are not correlated [138].
10 Conclusion and promising future directions
The Nanopore sequencing technology is based on the DNA translocation across a lipid-bilayer membrane through a pore, which is formed by Staphylococcus aureus alpha-hemolysin after applying the electrical fields [140]. This novel method has been used for mycosis detection in various studies. Ashikawa et al. [141] applied a nanopore sequencing system for the identification of five Candida species in positive blood-culture vials and their performance was compared with Sanger sequencing. This system provides rapid optimization of reagents and instruments. This system could further help to develop accurate and rapid point-of-care devices for clinical and field usage [142].
CRISPR-Cas9 (clustered regulatory interspersed short palindromic sequences-CRISPR associated protein 9) versatility has led to the development of an identification tool known as Specific Highly Sensitive Enzymatic Reporter UnLOCKING (SHERLOCK). SHERLOCK could successfully identify target nucleic acids in attomolar concentrations to distinguish closely-related viruses and genotypes up to the difference of a single base pair [143]. Furthermore, Next-generation (SHERLOCK version 2) is a quantitative multiplex analysis that could visualize the final results using lateral flow devices (LFDs) system [144]. SHERLOCK v2 could achieve high specificity and sensitivity in combination with the HUDSON method and rapid DNA extraction techniques. This combination could be successfully used for the diagnosis and identification of Dengue and Zika viruses. In short, the SHERLOCK v2 system coupled with an efficient DNA extraction tool provides a reliable portable platform to identify mycotic pathogens [145].
This review updated the current overview of Candida infections. Other studies have reported an alarming rise in Candida disease. This suggests that the current diagnostic methods of pathogen-related infections should be reviewed, and new strategies should be developed for the diagnosis of mixed Candida spp. infections.
Data availability
Data will be made available on reasonable request.
References
Pfaller MA, Andes DR, Diekema DJ, Horn DL, Reboli AC, Rotstein C, Franks B, Azie NE (2014) Epidemiology and outcomes of invasive candidiasis due to non-albicans species of Candida in 2,496 patients: data from the prospective antifungal therapy (PATH) registry 2004–2008. PLoS ONE 9:e101510. https://doi.org/10.1371/journal.pone.0101510
Matthaiou DK, Christodoulopoulou T, Dimopoulos G (2015) How to treat fungal infections in ICU patients. BMC Infect Dis 15:1–8. https://doi.org/10.1186/s12879-015-0934-8
Schmiedel Y, Zimmerli S (2016) Common invasive fungal diseases: an overview of invasive candidiasis, aspergillosis, cryptococcosis, and Pneumocystis pneumonia. Swiss Med Wkly 146:1–12. https://doi.org/10.4414/smw.2016.14281
Bongomin F, Gago S, Oladele RO, Denning DW (2017) Global and multi-national prevalence of fungal diseases-estimate precision. J Fungi 3:57. https://doi.org/10.3390/jof3040057
Atiencia-Carrera MB, Cabezas-Mera FS, Tejera E, Machado A (2022) Prevalence of biofilms in Candida spp. bloodstream infections: a meta-analysis. PLoS One 17:e0263522. https://doi.org/10.1371/journal.pone.0263522
Vallabhaneni S, Mody RK, Walker T, Chiller T (2016) The global burden of fungal diseases. Infect Dis Clin 30:1–11. https://doi.org/10.1016/j.idc.2015.10.004
Espinel-Ingroff A, Canton E, Peman J, Rinaldi MG, Fothergill AW (2009) Comparison of 24-hour and 48-hour voriconazole MICs as determined by the clinical and laboratory standards institute broth microdilution method (M27–A3 document) in three laboratories: results obtained with 2,162 clinical isolates of Candida spp and other yeasts. J Clin Microbiol 47:2766–2771. https://doi.org/10.1128/JCM.00654-09
Mayer FL, Wilson D, Hube B (2013) Candida albicans pathogenicity mechanisms. Virulence 4:119–128. https://doi.org/10.4161/viru.22913
Vázquez-González D, Perusquía-Ortiz AM, Hundeiker M, Bonifaz A (2013) Opportunistic yeast infections: candidiasis, cryptococcosis, trichosporonosis and geotrichosis. JDDG 11:381–394. https://doi.org/10.1111/ddg.12097
Seth R, Xess I, Jana M (2019) Diagnosis of invasive fungal infections in children. Indian Pediatr 56:229–236. https://doi.org/10.1007/s13312-019-1505-7
Deorukhkar SC, Saini S, Mathew S (2014) Non-albicans Candida infection: an emerging threat. Interdiscip Perspect Infect Dis 2014:1–7. https://doi.org/10.1155/2014/615958
Berkow EL, Lockhart SR (2017) Fluconazole resistance in Candida species: a current perspective. Infect Drug Resist 10:237–245. https://doi.org/10.2147/IDR.S118892
Kaushik A, Kest H (2018) The role of antifungals in pediatric critical care invasive fungal infections. Crit Care Res Pract 2018:1–9. https://doi.org/10.1155/2018/8469585
Ramos-Pardo A, Castro-Álvarez R, Quindós G, Eraso E, Sevillano E, Kaberdin VR (2022) Assessing pH-dependent activities of virulence factors secreted by Candida albicans. MicrobiologyOpen 12:e1342. https://doi.org/10.1002/mbo3.1342
Pfaller MA, Diekema DJ (2007) Epidemiology of invasive candidiasis: a persistent public health problem. Clin Microbiol Rev 20:133–163. https://doi.org/10.1128/CMR.00029-06
Golubev WI (2011) Xanthophllomyces Golubev. In: Kurtzman CP, Fell JW, Boekhou T (eds) The yeasts: a taxonomic study, 4th edn. Elsevier, New York, p 2354
Terças ALG, Marques SG, Moffa EB, Alves MB, de Azevedo CM, Siqueira WL, Monteiro CA (2017) Antifungal drug susceptibility of Candida species isolated from HIV-positive patients recruited at a public hospital in São Luís, Maranhão. Brazil Front Microbiol 8:298. https://doi.org/10.3389/fmicb.2017.00298
Dadar M, Tiwari R, Karthik K, Chakraborty S, Shahali Y, Dhama K (2018) Candida albicans-biology, molecular characterization, pathogenicity, and advances in diagnosis and control–an update. Microb Pathog 117:128–138. https://doi.org/10.1016/j.micpath.2018.02.028
Kaur J, Nobile CJ (2023) Antifungal drug-resistance mechanisms in Candida biofilms. Curr Opin Microbiol 71:102237. https://doi.org/10.1016/j.mib.2022.102237
Williams D, Silva SC, Malic S, Kuriyama T, Lewis MAO (2000) Candida biofilms and oral candidosis: treatment and prevention. Periodontol 55:250–265
Patil S, Rao RS, Majumdar B, Anil S (2015) Clinical appearance of oral Candida infection and therapeutic strategies. Front Microbiol 6:1391. https://doi.org/10.3389/fmicb.2015.01391
Barchiesi F, Orsetti E, Osimani P, Catassi C, Santelli F, Manso E (2016) Factors related to outcome of bloodstream infections due to Candida parapsilosis complex. BMC Infect Dis 16:1–7. https://doi.org/10.1186/s12879-016-1704-y
Mamali V, Siopi M, Charpantidis S, Samonis G, Tsakris A, Vrioni G, Network C-C (2022) Increasing incidence and shifting epidemiology of candidemia in Greece: results from the first nationwide 10-year survey. J Fungi 8:116. https://doi.org/10.3390/jof8020116
Pappas PG, Rex JH, Sobel JD, Filler SG, Dismukes WE, Walsh TJ, Edwards JE (2004) Guidelines for treatment of candidiasis. Clin Infect Dis 38:161–189. https://doi.org/10.1086/380796
Brown GD, Denning DW, Gow NA, Levitz SM, Netea MG, White TC (2012) Hidden killers: human fungal infections. Sci Transl Med 4:165rv13. https://doi.org/10.1126/scitranslmed.3004404
Rokas A (2022) Evolution of the human pathogenic lifestyle in fungi. Nat Microbiol 7:607–619. https://doi.org/10.1038/s41564-022-01112-0
Tamura NK, Negri MFN, Bonassoli LA, Svidzinski TIE (2007) Virulence factors for Candida spp recovered from intravascular catheters and hospital workers hands. Rev Soc Bras Med Trop 40:91–93
Czechowicz P, Nowicka J, Gościniak G (2022) Virulence factors of Candida spp and host immune response important in the pathogenesis of vulvovaginal candidiasis. Int J Mol Sci 23:5895. https://doi.org/10.3390/ijms23115895
Alenzi FQB (2016) Virulence factors of Candida species isolated from patients with urinary tract infection and obstructive uropathy. Pak J Med Sci 32:143–146. https://doi.org/10.12669/pjms.321.8559
Gow NA (2013) A developmental program for Candida commensalism. Nat Genet 45:967–968. https://doi.org/10.1038/ng.2737
Noble SM, Gianetti BA, Witchley JN (2017) Candida albicans cell-type switching and functional plasticity in the mammalian host. Nat Rev Microbiol 15:96–108. https://doi.org/10.1038/nrmicro.2016.157
Jimenez-Lopez C, Lorenz MC (2013) Fungal immune evasion in a model host–pathogen interaction: Candida albicans versus macrophages. PLoS Pathog 9:e1003741. https://doi.org/10.1371/journal.ppat.1003741
Li F, Svarovsky MJ, Karlsson AJ, Wagner JP, Marchillo K, Oshel P, Andes D, Palecek SP (2007) Eap1p, an adhesin that mediates Candida albicans biofilm formation in vitro and in vivo. Eukaryot Cell 6:931–939. https://doi.org/10.1128/EC.00049-07
Hashash R, Younes S, Bahnan W, El Koussa J, Maalouf K, Dimassi HI, Khalaf RA (2011) Characterisation of Pga1, a putative Candida albicans cell wall protein necessary for proper adhesion and biofilm formation. Mycoses 54:491–500. https://doi.org/10.1111/j.1439-0507.2010.01883.x
Moyes DL, Richardson JP, Naglik JR (2015) Candida albicans-epithelial interactions and pathogenicity mechanisms: scratching the surface. Virulence 6:338–346. https://doi.org/10.1080/21505594.2015.1012981
Dalle F, Wächtler B, L’Ollivier C, Holland G, Bannert N, Wilson D, Labruère C, Bonnin A, Hube B (2010) Cellular interactions of Candida albicans with human oral epithelial cells and enterocytes. Cell Microbiol 12:248–271. https://doi.org/10.1111/j.1462-5822.2009.01394.x
Naglik JR (2014) Candida immunity. New J Sci 2014:27. https://doi.org/10.1155/2014/390241
Brand A, Shanks S, Duncan VM, Yang M, Mackenzie K, Gow NA (2007) Hyphal orientation of Candida albicans is regulated by a calcium-dependent mechanism. Curr Biol 17:347–352. https://doi.org/10.1016/j.cub.2006.12.043
Thompson DS, Carlisle PL, Kadosh D (2011) Coevolution of morphology and virulence in Candida species. Eukaryot Cell 10:1173–1182. https://doi.org/10.1128/EC.05085-11
Cavalheiro M, Teixeira MC (2018) Candida biofilms: threats, challenges, and promising strategies. Front Med 5:28. https://doi.org/10.3389/fmed.2018.00028
Hamied AS (2021) Candida and candidiasis-a review of virulence factors. J Biotechnol Res Cent 15:13–20. https://doi.org/10.24126/jobrc.2021.15.2.606
Talapko J, Juzbašić M, Matijević T, Pustijanac E, Bekić S, Kotris I, Škrlec I (2021) Candida albicans-the virulence factors and clinical manifestations of infection. J Fungi 7:79. https://doi.org/10.3390/jof7020079
Aliyu A (2022) Effects of biofilm formation and plethora of Candida species causing ailments: a mini-review. Gadau J Pure Alli Sci 1:200–210. https://doi.org/10.54117/gjpas.v1i2.27
Nobile CJ, Johnson AD (2015) Candida albicans biofilms and human disease. Annu Rev Microbiol 69:71. https://doi.org/10.1146/annurev-micro-091014-104330
Tomee JFC, Kauffman HF (2000) Putative virulence factors of Aspergillus fumigatus. Clin Exp Allergy 30:476–484. https://doi.org/10.1046/j.1365-2222.2000.00796.x
Gulati M, Nobile CJ (2016) Candida albicans biofilms: development, regulation, and molecular mechanisms. Microbes Infect 18:310–321. https://doi.org/10.1016/j.micinf.2016.01.002
Araujo D, Henriques M, Silva S (2017) Portrait of Candida species biofilm regulatory network genes. Trends Microbiol 25:62–75. https://doi.org/10.1016/j.tim.2016.09.004
Galocha M, Pais P, Cavalheiro M, Pereira D, Viana R, Teixeira MC (2019) Divergent approaches to virulence in C. albicans and C. glabrata: two sides of the same coin. Int J Mol Sci 20:2345. https://doi.org/10.3390/ijms20092345
Molero G, Díez-Orejas R, Navarro-García F, Monteoliva L, Pla J, Gil C, Sánchez-Pérez M, Nombela C (1998) Candida albicans: genetics, dimorphism and pathogenicity. Internatl Microbiol 1:95–106
Pukkila-Worley R, Peleg AY, Tampakakis E, Mylonakis E (2009) Candida albicans hyphal formation and virulence assessed using a Caenorhabditis elegans infection model. Eukaryot Cell 8:1750–1758. https://doi.org/10.1128/EC.00163-09
Han TL, Cannon RD, Villas-Bôas SG (2011) The metabolic basis of Candida albicans morphogenesis and quorum sensing. Fungal Genet Biol 48:747–763. https://doi.org/10.1016/j.fgb.2011.04.002
Cannon RD, Chaffin WL (1999) Oral colonization by Candida albicans. Crit Rev Oral Biol Med 10:359–383. https://doi.org/10.1177/10454411990100030701
Calderone R, Suzuki S, Cannon R, Cho T, Boyd D, Calera J, Chibana H, Herman D, Holmes A, Jeng HW, Kaminishi H, Matsumoto T, Mikami T, O’Sullivan JM, Sudoh M, Suzuki M, Nakashima Y, Tanaka T, Tompkins GR, Watanabe T (2000) Candida albicans: adherence, signaling, and virulence. Med Mycol 38:125–137. https://doi.org/10.1080/mmy.38.s1.125.137
Tsuchimori N, Sharkey LL, Fonzi WA, French SW, Edwards JE Jr, Filler SG (2000) Reduced virulence of HWP1-deficient mutants of Candida albicans and their interactions with host cells. Infect Immun 68:1997–2002. https://doi.org/10.1128/IAI.68.4.1997-2002.2000
Yang YL (2003) Virulence factors of Candida species. J Microbiol Immunol Ifect 36:223–228
Naglik JR, Challacombe SJ, Hube B (2003) Candida albicans secreted aspartyl proteinases in virulence and pathogenesis. Microbiol Mol Biol Rev 67:400–428. https://doi.org/10.1128/MMBR.67.3.400-428.2003
Williams DW, Jordan RP, Wei XQ, Alves CT, Wise MP, Wilson MJ, Lewis MA (2013) Interactions of Candida albicans with host epithelial surfaces. J Oral Microbiol 5:22434. https://doi.org/10.3402/jom.v5i0.22434
Samaranayake YH, Dassanayake RS, Cheung BPK, Jayatilake JAMS, Yeung KWS, Yau JYY, Samaranayake LP (2006) Differential phospholipase gene expression by Candida albicans in artificial media and cultured human oral epithelium. APMIS 114:857–866. https://doi.org/10.1111/j.1600-0463.2006.apm_479.x
Ibrahim AS, Mirbod F, Filler SG, Banno Y, Cole GT, Kitajima Y, Edwards JE, Nozawa Y, Ghannoum MA (1995) Evidence implicating phospholipase as a virulence factor of Candida albicans. Infect Immun 63:1993–1998. https://doi.org/10.1111/j.1567-1364.2009.00570.x
Cuervo G, Garcia-Vidal C, Puig-Asensio M, Merino P, Vena A, Martín-Peña A, Montejo JM, Ruiz A, Lázaro-Perona F, Fortún J, Fernández-Ruiz M, Suarez AI, Castro C, Cardozo C, Gudiol C, Aguado JM, Paño JR, Pemán J, Salavert M, Garnacho-Montero J, Cisneros JM, Soriano A, Muñoz P, Almirante B, Carratalà J, For the REIPI, the GEMICOMED (SEIMC) and the Spanish CANDI-Bundle Group (2019) Usefulness of guideline recommendations for prognosis in patients with candidemia. Med Mycol 57:659–667. https://doi.org/10.1093/mmy/myy118
Koehler P, Stecher M, Cornely OA, Koehler D, Vehreschild MJ, Bohlius J, Wisplinghoff H, Vehreschild JJ (2019) Morbidity and mortality of candidaemia in Europe: an epidemiologic meta-analysis. Clin Microbiol Infect 25:1200–1212. https://doi.org/10.1016/j.cmi.2019.04.024
Galia L, Pezzani MD, Compri M, Callegari A, Rajendran NB, Carrara E, Tacconelli E, The Combacte magnet epi-net network (2022) Surveillance of antifungal resistance in candidemia fails to inform antifungal stewardship in European countries. J Fungi 8:249. https://doi.org/10.3390/jof8030249
Arendrup MC, Patterson TF (2017) Multidrug-resistant Candida: epidemiology, molecular mechanisms, and treatment. J Infect Dis 216:S445–S451. https://doi.org/10.1093/infdis/jix131
World Health Organization (2021) First meeting of the WHO antifungal expert group on identifying priority fungal pathogens. Available online: https://www.who.int/publications/i/item/9789240006355. Accessed 11 Dec 2021
World Health Organization (2021) GLASS early implementation protocol for inclusion of Candida spp. Available online: https://www.who.int/publications/i/item/WHO-WSI-AMR-2019.4. Accessed 11 Dec 2021
Segal E, Frenkel M (2018) Experimental in vivo models of candidiasis. J Fungi 4:21. https://doi.org/10.3390/jof4010021
Diekema D, Arbefeville S, Boyken L, Kroeger J, Pfaller M (2012) The changing epidemiology of healthcare-associated candidemia over three decades. Diagn Microbiol Infect Dis 73:45–48. https://doi.org/10.1016/j.diagmicrobio.2012.02.001
Challacombe SJ (1994) Immunologic aspects of oral candidiasis. Oral Surg Oral Med Oral Pathol 78:202–210
Pfaller MA, Diekema DJ, Gibbs DL, Newell VA, Ellis D, Tullio V, Rodloff A, Fu W, Ling TA (2010) Results from the Artemis disk global antifungal surveillance study, 1997 to 2007: a 10.5-year analysis of susceptibilities of Candida species to fluconazole and voriconazole as determined by CLSI standardized disk diffusion. J Clin Microbiol 48:1366–1377. https://doi.org/10.1128/JCM.02117-09
Wächtler B, Citiulo F, Jablonowski N, Förster S, Dalle F, Schaller M, Wilson D, Hube B (2012) Candida albicans-epithelial interactions: dissecting the roles of active penetration, induced endocytosis and host factors on the infection process. PLoS One 7:e36952. https://doi.org/10.1371/journal.pone.0036952
Dabas PS (2013) An approach to etiology, diagnosis and management of different types of candidiasis. J Yeast Fungal Res 4:63–74. https://doi.org/10.5897/JYFR2013.0113
Taverne-Ghadwal L, Kuhns M, Buhl T, Schulze MH, Mbaitolum WJ, Kersch L, Weig M, Bader O, Groß U (2022) Epidemiology and prevalence of oral candidiasis in HIV patients from Chad in the post-HAART Era. Front Microbiol 13:844069. https://doi.org/10.3389/fmicb.2022.844069
Singh A, Verma R, Murari A, Agrawal A (2014) Oral candidiasis: an overview. J Oral Maxillofac Pathol 18:S81–S85. https://doi.org/10.4103/0973-029X.141325
Millsop JW, Fazel N (2016) Oral candidiasis. Clin Dermatol 34:487–494. https://doi.org/10.1016/j.clindermatol.2016.02.022
Hellstein JW, Marek CL (2019) Candidiasis: red and white manifestations in the oral cavity. Head Neck Pathol 13:25–32. https://doi.org/10.1007/s12105-019-01004-6
Kamath P, Pais M, Nayak MG (2013) Risk of vaginal candidiasis among pregnant women. Int J Curr Microbiol App Sci 2:141–146
Cateau E, Rodier MH, Imbert C (2012) Candidoses associées aux cathéters-quelle place pour les verrous antifongiques? Med Sci 28:740–745. https://doi.org/10.1051/medsci/2012288016
Seleem D, Pardi V, Murata RM (2017) Review of flavonoids: a diverse group of natural compounds with anti-Candida albicans activity in vitro. Arch Oral Biol 76:76–83. https://doi.org/10.1016/j.archoralbio.2016.08.030
Magill SS, Edwards JR, Bamberg W, Beldavs ZG, Dumyati G, Kainer MA, Ruth L, Meghan M, Laura MH, Joelle N, Susan MR, Deborah LT, Lucy EW, Fridkin SK (2014) Multistate point-prevalence survey of health care–associated infections. N Engl J Med 370:1198–1208. https://doi.org/10.1056/NEJMoa1306801
Cleveland AA, Harrison LH, Farley MM, Hollick R, Stein B, Chiller TM, Lockhart SR, Park BJ (2015) Declining incidence of candidemia and the shifting epidemiology of Candida resistance in two US metropolitan areas, 2008–2013: results from population-based surveillance. PLoS ONE 10:e0120452. https://doi.org/10.1371/journal.pone.0120452
Kullberg BJ, Arendrup MC (2015) Invasive candidiasis. N Engl J Med 373:1445–1456. https://doi.org/10.1056/NEJMra1315399
McCarty TP, Pappas PG (2016) Invasive candidiasis. Infect Dis Clin 30:103–124. https://doi.org/10.1016/j.idc.2015.10.013
Hesstvedt L, Gaustad P, Andersen CT, Haarr E, Hannula R, Haukland HH, Hermansen NO, Larssen KW, Mylvaganam H, Ranheim TE, Sandven P, Nordøy I, Kanestrøm A, Grub C, Onken A, Thielsen C, Skaare D, Tofteland S, Sønsteby LJ, Hjetland R, Hide R, Vik E, Kümmel A, Åsheim S (2015) Twenty-two years of candidaemia surveillance: results from a Norwegian national study. Clin Microbiol Infect 21:938–945. https://doi.org/10.1016/j.cmi.2015.06.008
Miceli MH, Díaz JA, Lee SA (2011) Emerging opportunistic yeast infections. Lancet Infect Dis 11:142–151. https://doi.org/10.1016/S1473-3099(10)70218-8
Ericson EL, Klingspor L, Ullberg M, Özenci V (2012) Clinical comparison of the Bactec mycosis IC/F, BacT/Alert FA, and BacT/Alert FN blood culture vials for the detection of candidemia. Diagn Microbiol Infect Dis 73:153–156. https://doi.org/10.1016/j.diagmicrobio.2012.02.020
Coronado-Castellote L, Jiménez-Soriano Y (2013) Clinical and microbiological diagnosis of oral candidiasis. J Clin Exp Dent 5:e279–e286. https://doi.org/10.4317/jced.51242
Erdogan A, Rao SS (2015) Small intestinal fungal overgrowth. Curr Gastroenterol Rep 17:1–7. https://doi.org/10.1007/s11894-015-0436-2
Zaoutis TE, Argon J, Chu J, Berlin JA, Walsh TJ, Feudtner C (2005) The epidemiology and attributable outcomes of candidemia in adults and children hospitalized in the United States: a propensity analysis. Clin Infect Dis 41:1232–1239. https://doi.org/10.1086/496922
Kojic EM, Darouiche RO (2004) Candida infections of medical devices. Clin Microbiol Rev 17:255–267. https://doi.org/10.1128/CMR.17.2.255-267.2004
Peleg AY, Hogan DA, Mylonakis E (2010) Medically important bacterial–fungal interactions. Nat Rev Microbiol 8:340–349. https://doi.org/10.1038/nrmicro2313
Coad BR, Kidd SE, Ellis DH, Griesser HJ (2014) Biomaterials surfaces capable of resisting fungal attachment and biofilm formation. Biotechnol Adv 32:296–307. https://doi.org/10.1016/j.biotechadv.2013.10.015
Coad BR, Griesser HJ, Peleg AY, Traven A (2016) Anti-infective surface coatings: design and therapeutic promise against device-associated infections. PLoS Pathog 12:e1005598. https://doi.org/10.1371/journal.ppat.1005598
Ahmad S, Khan Z (2012) Invasive candidiasis: a review of nonculture-based laboratory diagnostic methods. Indian J Med Microbiol 30:264–269. https://doi.org/10.4103/0255-0857.99482
Hoppe JE, Frey P (1999) Evaluation of six commercial tests and the germ-tube test for presumptive identification of Candida albicans. Eur J Clin Microbiol Infect Dis 18:188–191. https://doi.org/10.1007/s100960050256
Moya-Salazar J, Rojas R (2018) Comparative study for identification of Candida albicans with germ tube test in human serum and plasma. Clin Microbiol Infect Dis 3:1–4. https://doi.org/10.15761/CMID.1000143
Shin JH, Nolte FS, Holloway BP, Morrison CJ (1999) Rapid identification of up to three Candida species in a single reaction tube by a 5′ exonuclease assay using fluorescent DNA probes. J Clin Microbiol 37:165–170. https://doi.org/10.1128/JCM.37.1.165-170.1999
Latouche GN, Daniel HM, Lee OC, Mitchell TG, Sorrell TC, Meyer W (2007) Comparison of use of phenotypic and genotypic characteristics for identification of species of the anamorph genus Candida and related teleomorph yeast species. J Clin Microbiol 35:3171–3180. https://doi.org/10.1128/jcm.35.12.3171-3180.1997
Roberts GD, Wang HS, Hollick GE (1976) Evaluation of the API 20 C microtube system for the identification of clinically important yeasts. J Clin Microbiol 3:302–305. https://doi.org/10.1128/jcm.3.3.302-305.1976
Freydiere AM, Guinet R, Boiron P (2001) Yeast identification in the clinical microbiology laboratory: phenotypical methods. Med Mycol 39:9–33. https://doi.org/10.1080/mmy.39.1.9.33
Salkin IF, Pruitt WR, Padhye AA, Sullivan D, Coleman D, Pincus DH (1998) Distinctive carbohydrate assimilation profiles used to identify the first clinical isolates of Candida dubliniensis recovered in the United States. J Clin Microbiol 36:1467–1467. https://doi.org/10.1128/JCM.36.5.1467-1467.1998
Liguori G, Di Onofrio V, Lucariello A, Gallé F, Signoriello G, Colella G, D’Amora M, Rossano F (2009) Oral candidiasis: a comparison between conventional methods and multiplex polymerase chain reaction for species identification. Oral Microbiol Immunol 24:76–78. https://doi.org/10.1111/j.1399-302X.2008.00447.x
Bernal S, Mazuelos EM, Chávez M, Coronilla J, Valverde A (1998) Evaluation of the new API Candida system for identification of the most clinically important yeast species. Diagn Microbiol Infect Dis 32:217–221. https://doi.org/10.1016/S0732-8893(98)00119-9
Campbell CK, Davey KG, Holmes AD, Szekely A, Warnock DW (1999) Comparison of the API Candida system with the AUXACOLOR system for identification of common yeast pathogens. J Clin Microbiol 37:821–823. https://doi.org/10.1128/JCM.37.3.821-823.1999
Sudhan SS, Sharma P, Sharma M (2016) Candidemia in a tertiary care hospital in Jammu (J&K)-A comparison of conventional methods and CHROMagar technique for speciation. Int J Sci Res 5:451–454
Lockhart SR, Lyman MM, Sexton DJ (2022) Tools for detecting a “Superbug”: updates on Candida auris testing. J Clin Microbiol 60:e00808-e821. https://doi.org/10.1128/jcm.00808-21
Willinger B, Hillowoth C, Selitsch B, Manafi M (2001) Performance of Candida ID, a new chromogenic medium for presumptive identification of Candida species, in comparison to CHROMagar Candida. J Clin Microbiol 39:3793–3795. https://doi.org/10.1128/JCM.39.10.3793-3795.2001
Bauters TG, Nelis HJ (2002) Comparison of chromogenic and fluorogenic membrane filtration methods for detection of four Candida species. J Clin Microbiol 40:1838–1839. https://doi.org/10.1128/JCM.40.5.1838-1839.2002
Buchaille L, Freydiere AM, Guinet R, Gille Y (1998) Evaluation of six commercial systems for identification of medically important yeasts. Eur J Clin Microbiol Infect Dis 17:479–488. https://doi.org/10.1007/BF01691130
Yücesoy M, Marol S (2003) Performance of CHROMAGAR Candida and BIGGY agar for identification of yeast species. Ann Clin Microbiol Antimicrob 2:1–7. https://doi.org/10.1186/1476-0711-2-8
Pfaller MA, Preston T, Bale M, Koontz FP, Body BA (1988) Comparison of the Quantum II, API Yeast Ident, and automicrobic systems for identification of clinical yeast isolates. J Clin Microbiol 26:2054–2058. https://doi.org/10.1128/jcm.26.10.2054-2058.1988
Graf B, Adam T, Zill E, Göbel UB (2000) Evaluation of the VITEK 2 system for rapid identification of yeasts and yeast-like organisms. J Clin Microbiol 38:1782–1785. https://doi.org/10.1128/JCM.38.5.1782-1785.2000
Mikulska M, Furfaro E, Del Bono V, Raiola AM, Ratto S, Bacigalupo A, Viscoli C (2012) Piperacillin/tazobactam (TazocinTM) seems to be no longer responsible for false-positive results of the galactomannan assay. J Antimicrob Chemother 67:1746–1748. https://doi.org/10.1093/jac/dks111
Fontana C, Gaziano R, Favaro M, Casalinuovo IA, Pistoia ES, Di Francesco P (2012) 1–3)-β-D-glucan vs galactomannan antigen in diagnosing invasive fungal infections (IFIs. Open Microbiol J 6:70–73. https://doi.org/10.2174/1874285801206010070
Singh S (2014) Molecular-genetic approaches for identification and typing of pathogenic Candida yeasts: a review. Int J Innov Res Sci Eng Technol 3:16199–16211
Arnavielhe S, De Meeüs T, Blancard A, Mallie M, Renaud F, Bastide JM (2000) Multicentric genetic study of Candida albicans isolates from non-neutropenic patients using MLEE typing: population structure and mode of reproduction. Mycoses 43:109–117. https://doi.org/10.1046/j.1439-0507.2000.00552.x
Boriollo MFG, Rosa EAR, Gonçalves RB, Höfling JF (2006) Parity among interpretation methods of MLEE patterns and disparity among clustering methods in epidemiological typing of Candida albicans. J Microbiol Methods 64:346–365. https://doi.org/10.1016/j.mimet.2005.05.012
Magee BB, Magee PT (1987) Eleetrophoretic karyotypes and chromosome numbers in Candida species. J Gen Microbiol 133:425–430. https://doi.org/10.1099/00221287-133-2-425
Santos PO, Melo JO, Ponzzes CMPBS, Alves JAB, De Melo DLFM, Botelho NDS, Yamada-Ogatta SF, Mann RS, Trindade RDC (2012) Multilocus enzyme electrophoresis analysis and exoenzymatic activity of Candida albicans strains isolated from women with vaginal candidiasis. Mycoses 55:64–72. https://doi.org/10.1111/j.1439-0507.2011.02043.x
Hahm BK, Maldonado Y, Schreiber E, Bhunia AK, Nakatsu CH (2003) Subtyping of foodborne and environmental isolates of Escherichia coli by multiplex-PCR, rep-PCR, PFGE, ribotyping and AFLP. J Microbiol Methods 53:387–399. https://doi.org/10.1016/S0167-7012(02)00259-2
Olive DM, Bean P (1999) Principles and applications of methods for DNA-based typing of microbial organisms. J Clin Microbiol 37:1661–1669. https://doi.org/10.1128/JCM.37.6.1661-1669.1999
Lehmann PF, Lin D, Lasker BA (1992) Genotypic identification and characterization of species and strains within the genus Candida by using random amplified polymorphic DNA. J Clin Microbiol 30:3249–3254. https://doi.org/10.1128/jcm.30.12.3249-3254.1992
Tabit FT (2016) Advantages and limitations of potential methods for the analysis of bacteria in milk: a review. J Food Sci Technol 53:42–49. https://doi.org/10.1007/s13197-015-1993-y
Ball LM, Bes MA, Theelen B, Boekhout T, Egeler RM, Kuijper EJ (2004) Significance of amplified fragment length polymorphism in identification and epidemiological examination of Candida species colonization in children undergoing allogeneic stem cell transplantation. J Clin Microbiol 42:1673–1679. https://doi.org/10.1128/JCM.42.4.1673-1679.2004
Levterova V, Panaiotov S, Brankova N, Tankova K (2010) Typing of genetic markers involved in stress response by fluorescent cDNA-amplified fragment length polymorphism technique. Mol Biotechnol 45:34–38. https://doi.org/10.1007/s12033-009-9236-y
Hsu MC, Chen KW, Lo HJ, Chen YC, Liao MH, Lin YH, Li SY (2003) Species identification of medically important fungi by use of real-time light cycler PCR. J Med Microbiol 52:1071–1076. https://doi.org/10.1099/jmm.0.05302-0
Magalhães J, Correia MJ, Silva RM, Esteves AC, Alves A, Duarte AS (2022) Molecular techniques and target selection for the identification of Candida spp in oral samples. Appl Sci 12:9204. https://doi.org/10.3390/app12189204
Morace G, Sanguinetti M, Posteraro B, Lo Cascio G, Fadda G (1997) Identification of various medically important Candida species in clinical specimens by PCR-restriction enzyme analysis. J Clin Microbiol 35:667–672. https://doi.org/10.1128/jcm.35.3.667-672.1997
Widjojoatmodjo MN, Borst A, Schukkink RAF, Box ATA, Tacken NMM, Gemen BV, Verhoef J, Top B, Fluit AC (1999) Nucleic acid sequence-based amplification (NASBA) detection of medically important Candida species. J Microbiol Methods 38:81–90. https://doi.org/10.1016/S0167-7012(99)00079-2
Kempf VAJ, Trebesius K, Autenrieth IB (2000) Fluorescent in situ hybridization allows rapid identification of microorganisms in blood cultures. J Clin Microbiol 38:830–838. https://doi.org/10.1128/JCM.38.2.830-838.2000
Sampaio P, Gusmao L, Correia A, Alves C, Rodrigues AG, Pina-Vaz C, Amorim A, Pais C (2005) New microsatellite multiplex PCR for Candida albicans strain typing reveals microevolutionary changes. J Clin Microbiol 43:3869–3876. https://doi.org/10.1128/JCM.43.8.3869-3876.2005
Garcia-Hermoso D, Cabaret O, Lecellier G, Desnos-Ollivier M, Hoinard D, Raoux D, Costa JM, Dromer F, Bretagne S (2007) Comparison of microsatellite length polymorphism and multilocus sequence typing for DNA-based typing of Candida albicans. J Clin Microbiol 45:3958–3963. https://doi.org/10.1128/jcm.01261-07
Bougnoux ME, Tavanti A, Bouchier C, Gow NAR, Magnier A, Davidson AD, Maiden MCJ, d’Enfert C, Odds FC (2003) Collaborative consensus for optimized multilocus sequence typing of Candida albicans. J Clin Microbiol 41:5265–5266. https://doi.org/10.1128/JCM.41.11.5265-5266.2003
Kurella M, Hsiao LL, Yoshida T, Randall JD, Chow G, Sarang SS, Jensen RV, Gullans SR (2001) DNA microarray analysis of complex biologic processes. J Am Soc Nephrol 12:1072–1078. https://doi.org/10.1681/ASN.V1251072
Kittichotirat W, Bumgarner RE, Asikainen S, Chen C (2011) Identification of the pangenome and its components in 14 distinct Aggregatibacter actinomycetemcomitans strains by comparative genomic analysis. PLoS ONE 6:e22420. https://doi.org/10.1371/journal.pone.0022420
Timmins EM, Howell SA, Alsberg BK, Noble WC, Goodacre R (1998) Rapid differentiation of closely related Candida species and strains by pyrolysis-mass spectrometry and fourier transform-infrared spectroscopy. J Clin Microbiol 36:367–374. https://doi.org/10.1128/JCM.36.2.367-374.1998
Barantsevich N, Barantsevich E (2022) Diagnosis and treatment of invasive candidiasis. Antibiotics 11:718. https://doi.org/10.3390/antibiotics11060718
Dujon B, Sherman D, Fischer G, Durrens P, Casaregola S, Lafontaine I, de Montigny J, Marck C, Neuvéglise C, Talla E, Goffard N, Goffard N, Frangeul L, Aigle M, Anthouard V, Babour A, Barbe V, Barnay S, Blanchin S, Beckerich JM, Beyne E, Bleykasten C, Boisramé A, Boyer J, Cattolico L, Confanioleri F, de Daruvar A, Despons L, Fabre E, Fairhead C, Ferry-Dumazet H, Groppi A, Hantraye F, Hennequin C, Jauniaux N, Joyet P, Kachouri R, Kerrest A, Koszul R, Lemaire M, Lesur I, Ma L, Muller H, Nicaud JM, Nikolski M, Oztas S, Ozier-Kalogeropoulos O, Pellenz S, Potier S, Richard GF, Straub ML, Suleau A, Swennen D, Tekaia F, Wésolowski-Louvel M, Westhof E, Wirth B, Zeniou-Meyer M, Zivanovic I, Bolotin-Fukuhara M, Thierry A, Bouchier C, Caudron B, Scarpelli C, Gaillardin C, Weissenbach J, Wincker P, Souciet JL (2004) Genome evolution in yeasts. Nature 430:35–44. https://doi.org/10.1038/nature02579
Butler G, Rasmussen MD, Lin MF, Santos MA, Sakthikumar S, Munro CA, Rheinbay E, Grabherr M, Forche A, Reedy JL, Agrafioti I, Arnaud MB, Bates S, Brown AJP, Brunke S, Costanzo MC, Fitzpatrick DA, de Groot PWJ, Harris D, Hoyer LL, Hube B, Klis FM, Kodira C, Lennard N, Logue ME, Martin R, Neiman AM, Nikolaou E, Quail MA, Quinn J, Santos MC, Schmitzberger FF, Sherlock G, Shah P, Silverstein KAT, Skrzypek MS, Soll D, Staggs R, Stansfield I, Stumpf MPH, Sudbery PE, Srikantha T, Zeng Q, Berman J, Berriman M, Heitman J, Gow NAR, Lorenz MC, Birren BW, Kellis M, Cuomo CA (2009) Evolution of pathogenicity and sexual reproduction in eight Candida genomes. Nature 459:657–662. https://doi.org/10.1038/nature08064
Zhang J, Hollis RJ, Pfaller MA (1997) Variations in DNA subtype and antifungal susceptibility among clinical isolates of Candida tropicalis. Diagn Microbiol Infect Dis 27:63–67. https://doi.org/10.1016/S0732-8893(97)00002-3
Kasianowicz JJ, Brandin E, Branton D, Deamer DW (1996) Characterization of individual polynucleotide molecules using a membrane channel. Proc Natl Acad Sci USA 93:13770–13773. https://doi.org/10.1073/pnas.93.24.1377
Ashikawa S, Tarumoto N, Imai K, Sakai J, Kodana M, Kawamura T, Ikebuchi K, Murakami T, Mitsutake K, Maesaki S, Maeda T (2018) Rapid identification of pathogens from positive blood culture bottles with the MinION nanopore sequencer. J Med Microbiol 67:1589–1595. https://doi.org/10.1099/jmm.0.000855
Gabaldón T (2019) Recent trends in molecular diagnostics of yeast infections: from PCR to NGS. FEMS Microbiol Rev 43:517–547. https://doi.org/10.1093/femsre/fuz015
Gootenberg JS, Abudayyeh OO, Lee JW, Essletzbichler P, Dy AJ, Joung J, Verdine V, Donghia N, Daringer NM, Freije CA, Myhrvold C, Bhattacharyya RP, Livny J, Regev A, Koonin EV, Hung DT, Sabeti PC, Collins JJ, Zhang F (2017) Nucleic acid detection with CRISPR-Cas13a/C2c2. Science 356:438–442. https://doi.org/10.1126/science.aam9321
Gootenberg JS, Abudayyeh OO, Kellner MJ, Joung J, Collins JJ, Zhang F (2018) Multiplexed and portable nucleic acid detection platform with Cas13, Cas12a, and Csm6. Science 360:439–444. https://doi.org/10.1126/science.aaq0179
Myhrvold C, Freije CA, Gootenberg JS, Abudayyeh OO, Metsky HC, Durbin AF, Kellner MJ, Tan AL, Paul LM, Parham LA, Garcia KF, Barnes KG, Chak B, Mondini A, Nogueira ML, Isern S, Michael SF, Lorenzana I, Yozwiak NL, Macinnis BL, Bosch I, Gehrke L, Zhang F, Sabeti PC (2018) Field-deployable viral diagnostics using CRISPR-Cas13. Science 360:444–448. https://doi.org/10.1126/science.aas8836
Funding
No funding was received to assist with the preparation of this manuscript.
Author information
Authors and Affiliations
Contributions
SHA: collecting resources, preparing figures, and writing up. GEHO: conceptualization. KE: review final draft of manuscript. HHA: language editing.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare no conflict of interest.
Ethical approval
Not applicable.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Arafa, S.H., Elbanna, K., Osman, G.E.H. et al. Candida diagnostic techniques: a review. J.Umm Al-Qura Univ. Appll. Sci. 9, 360–377 (2023). https://doi.org/10.1007/s43994-023-00049-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s43994-023-00049-2