Abstract
ASXL1 is one of the most frequently mutated genes in malignant myeloid diseases. In patients with myeloid malignancies, ASXL1 mutations are usually heterozygous frameshift or nonsense mutations leading to C-terminal truncation. Current disease models have predominantly total loss of ASXL1 or overexpressed C-terminal truncations. These models cannot fully recapitulate leukemogenesis and disease progression. We generated an endogenous C-terminal-truncated Asxl1 mutant in zebrafish that mimics human myeloid malignancies. At the embryonic stage, neutrophil differentiation was explicitly blocked. At 6 months, mutants initially exhibited a myelodysplastic syndrome-like phenotype with neutrophilic dysplasia. At 1 year, about 13% of mutants further acquired the phenotype of monocytosis, which mimics chronic myelomonocytic leukemia, or increased progenitors, which mimics acute myeloid leukemia. These features are comparable to myeloid malignancy progression in humans. Furthermore, transcriptome analysis, inhibitor treatment, and rescue assays indicated that asxl1-induced neutrophilic dysplasia was associated with reduced expression of bmi1a, a subunit of polycomb repressive complex 1 and a reported myeloid leukemia-associated gene. Our model demonstrated that neutrophilic dysplasia caused by asxl1 mutation is a foundation for the progression of myeloid malignancies, and illustrated a possible effect of the Asxl1-Bmi1a axis on regulating neutrophil development.
Similar content being viewed by others
Introduction
Myeloid malignancies are characterized by the proliferation and defective differentiation of myeloid progenitors [1, 2]. The main types of these malignancies are myelodysplastic syndrome (MDS), myeloproliferative neoplasms (MPNs), and acute myeloid leukemia (AML). Chronic myelomonocytic leukemia (CMML) exhibits features of both MPN and MDS [3]. MDS, MPNs, and CMML have an inherent risk of transforming to AML. Knowledge about the leukemogenic mechanisms of myeloid malignancies can be a basis for developing therapeutic options.
Patients with multiple myeloid malignancies [4] including MDS, CMML, and AML frequently harbor somatic mutations in addition of sex combs-like 1 (ASXL1). In patients, ASXL1 mutations are usually heterozygous frameshift or nonsense mutations leading to C-terminal truncation [4] and loss of the ASXM2 domain and a PHD finger. Truncated ASXL1 mutations independently predict poor outcomes and shorter overall survival for patients with CMML [5, 6].
To understand how ASXL1 participates in normal hematopoiesis and leukemogenesis, numerous models of aberrant ASXL1 expression have been described [7,8,9,10,11]. Mouse Asxl1 knockout (KO) models exhibit multilineage dysplasia [7]. Whereas loss of asxl1 in zebrafish is described as leading to the apoptosis of hematopoietic stem cells (HSCs), two 17-month adult asxl1 mutants developed an AML-like phenotype [12]. Overexpression of C-terminally truncated ASXL1 in mice inhibits myeloid differentiation and induces MDS-like diseases [8, 10, 11]. To obtain a model more closely resembling human disease, heterozygous Asxl1G643fs mutant knock-in mice were generated [13, 14]. These models have no obvious hematopoiesis disorders within 18 months, but show MDS/MPN-like disease by 18–24 months [13, 14]. Even with these models and their different phenotypes, the role of ASXL1 in hematopoiesis is still largely unknown.
Mechanistically, the ASXL1 protein is thought to mediate the balance between polycomb and trithorax functions by regulating histone modifications such as monoubiquitination of histone H2A at lysine 119 (H2AK119ub) [15, 16] and methylation of histone H3 at lysine 3 (H3K4me3) [10]. C-terminally truncated ASXL1 aberrantly enhances BAP1 function in deubiquitination of H2AK119ub [16,17,18], which is catalyzed by polycomb repressive complex 1 (PRC1) [19]. This hyperactive mutant ASXL1/BAP1 complex is thought to promote impairment of myeloid differentiation through inhibition of H2AK119ub at posterior HOXA genes and IRF8, encoding an essential transcription factor in myeloid lineages [18]. The ASXL1G643fs mutation disrupts interactions with BMI1, a subunit of PRC1, and downregulated BMI1-driven H2AK119ub is observed in the derepressed p16Ink4a, leading in an MDS-like phenotype [14]. Hence, precisely how ASXL1 regulates H2AK119ub remains unclear. Furthermore, no consistent evidence indicates which factor is more important in mutant ASXL1-induced leukemogenesis. We need a model that closely resembles disease development in patients to recapitulate potential leukemogenesis and disease progression with ASXL1 C-terminal mutations.
An increasing number of leukemia models have been developed in zebrafish (Danio rerio) [20]. With properties such as external fertilization and embryonic development, zebrafish are an excellent model to study hematopoiesis, especially in early embryogenesis. To elucidate the function of Asxl1 in hematopoiesis and the leukemogenic mechanisms of Asxl1-truncated mutants, we established an endogenous truncated mutant in zebrafish that is comparable to human disease. Our C-terminally truncated zebrafish asxl1 mutant, unlike mouse models, has explicitly impaired neutrophil development. At 1 year, in addition to neutrophilic dysplasia, around 13% of the mutants had the phenotype of monocytosis, which resembles CMML, or increased myeloblasts, similar to AML. This leukemogenesis induced by the truncated mutant Asxl1 is transplantable. Furthermore, we found that expression of bmi1a, a subunit of PRC1, was decreased in asxl1−/− fish and its reduction was associated with neutrophil dysplasia in our mutants. These results were demonstrated with transcriptome analysis, inhibitor treatment, and rescue assays. Our work showed that impaired neutrophil differentiation is fundamental for the progression of myeloid malignancies and illustrated the possible function of the Asxl1-Bmi1 axis in regulating neutrophil development.
Materials and methods
All information on oligos and target sequences is in Supplementary Table 1.
Zebrafish, embryo collection, and treatment
All experiments involving zebrafish were done in accordance with guidelines from the Institutional Animal Care and Use Committee of South China University of Technology. Wild-type AB strain zebrafish were raised and maintained under standard conditions [21]. Embryos were collected and staged according to Kimmel et al. [22].
Generation of asxl1-mutant lines and validation
CRISPR/Cas9 was used to create asxl1 mutants. Cas9 mRNA was synthesized using mMESSAGE mMACHINE mRNA transcription-synthesis kits (Ambion). Cas9 mRNA (0.1 ng/embryo) and guide RNA (0.05 ng/embryo) were injected into single-cell wild-type embryos. Mutant embryos were identified through genotyping (see Supplementary Methods) and allowed to develop to F0. We identified F0 fish with inheritable mutations. F1 progeny from F0 outcrosses was identified by genotyping and sequencing. Identified F1 and progeny were used for experiments. asxl1+/+ and asxl1−/− embryos were generated and genotyped from heterozygous intercrosses.
RNA extraction and qRT-PCR
RNA was extracted using TRIzol (Invitrogen) and reverse transcribed by M-MLV Reverse Transcriptase (Promega). qPCR was with FastStart SYBR Green Master (Roche) and fold-changes determined by ΔΔCt method. All reactions were normalized against ef1α. Melting-curve analysis confirmed the presence of single PCR products. Significance was determined using Student’s t tests with a significance threshold of p < 0.05.
RNA-sequencing (RNA-seq) and bioinformatics
At 3 days postfertilization (dpf), asxl1+/+ and asxl1−/− embryos from asxl1+/− intercrosses were collected, and tails including caudal hematopoietic tissue were used for total RNA extraction. Each sample library was from around 20 tails. Sequencing libraries were prepared from poly-A selected RNA using NGS RNA Library Prep kits (Novogene). Sequencing was by an Illumina NovaSeq sequencing platform to obtain 2 × 150-bp pair-end reads. Sequencing quality was assessed using FastQC [23]. Reads were mapped onto the GRCz11 zebrafish reference genome using STAR (version 2.5.1b) with default setting [24]. Raw read counts for each Ensembl-annotated gene were calculated by Featurecounts [25]. EdgeR (version 3.26.5, false discovery rate (FDR) < 0.05; exact test) was used to call differentially expressed genes [26, 27]. Gene ontology (GO) for biological processes was enriched using the Metascape online tool [28].
Inhibitor treatment
Embryos at 24 h postfertilization (hpf) were soaked in egg water containing UNC3866 (Selleckchem, S8359), PRT4165 (Selleckchem, S5315), CPI455 HCl (Selleckchem, S8287), or GSK J4 (Selleckchem, S7581).
Results
Zebrafish asxl1 is expressed in hematopoietic cells, including myeloid cells
The asxl1 and asxl2 genes are zebrafish homologs of the Drosophila Asx gene. Mammalian and zebrafish Asxl proteins are evolutionarily conserved. Phylogenetic analysis clustered zebrafish Asxl1 with mouse and human ASXL1, but separately from groups of ASXL2 or ASXL3 protein sequences (Supplementary Fig. 1A). The amino acid sequences of zebrafish Asxl1 and Asxl2 share an overall 33 and 42% identity with human ASXL1 and ASXL2 orthologs. They share a conserved common domain architecture, including ASXN, ASXH, ASXM1, ASXM2, and PHD domains (Supplementary Fig. 1B). These results indicated that asxl1/2 are authentic orthologs of mammalian ASXL1/2. The conserved structural domains suggest functional conservation of ASXL members among species during evolution.
Expression patterns of asxl1 during embryonic development were detected by WISH (Supplementary Fig. 1C). In embryos, asxl1 was expressed as early as the 1-cell stage, indicating the presence of maternal transcript. At 5.3 hpf, asxl1 was ubiquitously expressed and at 18 hpf, asxl1 was restricted to the developing brain, eye, anterior lateral mesoderm, and posterior lateral mesoderm. At 24 and 36 hpf, asxl1 was specifically expressed in brain and eye. At 2 dpf, asxl1 was mostly in the nervous system (Supplementary Fig. 1C).
To investigate if asxl1 participates in hematopoiesis, we first examined if asxl1 expresses in hematopoietic cells. We collected adult kidney marrow cells from wild type before further sorting myeloid and lymphoid cells by FACS. qRT-PCR confirmed asxl1 expression in adult hematopoietic cells, with higher levels in myeloid cells (Supplementary Fig. 1D). Hence, asxl1 was expressed in hematopoietic cells, including myeloid cells. The expression of asxl1 in myeloid cells supported the hypothesis that asxl1 participates in zebrafish hematopoiesis, including myelopoiesis.
asxl1 C-terminal truncation generates new zebrafish mutants
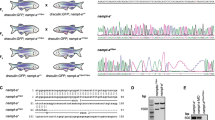
To examine if asxl1 participates in hematopoiesis, we generated an asxl1-mutant line. CRISPR/Cas9 methods were used at exon 12 of the zebrafish asxl1 gene to produce lines with premature stop codons, similar to mutations in myeloid malignancies (Fig. 1A). We obtained two independent asxl1 alleles with exon 12 deletions of 7 bp in asxl1e12 (−7) and 22 bp in asxl1e12 (−22). Both deletions predicted to lead to stop codons (in amino acids 919 and 914, respectively) before the ASXM2 and PHD domains (Fig. 1A). These two mutants have the same phenotypes so unless specified, asxl1e12 (−22) data are presented. qRT-PCR for asxl1 expression in mutants showed decreased, which may be caused by nonsense-mediated decay (Fig. 1B). Most surviving asxl1−/− fish displayed dwarfism and lower body weight (Fig. 1C). These results indicated that C-terminally truncated Asxl1 in zebrafish may lead to developmental abnormalities.
A Site-specific targeting for CRISPR/Cas9 cleavage within exon 12 of the zebrafish asxl1 gene. Alignment of nucleotide sequences from wild-type and mutant asxl1 alleles in asxl1e12 (−7) and asxl1e12 (−22) zebrafish lines. Dashes in DNA sequences are the nucleotides deleted during repair of CRISPR/Cas9-induced double-strand breaks. PAM protospacer adjacent motif. CRISPR/cas9-induced asxl1 frameshift mutations predicted to lead to C-terminally truncated proteins. Truncated proteins predicted from mutant alleles asxl1e12 (−7) and asxl1e12 (−22) lack the last two domains (ASXM2 and PHD). B qRT-PCR comparing expression of asxl1 in asxl1+/+ and mutant asxl1−/− in 3 days postfertilization (dpf) larvae and adult kidney marrow (3 dpf, 3 dpf larvae tails, n ≥ 10 per group, performed with four replicates; 6 m, 6-month kidney marrow, n = 4 per genotype; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± standard deviation (SD)). C Gross appearance of asxl1−/− zebrafish compared with asxl1+/+ littermates (at 1.5 years). Body weights of asxl1+/+ and asxl1−/− zebrafish (scale bar, 1 cm; asxl1+/+, n = 33; asxl1−/−, n = 24; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001, error bars, mean ± SD).
Zebrafish asxl1 mutants have impaired neutrophil differentiation as embryos
To investigate asxl1 in zebrafish hematopoiesis, we detected expression of lineage markers in mutants from intercrosses of asxl1 heterozygotes. First, the effects of asxl1 C-terminal truncated mutation on primitive hematopoiesis (<24 hpf) were examined. Erythropoiesis (gata1) and myeloid progenitor (pu.1) markers were indistinguishable between asxl1 mutants and siblings (Supplementary Fig. 2A). Additionally, a mature neutrophil marker (lyz) and intermediate neutrophilic progenitor marker for myeloperoxidase (mpx) were comparable between asxl1 mutant and siblings (Supplementary Fig. 2A). Finally, we examined expression of a gene associated with macrophages (mfap4) (Supplementary Fig. 2A) and found it was not impaired in asxl1 mutants. These data suggested that primitive hematopoiesis was not disrupted in asxl1 mutants.
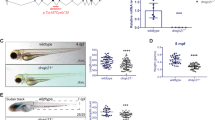
We next characterized the biological function of asxl1 in zebrafish definitive hematopoiesis by examining the effects of asxl1 C-terminal truncated mutation on hematopoietic lineages. WISH and qRT-PCR at 3 dpf embryos (Fig. 2A) showed significantly reduced lyz and mpx transcripts. These observations suggested that asxl1 may disrupt differentiation from myeloid progenitor to neutrophil. Supporting this, Sudan Black staining showed that SB+ granule containing mature neutrophils [29] were reduced in asxl1 mutants (Fig. 2A). Furthermore, FACS-sorted neutrophils from asxl1+/+ and asxl1−/− embryos were May–Grünwald–Giemsa stained to observe neutrophils morphology. Compared with wild type, the asxl1 mutant exhibited a lower proportion of mature neutrophils, whose signature is banded and segmented nuclei (Fig. 2B). Expression of HSC marker (c-myb) (Supplementary Fig. 2B) and a lymphocyte marker (rag1) has no obvious change between asxl1 mutants and siblings (Supplementary Fig. 2C). Expression of an erythrocyte (hbbe1) and a macrophage marker (mfap4) was not impaired in asxl1 mutants (Supplementary Fig. 2C). Collectively, these results suggested that asxl1 C-terminal truncation may specifically impair neutrophil differentiation. To determine if asxl1 mutation caused the impaired neutrophil development, we performed rescue experiments. After injecting wild-type asxl1 mRNA into 1-cell stage embryos, lyz+ cells increased in 3 dpf asxl1 mutants and siblings (Fig. 2C). This result suggested a crucial role for asxl1 in neutrophil differentiation.
A Decreased lyz and mpx expression and decreased SB+ cells in asxl1 mutants. WISH for lyz, mpx, and SB staining (3 dpf; scale bar, 200 μm; black boxes show enlarged details; lyz WISH, asxl1+/+, n = 16, asxl1−/−, n = 17; mpx WISH, asxl1+/+, n = 22, asxl1−/−, n = 17; SB staining, asxl1+/+, n = 19, asxl1−/−, n = 24; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD). qRT-PCR comparison of lyz and mpx expression between wild type and mutant (3 dpf larvae, n ≥ 10 per group, performed with four replicates; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD). B May–Grünwald–Giemsa staining of neutrophils from 4 dpf embryos and quantification of mature neutrophils by morphology. Neutrophils were collected from over 400 embryos of each genotype asxl1+/+ or asxl1−/−. After staining, 100 cells were randomly chosen for further calculation. This process was repeated three times for statistical tests (scale bars, 10 μm; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD). C Reduced lyz expression rescued by asxl1 expression. Left panel: WISH of lyz (3 dpf) after injection of asxl1 mRNA. Right panel: quantification of lyz+ cells in asxl1+/+ and asxl1−/− tails (0.1 ng mRNA/embryo; scale bar, 200 μm; black boxes show enlarged images; asxl1sib, n = 19, asxl1mut, n = 10, asxl1sib R, n = 27, asxl1mut = 6. R, mRNA injected; one-way ANOVA followed by LSD Fisher’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD).
Six-month zebrafish asxl1 mutants exhibit MDS-like phenotypes with neutrophilic dysplasia
The disruption of neutrophil development in asxl1−/− larvae may lead to MDS-like phenotypes. To investigate this possibility, hematopoietic cells from kidney marrow, the functional ortholog of bone marrow, were examined in surviving homozygous mutants. Cytological staining of 6-month-old mutants indicated a significant decrease in the proportion of mature neutrophils accompanied by an increase in the proportion of erythroblasts (Fig. 3A). Neutrophils were FACS-sorted from asxl1+/+ and asxl1−/− whole kidney marrow and May–Grünwald–Giemsa stained (Fig. 3B). Neutrophils from asxl1−/− had lower percentages of banded and segmented mature neutrophils than wild type (Fig. 3B). Expression of neutrophil markers (cebp1, mpx, lyz) was lower in asxl1 mutants than siblings, while expression of progenitor markers (cebpa) was higher (Fig. 3C). This finding indicated that asxl1-mutant neutrophils remained in an immature stage. The decrease and immaturity of neutrophils are typical of MDS with neutrophilic dysplasia [3].
A Proportion of hematopoietic cells in whole kidney marrow of 6-month-old asxl1+/+, asxl1−/− (n = 9 and n = 10, respectively; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD). Mye P myeloid progenitors, Mye mature myeloid cells, Ery erythrocytes, Lym lymphocytes, Neu neutrophils, Mac macrophages, Eos eosinophils. May–Grünwald–Giemsa staining of whole kidney marrow from 6-month-old asxl1+/+ and asxl1−/− (scale bar, 20 μm; black boxes show enlarged details of neutrophils; red arrows, neutrophils). B May–Grünwald–Giemsa staining of neutrophils from kidney marrow. Neutrophils were quantified by morphology. Neutrophils were collected from ten adult fish of each genotype asxl1+/+ or asxl1−/−. After staining, 500 cells were randomly chosen for further calculation. This process was repeated three times for statistical tests (scale bar, 20 μm; black boxes show enlarged details of neutrophils; red arrows, neutrophils; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001, error bars, mean ± SD). C qRT-PCR comparing expression of myeloid markers in sorted neutrophils from asxl1+/+ and asxl1−/−. Neutrophils were collected from ten adult fish of each genotype asxl1+/+ or asxl1−/−. qPCR was performed with three technical replicates (two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001, error bars, mean ± SD) (color figure online).
About 13% of 1-year-old zebrafish asxl1 mutants progress to CMML- or AML-like disease
Both loss of Asxl1 and transgenically expressed truncated Asxl1 induce severe hematopoietic diseases in mice [7,8,9,10,11]. We therefore tested if adult asxl1 zebrafish mutants developed severe myeloid malignancies. Kidney marrow from 55 1-year-old adult asxl1−/− mutants was collected for blood cell count analysis (Supplementary Fig. 3A–F and Supplementary Table 2). These samples showed a significantly decreased proportion of neutrophils, similar to the 6-month mutants (Supplementary Fig. 3A). We observed a significantly increased proportion of monocytes/macrophages in asxl1 mutants (Supplementary Fig. 3B) and a small increase in the proportion of myeloid progenitors (Supplementary Fig. 3C). Eosinophils and lymphocytes had no obvious change in asxl1 mutants (Supplementary Fig. 3D, E). The significant increase in the proportion of erythroblasts in 6-month asxl1 mutants was no longer observed in 1-year stage mutants (Supplementary Fig. 3F). This result may suggest the increased proportion of erythroblasts may be a temporary phenotype or found only in some samples. These data indicated that the asxl1-truncated mutation induced myeloid malignancies with neutrophilic dysplasia and monocytosis.
Among the 55 1-year adult asxl1 mutants, 5 had a severe monocytosis phenotype (defined as >20% of kidney marrow cells as monocytes/macrophages) with neutrophilic dysplasia and without excessive blasts (Supplementary Figs. 3A–C and 4A). These phenotypes are similar to CMML patients, who have increased numbers of monocytes with at least one lineage dysplasia, as classified in the World Health Organization (WHO) guidelines [3]. We also found two asxl1 mutants with increased numbers of blasts. In these two mutants, the proportion of myeloid progenitors/blasts was over 40% of whole kidney marrow (Supplementary Fig. 3C and Fig. 4A), similar to features associated with human AML, as defined by WHO [30]. These data suggested that some of the asxl1-truncated zebrafish mutants further developed CMML-like or AML-like disease (Fig. 4A).
A May–Grünwald–Giemsa staining of peripheral blood (PB) and whole kidney marrow (KM) cells presentative five fish with CMML-like and two fish with AML-like phenotypes (scale bar, 20 μm; black boxes show enlarged details of macrophages and myeloid progenitors; red arrows, macrophages; yellow arrows, myeloid progenitors). B Hematoxylin and eosin staining of paraffin-embedded sections of liver from representative asxl1+/+ (a, b) and asxl1−/− (c–f) (scale bar, 40 μm; black box, macrophage infiltration; green arrow, vacuolated cytoplasm; red arrow, gaps; yellow arrow, fibrosis) (color figure online).
Histologic analysis of liver sections was performed on zebrafish with myeloid malignancies. CMML-like zebrafish showed perivascular macrophage infiltration (Fig. 4B—c, d). Liver parenchyma appeared abnormal in these mutants, with poorly demarcated cells (Fig. 4B). Cells with vacuolated cytoplasm were frequently observed in asxl1−/− (Fig. 4B—e), similar to phenotypes as observed in by Gjini et al. [12]. Many gaps in the liver parenchyma were observed (Fig. 4B). Severe fibrosis was also observed in one of the CMML-like zebrafish (Fig. 4B—f). This phenomenon was similar to nonalcoholic fatty liver disease (NAFLD), where macrophages have a central role in progression [31]. The significantly increased proportion of monocytes/macrophages in our asxl1 mutant may be a contributor to NAFLD-like progression in our asxl1 mutants.
C-terminally truncated Asxl1 mutation-induced leukemia cells are transplantable
To confirm if C-terminally truncated asxl1 hematopoietic cells intrinsically induced leukemogenesis, we performed transplantation assays (Fig. 5A). asxl1+/+ and asxl1−/− kidney marrow cells were intracardially injected into sublethally irradiated 9-month wild-type recipient fish. At 18 days posttransplantation, 3 asxl1+/+ and 5 asxl1−/− recipients survived out of 19 and 35, respectively (Fig. 5A). The peripheral blood and kidney marrow cells from surviving recipients were collected for genotyping. Of five asxl1−/− surviving recipients, four successfully repopulated (Supplementary Fig. 4A, B). Kidney marrow from these four successfully repopulated recipients was collected for further blood cell count analysis (Fig. 5B, C). The asxl1−/− surviving recipients showed a significant decreased in the proportion of neutrophils (Fig. 5C). The neutrophilic dysplasia we observed in asxl1−/− recipients was similar to the phenotype of 6-month and 1-year asxl1 mutants (Fig. 3a and Supplementary Fig. 3A). A severe monocytosis phenotype (about 64% of kidney marrow cells were monocytes/macrophages) was observed in one asxl1−/− recipient, and the morphology of these increased macrophages was aberrant (Fig. 5B—5). This phenotype is comparable to the CMML-like phenotype. These results demonstrated that our asxl1 mutated cells intrinsically induced leukemogenesis and were transplantable.
A Schematic of the experimental procedure. Wild-type fish were irradiated at 24 Gy 2 days before transplantation. Whole kidney marrow cells were prepared from 1-year stage asxl1+/+ and asxl1−/− donors and were dilute to 105/μL. Each recipient was injected with 3 μL donor kidney marrow cells. Transplanted fish were raised in sterile water. After 18 days, surviving fish were collected for analysis (dpt, days posttransplantation). B May–Grünwald–Giemsa staining of whole kidney marrow of successful transplanted fish (asxl1+/+, n = 3; asxl1−/−, n = 4; scale bar, 20 μm; black boxes show enlarged details of neutrophils and macrophages/monocytes; green arrows, neutrophils; red arrow, macrophages). C Proportion of hematopoietic cells in whole kidney marrow of transplanted fish (asxl1+/+, n = 3; asxl1−/−, n = 4; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001, error bars, mean ± SD). Mye P myeloid progenitors, Mye mature myeloid cells, Ery erythrocytes, Lym lymphocytes, Neu neutrophils, Mac macrophages, Eos eosinophils (color figure online).
Transcriptome analysis shows disruption of neutrophil development
Our data showed that asxl1 mutations induced dysregulation of myeloid differentiation, mainly displayed as similarities to myeloid malignancies with neutrophilic dysplasia. To understand the basis for impaired neutrophil differentiation in zebrafish asxl1 mutants, we performed RNA-seq on 3 dpf asxl1 mutants and its littermate controls. Our data showed 294 differentially expressed genes (FDR < 0.05, 146 downregulated genes, 148 upregulated genes, Supplementary Table 3, analysis detailed in “Methods”). Similar to the observed phenotypes, expression of all neutrophil markers (lyz, mpx, npsn, srgn) was downregulated (Fig. 6A and Supplementary Fig. 5A). Expression of myeloid progenitor (pu.1 and cebpa) and macrophage (mfap4 and mpeg1) markers was not impaired (Supplementary Fig. 5B). GO analysis showed that wounding response-related genes were significantly enriched in asxl1 mutants, possibly because impaired neutrophils influenced the response to wounding (Fig. 6B). We also found that genes for an inflammatory cytokine (cxcl18b) and a matrix metalloproteinase (mmp13a) were upregulated in asxl1 mutants (Fig. 6A), which may be from induction by the deficiency of neutrophil granzymes [32, 33]. Our transcriptome analysis mainly showed disruption of neutrophil development, also suggesting that neutrophil deficiency may stimulate expression of some inflammatory cytokines and enhance the inflammatory response.
A Heat map of differentially expressed genes (false discovery rate < 0.05; exact test) in 3 dpf asxl1−/− compared to asxl1+/+ via RNA-seq. Neutrophil markers (green), PRC1 members (orange), and inflammatory response associated genes (purple) were highlighted. Rep replicate. B Gene ontology (GO) enrichment analysis of biological processes based on differentially expressed genes identified through RNA-seq (hypergeometric test). C Expression of bmi1a and cbx4 from RNA-seq (n = 20 per replicate, CPM count-per-million; exact test, *false discovery rate (FDR) < 0.05, **FDR < 0.01, ***FDR < 0.001; error bars, mean ± SD). qRT-PCR comparing expression of bmi1a and cbx4 in asxl1+/+ and asxl1−/− in 3 dpf larvae (n ≥ 10 per group, performed with four replicates; two-tailed Student’s t test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD) (color figure online).
Expression of bmi1a and cbx4 is disrupted in asxl1 mutants
ASXL1 is thought to regulate H2AK119ub, which is ubiquitinated by PRC1 [34, 35]. ASXL1/2 mainly serve as cofactors for the H2AK119ub deubiquitinase BAP1 [16, 17, 36, 37], and overexpression of truncated ASXL1 reduces H2AK119ub [10]. From our RNA-seq data, the expression of bmi1a and cbx4, which encode members of PRC1, was downregulated in asxl1 mutants (Fig. 6A, C). By qRT-PCR, we confirmed the reduced expression of bmi1a and cbx4 in asxl1 mutants (Fig. 6C). Our results indicated that the Asxl1 mutant may have disrupted expression of PRC1 complex components.
Treatment with bmi1a and cbx4 inhibitors impairs neutrophil development
Studies showed that bmi1a is associated with the proliferative activity of granulocyte/macrophage progenitor (GMP) [38] and is a biomarker for hematologic malignancies [39]. This finding indicates that bmi1a may participate in myelopoiesis. We investigated if disruption of myelopoiesis in our asxl1 mutant was associated with disruption of bmi1a and cbx4 expression by testing if their inhibitors can phenocopy the asxl1 mutation-caused neutrophilic dysplasia phenotype. In addition to H2AK119ub, H3K4me3 and H3K27me3 are also reported to be associated with ASXL1 [40,41,42]. Therefore, inhibitors targeting these demethylases were also employed. After testing for tolerance of the chemotherapeutics, the maximum doses were used: CPI455 (KDM5 H3K4me demethylase inhibitor, 40 μM), UNC3866 (CBX4/CBX7 inhibitor, 50 μM), PRT4165 (BMI1/RING1A inhibitor, 10 μM), and GSK J4 (JMJD3/UTX H3K27me demethylase inhibitor, 2 μM) (Supplementary Fig. 6A). After treatment of embryos with CBX4/CBX7 inhibitor or BMI1/RING1A inhibitor, lyz+ neutrophils decreased in asxl1+/+ compared with DMSO-treated controls (Fig. 7A). This result suggested that inhibition of Cbx4/Bmi1a phenocopied asxl1−/−. The KDM5 H3K4me demethylase inhibitor did not change the number of lyz+ cells in asxl1-mutant embryos (Supplementary Fig. 6B). The JMJD3/UTX H3K27me demethylase inhibitor increased the number of lyz+ cells in asxl1+/+ but not asxl1−/− (Supplementary Fig. 6C). This result indicated that induction of lyz+ cells after H3K27me demethylase JMJD3/UTX inhibitor treatment in wild-type embryos may involve asxl1. These results are consistent with the hypothesis that a reduction in expression of PRC1 members bmi1a and cbx4 may be associated with disruption of neutrophils in asxl1−/− embryos.
A WISH for lyz (3 dpf) after UNC3866 treatment showing lyz+ cells decreased after 50 μM UNC3866 treatment. Quantification of lyz+ cells in asxl1+/+ and asxl1−/− tails (scale bar, 200 μm; black boxes show enlarged images; asxl1+/+, n = 16, asxl1−/−, n = 18, asxl1+/+ T, n = 20, asxl1−/− T, n = 15; one-way ANOVA followed by LSD Fisher’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD). T treated, ns not significant. WISH of lyz (3 dpf) after 10 μM PRT4165 treatment showing lyz+ cells decreased after PRT4165 treatment. Quantification of lyz+ cells in asxl1+/+ and asxl1−/− tails (scale bar, 200 μm; black boxes show enlarged images; asxl1+/+, n = 12, asxl1−/−, n = 15, asxl1+/+ T, n = 10, asxl1−/− T, n = 14; one-way ANOVA followed by LSD Fisher’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD). T treated, ns not significant. B WISH for lyz (3 dpf) after bmi1a mRNA injection showing reduced lyz expression was rescued by bmi1a mRNA injection. Quantification of lyz+ cells in asxl1+/+ and asxl1−/− tails (about 0.1 ng mRNA/embryo; scale bar, 200 μm; black boxes show enlarged images; control: asxl1+/+, n = 12, asxl1−/−, n = 18; bmi1a: asxl1+/+, n = 18, asxl1−/−, n = 10; one-way ANOVA followed by LSD Fisher’s post hoc test, *p < 0.05, **p < 0.01, ***p < 0.001; error bars, mean ± SD; T treated, ns not significant). C Western blot of H2AK119ub, H3K4me3 and H3K27me3 in 2 dpf asxl1+/+ and asxl1−/− embryos and quantification of Western blot data (input embryos: about ten embryos/well, performed with three replicates; two-tailed Student’s t test, ns not significant; error bars, mean ± SD, internal control, H3).
Impaired neutrophil development from asxl1 mutations is rescued by bmi1a mRNA
To determine if impaired neutrophil differentiation was associated with decreased bmi1a/cbx4 expression, we overexpressed bmi1a and cbx4 in asxl1 mutants by injecting bmi1a/cbx4-3xFLAG-P2A-DsRed mRNA (Supplementary Fig. 7A). Fluorescence indicated bmi1a/cbx4 expression: DsRed was observed in embryos injected with bmi1a/cbx4-3xFLAG-P2A-DsRed mRNA compared to controls (Supplementary Fig. 7B). Expression of both lyz and SB+ neutrophils was significantly increased in asxl1 mutants after injecting bmi1a mRNA (Fig. 7B and Supplementary Fig. 7C). Injection with cbx4 mRNA led to almost complete embryonic lethality at 3 dpf and embryos exhibited an expanded heart field (Supplementary Fig. 7B). Our data showed that expression of bmi1a can rescue the asxl1 mutation-induced phenotype and indicated that decreased bmi1a expression may be a mechanism by which neutrophils were disrupted in asxl1−/− embryos.
BMI1 is reported to be required for the formation of PRC1 and for RING1A/1B ubiquitin ligase activity [43]. To check if the reduction in bmi1a also influenced H2AK119ub, we examined the global H2A ubiquitination state in asxl1-mutant zebrafish embryos. Western blots showed decreased global H2AK119ub (Fig. 7C). Global H3K4me3 and H3K27me3, which are reported to be associated with ASXL1 [8, 41, 42], were also detected. H3K4me3 was decreased in zebrafish asxl1 mutants, whereas global H3K27me3 did not obviously change. Our data showed that an endogenous asxl1-truncated mutant mainly disrupted H2AK119ub and this disruption may be associated with reduced expression of genes encoding PRC1 members, bmi1a and cbx4.
Discussion
How might ASXL1 facilitate myeloid malignancies?
Patients with multiple myeloid malignancies frequently harbor an ASXL1 mutation. Existing disease models mainly have total loss of ASXL1 [7, 9] or overexpression of a C-terminal truncation [8, 10, 11]. Loss of ASXL1 in mice is described to exhibit multilineage dysplasia [7, 9]. A zebrafish KO model is described to have reduced HSCs at 3 dpf [12]. In addition, C-terminally truncated ASXL1 can be detected in leukemia cells with an ASXL1 mutation [42] and expression of ASXL1 with a C-terminal truncation may confer a gain-of-function by promoting BAP1 activity [18]. These results raised the possibility that a ASXL1 mutation might confer a gain-of-function effect. Therefore, mouse models overexpressing C-terminally truncated ASXL1 were generated and showed dysregulation of multilineage differentiation [10, 11]. However, an overexpressing model may not fully recapitulate ASXL1 function because of protein overdose. Therefore, endogenous heterozygous Asxl1G643fs mutant knock-in mice were generated and had no obvious hematopoiesis disorders within 18 months, but showed MDS/MPN-like disease by 18–24 months [13, 14].
In contrast to mouse models, zebrafish have external fertilization and embryonic development, allowing us to observe phenotypes starting from the embryonic stage. Our zebrafish with endogenous truncated asxl1 mutants captured the most frequent C-terminal frameshift mutations in patients. This model had specifically impaired neutrophil differentiation without effects on other lineages including erythrocytes, macrophages, and lymphocytes. Furthermore, transcriptome analysis showed that zebrafish asxl1 mutants exhibited predominantly dysregulation of neutrophil-specific genes. Our work complements a recent study employing single-cell RNA-seq that suggests the involvement of ASXL1 in neutrophil differentiation [44].
Our data showed that C-terminally truncated asxl1 disrupted only neutrophil differentiation. The observed mild phenotype may be more comparable to clinical features since ASXL1 mutations frequently coexist with other mutations [45]. This phenomenon complements previous studies showing that Asxl1 mutation alone is not sufficient for leukemogenesis [13]. Moreover, we found that neutrophil disruption happened as early as the embryonic stage and persisted throughout the lifetime. At 6 months, mutants with Asxl1 truncations in our study mainly displayed MDS with neutrophilic dysplasia, which presented as refractory neutropenia. At 1 year, in addition to neutrophilic dysplasia, about 13% of mutants acquired other phenotypes of monocytosis that mimic CMML or increased myeloblasts similar to AML (Fig. 4). This result suggests a fundamental role for neutrophil disruption in myeloid malignancies progression.
However, further progression to CMML-like or AML-like phenotypes was observed only in some mutants, suggesting that asxl1 mutations alone are sufficient for MDS-SLD but not for severe myeloid malignancies, with other events responsible for progression. Clonal evolution and the alteration of the hematopoietic niche might contribute to disease progression [1]. In addition, inflammation was recently reported to contribute to clonal hematopoiesis of indeterminate potential [46]. Chronic inflammation may lead to HSC exhaustion and aging, which may promote mutation [47,48,49,50]. Our transcriptome analysis showed a possible inflammatory response in our mutants. Mutant asxl1-induced neutrophilic dysplasia may induce persistent inflammation and increase the risk of myeloid malignancy progression. Further studies need to be done to investigate the mechanism of progression, and our model could provide an opportunity to investigate additional contributors to myeloid malignancy progression.
How might bmi1a be an intermediate in neutrophil disruption?
Our data indicated that disruption of neutrophil development has a fundamental role in myeloid malignancy progression in asxl1 mutants. We explored the potential mechanism. Previous studies show that an ASXL1G643fs mutant interrupts its interaction with BMI1, leading to an MDS-like phenotype [14]. However, in our model, we observed reduced expression of bmi1a. Hence, ASXL1 mutants may not only block interaction with BMI1 but also disrupt its expression. Absence of Bmi1 affects the proliferative activity of GMP, and Bmi1−/− GMP has dispersed macrophage colonies [38]. This result indicates that Bmi1 may participate in myeloid differentiation. Using a bmi1a inhibitor, we mimicked a neutrophil dysplasia phenotype (Fig. 7A). Supplementing with bmi1a mRNA rescued asxl1-induced neutrophil dysregulation (Fig. 7B). Hence, our work illustrated an alternative mechanism of Asxl1-Bmi1a axis on regulating neutrophil development.
The reduction in H2AK119ub is important in mutant ASXL1-induced leukemogenesis. Recent studies show that overexpression of C-terminally truncated ASXL1 (ASXL1-MT) and BAP1 leads to a hyperactive complex that subsequently upregulates IRF8, which is an essential transcription factor for myeloid lineage development [51,52,53], through H2AK119 deubiquitination. In addition, disrupted interaction between ASXL1G643fs and BMI1 perturbs BMI1-driven H2AK119ub, further promoting an MDS-like phenotype. The reduced expression of PRC1 members, including BMI1 and CBX4, further contributes to the mutant asxl1-induced dysregulation of H2AK119ub. However, we are not suggesting that reduction of H2AK119ub blocks neutrophil differentiation. This H2AK119ub change could be a consequence of the reduced expression of bmi1a and cbx4. In any case, our data show a novel way for ASXL1 mutations to reduce H2AK119ub and raise the possibility that the Asxl1-Bmi1a axis is involved in neutrophil differentiation.
Taken together, results from this study show that endogenous truncated asxl1 mutations result in impaired neutrophil differentiation. Our model revealed that neutrophilic dysplasia caused by asxl1 mutations is a foundation for the progression of myeloid malignancies, and bmi1a dysregulation is associated with neutrophil dysplasia (Fig. 8). Zebrafish are an ideal model for large-scale chemical screening. Our endogenous truncated asxl1 zebrafish model provides a new opportunity for a high-throughput drug-screening platform using whole organisms.
In our model, C-terminally truncated with asxl1 mutations specifically blocked neutrophil differentiation. The mutants initially exhibited an MDS-like phenotype with neutrophilic dysplasia, and some mutants progressed to more severe phenotypes, similar to CMML/AML disease. Dysregulation of bmi1a may be a reason for leukemogenesis in asxl1 mutants. Asxl1 wild-type Asxl1 protein, tAsxl1 truncated Asxl1 protein.
Data availability
RNA-seq data are available at GEO under accession number GSE142214. For original data, please contact Xiao Fang: fangxiao@scut.edu.cn.
References
Korn C, Méndez-Ferrer S. Myeloid malignancies and the microenvironment. Blood. 2017;129:811–22.
Murati A, Brecqueville M, Devillier R, Mozziconacci M-J, Gelsi-Boyer V, Birnbaum D. Myeloid malignancies: mutations, models and management. BMC Cancer. 2012;12:304.
Arber DA, Orazi A, Hasserjian R, Thiele J, Borowitz MJ, Le Beau MM, et al. The 2016 revision to the World Health Organization classification of myeloid neoplasms and acute leukemia. Blood. 2016;127:2391–405.
Gelsi-Boyer V, Brecqueville M, Devillier R, Murati A, Mozziconacci MJ, Birnbaum D. Mutations in ASXL1 are associated with poor prognosis across the spectrum of malignant myeloid diseases. J Hematol Oncol. 2012;5:12.
Gelsi-Boyer V, Trouplin V, Roquain J, Adelaide J, Carbuccia N, Esterni B, et al. ASXL1 mutation is associated with poor prognosis and acute transformation in chronic myelomonocytic leukaemia. Br J Haematol. 2010;151:365–75.
Itzykson R, Kosmider O, Renneville A, Gelsi-Boyer V, Meggendorfer M, Morabito M, et al. Prognostic score including gene mutations in chronic myelomonocytic leukemia. J Clin Oncol. 2013;31:2428–36.
Abdel-Wahab O, Gao J, Adli M, Dey A, Trimarchi T, Chung YR, et al. Deletion of Asxl1 results in myelodysplasia and severe developmental defects in vivo. J Exp Med. 2013;210:2641–59.
Inoue D, Kitaura J, Togami K, Nishimura K, Enomoto Y, Uchida T, et al. Myelodysplastic syndromes are induced by histone methylation-altering ASXL1 mutations. J Clin Investig. 2013;123:4627–40.
Wang J, Li Z, He Y, Pan F, Chen S, Rhodes S, et al. Loss of Asxl1 leads to myelodysplastic syndrome-like disease in mice. Blood. 2014;123:541–53.
Nagase R, Inoue D, Pastore A, Fujino T, Hou HA, Yamasaki N, et al. Expression of mutant Asxl1 perturbs hematopoiesis and promotes susceptibility to leukemic transformation. J Exp Med. 2018;215:1729–47.
Yang H, Kurtenbach S, Guo Y, Lohse I, Durante MA, Li J, et al. Gain of function of ASXL1 truncating protein in the pathogenesis of myeloid malignancies. Blood. 2018;131:328–41.
Gjini E, Jing CB, Nguyen AT, Reyon D, Gans E, Kesarsing M, et al. Disruption of asxl1 results in myeloproliferative neoplasms in zebrafish. Dis Models Mech. 2019;12.
Hsu YC, Chiu YC, Lin CC, Kuo YY, Hou HA, Tzeng YS, et al. The distinct biological implications of Asxl1 mutation and its roles in leukemogenesis revealed by a knock-in mouse model. J Hematol Oncol. 2017;10:139.
Uni M, Masamoto Y, Sato T, Kamikubo Y, Arai S, Hara E, et al. Modeling ASXL1 mutation revealed impaired hematopoiesis caused by derepression of p16Ink4a through aberrant PRC1-mediated histone modification. Leukemia. 2019;33:191–204.
Dey A, Seshasayee D, Noubade R, French DM, Liu J, Chaurushiya MS, et al. Loss of the tumor suppressor BAP1 causes myeloid transformation. Science. 2012;337:1541–6.
Sahtoe DD, van Dijk WJ, Ekkebus R, Ovaa H, Sixma TK. BAP1/ASXL1 recruitment and activation for H2A deubiquitination. Nat Commun. 2016;7:10292.
Balasubramani A, Larjo A, Bassein JA, Chang X, Hastie RB, Togher SM, et al. Cancer-associated ASXL1 mutations may act as gain-of-function mutations of the ASXL1-BAP1 complex. Nat Commun. 2015;6:7307.
Asada S, Goyama S, Inoue D, Shikata S, Takeda R, Fukushima T, et al. Mutant ASXL1 cooperates with BAP1 to promote myeloid leukaemogenesis. Nat Commun. 2018;9:2733.
Simon JA, Kingston RE. Mechanisms of polycomb gene silencing: knowns and unknowns. Nat Rev Mol Cell Biol. 2009;10:697–708.
Konantz M, Schurch C, Hanns P, Muller JS, Sauteur L, Lengerke C. Modeling hematopoietic disorders in zebrafish. Dis Models Mech. 2019;12.
Westerfield M. The zebrafish book: a guide for the laboratory use of zebrafish (Danio Rerio). Eugene, University of Oregon Press, 2007.
Kimmel CB, Ballard WW, Kimmel SR, Ullmann B, Schilling TF. Stages of embryonic development of the zebrafish. Dev Dyn. 1995;203:253–310.
Andrews S. FastQC: a quality control tool for high throughput sequence data. http://www.bioinformatics.babraham.ac.uk/projects/fastqc. 2010.
Dobin A, Davis CA, Schlesinger F, Drenkow J, Zaleski C, Jha S, et al. STAR: ultrafast universal RNA-seq aligner. Bioinformatics. 2013;29:15–21.
Liao Y, Smyth GK, Shi W. featureCounts: an efficient general purpose program for assigning sequence reads to genomic features. Bioinformatics. 2014;30:923–30.
Robinson MD, Smyth GK. Moderated statistical tests for assessing differences in tag abundance. Bioinformatics. 2007;23:2881–7.
Robinson MD, McCarthy DJ, Smyth GK. edgeR: a Bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics. 2010;26:139–40.
Zhou Y, Zhou B, Pache L, Chang M, Khodabakhshi AH, Tanaseichuk O, et al. Metascape provides a biologist-oriented resource for the analysis of systems-level datasets. Nat Commun. 2019;10:1523.
Jin H, Li L, Xu J, Zhen F, Zhu L, Liu PP, et al. Runx1 regulates embryonic myeloid fate choice in zebrafish through a negative feedback loop inhibiting Pu.1 expression. Blood. 2012;119:5239–49.
Döhner H, Estey E, Grimwade D, Amadori S, Appelbaum FR, Büchner T, et al. Diagnosis and management of AML in adults: 2017 ELN recommendations from an international expert panel. Blood. 2017;129:424–47.
Kazankov K, Jørgensen SMD, Thomsen KL, Møller HJ, Vilstrup H, George J, et al. The role of macrophages in nonalcoholic fatty liver disease and nonalcoholic steatohepatitis. Nat Rev Gastroenterol Hepatol. 2018;16:145–59.
Wang K, Fang X, Ma N, Lin Q, Huang Z, Liu W, et al. Myeloperoxidase-deficient zebrafish show an augmented inflammatory response to challenge with Candida albicans. Fish Shellfish Immunol. 2015;44:109–16.
Di Q, Lin Q, Huang Z, Chi Y, Chen X, Zhang W, et al. Zebrafish nephrosin helps host defence against Escherichia coli infection. Open Biol. 2017;7:170040.
Wang H, Wang L, Erdjument-Bromage H, Vidal M, Tempst P, Jones RS, et al. Role of histone H2A ubiquitination in Polycomb silencing. Nature. 2004;431:873–8.
Piunti A, Shilatifard A. Epigenetic balance of gene expression by Polycomb and COMPASS families. Science. 2016;352:aad9780.
Scheuermann JC, de Ayala Alonso AG, Oktaba K, Ly-Hartig N, McGinty RK, Fraterman S, et al. Histone H2A deubiquitinase activity of the Polycomb repressive complex PR-DUB. Nature. 2010;465:243–7.
Daou S, Barbour H, Ahmed O, Masclef L, Baril C, Sen Nkwe N, et al. Monoubiquitination of ASXLs controls the deubiquitinase activity of the tumor suppressor BAP1. Nat Commun. 2018;9:4385.
Yuan J, Takeuchi M, Negishi M, Oguro H, Ichikawa H, Iwama A. Bmi1 is essential for leukemic reprogramming of myeloid progenitor cells. Leukemia. 2011;25:1335–43.
Sahasrabuddhe AA. BMI1: a biomarker of hematologic malignancies. Biomark Cancer. 2016;8:65–75.
Wang J, Li Z, He Y, Pan F, Chen S, Rhodes S, et al. Loss of Asxl1 leads to myelodysplastic syndrome–like disease in mice. Blood. 2014;123:541–53.
Abdel-Wahab O, Adli M, LaFave LM, Gao J, Hricik T, Shih AH, et al. ASXL1 mutations promote myeloid transformation through loss of PRC2-mediated gene repression. Cancer Cell. 2012;22:180–93.
Inoue D, Fujino T, Sheridan P, Zhang YZ, Nagase R, Horikawa S, et al. A novel ASXL1-OGT axis plays roles in H3K4 methylation and tumor suppression in myeloid malignancies. Leukemia. 2018;32:1327–37.
Cao R, Tsukada Y, Zhang Y. Role of Bmi-1 and Ring1A in H2A ubiquitylation and Hox gene silencing. Mol Cell. 2005;20:845–54.
Braun T, Lusardi T, Enright T, Schonrock Z, Coblentz C, Mohamed H, et al. Single cell RNA sequencing identifies a crucial role for ASXL1 in neutrophil development. Blood. 2019;134:212.
Asada S, Fujino T, Goyama S, Kitamura T. The role of ASXL1 in hematopoiesis and myeloid malignancies. Cell Mol life Sci. 2019;76:2511–23.
Craver BM, El Alaoui K, Scherber RM, Fleischman AG. The critical role of inflammation in the pathogenesis and progression of myeloid malignancies. Cancers. 2018;10.
Esplin BL, Shimazu T, Welner RS, Garrett KP, Nie L, Zhang Q, et al. Chronic exposure to a TLR ligand injures hematopoietic stem cells. J Immunol. 2011;186:5367–75.
Zhao Y, Ling F, Wang HC, Sun XH. Chronic TLR signaling impairs the long-term repopulating potential of hematopoietic stem cells of wild type but not Id1 deficient mice. PLoS ONE. 2013;8:e55552.
Matatall KA, Jeong M, Chen S, Sun D, Chen F, Mo Q, et al. Chronic infection depletes hematopoietic stem cells through stress-induced terminal differentiation. Cell Rep. 2016;17:2584–95.
Takizawa H, Fritsch K, Kovtonyuk LV, Saito Y, Yakkala C, Jacobs K, et al. Pathogen-Induced TLR4-TRIF innate immune signaling in hematopoietic stem cells promotes proliferation but reduces competitive fitness. Cell Stem Cell. 2017;21:225–40.e225.
Li L, Jin H, Xu J, Shi Y, Wen Z. Irf8 regulates macrophage versus neutrophil fate during zebrafish primitive myelopoiesis. Blood. 2011;117:1359–69.
Kurotaki D, Yamamoto M, Nishiyama A, Uno K, Ban T, Ichino M, et al. IRF8 inhibits C/EBPalpha activity to restrain mononuclear phagocyte progenitors from differentiating into neutrophils. Nat Commun. 2014;5:4978.
Shiau CE, Kaufman Z, Meireles AM, Talbot WS. Differential requirement for irf8 in formation of embryonic and adult macrophages in zebrafish. PLoS ONE. 2015;10:e0117513.
Acknowledgements
This work was supported by the National Key Research and Development Program of China (2018YFA0801003; 2018YFA0800200) and the National Natural Science Foundation of China (31522031; 31571526; 81770167; 31872843).
Author information
Authors and Affiliations
Contributions
XF, WZ, and KY designed the research; XF generated asxl1 mutants and identified their phenotype; XF and SX performed cell sorting, cell staining, and transplantation assay; XF performed RNA-seq library preparation and analysis; XF generated constructs and performed rescue assays (asxl1 and bmi1a) and drug treatment. XF analyzed results, made figures, and wrote the paper. WZ, KY, YZ, JX, ZH, and WL discussed the results and commented on the paper.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Fang, X., Xu, S., Zhang, Y. et al. Asxl1 C-terminal mutation perturbs neutrophil differentiation in zebrafish. Leukemia 35, 2299–2310 (2021). https://doi.org/10.1038/s41375-021-01121-8
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41375-021-01121-8
- Springer Nature Limited
This article is cited by
-
Atypical chronic myeloid leukemia found in a patient with eosinophilia for six years: a case report
BMC Geriatrics (2024)
-
Transgenic IDH2R172K and IDH2R140Q zebrafish models recapitulated features of human acute myeloid leukemia
Oncogene (2023)
-
ASXL1/2 mutations and myeloid malignancies
Journal of Hematology & Oncology (2022)