Abstract
Forkhead (Fkh/Fox) domain transcription factors (TFs) mediate multiple cardiogenic processes in both mammals and Drosophila. We showed previously that the Drosophila Fox gene jumeau (jumu) controls three categories of cardiac progenitor cell division—asymmetric, symmetric, and cell division at an earlier stage—by regulating Polo kinase activity, and mediates the latter two categories in concert with the TF Myb. Those observations raised the question of whether other jumu-regulated genes also mediate all three categories of cardiac progenitor cell division or a subset thereof. By comparing microarray-based expression profiles of wild-type and jumu loss-of-function mesodermal cells, we identified nebbish (neb), a kinesin-encoding gene activated by jumu. Phenotypic analysis shows that neb is required for only two categories of jumu-regulated cardiac progenitor cell division: symmetric and cell division at an earlier stage. Synergistic genetic interactions between neb, jumu, Myb, and polo and the rescue of jumu mutations by ectopic cardiac mesoderm-specific expression of neb demonstrate that neb is an integral component of a jumu-regulated subnetwork mediating cardiac progenitor cell divisions. Our results emphasize the central role of Fox TFs in cardiogenesis and illustrate how a single TF can utilize different combinations of other regulators and downstream effectors to control distinct developmental processes.
Similar content being viewed by others
Introduction
While at least eight Forkhead (Fkh/Fox) domain transcription factors (TFs) are required for proper cardiac development in mammals1,2,3,4,5,6,7,8,9,10,11,12,13,14,15,16, and mutations in four Fox genes have been linked to human congenital heart defects17,18,19,20,21,22,23,24, relatively little is known about the molecular mechanisms or the downstream target genes by which these Fox TF-mediated developmental functions are brought about25. Our previous work had identified the conserved cardiogenic roles of two Fox genes, jumeau (jumu) and Checkpoint suppressor homologue (CHES-1-like) in both the specification and subsequent division of progenitor cells in the Drosophila heart26,27,28.
We showed previously that both jumu and CHES-1-like function in a mutually redundant manner to transcriptionally activate the expression of heartless (htl) and frizzled (fz), which respectively encode the receptors in a FGF-signaling pathway and a Wnt-signaling pathway required for cardiac progenitor specification; consequently, lacking both Fox genes results in a failure of much of the heart to be specified26. We also demonstrated that jumu and CHES-1-like mediate three distinct categories of cell division that generate the correct subtypes and numbers of cells constituting the heart—asymmetric cardiac progenitor cell divisions, symmetric cardiac progenitor cell divisions, and cell divisions at an earlier stage that produced the cardiac precursor cells—by regulating the activity of the conserved kinase Polo28. In addition, our investigations revealed that the TF Myb acts synergistically with Jumu, CHES-1-like, and Polo to mediate only two of these three categories of cardiac progenitor cell division: symmetric and cell division at an earlier stage27. In contrast, asymmetric cardiac progenitor cell division involved Jumu and CHES-1-like utilizing Polo to phosphorylate Partner of numb (Pon) and thereby bring about the localization of membrane-associated Numb protein to one pole of the dividing cell28,29.
Additionally, we found that Fox TF binding sites were significantly enriched in the enhancers of genes expressed in the heart27,30, and that loss-of-function mutations in jumu or CHES-1-like significantly altered the expression levels of many known cardiac genes26. Taken together, these data indicate that the Fox TFs regulate a large number of genes in addition to polo, htl, and fz in order to mediate cardiogenesis.
Collectively, these observations raise three intriguing developmental questions. First, are additional Fox TF-regulated genes besides polo, htl, and fz involved in the two distinct cardiogenic processes—cardiac progenitor specification and cardiac progenitor cell division—that we have discovered as being mediated by the Fox TFs to date?
Second, jumu and CHES-1-like were both seen to be simultaneously regulating the activity of each relevant downstream target gene (htl, fz, or polo) whenever any one of these target genes was characterized as mediating a cardiogenic process. Thus, a particularly germane question is whether there are any downstream cardiogenic targets and, by extension, associated pathways or subnetworks, that are exclusively regulated and mediated by one, but not the other, of these two Fox TFs.
Third, while Myb itself acts in concert with both Fox genes at the same hierarchical level to mediate only two of the three categories of cardiac progenitor cell divisions, polo, the downstream target of both jumu and CHES-1-like, is utilized for all three. Thus, another pertinent question is whether every Fox TF-regulated gene utilized in cardiac progenitor cell division also plays critical roles in all three categories of cell division or is necessary for only certain specific subsets.
Here we show that the kinesin-encoding gene nebbish (neb) is activated by only one Fox gene, jumu; that neb is required in conjunction with polo and Myb to mediate cardiac progenitor cell divisions; and that neb is necessary for only two of the three categories of cardiac progenitor cell division—symmetric and cell division at an earlier stage—thereby extending our understanding of the Fox-regulated subnetworks mediating cardiogenesis.
Results
jumu, but not CHES-1-like, activates mesodermal expression of neb
One of our goals was to discover downstream genes mediating cardiogenic processes, if present, that were regulated by one, but not the other, of the twin Fox TFs Jumu and CHES-1-like. We had previously used flow cytometry and Affymetrix microarrays to obtain genome-wide transcription expression profiles of purified mesodermal cells from wild-type embryos, embryos homozygous for a null mutation in jumu, and embryos homozygous for a null mutation in CHES-1-like at embryonic stages 11 and early 12, i.e. when cardiac progenitor specification and cell division occur in Drosophila26. Reasoning that cardiogenic genes regulated by a Fox TF are likely to exhibit differential expression between the heart-producing mesoderm in wild-type and Fox loss-of-function mutant embryos, we compared the transcription expression profiles between mesodermal cells from wild-type embryos and the Fox gene mutants. We found that neb had its expression levels significantly downregulated in jumu null mutants [log2(fold change) = − 0.69058; adjusted P-value = 1.65 × 10−5 after correction for multiple hypothesis testing] but remained essentially unaffected in CHES-1-like null mutants [log2(fold change) = 0.14551; adjusted P-value = 0.195].
In order to independently verify the effects of these two Fox genes on neb expression, we used the pan-mesodermal twi-GAL4 driver to knock down either jumu or CHES-1-like via RNA interference (RNAi) exclusively in the mesoderm. We then assessed and compared the resulting neb transcript levels at stages 11–12 with that in control embryos by reverse transcription quantitative real-time PCR (RT-qPCR). Since jumu and CHES-1-like are also expressed in tissues other than the mesoderm31, null mutations in the Fox genes could potentially also affect neb expression levels in these non-mesodermal tissues. Utilizing exclusively mesoderm-targeted RNAi knockdowns of these Fox genes instead of null mutations in jumu or CHES-1-like thus ensured that any detected alterations in neb transcript levels in total RNA were due solely to the effects of the Fox genes in the mesoderm. Our RT-qPCR assays revealed a decrease in neb expression levels in mesoderm-targeted jumu knockdowns compared to that in control embryos containing only the twi-GAL4 driver, but detected no such reduction in mesoderm-targeted CHES-1-like knockdowns (Fig. 1).
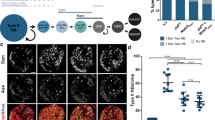
jumu, but not CHES-1-like, activates mesodermal expression of neb. Relative jumu, CHES-1-like, and neb gene expression levels in jumu or CHES-1-like RNAi knockdown embryos assessed and compared to control embryos via RT-qPCR and the 2−ΔΔCT method. Error bars indicate standard deviations. Mesoderm-targeted jumu and CHES-1-like RNAi do indeed reduce jumu and CHES-1-like gene expression levels compared to controls, respectively, indicating that these RNAi constructs result in the relevant Fox gene knockdowns. However, while mesoderm-targeted jumu RNAi knockdown also decreases neb transcript expression levels relative to the control, no such reduction is detected in mesoderm-targeted CHES-1-like knockdowns.
Collectively, both our microarray-based expression profiling data and RT-qPCR-mediated assessment of Fox gene knockdowns indicated that jumu, but not CHES-1-like, regulates neb in the mesoderm by activating its expression.
neb is expressed in the cardiac mesoderm during cardiac progenitor specification and cell division
neb had previously been described as being expressed in the nervous tissue during mid-embryogenesis31. Using in situ hybridization, we found that the neb transcript is also expressed, albeit at lower levels, in the cardiac mesoderm and in subsets of the somatic mesoderm at embryonic stages 11 and 12, i.e. when the cardiac progenitor cells are specified and undergo cell division (Fig. 2a,b). However, the expression pattern of the transcript made it difficult to determine unequivocally whether the expression level of neb is lower specifically in the cardiac mesoderm of jumu loss-of-function mutants by in situ hybridization assays. Regardless, the observations that neb is both expressed in the cardiac mesoderm and is transcriptionally activated mesodermally by jumu, collectively raise the question of whether neb has a critical a role in cardiogenesis.
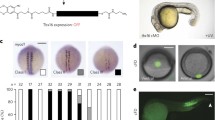
neb mRNA expression in the cardiac mesoderm. (a) Antisense riboprobe hybridizing to the mRNA shows that the neb transcript is expressed in the cardiac mesoderm in stage 11 wild-type embryos. (b) Control in situ hybridization with a sense riboprobe complementary to that used in (a) developed under identical conditions for the same period. The absence of similar staining in (b) indicates that the staining in (a) is neb-specific and not an artifact. (c) In situ hybridization of stage 11 embryos of the genotype TinD-GAL4/+; jumuDf(3R)Exel6157 svp-lacZ/jumuDf(3R)Exel6157 UAS-neb with the same antisense neb riboprobe as in (a) developed using identical conditions but for a much shorter period. The specific TinD-GAL4-targeted expression of neb to the cardiac mesoderm is clearly observed while the staining for endogenous neb expression is considerably fainter due to the shorter period used. The arrows indicate cells of the cardiac mesoderm in all three panels (a–c).
neb is required in the cardiac mesoderm for two distinct categories of cardiac progenitor cell divisions
The metamerically repeated and stereotyped pattern of distinct cardiac cell types in the embryonic Drosophila heart makes it particularly amenable to discovering whether neb has one or more crucial cardiogenic roles, and what those roles might be, by examining the phenotypes of neb loss-of-function mutations. The heart itself is comprised of two major cell types: an inner tube of Myocyte enhancer factor 2 (Mef-2)-expressing contractile cardial cells (CCs) and an outer sheath of Zn finger homeodomain 1 (zfh1)-expressing nephrocytic pericardial cells. From anterior to posterior, the cardial cells consist of two Seven up-expressing CCs (Svp-CCs) followed by four Tinman-expressing CCs (Tin-CCs) in every repeating hemisegment. These cardial cells are surrounded by a larger number of pericardial cells, which includes two Seven up-expressing PCs (Svp-PCs) that are located laterally to the CCs in each and every hemisegment. The different cellular subtypes comprising these cardial and pericardial cells arise through a stereotyped series of cardiac progenitor cell divisions32,33,34,35. Specifically, cell division at an earlier stage ensures that there are two Svp precursor cells per hemisegment. Subsequently, the two Svp-CCs in wild-type cardiac hemisegments arise by two asymmetric progenitor cell divisions from these two Svp precursor cells, with each cell division generating one Svp-CC and one Svp-PC (Fig. 3a, yellow and red cells, respectively). In contrast, two symmetric cell divisions give rise to the four Tin-CCs (Fig. 3a, green cells) per hemisegment from two other precursor cells.
Schematic showing cell lineage relationships in a wild-type Drosophila embryonic heart and the expected phenotypes due to defects in cardiac progenitor specification or cell division. (a) Lines connect daughter cells arising from the division of each precursor cell. (b) Defects in cardiac progenitor cell specification would be expected to result in the absence of all the cardial cells in one or more hemisegments. Defects in asymmetric cell division would result in an increase or reduction in the number of Svp-CCs accompanied by a corresponding decrease or increase in the number of Svp-PCs, or larger Svp-CC nuclei with missing corresponding Svp-PCs due to errors in karyokinesis. Defective symmetric cell division would result in an alteration in the expected number of four Tin-CCs per hemisegment, while errors at the earlier stage of cell division producing the two Svp precursors would result in hemisegments with either one or three pairs of Svp-CCs and Svp-PCs instead of the customary two pairs.
These lineage relationships allow us to use the numbers of the Tin-CCs, Svp-CCs, and Svp-PCs in individual hemisegments in neb loss-of-function mutants to determine whether neb is mediating one or more of these three known categories of cardiac progenitor cell divisions (Fig. 3b). If neb mutations cause asymmetric cell division defects, we would expect to see an increase or reduction in the number of Svp-CCs accompanied by a corresponding decrease or increase in the number of Svp-PCs, or larger Svp-CC nuclei with missing corresponding Svp-PCs due to errors in karyokinesis. Defective symmetric cell division, on the other hand, would be identified as any alteration in the expected number of four Tin-CCs per hemisegment, while errors at the earlier stage of cell division producing the two Svp precursors would result in hemisegments with either one or three pairs of Svp-CCs and Svp-PCs instead of the customary two pairs28. Our examination found that embryos homozygous for the nebk05702 loss-of-function allele exhibited a significant increase over wild-type both in the fraction of hemisegments with excess or fewer Tin-CCs (P = 2.00 × 10−6), corresponding to symmetric cell division defects, and in the fraction of hemisegments with either one or three pairs of Svp-CCs and Svp-PCs (P = 0.016), corresponding to earlier cell division defects affecting the number of Svp precursors (Fig. 4a–c, Supplementary Table S1). The fraction of hemisegments exhibiting asymmetric cell division defects, in contrast, was miniscule and not significantly different (P = 0.481) from that in wild-type embryos (Fig. 4a–c, Supplementary Table S1). Collectively, our results demonstrate that neb is necessary to mediate both symmetric cell divisions and the earlier round of cell divisions that give rise to the Svp precursors, but is not required for asymmetric cell divisions. Of note, we did not detect any instance where all the cardial cells in one or more entire hemisegments were missing, as would be expected in the case of cardiac progenitor specification defects, and were seen when htl or fz functions were disrupted26, 36,37,38.
Cardiac progenitor cell division defects associated with neb loss of function. (a) A heart from an otherwise wild-type embryo bearing one copy of the svp-lacZ enhancer trap showing hemisegments consisting of four Tin-CCs (green), two Svp-CCs (yellow), and two Svp-PCs (red). (b) A heart from an embryo homozygous for the nebk05702 null mutation (and also carrying one copy of the svp-lacZ enhancer trap) exhibiting both cardiac progenitor symmetric cell division defects (arrowheads) and defects at an earlier round of cell division specifying the number of Svp precursors (arrows). In these images (derived by flattening z-stacks), a few Svp-PCs are hidden underneath the CCs or appear quite faint in certain hemisegments, but all were clearly discernible in the individual planes of the z-stacks from which the images were obtained. (c) Percentage of hemisegments exhibiting each type of cardiac progenitor cell division defect in embryos that are wild-type (n = 196 hemisegments) or homozygous for the nebk05702 mutation (n = 186 hemisegments). The significance of each type of cell division defect in the neb mutants compared with wild-type is shown. (d) A wild-type heart stained to show only Tin-CCs (green) and Svp-CCs (yellow). Each hemisegment contains four Tin-CCs and two Svp-CCs. (e) Similar staining in embryos where neb has been knocked down specifically in the cardiac mesoderm exhibits changes in Tin-CC numbers and Svp-CC numbers consistent with symmetric (arrowheads) and earlier (arrows) cell division defects, respectively.
We next attempted to ascertain whether this neb function, necessary for mediating both symmetric cell divisions and divisions at the earlier stage affecting the number of Svp precursor cells, is required specifically in the cardiac progenitors. We thus knocked down neb activity specifically in the cardiac progenitors by cardiac mesoderm and heart-targeted RNA interference (RNAi) directed by the TinD-GAL439 and Hand-GAL440 drivers and examined the resulting embryos for alterations in expected cardial cell numbers consistent with errors in cardiac progenitor cell divisions. Staining with appropriate antibodies showed hemisegments with localized changes in the number of Tin-CCs corresponding to symmetric cell division defects and Svp-CCs corresponding to defective cell division at an earlier stage (Fig. 4d,e, Supplementary Fig. S1, Supplementary Table S2).
Taken together, these results demonstrate that the requirement of neb for mediating two distinct categories of cardiac progenitor cell division is autonomous to the cardiac mesoderm and raise the question of which cell division pathway or subnetwork neb is involved in, one that we address in the following experiments.
Synergistic genetic interactions between neb, jumu, polo, and Myb
We had previously shown that jumu and CHES-1-like regulate polo to mediate all three categories of cardiac progenitor cell divisions28; and that Myb functions synergistically with jumu, CHES-1-like, and polo to control both symmetric and earlier cardiac progenitor cell divisions27, the same two cardiogenic processes mediated by neb. These observations raise the questions of whether neb functions through the same genetic pathways as Myb, jumu, and polo, and, if it does, for which specific categories of cardiac progenitor cell divisions. Operating on the premise that we may expect to see synergistic, i.e. more than merely additive, genetic interactions between mutant alleles of neb and another gene if they both function together in the same genetic pathway, we quantified and compared the phenotypes of single heterozygotes of mutations in neb and single heterozygotes of mutations in jumu, polo, or Myb, with those of embryos that were doubly heterozygous for mutations in both neb and jumu, neb and polo, and neb and Myb, respectively.
Double heterozygotes for both neb and jumu mutations exhibit symmetric cell division defects and earlier Svp precursor-determining cell division defects at frequencies that are significantly more severe (P = 1.80 × 10−5 and P = 8.23 × 10−4, respectively) than the additive effects of both the neb single heterozygotes and the jumu single heterozygotes (Figs. 5a–c, 6a, Supplementary Table S1). However, the increase in the frequency of asymmetric cell division defects in nebMI0225/+; jumuDf(3R)Exel6157/+double heterozygotes over the additive sum of those in the component nebMI0225/+and jumuDf(3R)Exel6157/+single heterozygotes is barely significant (P = 0.0429) (Figs. 5a–c, 6a, Supplementary Table S1). Our results thus indicate clearly that jumu and neb do indeed mediate both symmetric and earlier Svp precursor-determining cardiac progenitor cell divisions through the same genetic pathways, but the evidence for neb functioning in concert with jumu to bring about asymmetric cardiac progenitor cell divisions as well is not as strong. Note that these genetic interaction results are also consistent with our initial finding of neb being required for symmetric and earlier, but not asymmetric, cardiac progenitor cell divisions.
neb exhibits synergistic genetic interactions with jumu, polo, and Myb, but not with CHES-1-like. Representative hearts from embryos (a) heterozygous for the nebMI02225 allele, (b) heterozygous for the jumuDf(3R)Exel6157 allele, (c) doubly heterozygous for both the nebMI02225 and jumuDf(3R)Exel6157 alleles, (d) heterozygous for the polo10 allele, (e) doubly heterozygous for both the nebMI02225 and polo10 alleles, (F) heterozygous for the MybMH107 allele, (g) doubly heterozygous for both the nebMI02225 and MybMH107 alleles, (h) heterozygous for the CHES-1-like1 allele, and (i) doubly heterozygous for both the nebMI02225 and CHES-1-like1 alleles. All of these embryos carry one copy of the svp-lacZ enhancer trap, thereby allowing the identification of Tin-CCs (green), Svp-CCs (yellow), and Svp-PCs (red). In these images (derived by flattening z-stacks), a few Svp-PCs are hidden underneath the CCs or appear quite faint in certain hemisegments, but all were clearly discernible in the individual planes of the z-stacks from which the images were obtained. Asymmetric cell division defects are denoted by asterisks, symmetric cell division defects by arrowheads, and earlier Svp precursor-determining cell division defects by arrows.
Quantification and significance of cardiac progenitor cell division defects in genetic interaction assays. (a) Percentage of hemisegments exhibiting each type of cardiac progenitor cell division defect in embryos that are heterozygous for the nebMI02225 allele (n = 188 hemisegments), heterozygous for the jumuDf(3R)Exel6157 allele (n = 210 hemisegments), or doubly heterozygous for both the nebMI02225 and jumuDf(3R)Exel6157 alleles (n = 238 hemisegments). (b) Percentage of hemisegments exhibiting each type of cardiac progenitor cell division defect in embryos that are heterozygous for the nebMI02225 allele, heterozygous for the polo10 allele (n = 196 hemisegments), or doubly heterozygous for both the nebMI02225 and polo10 alleles (n = 210 hemisegments). (c) Percentage of hemisegments exhibiting each type of cardiac progenitor cell division defect in embryos that are heterozygous for the nebMI02225 allele, heterozygous for the MybMH107 allele (n = 193 hemisegments), or doubly heterozygous for both the nebMI02225 and MybMH107 alleles (n = 179 hemisegments). (d) Percentage of hemisegments exhibiting each type of cardiac progenitor cell division defect in embryos that are heterozygous for the nebMI02225 allele, heterozygous for the CHES-1-like1 allele (n = 210 hemisegments), or doubly heterozygous for both the nebMI02225 and CHES-1-like1 alleles (n = 192 hemisegments).
Since jumu brings about all three categories of cardiac progenitor cell divisions by regulating the activity of Polo kinase28, we used similar genetic interaction assays next to assess whether neb and polo also function through the same genetic pathways. Our results revealed highly significant synergistic genetic interactions between neb and polo for both symmetric cell divisions and earlier cell divisions (P = 3.89 × 10−4 and P = 1.00 × 10−6, respectively), but no significant interactions in the case of asymmetric cardiac progenitor cell divisions (P = 0.117) (Figs. 5a,d,e, 6b, Supplementary Table S1).
Since Myb has been shown to regulate polo expression41, and act synergistically with jumu to mediate only symmetric and earlier Svp precursor-determining cardiac progenitor cell divisions27, we next addressed whether neb works in concert with Myb via genetic interaction assays, and in which classes of cardiac progenitor cell divisions. Significant synergistic genetic interactions between neb and Myb were also detected in the case of both symmetric cell divisions and earlier cell divisions (P = 7.60 × 10−5 and P = 7.43 × 10−4, respectively), but not for asymmetric cell divisions (P = 0.340) (Figs. 5a,f,g, 6c, Supplementary Table S1).
Taken together, these results and our initial findings show that neb is necessary for only two of the three categories of cardiac progenitor cell divisions, symmetric and earlier, and that it mediates these two types of cell divisions by working in concert with jumu, Myb, and polo through the same genetic pathways.
Similar synergistic genetic interactions are not detected between neb and CHES-1-like
As described previously, both of the jumu-regulated cardiogenic processes that we had discovered, cardiac progenitor specification and cell division, are also regulated by another Fox TF, CHES-1-like. Furthermore, the downstream genes used for each of these processes, htl and fz in the case of cadiac mesoderm specification, and polo in the case of cardiac progenitor cell divisions, are each also regulated by both jumu and CHES-1-like27,28. neb, however, provides an exception to this trend because, based on our expression profiling data, it is transcriptionally regulated only by jumu, and not by CHES-1-like. Does the observation that CHES-1-like does not regulate neb imply that neb functions completely independently of CHES-1-like, or are they both components of the same genetic subnetwork?
In an attempt to answer this question, we assessed potential genetic interactions between neb and CHES-1-like in mediating cardiac progenitor cell divisions, based on the premise that such interactions are likely to be synergistic if neb and CHES-1-like function in concert in the same pathways. The frequency of cardiac progenitor cell division defects in CHES-1-like1/+; nebMI0225/+ double heterozygotes was not significantly different from the additive sum of those of the CHES-1-like1/+ and nebMI0225/+ single heterozygotes during asymmetric (P = 0.266), symmetric (P = 0.721), or earlier Svp precursor-determining cell divisions (P = 0.530) (Figs. 5a,h,i, 6d, Supplementary Table S1). The lack of synergistic genetic interactions between neb and CHES-1-like in any of the three cardiac progenitor cell division categories suggests strongly that these two genes function independently of each other in these cardiogenic processes.
Cardiac mesoderm-targeted ectopic expression of neb partially rescues both symmetric and earlier cardiac progenitor cell division defects in embryos lacking jumu function
Our results so far show that jumu activates neb and that neb is specifically required in the cardiac mesoderm for mediating symmetric and earlier Svp precursor-determining cell divisions, i.e. two of the three categories of jumu-regulated cardiac progenitor cell divisions. The most parsimonious hypothesis that can explain these observations is that jumu transcriptionally upregulates neb in the cardiac mesoderm to bring about symmetric and earlier cardiac progenitor cell divisions, and that in jumu loss-of-function mutations the ensuing reduction in neb expression levels in this tissue results in defects in only these two types of cell divisions. However, the nature of neb expression prevented us from qualitatively assessing via in situ hybridization assays whether the reduction of neb transcript levels in the entire mesoderm in jumu mutants reflected a reduction of neb levels specifically in the cardiac mesoderm.
Therefore, to test this hypothesis, we ectopically expressed full-length neb specifically in the cardiac mesoderm in jumu loss-of-function mutants under the control of the TinD-GAL4 driver (Fig. 2c). If our hypothesis that neb acts downstream of jumu specifically in the cardiac mesoderm is correct, then the cardiac mesoderm-specific ectopic expression of neb should be able to partially restore neb levels and rescue the symmetric and earlier cardiac progenitor cell division defects in jumu mutants. We found that ectopic expression of neb throughout the cardiac mesoderm does indeed significantly reduce the severity of defects of jumu mutants in symmetric cardiac progenitor cell divisions (P = 0.0144). Defects in Svp precursor-determining cell divisions are also rescued, albeit with borderline significance (P = 0.0632). In contrast, no significant difference is detected in asymmetric cell division defects between jumu mutants alone and jumu mutants expressing full-length neb (P = 0.781) (Fig. 7a–c, Supplementary Table S1). Collectively, our results thus indicate a downstream requirement of neb in the cardiac mesoderm for correct symmetric and earlier cell divisions mediated by jumu.
Cardiac mesoderm-targeted ectopic expression of neb partially rescues both symmetric and earlier Svp precursor-determining cardiac progenitor cell division defects in embryos lacking jumu function. (a) Representative heart from an embryo homozygous for the jumuDf(3R)Exel6157 null mutation exhibiting asymmetric cell division defects (asterisks), symmetric cell division defects (arrowheads), and earlier Svp precursor-determining cell division defects (arrow). (b) Representative heart from an embryo homozygous for the jumuDf(3R)Exel6157 null mutation but ectopically expressing neb in the cardiac mesoderm exhibiting only asymmetric cell division defects. All of these embryos possess one copy of the svp-lacZ enhancer trap, allowing the identification of Tin-CCs (green), Svp-CCs (yellow), and Svp-PCs (red). (c) Quantification and significance of each type of cardiac progenitor cell division defect (n = 193 hemisegments for embryos homozygous for the jumuDf(3R)Exel6157 null mutation alone, and n = 205 hemisegments for embryos homozygous for the jumuDf(3R)Exel6157 null mutation but ectopically expressing neb in the cardiac mesoderm).
Discussion
Our previous work had identified the roles of htl and fz as Fox TF-regulated genes that mediate cardiac progenitor specification and polo as yet another Fox TF-regulated gene that mediates three distinct categories of cardiac progenitor cell division26,28. In this study, we identify another Fox TF-regulated downstream gene, neb, which also mediates cardiac progenitor cell divisions. The previously characterized Fox downstream targets htl, fz, and polo are each regulated by both jumu and CHES-1-like simultaneously. Our expression profiling results indicate that neb, in contrast, is transcriptionally regulated only by Jumu, and not by CHES-1-like. This independence from CHES-1-like regulatory control is also supported by neb exhibiting synergistic genetic interactions with jumu in cardiac progenitor cell division subnetworks, but not with CHES-1-like.
We showed further that it is the activation of neb by Jumu specifically in the cardiac mesoderm that is necessary for proper cardiac progenitor cell divisions. It is possible that Jumu may also activate neb expression at other mesodermal domains besides the cardiac mesoderm: our expression profiling was based on the entire mesoderm, and in situ hybridization assays showed neb expression in subsets of the somatic mesoderm in addition to the cardiac mesoderm. However, the twin observations that cardiac mesoderm-specific knockdown of neb partially phenocopies jumu loss-of-function cardiac progenitor cell division defects and that cardiac mesoderm-specific ectopic expression of neb partially rescues jumu loss-of-function cell division defects demonstrate clearly that jumu mediates these cardiac progenitor cell division defects by activating neb expression specifically in the cardiac mesoderm.
Our previous examinations of cardiac progenitor cell divisions had shown that while jumu, CHES-1-like, and polo are required for all three categories of cardiac progenitor cell division—asymmetric cell divisions, symmetric cell divisions, and cell divisions at an earlier stage that produce the Svp precursor cells—Myb is required for only the two latter classes27,28. In this study, we discovered that neb, like Myb, is also required for only the symmetric and earlier cardiac progenitor cell divisions, not asymmetric cell divisions. Furthermore, while neb exhibits no synergistic genetic interactions with CHES-1-like in any of the three classes of cardiac progenitor cell divisions, it displays synergistic genetic interactions with Myb and polo exclusively in symmetric and earlier Svp precursor-determining cell divisions. And while neb and jumu do show synergistic genetic interactions in all three categories of cardiac progenitor cell divisions, the interaction in the case of asymmetric cell divisions is barely significant. Finally, we find that ectopic cardiac mesoderm-specific expression of neb partially rescues symmetric and earlier cardiac progenitor cell division defects in jumu loss-of-function mutants, but not asymmetric cell division defects. Given also prior studies showing synergistic genetic interactions between Myb, jumu, and polo in these same two classes of cardiac progenitor cell divisions27, and the regulation of polo by both jumu and Myb28,41, our data thus suggests the presence of an additional CHES-1-like-independent, jumu-regulated pathway involving Myb, polo, and neb that mediates only symmetric and earlier cell divisions. This pathway both expands our prior model of Fox TF-regulated cardiogenic processes (Fig. 8) and emphasizes the central role of jumu in heart development by illustrating how it utilizes different combinations of other regulators and downstream effectors to mediate each of these processes.
Model for jumu and CHES-1-like-regulated cardiogenic processes based on previous work and this current study. jumu and CHES-1-like initially transcriptionally activate htl and fz to mediate the specification of cardiac progenitors from the dorsal mesoderm by the FGF and Wnt-signaling pathways26. Polo, activated by the synergistic functions of jumu and CHES-1-like, then brings about the phosphorylation of Pon necessary for the localization of Numb protein during asymmetric cell division of the Svp precursor cells28. Additionally, jumu, CHES-1-like, and Myb also act synergistically to activate Polo to mediate both symmetric and earlier Svp precursor-determining cell divisions27,28,41. The results of the current study suggest the presence of an additional jumu-regulated pathway (highlighted in peach) that acts independently of CHES-1-like, involves Myb, polo, and neb, and mediates only symmetric and earlier Svp precursor-determining cardiac progenitor cell divisions. Lines without arrowheads connecting neb with polo and with Myb indicate that while these three genes do indeed function synergistically to mediate symmetric and earlier cardiac progenitor cell divisions, our data at present cannot determine whether Myb or polo are acting upstream or downstream of neb. Since both polo and neb are regulated by jumu, and since it is not yet known whether polo acts upstream or downstream of neb, these two genes have been drawn at the same level.
neb encodes a kinesin, the Drosophila ortholog of the mammalian Kif14, that is required for chromatin condensation, chromosome segregation, and cytokinesis31,42,43,44; thus errors in one or more of these processes could result in the symmetric or earlier cardiac progenitor cell division defects seen in neb mutants. In particular, Neb forms a conserved physical complex with another kinesin, Pavarotti, and the Citron kinase Sticky at the nexus of the central spindle and the actomyosin contractile ring to bring about the cleavage furrow ingression necessary for cytokinesis45. Thus it is particularly intriguing to learn from our expression profiling data that both sticky (sti) and pavarotti (pav) are also activated by Jumu in the mesoderm [log2(fold change) = − 0.77505; adjusted P-value = 8.27 × 10−8 and log2(fold change) = − 0.54834; adjusted P-value = 2.51 × 10−6 respectively in jumu mutants] (Supplementary Table S3). These data raise the possibility that Jumu may be mediating one aspect of cardiac progenitor cell divisions by regulating not just one but multiple components of this cytokinesis subnetwork and will be the focus of future investigations.
A particularly interesting question raised by our observations is why loss-of-function mutations in neb affect symmetric cell divisions of cardiac progenitor cells, but not asymmetric cell divisions. One can easily conceive of the converse: mutations in numb or pon, for instance, that disrupt asymmetric cell division without affecting symmetric cell division by failing to appropriately localize Numb to one pole of the dividing cell, and thus disproportionately to one of the resulting daughter cells27,28,29. Proper Numb localization is required only for asymmetric cell division. But why would mutations in neb, a gene purportedly involved in more general cell division phenomena such as chromosome condensation, chromosome segregation, and cytokinesis, have no effect on cardiac progenitor asymmetric cell division? One possible explanation could be the presence of other redundant genes and/or pathways that may compensate for the lack of neb function during asymmetric cell division. These potential redundant genes and pathways could also be involved in symmetric cell division and may explain why null mutations in neb did not result in symmetric cell division defects in each and every hemisegment.
Another intriguing question is whether the activation of neb by Jumu is mediated by direct transcriptional regulation, requiring the functional binding of the Fox TF itself to cis-regulatory regions of the neb gene to stimulate its transcription; or indirect, involving regulation by other, Jumu-activated TFs. While this question cannot be completely answered without the identification and functional analysis of the cis-regulatory regions mediating neb transcription in the cardiac mesoderm, we note that ChIP-seq analyses compiled in the modERN and modENCODE databases identify three in vivo Jumu binding regions in the immediate neighborhood of the neb gene, one overlapping the promoter, the other two in the intron, and all three coinciding with the in vivo binding sites of Myb as well as those of the cardiogenic TFs Tinman, Tailup, and Twist46,47 (Supplementary Fig. S2). These observations raise the possibility that one or more of these three Jumu binding sites may be the cis-regulatory region(s) mediating neb transcription and suggest that neb might be a direct transcriptional target of Jumu as well as of Myb.
We have previously suggested28 that FoxM1 is the functional ortholog of jumu in mammals because FoxM1 regulates Plk1, the mammalian ortholog of polo48, because FoxM1 and Myb function as synergistic master regulators of key genetic programs of human B cell proliferation in the germinal center49, and because mutations in FoxM1 exhibit cardiac phenotypes—highly irregular orientation of cardiomyocytes with enlarged polyploid nuclei—that are similar to those in jumu mutants10. Given our discovery of a jumu-regulated pathway for cell division that involves Myb, polo, and neb in Drosophila, and possible cis-regulatory regions of neb that bind both Jumu and Myb, it was very interesting to find that both mammalian FoxM1 and B-Myb bind to the promoter of the neb ortholog Kif14 and activate its transcription in breast cancer cell lines50,51. And, not surprisingly, Kif14 is considered to play a role as an oncogene in cancer types as varied as breast, lung, liver, gastric, colorectal, ovarian, cervical, and prostate cancers in addition to glioma, medulloblastoma, and retinoblastoma52,53,54,55,56,57,58,59,60,61,62,63,64.
Taken together, these data suggest that the evolutionary role of neb and its ortholog Kif14 is quite crucial, involving not only a conserved complex mediating cytokinesis in both flies and mammals45,65, but also its potentially conserved regulation by Fox and Myb TFs. It remains to be seen whether the Fox TF-regulated role of this kinesin is also conserved in cardiac progenitor cell divisions in mammals. The observation that the Meckel–Gruber syndrome linked to some mutations of human Kif14 is occasionally found to be also associated with congenital heart defects66 suggests that this may indeed be the case.
Methods
Drosophila strains and genetics
The following mutant alleles, deficiencies, and transgenes were used for counting cell division defects: jumuDf(3R)Exel6157 [also known as Df(3R)Exel6157; FlyBase ID: FBab0038212] and CHES-1-like1 [also known as Df(1)CHES-1-like1; FlyBase ID: FBab0047384]28; nebk05702 [FlyBase ID: FBal0043429]42; nebMI0225 [FlyBase ID: FBal0249290]; UAS-nebRNAi [nebGD7072; FlyBase ID: FBal0210181]; UAS-neb [nebUAS.Tag:HA; FlyBase ID: FBal0197607]; svp-lacZ [svp3; FlyBase ID: FBal0016610]33; polo10 [poloS132408; FlyBase ID: FBal0099891]67; MybMH107 [FlyBase ID: FBal0141652]68; UAS-Dcr-2 [Dcr-2UAS.cDa; FlyBase ID: FBal0211026]69; TinD-GAL4 [GAL4tin.D; FlyBase ID: FBal0267971]30,39; and Hand-GAL4 [GAL4HCH.Hand; FlyBase ID: FBal0190607]40. They were maintained over FM7c, ftz-lacZ, CyO, ftz-lacZ, and TM3, ftz-lacZ balancers when they could not be homozygosed before crossing. Relevant embryos resulting from these crosses were distinguished from their siblings by the absence of anti-β-galactosidase staining in the ftz-lacZ pattern of the balancers. CHES-1-like and Myb heterozygotes were distinguished from hemizygotes by both positive staining for anti-Sxl since they were female and the absence of anti-β-galactosidase staining in the ftz-lacZ pattern. The same anti-β-galactosidase staining, along with anti-Mef-2 staining, was also used to distinguish the Svp-CCs and Svp-PCs except in the case of RNA interference assays. The nebMI0225 allele, a weak hypomorphic mutation that fails to complement the nebk05702 null allele, was used in genetic interaction assays because it allowed cell division defects to be assessed with increased sensitivity. The genotypes of all the embryos used for counting cell division defects (Supplementary Table S1) and presented in the figures are described below.
svp-lacZ/+ | Figure 4a |
nebk05702; svp-lacZ/+ | Figure 4b |
nebMI02225/+; svp-lacZ/+ | Figure 5b |
jumuDf(3R)Exel6157 svp-lacZ/+ | Figure 5a |
nebMI02225/+; jumuDf(3R)Exel6157 svp-lacZ/+ | Figure 5c |
polo10 svp-lacZ/+ | Figure 5d |
nebMI02225/+; polo10 svp-lacZ/+ | Figure 5e |
MybMH107/+; svp-lacZ/+ | Figure 5f |
MybMH107/+; nebMI02225/+; svp-lacZ/+ | Figure 5g |
CHES-1-like1/+; svp-lacZ/+ | Figure 5h |
CHES-1-like1/+; nebMI02225/+; svp-lacZ/+ | Figure 5i |
jumuDf(3R)Exel6157 svp-lacZ/jumuDf(3R)Exel6157 | Figure 7a |
TinD-GAL4/+; jumuDf(3R)Exel6157 svp-lacZ/jumuDf(3R)Exel6157 UAS-neb | Figure 7b |
+/+ | Figure 4d |
UAS-nebRNAi/+ | Suppl. Fig. S1 |
Hand-GAL4/+; TinD-GAL4 UAS-Dcr-2/+ | Suppl. Fig. S1 |
Hand-GAL4/+; TinD-GAL4 UAS-Dcr-2/UAS-nebRNAi | Figure 3e |
The following transgenes were used for RT-qPCR: twi-GAL4 [GAL4twi.PG; FlyBase ID: FBal0040491]70,71; UAS-Dcr-2 [described earlier]; UAS-jumuRNAi [jumuGD4099; FlyBase ID: FBal0209759]28,69; and UAS-CHES-1-likeRNAi [CHES-1-likeGD4327; FlyBase ID: FBal0206447]69,72. The genotypes of all the embryos used for the RT-qPCR assays are described below:
twi-GAL4 UAS-Dcr-2/+ |
twi-GAL4 UAS-Dcr-2/+; UAS-jumuRNAi/+ |
twi-GAL4 UAS-Dcr-2/+; UAS-CHES-1-likeRNAi/+ |
RNA interference assays
RNA interference knockdown of neb was performed using the UAS-nebRNAi (nebGD7072) construct targeted to the cardiac mesoderm using both TinD-GAL4 and Hand-GAL4 drivers simultaneously. The efficiency of knockdowns was enhanced with UAS-Dcr-2. Cell division defects were counted in the neb knockdown and controls (wild-type, UAS-nebRNAi alone, and both GAL4 drivers with UAS-Dcr-2 but no UAS-nebRNAi) by staining with anti-Mef2 and anti-Svp to identify Tin-CCs and Svp-CCs.
Reverse transcription quantitative real-time PCR (RT-qPCR) assays
Details of the RT-qPCR methodology and the mesoderm-targeted RNA interference knockdown of jumu and CHES-1-like used in these assays are included in Supplementary Method S1.
In situ hybridization, immunohistochemistry, microscopy, and cell counting
Embryo fixation, riboprobe synthesis, in situ hybridization, and fluorescent immunohistochemistry were performed as described previously26,28. The following primary antibodies were used: rabbit anti-Mef2 (1:1000, gift from B. Paterson; 1:7500, gift from J. Jacobs), mouse anti-β-galactosidase (1:500, Catalog no. Z3783 from Promega), chicken anti-β-galactosidase (1:500, Catalog no. ab9631 from Abcam, Inc), mouse anti-Svp (1:5, monoclonal 5B11 from the Developmental Studies Hybridoma Bank), and mouse anti-Sxl (1:20, monoclonal M18 from the Developmental Studies Hybridoma Bank). Fluorescent microscopy was performed on a Zeiss AxioImager with Apotome. Z-stacks of entire stage 16 embryonic hearts were scanned with a 40X objective and 0.31 µm steps, and all planes for each z-stack were examined to count cells and determine cell division defects. Cell counting and assessment of cell division defects were performed blind (i.e. the individual evaluating cardiac progenitor cell division defects did not know the genotypes of the embryos being assessed) to avoid any potential bias. For all quantitative studies, cells in 179 or more hemisegments were counted for each genotype.
Statistical methods
Comparison of cell division error rates between genotypes was done using regression models with the response variable being the proportion of hemisegment errors for each embryo. Due to violation of regression assumptions, e.g., non-normality and heteroscedasticity, permutation (randomization) tests were used to obtain reliable p-values73.
For comparing rates between two genotypes, for example nebk05702 and wild-type, the following general linear model was used:
where \({Y}_{j}\) is the proportion of hemisegment errors for embryo j and indicator variable \({I}_{j}\) is 1 if embryo j has phenotype nebk05702 and 0 otherwise. To obtain a permutation p-value for testing \({H}_{0}: {\beta }_{1}=0\), the estimate of \({\beta }_{1}\) for the actual data is compared with the estimates obtained when the genotypes of the embryos are permuted, i.e., the phenotype labels are randomly shuffled among the embryos in the sample. The permutation test p-value is then \(p=(n+1)/(N+1)\) where \(n\) is the number of permutation estimates for \({\beta }_{1}\) which exceed the estimate for the actual data and \(N\) is the number of permutations74. In order to obtain highly reproducible p-values, \(N={10}^{6}\) permutations were used for all permutation tests.
For determining if cell division error rates are non-additively related to two gene mutations, for example, to detect synergistic interaction between neb and Myb, a general linear model allowing for interaction was used:
where \({I}_{n,j}\) is 1 if the jth embryo is heterozygous for the nebMI02225 mutation and \({I}_{M,j}=1\) if it is heterozygous for the MybMH107 mutation. Since synergism is present only if \({\beta }_{3}\ne 0\), to detect it \({H}_{0}: {\beta }_{3}=0\) was tested using permutation. Since this is a multiple regression model, a somewhat more sophisticated permutation procedure, the Smith procedure (orthogonalization) was employed75.
Data availability
The Microarray-based expression profiling data used in this study are available from NCBI’S Gene Expression Omnibus through GEO Series accession number GSE65439 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=gse65439). Other data generated and analysed in this study are included in this article and its supplementary information files. All fly strains used in this study are available upon request.
References
Filosa, S. et al. Goosecoid and HNF-3beta genetically interact to regulate neural tube patterning during mouse embryogenesis. Development 124, 2843–2854 (1997).
Yamagishi, H. et al. Tbx1 is regulated by tissue-specific forkhead proteins through a common Sonic hedgehog-responsive enhancer. Genes Dev. 17, 269–281. https://doi.org/10.1101/GAD.1048903 (2003).
Hu, T. et al. Tbx1 regulates fibroblast growth factors in the anterior heart field through a reinforcing autoregulatory loop involving forkhead transcription factors. Development 131, 5491–5502. https://doi.org/10.1242/DEV.01399 (2004).
Harrelson, Z., Kaestner, K. H. & Evans, S. M. Foxa2 mediates critical functions of prechordal plate in patterning and morphogenesis and is cell autonomously required for early ventral endoderm morphogenesis. Biol. Open 1, 173–181. https://doi.org/10.1242/BIO.2012040 (2012).
Tamplin, O. J. et al. Microarray analysis of Foxa2 mutant mouse embryos reveals novel gene expression and inductive roles for the gastrula organizer and its derivatives. BMC Genom. 9, 511. https://doi.org/10.1186/1471-2164-9-511 (2008).
Kume, T., Jiang, H., Topczewska, J. M. & Hogan, B. L. The murine winged helix transcription factors, Foxc1 and Foxc2, are both required for cardiovascular development and somitogenesis. Genes Dev. 15, 2470–2482. https://doi.org/10.1101/GAD.907301 (2001).
Seo, S. & Kume, T. Forkhead transcription factors, Foxc1 and Foxc2, are required for the morphogenesis of the cardiac outflow tract. Dev. Biol. 296, 421–436. https://doi.org/10.1016/J.YDBIO.2006.06.012 (2006).
Winnier, G. E. et al. Roles for the winged helix transcription factors MF1 and MFH1 in cardiovascular development revealed by nonallelic noncomplementation of null alleles. Dev. Biol. 213, 418–431. https://doi.org/10.1006/DBIO.1999.9382 (1999).
Von Both, I. et al. Foxh1 is essential for development of the anterior heart field. Dev. Cell 7, 331–345. https://doi.org/10.1016/J.DEVCEL.2004.07.023 (2004).
Korver, W. et al. Uncoupling of S phase and mitosis in cardiomyocytes and hepatocytes lacking the winged-helix transcription factor Trident. Curr. Biol. 8, 1327–1330 (1998).
Bolte, C. et al. Expression of Foxm1 transcription factor in cardiomyocytes is required for myocardial development. PLoS ONE 6, e22217. https://doi.org/10.1371/journal.pone.0022217 (2011).
Hosaka, T. et al. Disruption of forkhead transcription factor (FOXO) family members in mice reveals their functional diversification. Proc. Natl. Acad. Sci. USA 101, 2975–2980. https://doi.org/10.1073/PNAS.0400093101 (2004).
Sengupta, A., Chakraborty, S., Paik, J., Yutzey, K. E. & Evans-Anderson, H. J. FoxO1 is required in endothelial but not myocardial cell lineages during cardiovascular development. Dev. Dyn. 241, 803–813. https://doi.org/10.1002/DVDY.23759 (2012).
Wang, B. et al. Foxp1 regulates cardiac outflow tract, endocardial cushion morphogenesis and myocyte proliferation and maturation. Development 131, 4477–4487. https://doi.org/10.1242/DEV.01287 (2004).
Zhang, Y. et al. Foxp1 coordinates cardiomyocyte proliferation through both cell-autonomous and nonautonomous mechanisms. Genes Dev. 24, 1746–1757. https://doi.org/10.1101/GAD.1929210 (2010).
Li, S., Zhou, D., Lu, M. M. & Morrisey, E. E. Advanced cardiac morphogenesis does not require heart tube fusion. Science 305, 1619–1622. https://doi.org/10.1126/science.1098674 (2004).
Swiderski, R. E. et al. Expression of the Mf1 gene in developing mouse hearts: implication in the development of human congenital heart defects. Dev. Dyn. 216, 16–27. https://doi.org/10.1002/(SICI)1097-0177(199909)216:1%3c16::AID-DVDY4%3e3.0.CO;2-1 (1999).
Gripp, K. W., Hopkins, E., Jenny, K., Thacker, D. & Salvin, J. Cardiac anomalies in Axenfeld-Rieger syndrome due to a novel FOXC1 mutation. Am. J Med. Genet. A 161A, 114–119. https://doi.org/10.1002/AJMG.A.35697 (2013).
Du, R. F. et al. A novel mutation of FOXC1 (R127L) in an Axenfeld-Rieger syndrome family with glaucoma and multiple congenital heart diseases. Ophthalm. Genet. 37, 111–115. https://doi.org/10.3109/13816810.2014.924016 (2016).
Topf, A. et al. Functionally significant, rare transcription factor variants in tetralogy of Fallot. PLoS ONE 9, E95453. https://doi.org/10.1371/JOURNAL.PONE.0095453 (2014).
Yu, S., Shao, L., Kilbride, H. & Zwick, D. L. Haploinsufficiencies of FOXF1 and FOXC2 genes associated with lethal alveolar capillary dysplasia and congenital heart disease. Am. J. Med. Genet. A 152A, 1257–1262. https://doi.org/10.1002/AJMG.A.33378 (2010).
Roessler, E. et al. Reduced NODAL signaling strength via mutation of several pathway members including FOXH1 is linked to human heart defects and holoprosencephaly. Am. J. Hum. Genet. 83, 18–29. https://doi.org/10.1016/J.AJHG.2008.05.012 (2008).
Wang, B. et al. Forkhead box H1 (FOXH1) sequence variants in ventricular septal defect. Int. J. Cardiol. 145, 83–85. https://doi.org/10.1016/J.IJCARD.2009.05.030 (2010).
Chang, S. W. et al. Genetic abnormalities in FOXP1 are associated with congenital heart defects. Hum. Mutat. 34, 1226–1230. https://doi.org/10.1002/HUMU.22366 (2013).
Zhu, H. Forkhead box transcription factors in embryonic heart development and congenital heart disease. Life Sci. 144, 194–201. https://doi.org/10.1016/J.LFS.2015.12.001 (2016).
Ahmad, S. M., Bhattacharyya, P., Jeffries, N., Gisselbrecht, S. S. & Michelson, A. M. Two Forkhead transcription factors regulate cardiac progenitor specification by controlling the expression of receptors of the fibroblast growth factor and Wnt signaling pathways. Development 143, 306–317. https://doi.org/10.1242/DEV.122952 (2016).
Ahmad, S. M. et al. Machine learning classification of cell-specific cardiac enhancers uncovers developmental subnetworks regulating progenitor cell division and cell fate specification. Development 141, 878–888. https://doi.org/10.1242/DEV.101709 (2014).
Ahmad, S. M. et al. Two forkhead transcription factors regulate the division of cardiac progenitor cells by a Polo-dependent pathway. Dev. Cell 23, 97–111. https://doi.org/10.1016/J.DEVCEL.2012.05.011 (2012).
Wang, H., Ouyang, Y., Somers, W. G., Chia, W. & Lu, B. Polo inhibits progenitor self-renewal and regulates Numb asymmetry by phosphorylating Pon. Nature 449, 96–100. https://doi.org/10.1038/NATURE06056 (2007).
Zhu, X. et al. Differential regulation of mesodermal gene expression by Drosophila cell type-specific Forkhead transcription factors. Development 139, 1457–1466. https://doi.org/10.1242/DEV.069005 (2012).
Molina, I. et al. A chromatin-associated kinesin-related protein required for normal mitotic chromosome segregation in Drosophila. J. Cell Biol. 139, 1361–1371. https://doi.org/10.1083/JCB.139.6.1361 (1997).
Gajewski, K., Choi, C. Y., Kim, Y. & Schulz, R. A. Genetically distinct cardial cells within the Drosophila heart. Genesis 28, 36–43. https://doi.org/10.1002/1526-968X(200009)28:1%3c36::AID-GENE50%3e3.0.CO;2-4 (2000).
Ward, E. J. & Skeath, J. B. Characterization of a novel subset of cardiac cells and their progenitors in the Drosophila embryo. Development 127, 4959–4969 (2000).
Han, Z. & Bodmer, R. Myogenic cells fates are antagonized by Notch only in asymmetric lineages of the Drosophila heart, with or without cell division. Development 130, 3039–3051 (2003).
Bodmer, R. & Frasch, M. Development and aging of the Drosophila heart. In Heart Development and Regeneration (eds N. Rosenthal & R.P. Harvey) Ch. 1.2, 47–86 (Academic Press, 2010).
Michelson, A. M., Gisselbrecht, S., Zhou, Y., Baek, K. H. & Buff, E. M. Dual functions of the heartless fibroblast growth factor receptor in development of the Drosophila embryonic mesoderm. Dev. Genet. 22, 212–229 (1998).
Bhanot, P. et al. Frizzled and Dfrizzled-2 function as redundant receptors for Wingless during Drosophila embryonic development. Development 126, 4175–4186 (1999).
Chen, C. M. & Struhl, G. Wingless transduction by the Frizzled and Frizzled2 proteins of Drosophila. Development 126, 5441–5452 (1999).
Yin, Z., Xu, X. L. & Frasch, M. Regulation of the twist target gene tinman by modular cis-regulatory elements during early mesoderm development. Development 124, 4971–4982 (1997).
Han, Z. & Olson, E. N. Hand is a direct target of Tinman and GATA factors during Drosophila cardiogenesis and hematopoiesis. Development 132, 3525–3536. https://doi.org/10.1242/DEV.01899 (2005).
Wen, H., Andrejka, L., Ashton, J., Karess, R. & Lipsick, J. S. Epigenetic regulation of gene expression by Drosophila Myb and E2F2-RBF via the Myb-MuvB/dREAM complex. Genes Dev. 22, 601–614. https://doi.org/10.1101/GAD.1626308 (2008).
Ruden, D. M., Cui, W., Sollars, V. & Alterman, M. A Drosophila kinesin-like protein, Klp38B, functions during meiosis, mitosis and segmentation. Dev. Biol. 191, 284–296. https://doi.org/10.1006/DBIO.1997.8726 (1997).
Ohkura, H. et al. Mutation of a gene for a Drosophila kinesin-like protein, Klp38B, leads to failure of cytokinesis. J. Cell Sci. 110(PT 8), 945–954 (1997).
Alphey, L. et al. KLP38B: a mitotic kinesin-related protein that binds PP1. J Cell Biol. 138, 395–409. https://doi.org/10.1083/JCB.138.2.395 (1997).
Bassi, Z. I., Audusseau, M., Riparbelli, M. G., Callaini, G. & D’Avino, P. P. Citron kinase controls a molecular network required for midbody formation in cytokinesis. Proc. Natl. Acad. Sci. USA 110, 9782–9787. https://doi.org/10.1073/PNAS.1301328110 (2013).
Kudron, M. M. et al. The modERN resource: genome-wide binding profiles for hundreds of Drosophila and Caenorhabditis elegans transcription factors. Genetics 208, 937–949. https://doi.org/10.1534/GENETICS.117.300657 (2018).
Negre, N. et al. A cis-regulatory map of the Drosophila genome. Nature 471, 527–531. https://doi.org/10.1038/NATURE09990 (2011).
Laoukili, J. et al. FoxM1 is required for execution of the mitotic programme and chromosome stability. Nat. Cell Biol. 7, 126–136. https://doi.org/10.1038/NCB1217 (2005).
Lefebvre, C. et al. A human B-cell interactome identifies MYB and FOXM1 as master regulators of proliferation in germinal centers. Mol. Syst. Biol. 6, 377. https://doi.org/10.1038/MSB.2010.31 (2010).
Wolter, P., Hanselmann, S., Pattschull, G., Schruf, E. & Gaubatz, S. Central spindle proteins and mitotic kinesins are direct transcriptional targets of MuvB, B-MYB and FOXM1 in breast cancer cell lines and are potential targets for therapy. Oncotarget 8, 11160–11172. https://doi.org/10.18632/ONCOTARGET.14466 (2017).
Fischer, M. & Muller, G. A. Cell cycle transcription control: DREAM/MuvB and RB-E2F complexes. Crit. Rev. Biochem. Mol. Biol. 52, 638–662. https://doi.org/10.1080/10409238.2017.1360836 (2017).
Huang, W. et al. Inhibition of KIF14 suppresses tumor cell growth and promotes apoptosis in human glioblastoma. Cell. Physiol. Biochem. Int. J. Exp. Cell. Physiol. Biochem. Pharmacol. 37, 1659–1670. https://doi.org/10.1159/000438532 (2015).
Hung, P. F. et al. The motor protein KIF14 inhibits tumor growth and cancer metastasis in lung adenocarcinoma. PLoS ONE 8, E61664. https://doi.org/10.1371/JOURNAL.PONE.0061664 (2013).
Li, K. K. et al. The kinesin KIF14 is overexpressed in medulloblastoma and downregulation of KIF14 suppressed tumor proliferation and induced apoptosis. Lab. Investig. J. Tech. Methods Pathol. 97, 946–961. https://doi.org/10.1038/LABINVEST.2017.48 (2017).
O’Hare, M. et al. Kif14 overexpression accelerates murine retinoblastoma development. Int. J. Cancer 139, 1752–1758. https://doi.org/10.1002/IJC.30221 (2016).
Qiu, H. L. et al. High expression of KIF14 is associated with poor prognosis in patients with epithelial ovarian cancer. Eur. Rev. Med. Pharmacol. Sci. 21, 239–245 (2017).
Singel, S. M. et al. KIF14 promotes AKT phosphorylation and contributes to chemoresistance in triple-negative breast cancer. Neoplasia (New York, N.Y.) 16, 247–256. https://doi.org/10.1016/J.NEO.2014.03.008 (2014).
Thériault, B. L. et al. Transcriptional and epigenetic regulation of KIF14 overexpression in ovarian cancer. PLoS ONE 9, E91540. https://doi.org/10.1371/JOURNAL.PONE.0091540 (2014).
Wang, W., Shi, Y., Li, J., Cui, W. & Yang, B. Up-regulation of KIF14 is a predictor of poor survival and a novel prognostic biomarker of chemoresistance to paclitaxel treatment in cervical cancer. Biosci. Rep. https://doi.org/10.1042/BSR20150314 (2016).
Wang, Z. Z. et al. KIF14 promotes cell proliferation via activation of Akt and is directly targeted by miR-200c in colorectal cancer. Int. J. Oncol. 53, 1939–1952. https://doi.org/10.3892/IJO.2018.4546 (2018).
Xu, H. et al. Silencing of KIF14 interferes with cell cycle progression and cytokinesis by blocking the p27(Kip1) ubiquitination pathway in hepatocellular carcinoma. Exp. Mol. Med. 46, E97. https://doi.org/10.1038/EMM.2014.23 (2014).
Yang, T., Zhang, X. B. & Zheng, Z. M. Suppression of KIF14 expression inhibits hepatocellular carcinoma progression and predicts favorable outcome. Cancer Sci. 104, 552–557. https://doi.org/10.1111/CAS.12128 (2013).
Yang, Z. et al. KIF14 promotes tumor progression and metastasis and is an independent predictor of poor prognosis in human gastric cancer. Biochim. Biophys. Acta Mol. Basis Dis. 181–192, 2019. https://doi.org/10.1016/J.BBADIS.2018.10.039 (1865).
Zhang, Y. et al. Overexpression of a novel candidate oncogene KIF14 correlates with tumor progression and poor prognosis in prostate cancer. Oncotarget 8, 45459–45469. https://doi.org/10.18632/ONCOTARGET.17564 (2017).
Gruneberg, U. et al. KIF14 and citron kinase act together to promote efficient cytokinesis. J. Cell. Biol. 172, 363–372. https://doi.org/10.1083/JCB.200511061 (2006).
Hartill, V., Szymanska, K., Sharif, S. M., Wheway, G. & Johnson, C. A. Meckel-Gruber syndrome: an update on diagnosis, clinical management and research advances. Front. Pediatr. 5, 244. https://doi.org/10.3389/FPED.2017.00244 (2017).
Donaldson, M. M., Tavares, A. A., Ohkura, H., Deak, P. & Glover, D. M. Metaphase arrest with centromere separation in polo mutants of Drosophila. J. Cell Biol. 153, 663–676 (2001).
Manak, J. R., Mitiku, N. & Lipsick, J. S. Mutation of the Drosophila homologue of the Myb protooncogene causes genomic instability. Proc. Natl. Acad. Sci. USA 99, 7438–7443. https://doi.org/10.1073/PNAS.122231599 (2002).
Dietzl, G. et al. A genome-wide transgenic RNAi library for conditional gene inactivation in Drosophila. Nature 448, 151–156. https://doi.org/10.1038/NATURE05954 (2007).
Greig, S. & Akam, M. Homeotic genes autonomously specify one aspect of pattern in the Drosophila mesoderm. Nature 362, 630–632. https://doi.org/10.1038/362630A0 (1993).
Buff, E., Carmena, A., Gisselbrecht, S., Jimenez, F. & Michelson, A. M. Signalling by the Drosophila epidermal growth factor receptor is required for the specification and diversification of embryonic muscle progenitors. Development 125, 2075–2086 (1998).
Mummery-Widmer, J. L. et al. Genome-wide analysis of Notch signalling in Drosophila by transgenic RNAi. Nature 458, 987–992. https://doi.org/10.1038/NATURE07936 (2009).
Manly, B. F. J. Randomization, Bootstrap, and Monte Carlo Methods in Biology 3rd edn. (Chapman & Hall/ CRC, Boca Raton, 2007).
Phipson, B. & Smyth, G. K. Permutation p-values should never be zero: calculating exact p-values when permutations are randomly drawn. Stat. Appl. Genet. Mol. Biol. https://doi.org/10.2202/1544-6115.1585 (2010).
Winkler, A. M., Ridgway, G. R., Webster, M. A., Smith, S. M. & Nichols, T. E. Permutation inference for the general linear model. Neuroimage 92, 381–397. https://doi.org/10.1016/J.NEUROIMAGE.2014.01.060 (2014).
Acknowledgements
This work is supported by the American Heart Association (grant 16SDG31390005 to S.M.A.) and the Indiana Academy of Science (Senior Research Grant to A.J.K.). We thank the Bloomington Drosophila Stock Center (NIH P4OOD018537) and the Vienna Drosophila Resource Center for providing fly strains; B. Paterson, J. Jacobs, and the Developmental Studies Hybridoma Bank Created by the NIHCD of the NIH for providing antibodies; and S. Hall for assistance in illustrating our models with biorender.com.
Author information
Authors and Affiliations
Contributions
S.M.A. and A.J.K. conceived and designed the overall research project and wrote the manuscript. A.J.K., M.P., K.R.S., and S.M.A. designed and performed individual experiments and interpreted the results. M.H.I. performed all statistical analyses.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Kump, A.J., Panta, M., Schwab, K.R. et al. The Drosophila Forkhead/Fox transcription factor Jumeau mediates specific cardiac progenitor cell divisions by regulating expression of the kinesin Nebbish. Sci Rep 11, 3221 (2021). https://doi.org/10.1038/s41598-021-81894-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-021-81894-1
- Springer Nature Limited