Abstract
Rheumatoid arthritis (RA) is a chronic, systemic, autoimmune disease that may lead to joint damage, deformity, and disability, if not treated effectively. Hedyotis diffusa Willd (HDW) and its main components have been widely used to treat a variety of tumors and inflammatory diseases. The present study utilized a network pharmacology approach, microarray data analysis and molecular docking to predict the key active ingredients and mechanisms of HDW against RA. Eleven active ingredients in HDW and 180 potential anti-RA targets were identified. The ingredients-targets-RA network showed that stigmasterol, beta-sitosterol, quercetin, kaempferol, and 2-methoxy-3-methyl-9,10-anthraquinone were key components for RA treatment. KEGG pathway results revealed that the 180 potential targets were inflammatory-related pathways with predominant enrichment of the AGE-RAGE, TNF, IL17, and PI3K-Akt signaling pathways. Screened through the PPI network and with Cytoscape software, RELA, TNF, IL6, TP53, MAPK1, AKT1, IL10, and ESR1 were identified as the hub targets in the HDW for RA treatment. Molecular docking was used to identify the binding of 5 key components and the 8 related-RA hub targets. Moreover, the results of network pharmacology were verified by vitro experiments. HDW inhibits cell proliferation in MH7A cells in a dose and time-dependent manner. RT-qPCR and WB results suggest that HDW may affect hub targets through PI3K/AKT signaling pathway, thereby exerting anti-RA effect. This study provides evidence for a clinical effect of HDW on RA and a research basis for further investigation into the active ingredients and mechanisms of HDW against RA.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA), one of the most common autoimmune diseases, is characterized by invasive joint synovitis, pannus formation, deterioration of joints, and loss of joint function1,2. RA occurs in 0.5 to 1.0% of the population worldwide3. In China, up to 5 million people suffer from pain and recurrence of RA4. Due to its high prevalence, debilitating nature, and disabling consequences, RA has generated a substantial clinical, economic, and social burden. The pathogenesis of RA is complicated, incompletely understood, and considered to be mediated by various mechanisms. At present, the typical events in the pathogenesis of RA are considered to be hyperplasia of cells in the synovial membrane consisting of synovial fibroblasts, macrophages, and lymphocytes5. Some studies have shown that fibroblast-like synoviocytes (FLS) are the dominant cell type and are considered to play a critical role in the pathogenesis of RA6,7. Disease-modifying antirheumatic drugs (DMARDs) and non-steroidal anti-inflammatory medications (NSAIDs) have been widely used in RA therapy8. Conventional DMARDs, including hydroxychloroquine, methotrexate (MTX), sulfasalazine, and leflunomide, have been approved by the US Food and Drug Administration (FDA) as a first-line therapy for RA patients9; however, there are no truly effective pharmacotherapies for the treatment of RA. Most of these drugs have frequent side effects, including gastrointestinal irritation, kidney injury, and cardiovascular risk10. There is therefore an urgent need for safe and effective medical treatments for RA.
In recent years, with the understanding of pathogenesis of RA in Chinese medicine, some progress has been made in the treatment of RA with traditional Chinese medicine. Hedyotis Diffusa Willd (HDW) is a member of the Rubiaceae family of Chinese herbal remedies and is mainly found in the southeastern provinces of China11. Modern pharmacological studies have shown that HDW exhibits multiple pharmacological effects, including anti-tumor, anti-inflammatory, anti-oxidation, anti-fibroblastic, hepatoprotective and immunomodulatory12,13. It has been widely studied as a potential therapeutic drug for treatment of malignant tumors of the breast, stomach, colon, rectum, cervix, and ovary14,15,16. It has also been used in the treatment of inflammation-related diseases, including urinary tract infection, colitis, tonsillitis, appendicitis, pharyngitis, hepatitis, dysentery, diarrhea, and snake bites17,18. Studies have shown that the certain chemical constituents of HDW (scandoside, asperuloside and asperulosidic acid) exerted an anti-inflammatory effect on LPS-induced RAW 264.7 macrophages by suppressing the NF-κB and MAPK signaling pathways19,20. In a complete Freund's adjuvant (CFA)-induced arthritis model in rats, 12 days of oral treatment with HDW extract ursolic acid (50 mg/kg/day) was demonstrated to suppress paw swelling, plasma PGE (2) production, spinal Fos expression, and arthritis-induced mechanical and thermal hyperalgesia21. Zhu et al.22,23 also revealed that HDW compounds ferulic acid and p-coumaric acid demonstrated an anti-inflammatory effect on collagen-induced arthritis as indicated by decreased numbers of inflammatory cells and reduced levels of IL-1β and TNF-α. Intriguingly, unfractionated HDW had a better therapeutic outcome than ferulic acid, although a poorer one than p-coumaric acid alone. A recent study has shown that HDW effectively suppressed the progression of disease in a collagen-induced arthritis (CIA) model by reducing the arthritis index, by reducing levels of IL-lβ, TNF-α, PGE2, RANKL, OPG, and RANKL/OPG, and by increasing the pain threshold24. Nevertheless, despite extensive studies on the pharmacological effect of HDW, the potential targets and underlying molecular mechanism(s) of HDW in RA remain unclear.
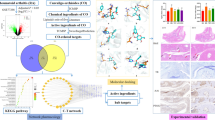
Network pharmacology is a novel approach that combines system network analysis and pharmacology25. Through network pharmacological analyses, we can investigate TCM systematically, identify the active components, predict potential targets and mechanisms, and provide the opportunity for modernization of TCM26. Therefore, our study aimed to investigate the active ingredients, potential targets, and the underlying mechanism of HDW for the treatment of RA by adopting a network pharmacology approach, molecular docking and cell experiments. The main scheme of this study is presented in Fig. 1. We screened public databases (TCMSP) and published literature to identify the active ingredients of HDW. Then, network pharmacology was used to analyze ingredients targets, drug targets, biological processes and pathways in RA treatment. In addition, we also used cell experiments to identify the results of network pharmacology.
Results
Active components and potential targets of HDW
Our study performed network pharmacology prediction based on network pharmacology evaluation method guidance‐Draft27. A total of 142 related components of HDW was retrieved from TCMSP and the published literature. According to pharmacokinetic characteristics (OB ≥ 30% and DL ≥ 0.18) and ADME information, 11 active components were selected from 142 ingredients of HDW. The TCMSP and Swiss Target Prediction databases were used to determine the pharmacological targets of the HDW components. Table 1 shows active components and the number of the corresponding potential targets of HDW. Detailed information of these components and targets is listed in Supplementary Table 1. Eventually, 180 potential targets were identified (after removing duplicates) using the Uniprot database.
Identification of the potential targets of RA
Using “Rheumatoid Arthritis” as the search term, 42, 192, 141, 174, and 623 disease-targets were obtained from the OMIM, DrugBank, TTD, GeneCards, and DisGeNET databases, respectively. Merging all results from the five databases and removing duplicates, 942 related-RA potential targets were finally collected.
Construction of the active components-common targets-RA network
Applying a Venn diagram, 85 common targets were found overlapped between HDW compound targets and RA-related targets (Fig. 2A). We imported 11 active components and 85 common targets into Cytoscape 3.9.0 software to construct a components-targets-RA network. Among these, the active ingredients with the highest degree value were stigmasterol, β-sitosterol, quercetin, kaempferol and 2-methoxy-3-methyl-9,10-anthraquinone. However, poriferasterol and scopoletin were removed since they lacked common targets in the network. The active ingredients-targets-RA network is shown in Fig. 2B. Results indicate that 5 components may provide the key to successful treatment of RA.
GO and KEGG enrichment analysis
In total, GO analysis identified 1542 significantly enriched GO terms (P.adjusted < 0.01 adjusted with Benjamini–Hochberg), consisting mainly of 1459 biological processes, 18 cellular components, and 65 molecular functions. We screened the top 10 ranked GO terms shown in Fig. 3A. In the biological process (GO:BP) category, the top terms were involved in responses to lipopolysaccharides, molecules of bacterial origin, and reactive oxygen species metabolic processes. In the cellular component (GO:CC) category, the top terms included membrane rafts, membrane microdomains, and membrane regions. In the molecular function (GO:MF) category, the top terms consisted of nuclear receptor activity, transcription factor activity, and cytokine receptor binding. To further identify underlying signaling pathways, we analyzed KEGG pathways. The top 20 significantly enriched pathways (P.adjusted value < 0.01) are shown in Fig. 3B. A list of genes contributing to the 20 selected pathways is provided in Supplementary Table 2. Numerous targets were found associated with the AGE-RAGE, TNF, IL17, and PI3K-Akt signaling pathways, all of which are associated with the prognosis and onset of RA.
GO and KEGG analysis of potential targets. (A) Different colors represent different categories. The height of the column represents the P.adjust value: the higher the value, the higher the reliability of the GO categories (P.adjust < 0.01). (B) Dot size indicates the number of target genes in the pathway, and dot color reflects the different P.adjsted value ranges.
Network visualization and identification of hub targets
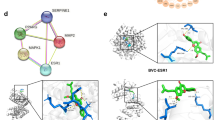
Next, we analyzed 85 potential therapeutic targets by using the STRING database to obtain a PPI network to explore the relationship between RA-related targets. The PPI relationship network, with a total of 85 nodes, 238 edges and an average node degree of 5.6 was generated with a confidence of 0.9 (Supplementary Fig. 1). PPI network diagrams were imported into Cytoscape 3.9.0 software for visualization (Fig. 4A). We further identified the subnetwork and hub targets from the PPI network using the CytoNCA plug-in (Fig. 4B). As shown in Fig. 4C, a subnetwork was identified, including 8 nodes and 27 edges. Moreover, RELA, TNF, IL6, TP53, MAPK1, AKT1, IL10, and ESR1 were identified as the hub targets in the HDW for RA treatment (Supplementary Table 3).
Construction of the PPI network and screening hub targets. (A) PPI network of 85 common targets. (B) Hub genes were screened from the PPI network using the Betweenness (BC), Closeness (CC), Degree (DC), and Network (NC) methods. (C) Subnetwork of the PPI network of 8 hub targets. The color and size of the nodes reflect the degree value for each protein target: the larger and darker the node, the greater the degree value. The different colored lines in the figure represent known interactions and predicted interactions (light blue: from curated databases; dark purple: experimentally determined; green: gene neighborhood; red: gene fusions; dark blue: gene co-occurrence; yellow: textmining; black: co-expression; light purple: protein homology).
Molecular docking
Candidate compounds stigmasterol, β-sitosterol, quercetin and kaempferol, and 2-methoxy-3-methyl-9,10-anthraquinone, are the top 5 (ranked by degree) in the compounds-targets-RA network. The hub targets, RELA, TNF, IL6, TP53, MAPK1, AKT1, IL10, and ESR1, play a significant role in the action of HDW against RA. Molecular docking of the 5 compounds and 8 hub genes revealed binding energies shown in Fig. 5. Five components of HDW exhibited strong binding to the 8 core targets with β-sitosterol showing the highest binding energy. these results imply that treatment with HDW may affect all the Figure targets in RA patients. The target proteins and the small molecules with strong binding affinity were visualized by PyMoL software (Fig. 6).
Docking complexes of ligand and receptor proteins and their binding residues are shown using PYMOL software. (A) RELA and β-sitosterol. (B) TNF and quercetin. (C) IL6 and quercetin. (D) TP53 and stigmasterol. (E) MAPK1 and β-sitosterol. (F) AKT1 and quercetin. (G) IL10 and β-sitosterol. (H) ESR1 and β-sitosterol.
Cell experiments
Overproliferation of fibroblast-like synoviocytes (FLS) are important pathogenesis of RA. Therefore, we investigated the effect of HDW at different concentrations (0, 0.5, 1, 2 mg/mL) on the proliferation of MH7A cells after 48 h. CCK-8 assay showed that HDW inhibited the proliferation of MH7A cells in a dose-dependent (Fig. 7A). According to the results, we chose to perform subsequent experiments with the dose of 0.5, 1, and 2 mg/mL. The WB result showed that HDW could inhibit the PI3K/AKT pathway by reducing phosphorylation of AKT in MH7A cells (P < 0.05, Fig. 7B). Hub targets in our work were verified by RT-qPCR and results showed that RELA, TNF, and IL6 were up-regulated while IL10 was down-regulated (Fig. 7C). These results validated our network pharmacology analysis, suggesting that HDW can play a role in treating RA by regulating PI3K/AKT signaling pathway and RA-related targets.
Cell experiments validate results of network pharmacology. (A) CCK8 assays of different HDW concentrations (0, 0.5, 1, 2 mg/mL) incubated MH7A cells for 48 h. (B) The expression levels of AKT and p-AKT were measured using western blotting. Original blots are presented in Supplementary Fig. 2. (C) The effect of HDW on the mRNA levels of RELA in MH7A cells. Compared with control (0 mg/mL), *P < 0.05, **P < 0.01, ***P < 0.001, ****P < 0.0001.
Discussion
Given their potent anti-inflammatory, anti-fibroblastic, and immunomodulatory actions, HDW and its components have been widely used in animal models over the last several years as an intervention for RA12,13,19,20,21,22,23. Unfortunately, the potential targets and molecular mechanism of HDW against RA are still inadequately understood. TCM network pharmacology emerging recently has become a flourishing field in TCM modern studies along with the rapid progress of bioinformatics. The study of TCM and biological network appeared for the first time in 200728, and some applications of traditional medicine network pharmacology for herbs or herbal formulae in RA29. In the present network pharmacological analysis, a total of 142 compounds of HDW were identified from TCMSP and published literature, and 11 compounds were selected by TCMSP and ADME criteria screening. A total of 180 targets related to potential compounds and 942 targets associated with RA were identified, and 85 common target genes were obtained from the overlapping part of identified compounds and RA. The components-targets-RA network analysis visualized the interaction of multi-components and multitargets about HDW on RA. The compounds targets network analysis indicated that the 5 compounds, including stigmasterol, β-sitosterol, quercetin, kaempferol, and 2-methoxy-3-methyl-9,10-anthraquinone, were linked to ≥ 10 target genes, and the 8 target genes (RELA, TNF, IL6, TP53, MAPK1, AKT1, IL10, and ESR1) were core target genes in the network. GO enrichment analysis indicated that numerous targets are involved in response to lipopolysaccharides and molecules of bacterial origin in BP, are localized to membrane rafts and membrane microdomains in CC, and are associated with nuclear receptor and transcription factor activities in MF. KEGG pathway analysis indicated that numerous targets are associated with certain inflammatory events and cancer. Molecular docking showed that stigmasterol, β-sitosterol, quercetin, kaempferol, and 2-methoxy-3-methyl-9,10-anthraquinone have good binding activity with RELA, TNF, IL6, TP53, MAPK1, AKT1, IL10, and ESR1 targets. Finally, the molecular mechanisms of HDW predicted by network pharmacology approach against RA were validated by in vitro experiments.
A potential components-targets-RA target network indicated that stigmasterol, β-sitosterol, quercetin, kaempferol, and 2-methoxy-3-methyl-9,10-anthraquinone, are likely to play vital roles in the process of RA treatment (Table 2). Indeed, apart from 2-methoxy-3-methyl-9,10-anthraquinone, all these components have previously been reported to exhibit potential antirheumatic therapeutic activity. For example, stigmasterol has been shown to protect CIA rats by suppressing proinflammatory mediators (TNF-α, IL-6, IL-1β, iNOS and COX-2) and increasing anti-inflammatory cytokine IL-1030. β-sitosterol exerts an inhibitory influence on synovial angiogenesis by suppressing endothelial cell proliferation and migration, thereby alleviating joint swelling and bone destruction in CIA mice31, and quercetin inhibits the release of proinflammatory cytokines (IL-6, TNF-α, IL-1β, IL-8, IL-13, IL-17) by activating SIRT1, thereby becoming a potential effector of RA32,33. Kaempferol inhibits the proliferation and migration of RA-FLSs and the release of activated T-cell-mediated inflammatory cytokines by suppressing fibroblast growth factor receptor 3-ribosomal S6 kinase 2 (FGFR3-RSK2) signaling34. Collectively, these active components exhibit antirheumatic activity by various mechanisms, including anti-inflammatory, immunoregulatory, and reduction of bone destruction. Notably, however, there have been few previous studies on the treatment of RA with stigmasterol and β-sitosterol.
KEGG pathway analysis indicated that the 85 therapeutic targets were enriched in viral infection and cancer, such as Kaposi’s sarcoma-associated herpesvirus infection, human cytomegalovirus infection, human papillomavirus infection, and prostate cancer. The evidence that viral infection such as human cytomegalovirus infection35 contributes to RA is strong, and that RA is associated with an increased risk of cancer36. Additionally, the KEGG analysis results also indicated that the AGE-RAGE, TNF, IL-17, and PI3K-Akt signaling pathways may be critical in the network pharmacology. It has been shown that the PI3K/AKT signaling pathway, the key molecular mechanism of the occurrence and development of RA, affects synovial inflammation37, synovial angiogenesis38, chondrocyte proliferation, apoptosis and autophagy39, and migration and invasion of fibroblast-like synoviocytes in RA40. The WB result showed that HDW could inhibit the PI3K/AKT pathway by reducing phosphorylation of AKT in MH7A cells.
Construction of the common target-related PPI network permitted screening of the top 8 core genes: RELA (transcription factor p65), TNF (tumor necrosis factor), IL6 (interleukin-6), TP53 (cellular tumor antigen p53), MAPK1 (mitogen-activated protein kinase 1), AKT1(RAC-α serine/threonine-protein kinase), IL10 (interleukin-10), and ESR1(estrogen receptor). RELA, ranking first with the highest connection in the PPI network, is a subunit of nuclear factor (NF)-κB and is regarded as an important member of the NF-κB pathway41. RELA participates in essential life activities, including cell proliferation, transformation, ap-optosis, inflammation and immune response and has been confirmed as an important component of the pathogenesis of RA42. TNF is an important therapeutic target for RA and TNF agents were the first molecular targeting drugs developed for the treatment of RA43,44. IL6 and IL 10 are critical for the development and progression of RA by affecting the inflammatory process, osteoclast-mediated bone resorption and pannus development45,46. RT‐qPCR results showed that RELA, TNF, and IL6 were up‐regulated while IL10 was down‐regulated with HDW treatment. However, there were no significant differences in other four targets, including TP53, MAPK1, AKT1, and ESR1. Studies with TP53, MAPK1, AKT1, and ESR1 have focused on cancer, whereas those on RA have focused on deficiency. Of course, it is possible that the level of mRNA observed may not accurately reflect the level of protein present. As such, whether HDW can directly influence these hub genes requires further study. The antirheumatic effect of compounds such as quercetin and kaempferol is partially associated with these target genes. For example, quercetin, a natural substance, can induce mitochondrial apoptosis of RA-FLSs through the p53 mechanism47. These results therefore illustrate the range of interactions between the multi-components and multi-targets of HDW in the treatment of RA.
As noted above, RELA, TNF, IL6, TP53, MAPK1, AKT1, IL10, and ESR1 are the top 8 hub genes in the PPI network. Additionally, however, potential targets were found to be involved in multiple pathways in the KEGG enrichment analysis. This indicates that HDW may exert its anti-RA effects by the combined interaction of multi-pathways and multi-targets. Also, the molecular docking analysis was carried out to investigate the interaction of 5 compounds and 8 targets. For example, the absolute value of binding energy about β-sitosterol and ESR1 is the highest in each group, indicating that β-sitosterol has a higher binding affinity than other target genes. Although there have been relatively few studies with stigmasterol and RA, the docking results indicated that stigmasterol performed good binding activity with TP53, ESR1, and IL10. In brief, the higher the binding affinity of components and targets, the more likely HDW treatment of RA will be achieved by modulating several related targets. However, further experiments are needed to clarify active compounds and mechanisms of action analysis of HDW for RA.
Methods
Active components and potential targets of HDW
Active components and potential targets of HDW were selected from the Traditional Chinese Medicine Systems Pharmacology Database and Analysis Platform (TCMSP, https://old.tcmsp-e.com/tcmsp.php)55. As recommended by TCMSP, the compounds with OB ≥ 30% have good absorption and slow metabolism after oral administration, while the compounds with DL ≥ 0.18 were chemically suitable for drug development. We therefore used OB ≥ 30% and DL ≥ 0.18 as filtering for thresholds56. An exhaustive search for components was performed using PubMed (https://pubmed.ncbi.nlm.nih.gov/) and CNKI (https://www.cnki.net/). The active components of HDW were determined by SwissADME database (http://www.swissadme.ch/). Afterward, the targets (probability ≥ 0.9) corresponding to the components were screened from the SwissTargetPrediction database (http://swisstargetprediction.ch/)57. Duplicates were deleted to obtain potential targets for further analysis.
Potential targets of rheumatoid arthritis
Potential RA-related therapeutic targets were collected using keywords “rheumatoid arthritis” as a query from the OMIM (https://www.omim.org/)58, DrugBank (https://go.drugbank.com/)59, TTD (http://db.idrblab.net/ttd/)60, GeneCards (https://www.genecards.org/)61 and DisGeNET (https://www.disgenet.org/) databases62. All acquired targets were imported into UniProt (https://www.uniprot.org/) for normalization and removal of duplicate and erroneous targets.
Common targets and construction of the compound-target-RA network
Targets related to the active components of HDW were mapped to RA-related targets to identify ones common to both. These were considered as targets of HDW on treating RA. Following compilation of these, the “compound-target-RA” network was established using Cytoscape 3.9.0 software to determine the interaction relationships63 and to further screen for key active components by calculating the extent of these.
Gene ontology and pathway enrichment analysis
Gene Ontology (GO) analysis and KEGG pathway analysis were performed using the R 3.6.3 and related R packages (clusterProfiler, Org.Hs.eg.db, and ggplot2)64. After adjusting the p-value using the Benjamini–Hochberg (BH) method, P.adjusted values < 0.01 were considered to be statistically significant.
Network visualization and identification of hub targets
To further identify interactive relationships among the common target proteins, the protein list was mapped to the STRING database (https://cn.string-db.org/)65. To ensure reliability, we used a cutoff ≥ 0.9 (high-confidence interaction score) to obtain the significant PPIs in network visualization. The CytoNCA plug-in was used to explore hub targets, and the top 8 were generated using Betweenness (BC), Closeness (CC), Degree (DC), and Network (NC) methods.
Molecular docking
To investigate the associations between the Active Components and hub targets, we applied molecular docking analysis. The mol2 structure files of the 5 main components were downloaded from TCMSP. The crystal structure of the 8 hub targets were obtained from the Protein Data Bank (PDB, https://www.rcsb.org/). Molecular docking studies were carried out with AutoDock 4.0 software. After docking, the results were carried out by sorting the binding energy predicted by docking conformations. The lower the affinity score is, the better the binding effect with a binding energy of less than − 5.0 kcal/mol defined as dependable binding. Binding modes were visualized with the program Pymol.
Cell culture and treatment
MH7A, a human RA-FLS cell line, was purchased from Jennio biological technology (Guangzhou, China). The MH7A cell was maintained in DMEM with 10% fetal bovine serum (Gibco, USA), supplemented with 1% penicillin-streptomycin (Hyclone, USA). HDW (Lot. Number: R02S11Y123184) was purchased from Shanghai Yuanye Bio-Technology.
Cell counting kit-8 (CCK-8) assay
CCK‐8 (Biyuntian, Shanghai, China) assay was used to measure cell proliferation. MH7A cells were seeded into 96-well plates at a cell density of 5 × 103 cells/well and incubated in a 5% CO2 incubator at 37 °C. After adhering to the wall, cells were treated with Bavachinin of different concentrations (0, 0.5, 1, and 2 mg/mL) for 48 h. Optical density at 450 nm was measured with a microplate reader after adding the CCK-8 solution and incubated for additional 3 h.
Quantitative real-time polymerase chain reaction (qRT-PCR)
MH7A cells were seeded in 6-well plates at 5 × 106 cells/well. After 48 h of treatment, cells were collected by trypsinization. Total RNA from MH7A cells and ankle joint tissues were extracted by employing the TRIzol reagent, and cDNA was synthesized using a reverse transcription kit. qPCR was performed using SYBR Green PCR Master Mix (Bio-Rad, USA). Finally, the relative quantity of mRNA was calculated using the 2−∆∆Ct method. The primers (synthesized in Qingke, China) used are listed in the Supplementary Table S4.
Western blotting (WB)
MH7A cells were cultured in T75 cell culture flask. After 48 h of treatment, cells were lysed with 1 × RIPA lysis buffer containing 1% of PMSF and 1% of phosphotransferase inhibitor. Protein concentration was detected by the BCA Protein Quantitative Kit (Thermo Fisher Scientific, USA). Subsequently, proteins (25 μg) were denatured by employing heating followed by electrophoresis with 10% SDS-PAGE, and were transferred to polyvinylidene fluoride membranes (Millipore, USA). The membranes were sealed with 5% skimmed milk (or 5% BSA) for 2 h at 37 °C, and incubated at 4 °C overnight with the following primary antibodies: protein kinase B (AKT), phospho-Akt (p‐AKT) and β-actin. After incubation, the membrane was further incubated with horseradish peroxidase (HRP)-conjugated secondary antibodies for 1.5 h at 37 °C. Lastly, specific bands were detected using enhanced chemiluminescence reagent (Thermo Fisher Scientific) and calculated by Image J software for the quantitative analysis of proteins.
Conclusions
In summary, this study explored the therapeutic effect and mechanism of HDW on RA through a strategy combining network pharmacology and in vivo experiment verification. Five active components and 8 core targets of HDW were finally screened. PI3K/AKT signaling pathways were enriched by KEGG analysis in addition to virus and cancer-related pathways. Molecular docking showed that the active ingredients successfully docked with related targets. Through experimental validation, we found that HDW may affect target genes RELA, TNF, IL6, and IL10 through multiple signaling pathways PI3K/AKT signaling pathways, thereby affecting the pathological process of RA and ultimately inhibiting the occurrence and development of RA. Our findings provide a theoretical basis for the use of HDW as a therapeutic for RA.
Data availability
All data generated or analysed during this study are included in its supplementary information files.
References
McInnes, I. B. & Schett, G. The pathogenesis of rheumatoid arthritis. N. Engl. J. Med. 365, 2205–2219. https://doi.org/10.1056/NEJMra1004965 (2011).
Basile, M. et al. Cognitive decline in rheumatoid arthritis: Insight into the molecular pathogenetic mechanisms. Int J. Mol. Sci. 22, 1185. https://doi.org/10.3390/ijms22031185 (2021).
Guo, Y. et al. Tryptase is a candidate autoantigen in rheumatoid arthritis. Immunology 142, 67–77. https://doi.org/10.1111/imm.12197 (2014).
Jin, S. et al. Chinese Registry of rheumatoid arthritis (CREDIT): II. Prevalence and risk factors of major comorbidities in Chinese patients with rheumatoid arthritis. Arthritis Res. Ther. 19, 251. https://doi.org/10.1186/s13075-017-1457-z (2017).
Matsumoto, T. et al. Soluble Siglec-9 suppresses arthritis in a collagen-induced arthritis mouse model and inhibits M1 activation of RAW264.7 macrophages. Arthritis Res. Ther. 18, 133. https://doi.org/10.1186/s13075-016-1035-9 (2016).
Huber, L. C. et al. Synovial fibroblasts: Key players in rheumatoid arthritis. Rheumatology (Oxford) 45, 669–675 (2006).
Noss, E. H. & Brenner, M. B. The role and therapeutic implications of fibroblast-like synoviocytes in inflammation and cartilage erosion in rheumatoid arthritis. Immunol. Rev. 223, 252–270. https://doi.org/10.1111/j.1600-065X.2008.00648.x (2008).
Olsen, N. J. & Stein, C. M. New drugs for rheumatoid arthritis. N. Engl. J. Med. 350, 2167–2179 (2004).
Shetty, A. et al. Tocilizumab in the treatment of rheumatoid arthritis and beyond. Drug Des. Dev. Ther. 8, 349–364. https://doi.org/10.2147/DDDT.S41437 (2014).
Obiri, D. D. et al. Xylopia aethiopica (Annonaceae) fruit extract suppresses Freund’s adjuvant-induced arthritis in Sprague–Dawley rats. J. Ethnopharmacol. 152, 522–531. https://doi.org/10.1016/j.jep.2014.01.035 (2014).
Pu, F. et al. The synergistic anticancer effect of cisplatin combined with Oldenlandia diffusa in osteosarcoma MG-63 cell line in vitro. Onco Targets Ther. 9, 255–263. https://doi.org/10.2147/OTT.S90707 (2016).
Zhang, R. et al. Isolation, purification, structural characteristics, pharmacological activities, and combined action of Hedyotis diffusa polysaccharides: A review. Int. J. Biol. Macromol. 183, 119–131. https://doi.org/10.1016/j.ijbiomac.2021.04.139 (2021).
Chen, R. et al. The Hedyotis diffusa Willd. (Rubiaceae): A review on phytochemistry, pharmacology quality control and pharmacokinetics. Molecules 21, 710. https://doi.org/10.3390/molecules21060710 (2016).
Wei, Z. et al. Preparation of graphene-multi-walled carbon nanotube composite for quantitative determination of 2-hydroxy-3-methylanthraquinone in Hedyotis diffusa. Int. J. Electrochem. Sci. 12, 629–638 (2017).
Qian, K. et al. Mechanism of Hedyotis diffusa in the treatment of cervical cancer. Front. Pharmacol. 12, 808144. https://doi.org/10.3389/fphar.2021.808144 (2021).
Feng, J. et al. Extract suppresses colorectal cancer growth through multiple cellular pathways. Oncol. Lett. 14, 8197–8205. https://doi.org/10.3892/ol.2017.7244 (2017).
Wang, C. et al. The antitumor constituents from Hedyotis diffusa Willd. Molecules 22, 2101. https://doi.org/10.3390/molecules22122101 (2017).
Kim, S.-J. et al. Oldenlandia diffusa ameliorates dextran sulphate sodium-induced colitis through inhibition of NF-κB activation. Am. J. Chin. Med. 39, 957–969. https://doi.org/10.1142/S0192415X11009330 (2011).
He, J. et al. Asperuloside and asperulosidic acid exert an anti-inflammatory effect via suppression of the NF-κB and MAPK signaling pathways in LPS-induced RAW 264.7 macrophages. Int. J. Mol. Sci. 19, 2027. https://doi.org/10.3390/ijms19072027 (2018).
He, J. et al. Scandoside exerts anti-inflammatory effect via suppressing NF-κB and MAPK signaling pathways in LPS-induced RAW 264.7 macrophages. Int. J. Mol. Sci. 19, 457. https://doi.org/10.3390/ijms19020457 (2018).
Kang, S.-Y. et al. The anti-arthritic effect of ursolic acid on zymosan-induced acute inflammation and adjuvant-induced chronic arthritis models. J. Pharm. Pharmacol. 60, 1347–1354. https://doi.org/10.1211/jpp/60.10.0011 (2008).
Zhu, H. et al. Anti-inflammatory effects of p-coumaric acid, a natural compound of, on arthritis model rats. Evid. Based Complement. Altern. Med. ECAM. 5198594, 2018. https://doi.org/10.1155/2018/5198594 (2018).
Zhu, H. et al. Anti-inflammatory effects of the bioactive compound ferulic acid contained in Oldenlandia diffusa on collagen-induced arthritis in rats. Evid. Based Complement. Altern. Med. ECAM. 2014, 573801. https://doi.org/10.1155/2014/573801 (2014).
Jia, P. et al. Therapeutic effects of Hedyotis diffusa Willd. on type II collagen-induced rheumatoid arthritis in rats. Zhongguo Ying Yong Sheng Li Xue Za Zhi = Chin. J. Appl. Physiol. 34, 558–561. https://doi.org/10.12047/j.cjap.5665.2018.125 (2018).
Ning, K. et al. Computational molecular networks and network pharmacology. Biomed. Res. Int. 2017, 7573904. https://doi.org/10.1155/2017/7573904 (2017).
Niu, B. et al. Network pharmacology study on the active components of and the mechanism of their effect against cerebral ischemia. Drug Des. Dev. Ther. 13, 3009–3019. https://doi.org/10.2147/DDDT.S207955 (2019).
Li, S. et al. Network pharmacology evaluation method guidance-Draft. World J. Tradit. Chin. Med. 7, 146–154 (2021).
Li, S. et al. Understanding ZHENG in traditional Chinese medicine in the context of neuro-endocrine-immune network. IET Syst. Biol. 1, 51–60 (2007).
Li, S. & Zhang, B. Traditional Chinese medicine network pharmacology: Theory, methodology and application. Chin. J. Nat. Med. 11, 110–120. https://doi.org/10.1016/S1875-5364(13)60037-0 (2013).
Ahmad Khan, M. et al. Stigmasterol protects rats from collagen induced arthritis by inhibiting proinflammatory cytokines. Int. Immunopharmacol. 85, 106642. https://doi.org/10.1016/j.intimp.2020.106642 (2020).
Liu, R. et al. β-Sitosterol modulates macrophage polarization and attenuates rheumatoid inflammation in mice. Pharm. Biol. 57, 161–168. https://doi.org/10.1080/13880209.2019.1577461 (2019).
Goyal, A. & Agrawal, N. Quercetin: A potential candidate for the treatment of arthritis. Curr. Mol. Med. https://doi.org/10.2174/1566524021666210315125330 (2021).
Kim, H.-R. et al. Quercetin, a plant polyphenol, has potential for the prevention of bone destruction in rheumatoid arthritis. J. Med. Food 22, 152–161. https://doi.org/10.1089/jmf.2018.4259 (2019).
Lee, C.-J. et al. Kaempferol targeting on the fibroblast growth factor receptor 3-ribosomal S6 kinase 2 signaling axis prevents the development of rheumatoid arthritis. Cell Death Dis. 9, 401. https://doi.org/10.1038/s41419-018-0433-0 (2018).
Davignon, J.-L. et al. Cytomegalovirus infection: Friend or foe in rheumatoid arthritis?. Arthritis Res. Ther. 23, 16. https://doi.org/10.1186/s13075-020-02398-3 (2021).
Bhandari, B. et al. Prevalence of cancer in rheumatoid arthritis: Epidemiological study based on the national health and nutrition examination survey (NHANES). Cureus 12, e7870. https://doi.org/10.7759/cureus.7870 (2020).
Tu, Y. et al. Glytabastan B, a coumestan isolated from Glycine tabacina, alleviated synovial inflammation, osteoclastogenesis and collagen-induced arthritis through inhibiting MAPK and PI3K/AKT pathways. Biochem. Pharmacol. 197, 114912. https://doi.org/10.1016/j.bcp.2022.114912 (2022).
Ao, L. et al. Matrine inhibits synovial angiogenesis in collagen-induced arthritis rats by regulating HIF-VEGF-Ang and inhibiting the PI3K/Akt signaling pathway. Mol. Immunol. 141, 13–20. https://doi.org/10.1016/j.molimm.2021.11.002 (2022).
Feng, F.-B. & Qiu, H.-Y. Effects of Artesunate on chondrocyte proliferation, apoptosis and autophagy through the PI3K/AKT/mTOR signaling pathway in rat models with rheumatoid arthritis. Biomed. Pharmacother. 102, 1209–1220. https://doi.org/10.1016/j.biopha.2018.03.142 (2018).
Ma, J.-D. et al. A novel function of artesunate on inhibiting migration and invasion of fibroblast-like synoviocytes from rheumatoid arthritis patients. Arthritis Res. Ther. 21, 153. https://doi.org/10.1186/s13075-019-1935-6 (2019).
Chen, S. et al. RelA/p65 inhibition prevents tendon adhesion by modulating inflammation, cell proliferation, and apoptosis. Cell Death Dis. 8, e2710. https://doi.org/10.1038/cddis.2017.135 (2017).
Giridharan, S. & Srinivasan, M. Mechanisms of NF-κB p65 and strategies for therapeutic manipulation. J. Inflamm. Res. 11, 407–419. https://doi.org/10.2147/JIR.S140188 (2018).
Yamanaka, H. TNF as a target of inflammation in rheumatoid arthritis. Endocr. Metab. Immune Disord. Drug Targets 15, 129–134 (2015).
Feldmann, M. Development of anti-TNF therapy for rheumatoid arthritis. Nat. Rev. Immunol. 2, 364–371. https://doi.org/10.1038/nri802 (2002).
Mollazadeh, H. et al. Immune modulation by curcumin: The role of interleukin-10. Crit. Rev. Food Sci. Nutr. 59, 1358139. https://doi.org/10.1080/10408398.2017.1358139 (2019).
Pandolfi, F. et al. Interleukin-6 in rheumatoid arthritis. Int. J. Mol. Sci. 21, 5238. https://doi.org/10.3390/ijms21155238 (2020).
Xiao, P. et al. p53 contributes to quercetin-induced apoptosis in human rheumatoid arthritis fibroblast-like synoviocytes. Inflammation 36, 272–278. https://doi.org/10.1007/s10753-012-9543-5 (2013).
Qian, K. et al. β-Sitosterol inhibits rheumatoid synovial angiogenesis through suppressing VEGF signaling pathway. Front. Pharmacol. 12, 816477. https://doi.org/10.3389/fphar.2021.816477 (2021).
Shen, P. et al. Quercetin-mediated SIRT1 activation attenuates collagen-induced mice arthritis. J. Ethnopharmacol. 279, 114213. https://doi.org/10.1016/j.jep.2021.114213 (2021).
Yang, Y. et al. Quercetin attenuates collagen-induced arthritis by restoration of Th17/Treg balance and activation of Heme Oxygenase 1-mediated anti-inflammatory effect. Int. Immunopharmacol. 54, 153–162. https://doi.org/10.1016/j.intimp.2017.11.013 (2018).
Saccol, R. D. S. P. et al. Effect of quercetin on E-NTPDase/E-ADA activities and cytokine secretion of complete Freund adjuvant-induced arthritic rats. Cell Biochem. Funct. 37, 474–485. https://doi.org/10.1002/cbf.3413 (2019).
Pan, D. et al. Kaempferol inhibits the migration and invasion of rheumatoid arthritis fibroblast-like synoviocytes by blocking activation of the MAPK pathway. Int. Immunopharmacol. 55, 174–182. https://doi.org/10.1016/j.intimp.2017.12.011 (2018).
Lin, F. et al. Kaempferol enhances the suppressive function of Treg cells by inhibiting FOXP3 phosphorylation. Int. Immunopharmacol. 28, 859–865. https://doi.org/10.1016/j.intimp.2015.03.044 (2015).
Yoon, H.-Y. et al. Kaempferol inhibits IL-1β-induced proliferation of rheumatoid arthritis synovial fibroblasts and the production of COX-2, PGE2 and MMPs. Int. J. Mol. Med. 32, 971–977. https://doi.org/10.3892/ijmm.2013.1468 (2013).
Ru, J. et al. TCMSP: A database of systems pharmacology for drug discovery from herbal medicines. J. Cheminform. 6, 13. https://doi.org/10.1186/1758-2946-6-13 (2014).
Liu, H. et al. Systems approaches and polypharmacology for drug discovery from herbal medicines: An example using licorice. J. Ethnopharmacol. 146, 773–793. https://doi.org/10.1016/j.jep.2013.02.004 (2013).
Daina, A. et al. SwissTargetPrediction: Updated data and new features for efficient prediction of protein targets of small molecules. Nucleic Acids Res. 47, W357–W364. https://doi.org/10.1093/nar/gkz382 (2019).
Hamosh, A. et al. Online mendelian inheritance in man (OMIM). Hum. Mutat. 15, 57–61 (2000).
Wishart, D. S. et al. DrugBank 5.0: A major update to the DrugBank database for 2018. Nucleic Acids Res. 46, D1074–D1082. https://doi.org/10.1093/nar/gkx1037 (2018).
Li, Y. H. et al. Therapeutic target database update 2018: Enriched resource for facilitating bench-to-clinic research of targeted therapeutics. Nucleic Acids Res. 46, D1121–D1127. https://doi.org/10.1093/nar/gkx1076 (2018).
Stelzer, G. et al. The GeneCards Suite: From gene data mining to disease genome sequence analyses. Curr. Protoc. Bioinform. 54, 1.30.1-1.30.33. https://doi.org/10.1002/cpbi.5 (2016).
Piñero, J. et al. The DisGeNET knowledge platform for disease genomics: 2019 update. Nucleic Acids Res. 48, D845–D855. https://doi.org/10.1093/nar/gkz1021 (2020).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498–2504 (2003).
Yu, G. et al. clusterProfiler: An R package for comparing biological themes among gene clusters. OMICS 16, 284–287. https://doi.org/10.1089/omi.2011.0118 (2012).
Szklarczyk, D. et al. The STRING database in 2017: Quality-controlled protein-protein association networks, made broadly accessible. Nucleic Acids Res. 45, D362–D368. https://doi.org/10.1093/nar/gkw937 (2017).
Acknowledgements
The authors wish to thank Chengdu Medical College for assistance during the present study.
Funding
This work was supported by grants from the National Natural Science Foundation of China (31701104), 2022 graduate innovation fund project (YCX2022-03-20) and National Undergraduate Innovation and Entrepreneurship Program (202213705017).
Author information
Authors and Affiliations
Contributions
Conceptualization, H.-D. and W.-K.S.; Investigation, H.-D.; Methodology and experiment validation, H.-D. and S.-S.Z.; Supervision, W.-K.S.; Visualization, H.-D. and J.J.; Writing—Original draft, H.-D.; Writing—Review and editing, W.-K.S., Q.-L.Z., and L.-J.W. All authors have read and agreed to the published version of the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Deng, H., Jiang, J., Zhang, S. et al. Network pharmacology and experimental validation to identify the potential mechanism of Hedyotis diffusa Willd against rheumatoid arthritis. Sci Rep 13, 1425 (2023). https://doi.org/10.1038/s41598-022-25579-3
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-022-25579-3
- Springer Nature Limited