Abstract
Nasopharyngeal carcinoma (NPC) is a cancer with a high metastatic rate and poor prognosis. Growing studies suggest that ferroptosis take part in the development of tumours. At the same time, the connection between ferroptosis-related genes (FRGs) and the prognosis of NPC remains unclear. In this study, we explored the dysregulated FRGs between normal control and tumour samples of NPC. Firstly, 14 of 36 differentially expressed FRGs were identified in NPC tissues compared to normal tissues, among which ABCC1, GLS2, CS and HMGCR were associated with poor prognosis for patients. The four ferroptosis genes were used for consensus cluster analysis and two risk-related FRGs (ABCC1 and GLS2) were used in a risk model. The ROC curve revealed the good predictive performance of this risk signature. Multivariate analysis revealed that risk score and intratumoral TILs were independent risk factors linked to prognosis. Additionally, our results suggested that the risk signature was attached to the immune microenvironment. Moreover, the NPC patients with high risk were sensitive to chemotherapeutic drugs including axitinib, docetaxel, embelin, epothilone.B, parthenolide, thapsigargin, tipifarnib, vinorelbine. Finally, the expression of ABCC1 and GLS2 was validated in NPC tissues using immunohistochemistry. Together, these results revealed ferroptosis may be a potential biomarker in NPC and representing a promising future direction in prognosis and therapeutic strategy for the treatment of NPC.
Similar content being viewed by others
Introduction
Nasopharyngeal carcinoma (NPC) is a cancer with a poor prognosis and high metastasis rate that frequently occurs in southern China1,2. Every year, there are over 133,000 new cases, resulting in 80,000 deaths from NPC in 20203. The incidence of NPC has been increasing annually. Epstein‒Barr virus (EBV) infection, preserved foods and alcohol consumption are important risk factors for developing NPC4,5. Despite the availability of conventional treatments, such as surgery, radiotherapy and chemotherapy, the risk of recurrence and metastasis remains high6. Therefore, it is vital to understand its pathogenesis and identify novel and reliable targets related to prognostic risk stratification for treating NPC.
Ferroptosis is a new cell death mode, characterized as an oxidative cell death induced by small molecules in certain circumstances, dependent on iron ions7. Ferroptosis is caused by an imbalance in the generation and degradation of intracellular reactive oxygen species (ROS)8. Several studies have confirmed that FRGs are associated with the clinical diagnosis, prognosis and progression of various cancers, including breast carcinoma, colorectal carcinoma, lung adenocarcinoma and glioma9,10,11. ABCC1 is involved in tumor progression, drug resistance and poor patient survival in classical Hodgkin lymphoma (CHL)12. GLS2 participates in p53-mediated ferroptosis and is related to hepatocellular carcinoma, cervical carcinoma and lung adenocarcinoma. Given the high iron levels of cancer cells and their ferroptosis sensitivity, ferroptosis has been proposed to be promising for tumor therapeutics13.
The tumor microenvironment (TME) is a unique internal environment that includes macrophages, stromal cells, tumor-infiltrating lymphocytes, and some immune cells14. It plays a vital role in tumorigenesis and therapeutic strategies15,16. Immunotherapy is a recently developed modality for treating recurrent or metastatic disease and has excellent clinical value17. Recent studies have shown cooperation between tumor ferroptosis and the TME. Wang et al. found that CD8 + T cells participate in antitumor activity by promoting tumor ferroptosis, which may be a potential therapeutic strategy18,19. However, the relationship between ferroptosis and the immune microenvironment/immunotherapy of NPC patients is unclear.
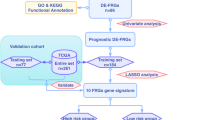
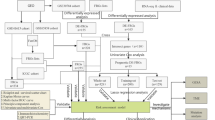
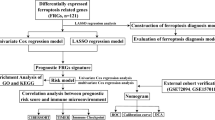
In this study, we comprehensively explored the expression and prognosis of FRGs from 3 NPC Gene Expression Omnibus (GEO) datasets and designed a risk signature containing 2 FRGs. Next, we researched the communication between the FRG risk signature and immune infiltration. We then investigated the correlation between the risk score and drug sensitivity for NPC therapy. Finally, the expression of ABCC1 and GLS2 was validated in tissues using immunohistochemistry (IHC).
Results
FRGs related to survival in NPC
To investigate the regulation of FRGs, we downloaded GSE53819 and GSE12452 from the GEO database. As shown in Fig. 1A, 25 differentially expressed FRG genes (DE_FRGs) were observed in GSE12452. In addition, 31 DE_FRGs were obtained in GSE53819 (Fig. 1B). Moreover, 14 DE_FRGs overlapped in both GSE12452 and GSE53819 (Fig. 2A). Next, we analyzed the potential prognostic role of 14 FRGs in GSE102349. We determined that 4 FRGs (ABCC1, CS, GLS2 and HMGCR) were associated with poor PFS in NPC (Fig. 2B).
The expression profiles of FRGs in NPC. (A) The heatmap of DE_FRGs in GSE53819 and GSE12452 using “edgeR” package (version 4.0.5, https://cran.r-project.org/). (B) The gene expression levels of FRGs in GSE53819 and GSE12452 from GEO (https://www.ncbi.nlm.nih.gov/geo/).
Estimation of consensus clustering of ferroptosis genes
Next, consensus clustering was performed to analyze four prognosis-related ferroptosis genes (ABCC1, HMGCR, CS and GLS2), and k = 3 may be the best choice in the GSE102349 datasets (Fig. 3A–C). The PCA and Kaplan‒Meier curve analysis of poor progression-free survival suggested that three clusters could be distinguished (Fig. 3D,E). The heatmap revealed that HMGCR and GLS2 were upregulated in the Cluster 2 and 3 subgroups, ABCC1 was upregulated in the Cluster 2 subgroup, and CS was upregulated in the Cluster 3 subgroup (Fig. 3F). Moreover, the correlation analysis of clustering and risk classification was performed using ggalluvial (Fig. 3G), and the results revealed that most patients in Cluster 2 and Cluster 3 were grouped into the high-risk group and were associated with a high proportion of NPC deaths. These results indicated that the expression of ferroptosis genes was associated with cluster subgroups and clinical characteristics.
Consensus Clustering of FRGs of NPC. (A) Consensus clustering analysis using the R package “ConsensusClusterPlus” (https://cran.r-project.org/); (B,C) Consensus clustering cumulative distribution function; (D) PCA analysis for three clusters; (E) K-M curves of poor progression-free survival for three clusters; (F) The heatmap show the four clusters’ clinical characteristics of FRG. (G) Correlation analysis of clustering and risk classification using the R package “ggalluvial” and “ggplot2” (https://cran.r-project.org/).
Prognostic risk model of FRGs in NPC
Next, LASSO and Cox regression analyses were performed to identify an FRG signature for predicting the prognosis of NPC. As shown in Fig. 4A and B, two FRGs (ABCC1 and GLS2) were used to establish the risk model, and the risk score was ABCC1 × 0.02775 + GLS2 × 0.0592. The distributions of risk scores, survival times and expression profiles were based on the two FRGs (Fig. 4C,D). The ROC curve analysis showed that the area under the curve (AUC) of 3-year OS was 0.837 (Fig. 4E). The results indicated that the risk score model exhibited stable performance.
Risk model from three FRGs. (A) Univariate Cox analysis between four FRGs and poor progression-free survival of NPC using the R package “ConsensusClusterPlus”. (B) LASSO calculated the risk model using the R package “glmnet” (https://cran.r-project.org/src/contrib/Archive/glmnet/). (C) The distributions from two prognostic FRGs. (D) Kaplan–Meier poor progression-free survival curves for patients in risk score. (E) ROC curve showed the efficiency of the risk score model using the R package “survivalROC”.
Prognostic signature of FRGs in NPC
To explore the independent prognostic signature in NPC, we evaluated the predictive value of clinical stage, stromal TILs, intratumoral TILs, differentiation and risk scores by performing univariate and multivariate analyses. Univariate analysis showed that stromal TILs and the risk score were significantly associated with progression-free survival (PFS). Multivariate Cox analysis showed that the risk score and intratumoral TILs were independent prognostic signatures (Fig. 5A). The ROC analysis showed that the AUC values were 0.92, 0.847, and 0.817 for 1-, 2-, and 3-year OS, respectively (Fig. 5B). Then, we constructed a nomogram to integrate risk scores, stromal TILs and intratumoral TILs, which can predict the probability of survival for NPC patients (Fig. 5C).
The correlation between risk score and clinical features
To reveal the communication between the risk score and clinical data of NPC, the correlation between clinical features (including the expression-based (EB) subtype, stage, survival status and differentiation) and the risk score was studied. The results revealed that patients with EB subtype I/II had a higher risk scores than those with subtype III, while patients in the advanced stage (III/IV) had a higher risk scores than those in the early stage (I/II). The risk score in deceased patients was higher than that in living patients. In contrast, the risk score between the differentiated and undifferentiated groups showed no significant differences. Together, the results indicated that the risk score was related to EB subtype, clinical stage and survival status (Fig. 6).
Risk-related signaling pathways
To investigate the potential functions of DEGs in risk-related NPC, a total of 201 upregulated (red) and 86 downregulated (green) DEGs were identified in the high-risk group (Fig. 7A). The heatmap of 287 DEGs was identified between high/low risk (Fig. 7B). The GO functional analysis suggested that the DEGs are mainly related to immune cell development, including leukocyte differentiation, lymphocyte differentiation, B-cell activation, lymphocyte proliferation, and B-cell proliferation (Fig. 7C,E). The KEGG functional enrichment analysis indicated that the DEGs were enriched in the B-cell receptor signaling pathway, PI3K-Akt signaling pathway, primary immunodeficiency pathway, Ras signaling pathway, and PPAR signaling pathway (Fig. 7D,F).
Relationship of the risk score with immune infiltration in NPC
We used the CIBERSORT method to investigate immune infiltration in NPC patients in the low- and high-risk groups. The results revealed different immune infiltration in the high- and low-risk groups, including resting DCs and follicular helper T cells, while higher infiltration scores of naive B cells and memory B cells were observed in the low-risk patients (Fig. 8A). Then, we analyzed the relationship between the risk score and the infiltration of several immune cells (Fig. 8B). The results suggested that the risk score was significantly positively correlated with plasma cells, follicular helper T cells, and resting DCs but negatively associated with memory B cells and naive B cells. Moreover, ABCC1 was positively associated with plasma cells, follicular helper T cells, resting DCs, and resting mast cells but negatively associated with memory B cells, naive B cells, and gamma delta T cells. GLS2 was strongly positively correlated with activated NK and follicular helper T cells, while it was negatively correlated with memory B cells. These results revealed that the ABCC1 and GLS2 prognostic signatures for NPC patients are associated with immune infiltration.
The relationship between risk score and chemotherapeutic drugs
To select effective anticancer drugs for NPC patients, we explored the relationships between the risk score and chemotherapeutic agents. Our study showed that high-risk NPC patients were more sensitive to agents including docetaxel, embelin, and epothilone B, parthenolide, thapsigargin, tipifarnib, axitinib, and vinorelbine, suggesting that the risk scores may be a potential indicator of chemical sensitivity in the treatment of NPC (Fig. 9).
Validation of the expression and prognosis of ABCC1 and GLS2 in NPC
We analyzed the expression of ABCC1 and GLS2 in pancancers using GEPIA2 (Figs. S1, S2). The results demonstrated that the expression levels of ABCC1 and GLS2 were diverse in various types of tumors. Since there is no NPC data in The Cancer Genome Atlas (TCGA), HNSC data were used to evaluate NPC in some articles20,21,22. Here, we found that ABCC1 was overexpressed in HNSC, which was consistent with the expression of ABCC1 in NPC. Nevertheless, contrary to our results, GLS2 expression was decreased in HNSC. Given that HNSC data are not exclusively derived from NPC samples, some variation is possible (Supplementary Figs. S1, S2).
To verify the potential effect of ABCC1 and GLS2 expression in NPC, IHC was performed to analyze ABCC1 and GLS2 expression in NPC tissues (Fig. 10A,B), and the clinical information related to 11 samples, such as sex, age, histological type, and fustat (Supplementary Table S1), was assessed. Compared with normal nasopharyngeal mucosa (NNM), the expression of ABCC1 and GLS2 was upregulated in NPC tissues (*p < 0.05; **p < 0.01) (Fig. 10C,D). This result indicated that ABCC1 and GLS2 were overexpressed in NPC. In addition, we investigated the relationship between high/low expression of ABCC1 and GLS2 and poor survival in tissues. The results showed that the patients with high expression of ABCC1 and GLS2 had a higher mortality rate than those with low expression of ABCC1 and GLS2, which indicated that a high expression of ABCC1 and GLS2 was related to poor survival, as shown in Fig. 10E.
The expression and prognosis of ABCC1 and GLS2 in NPC tissues. (A,B) The expression of ABCC1 and GLS2 in 3 nasopharyngeal carcinomas and adjacent non-tumor tissues by IHC. The scale bar was 20 μm. (C,D) Statistical analysis to assess the expression level of ABCC1 and GLS2 in NPC tissues. (E) Mortality rate testing assessed the correlation between ABCC1 and GLS2 low/high expression with the poor survival of the patients. *p < 0.05, **p < 0.01.
Discussion
Iron dependency is a characteristic of cancer cells; excessive ionic iron causes “iron enrichment” and leads to ferroptosis, which is a new cell death mode characterized by GPX4 inactivation and ROS accumulation23. Ferroptosis is involved in multiple biological processes, including degenerative diseases, carcinoma and T-cell immunity. With the continuous progress of basic and clinical research, ferroptosis has been applied to anticancer therapies, including chemotherapy, radiotherapy and immunotherapy24,25. Tumor-based ferroptosis may be a potential biomarker of cancer immunotherapy26,27. Consequently, exploring the role of ferroptosis is vital to enhancing the efficacy of cancer therapy.
This study identified four FRGs, ABCC1, GLS2, CS and HMGCR, in NPC. ABCC1 was associated with increased aggressiveness and poor survival in various tumors28,29,30. ABCC1 has been shown to be overexpressed in breast carcinoma and was correlated with poor prognosis and chemotherapeutic sensitivity31,32. Moreover, ABCC1 is associated with an increased risk of tumor progression, treatment resistance or death in CHL patients12. Our study revealed that ABCC1 was upregulated and correlated with prognosis. These results indicate that the ferroptosis-related gene ABCC1 is a risk factor and may be a promising therapeutic target for NPC. GLS2 is a mitochondrial glutaminase. It catabolizes glutamine to glutamate, which plays a vital role in cancer progression33. As a direct target gene of p53, GLS2 is associated with ferroptosis34. Previous studies suggested that GLS2 may play different roles depending on tumor type35,36. A few studies have suggested that GLS2 expression is associated with good prognosis; for example, high expression of GLS2 in HCC is associated with a long survival time of patients37. However, most studies have shown that overexpression or amplification of GLS2 was associated with poor survival prognosis, such as in breast cancer38,39, glioblastoma40, ovarian cancer41, and lung cancer34,42. In addition, GLS2 was more highly expressed and regulated by METTL3 by m6A modification, which was significantly associated with poor prognosis in esophageal squamous cell carcinoma43. These results are consistent with our results, and further studies are needed to confirm the molecular mechanisms of GLS2 in NPC. In this study, two FRGs (ABCC1 and GLS2) were used in the prognostic risk model, which was an independent prognostic biomarker related to EB subtype, clinical stage and survival status in NPC.
The tumor microenvironment (TME) is a unique environment with infiltrating immune and nonimmune cells44. Immune components can modulate the occurrence and development of cancers in the TME45,46,47. In this study, functional enrichment analyses demonstrated that the risk model was involved in immune-based pathways. Immune-infiltrating cells (T follicular helper cells, resting DCs) were markedly upregulated, while immune-infiltrating cells (naive B cells, memory B cells) were downregulated in the high-risk group. Consistent with previous studies, B-cell and memory B-cell expression was markedly lower than that in noncancerous tissues and was correlated with PFS in NPC48. These results revealed that FRG-based risk score models were linked to the prognosis of NPC, perhaps partly by regulating the TME.
Proper selection of the correct medications may increase the effectiveness of anticancer drugs for treated patients49. The correct diagnosis is critical to determining the correct treatment50. To explore the potential role of ferroptosis in drug therapy, our results suggested that the high-risk group was sensitive to some chemotherapy drugs, including docetaxel, embelin, and epothilone B, parthenolide, thapsigargin, tipifarnib, axitinib, and vinorelbine, indicating that these chemotherapy drugs may benefit high-risk NPC patients.
In this study, we analyzed the prognostic role of ferroptosis-related genes in NPC. Two FRG signatures were established to predict overall survival and immunotherapy in NPC patients. In conclusion, these results revealed a ferroptosis-related signature as a potential biomarker in NPC and provided deeper insights into the prognosis and therapeutic targets of NPC.
Materials and methods
Analyzing the differential expression of FRGs
The raw data for NPC patients were searched in GEO (https://www.ncbi.nlm.nih.gov/geo/). Differentially expressed genes (DEGs) were obtained with |log2FC|> 1 and an adjusted p < 0.05 between NPC tissue and control tissue from GSE53819 and GSE12452 using the "edgeR" package (version 4.0.5, https://cran.r-project.org/)51.
Estimation of consensus clustering
Consensus clustering (the R package “ConsensusClusterPlus”, https://cran.r-project.org/) was used to analyze four prognosis-related ferroptosis genes (ABCC1, HMGCR, CS and GLS2), and k = 3 may be the best choice in the GSE102349 datasets. Kaplan‒Meier (K-M) survival curves were used to calculate the prognostic signature. The correlation analysis of clustering and risk classification was performed using the R packages “ggalluvial” (version 0.12.2, https://cran.r-project.org/src/contrib/Archive/ggalluvial/) and “ggplot2” (version 3.0.0, https://cran.r-project.org/src/contrib/Archive/ggplot2/).
Investigation and prediction of prognostic signatures
Univariate Cox regression investigated the PFS from the GSE102349 dataset of NPC and four ferroptosis genes (ABCC1, HMGCR, CS, GLS2), for which hazard ratios (HRs) were calculated. LASSO regression was used to calculate the risk score using the R package “glmnet” (version 4.0, https://cran.r-project.org/src/contrib/Archive/glmnet/). ROC curve was used to evaluate the prognostic model using the R package “survivalROC” (version 1.0.2, https://cran.r-project.org/src/contrib/Archive/survivalROC/). Univariate/multivariate PFS analyses assessed the prognostic score. The nomogram was used to integrate poor progression-free survival using the R package “rms” (version 6.0, https://cran.r-project.org/src/contrib/Archive/rms/). p < 0.05 indicates statistical significance.
Identifying immune infiltration
GO and KEGG analyses (https://www.kegg.jp/kegg/kegg1.html) were used for the enrichment of DEGs obtained with |log2FC|> 1 and adjusted p < 0.05 using the R packages “clusterProfiler” and “enrichplot”52. CIBERSORT53 (https://cibersort.stanford.edu/) was used to detect the distribution of immune cells in GSE102349 (containing 113 NPC patients, 54 of whom had survival data)54, this data set (GSE102349) contains annotations about the distribution of intratumoral and stromal TILs.
Predicting the drug response of patients with high/low risk
The half-maximal inhibitory concentration (IC50) values of compounds/inhibitors for high/low risk nasopharyngeal carcinoma patients were predicted using the R “pRRophetic” package as previously described55.
Immunohistochemistry
Eleven nasopharyngeal carcinoma and three NNM collected from the Second Xiangya Hospital of Central South University from January 2018 to December 2018 were examined and included in this study, and the ethics committee approved this study. The patients obtained informed consent. The following primary antibodies were used: rabbit polyclonal anti-ABCC1 (AF7503, 1:500, Beyotime, China) and anti-GLS2 (bs-13376R, 1:500, Bioss, China). The secondary antibody was HRP-conjugated goat anti-rabbit (ab97051, 1:500, Sigma, USA). Sections were photographed by a microscope. The scores of staining intensity of tumor cells were as follows: 0 (no coloring); 1 (≤ 30%); 2 (31–60%); and 3 (61–100%). IHC was performed as previously described56.
Statistics
SPSS software 20.0 (SPSS, Inc., USA) and GraphPad Prism 7.0 (La Jolla, CA, USA) were used to analyze the data. p < 0.05 indicates statistical significance.
Ethics statement
The studies involving human participants were reviewed and approved by the ethics committee of the Second Xiangya Hospital, Central South University. The patients/participants provided their written informed consent to participate in this study. All methods were carried out in accordance with relevant guidelines and regulations.
Data availability
Publicly available datasets were analyzed in this study. This data can be found here: The raw data and corresponding clinical information were downloaded from Gene Expression Omnibus Database (GEO; https://www.ncbi.nlm.nih.gov/geo/). The expression array data GSE53819 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE53819) containing 18 control tissue and 18 NPC tissue and the throughput sequencing data GSE12452 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE12452) containing 10 control tissue and 31 NPC tissue and the GSE102349 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE102349) dataset containing 113 NPC patients used for subsequent analysis. All supporting data are included within the article, and all the data generated in this article are available from the first author on reasonable request.
References
Lin, X. et al. Silencing Op18/stathmin by RNA interference promotes the sensitivity of nasopharyngeal carcinoma cells to taxol and high-grade differentiation of xenografted tumours in nude mice. Basic Clin. Pharmacol. Toxicol. 119, 611–620 (2016).
Guo, Z., Wang, Y. J., He, B. S. & Zhou, J. Linc00312 single nucleotide polymorphism as biomarker for chemoradiotherapy induced hematotoxicity in nasopharyngeal carcinoma patients. Dis. Markers 2022, 6707821 (2022).
Sung, H. et al. Global cancer statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 Countries. CA Cancer J. Clin. 71, 209–249 (2021).
Chen, Y. P. et al. Nasopharyngeal carcinoma. Lancet 394, 64–80 (2019).
Liu, Z. et al. Quantification of familial risk of nasopharyngeal carcinoma in a high-incidence area. Cancer 123, 2716–2725 (2017).
Lee, A. W., Ma, B. B., Ng, W. T. & Chan, A. T. Management of nasopharyngeal carcinoma: Current practice and future perspective. J. Clin. Oncol. 33, 3356–3364 (2015).
Mou, Y. et al. Ferroptosis, a new form of cell death: Opportunities and challenges in cancer. J. Hematol. Oncol. 12, 34 (2019).
Gao, M. et al. Ferroptosis is an autophagic cell death process. Cell Res. 26, 1021–1032 (2016).
Tang, B. et al. The ferroptosis and iron-metabolism signature robustly predicts clinical diagnosis, prognosis and immune microenvironment for hepatocellular carcinoma. Cell Commun. Signal. 18, 174 (2020).
Immunotherapy activates unexpected cell death mechanism. Cancer Discov. 9, OF2 (2019).
Liu, H. J. et al. Ferroptosis-related gene signature predicts glioma cell death and glioma patient progression. Front. Cell Dev. Biol. 8, 538 (2020).
Greaves, W. et al. Detection of ABCC1 expression in classical Hodgkin lymphoma is associated with increased risk of treatment failure using standard chemotherapy protocols. J. Hematol. Oncol. 5, 47 (2012).
Hassannia, B., Vandenabeele, P. & Vanden Berghe, T. Targeting ferroptosis to iron out cancer. Cancer Cell 35, 830–849 (2019).
Cheng, Y. et al. Immune microenvironment related competitive endogenous RNA network as powerful predictors for melanoma prognosis based on WGCNA analysis. Front. Oncol. 10, 577072 (2020).
Zhao, P. et al. Dual-targeting biomimetic delivery for anti-glioma activity via remodeling the tumor microenvironment and directing macrophage-mediated immunotherapy. Chem. Sci. 9, 2674–2689 (2018).
He, W. et al. The high level of tertiary lymphoid structure is correlated with superior survival in patients with advanced gastric cancer. Front. Oncol. 10, 980 (2020).
Zhang, K. et al. A ceRNA network and a potential regulatory axis in gastric cancer with different degrees of immune cell infiltration. Cancer Sci. 111, 4041–4050 (2020).
Wang, W. et al. CD8(+) T cells regulate tumour ferroptosis during cancer immunotherapy. Nature 569, 270–274 (2019).
Jiang, M. et al. Targeting ferroptosis for cancer therapy: Exploring novel strategies from its mechanisms and role in cancers. Transl. Lung Cancer Res. 9, 1569–1584 (2020).
Wu, S., Zhang, C., Xie, J., Li, S. & Huang, S. A five-microRNA signature predicts the prognosis in nasopharyngeal carcinoma. Front. Oncol. 11, 723362 (2021).
Zhu, Q. et al. MIR106A-5p upregulation suppresses autophagy and accelerates malignant phenotype in nasopharyngeal carcinoma. Autophagy 17, 1667–1683 (2021).
Li, H. et al. Radiation-enhanced expression of CCL22 in nasopharyngeal carcinoma is associated with CCR4(+) CD8 T cell recruitment. Int. J. Radiat. Oncol. Biol. Phys. 108, 126–139 (2020).
Shi, Z. Z. et al. Ferroptosis in carcinoma: Regulatory mechanisms and new method for cancer therapy. Onco Targets Ther. 12, 11291–11304 (2019).
Lang, X. et al. Radiotherapy and immunotherapy promote tumoral lipid oxidation and ferroptosis via synergistic repression of SLC7A11. Cancer Discov. 9, 1673–1685 (2019).
Liu, Y. et al. Development and validation of a combined ferroptosis and immune prognostic classifier for hepatocellular carcinoma. Front. Cell Dev. Biol. 8, 596679 (2020).
Shen, L. et al. Ferroptosis in acute central nervous system injuries: The future direction?. Front. Cell Dev. Biol. 8, 594 (2020).
Lu, B. et al. The role of ferroptosis in cancer development and treatment response. Front. Pharmacol. 8, 992 (2017).
Yamada, A. et al. ABCC1-exported sphingosine-1-phosphate, produced by sphingosine kinase 1, shortens survival of mice and patients with breast cancer. Mol. Cancer Res. 16, 1059–1070 (2018).
Fan, C. C. et al. EFHD2 contributes to non-small cell lung cancer cisplatin resistance by the activation of NOX4-ROS-ABCC1 axis. Redox Biol. 34, 101571 (2020).
Yu, D. M., Huynh, T., Truong, A. M., Haber, M. & Norris, M. D. ABC transporters and neuroblastoma. Adv. Cancer Res. 125, 139–170 (2015).
Kunicka, T. et al. Non-coding polymorphisms in nucleotide binding domain 1 in ABCC1 gene associate with transcript level and survival of patients with breast cancer. PLoS One 9, e101740 (2014).
Balaji, S. A., Udupa, N., Chamallamudi, M. R., Gupta, V. & Rangarajan, A. Role of the drug transporter ABCC3 in breast cancer chemoresistance. PLoS One 11, e0155013 (2016).
Cox, A. G. et al. Yap reprograms glutamine metabolism to increase nucleotide biosynthesis and enable liver growth. Nat. Cell Biol. 18, 886–896 (2016).
Ma, C., Li, F. & Luo, H. Prognostic and immune implications of a novel ferroptosis-related ten-gene signature in lung adenocarcinoma. Ann. Transl. Med. 9, 1058 (2021).
Zhuo, S. et al. Clinical and biological significances of a ferroptosis-related gene signature in glioma. Front. Oncol. 10, 590861 (2020).
Singh, K. S. et al. African-centric TP53 variant increases iron accumulation and bacterial pathogenesis but improves response to malaria toxin. Nat. Commun. 11, 473 (2020).
Kuo, T. C. et al. Glutaminase 2 stabilizes Dicer to repress Snail and metastasis in hepatocellular carcinoma cells. Cancer Lett. 383, 282–294 (2016).
Dias, M. M. et al. GLS2 is protumorigenic in breast cancers. Oncogene 39, 690–702 (2020).
Wang, C. Y. et al. Gene signatures and potential therapeutic targets of amino acid metabolism in estrogen receptor-positive breast cancer. Am. J. Cancer Res. 10, 95–113 (2020).
Wang, D., Liu, J., Chen, Q., Yang, R. & Jiang, Q. Upregulation of glutaminase 2 and neutrophil cytosolic factor 2 is associated with the poor prognosis of glioblastoma. Biomark. Med. 14, 1585–1597 (2020).
Saha, S. K. et al. Multiomics analysis reveals that GLS and GLS2 differentially modulate the clinical outcomes of cancer. J. Clin. Med. 8, 355 (2019).
Zou, J. et al. Glutamine metabolism regulators associated with cancer development and the tumor microenvironment: A pan-cancer multi-omics analysis. Genes (Basel) 12, 1305 (2021).
Chen, X. et al. METTL3 promotes esophageal squamous cell carcinoma metastasis through enhancing GLS2 expression. Front. Oncol. 11, 667451 (2021).
Zhao, Y., Shao, Q. & Peng, G. Exhaustion and senescence: Two crucial dysfunctional states of T cells in the tumor microenvironment. Cell Mol. Immunol. 17, 27–35 (2020).
McCoach, C. E. & Bivona, T. G. Engineering multidimensional evolutionary forces to combat cancer. Cancer Discov. 9, 587–604 (2019).
Nersesian, S., Glazebrook, H., Toulany, J., Grantham, S. R. & Boudreau, J. E. Naturally killing the silent killer: NK cell-based immunotherapy for ovarian cancer. Front. Immunol. 10, 1782 (2019).
Langsten, K. L., Kim, J. H., Sarver, A. L., Dewhirst, M. & Modiano, J. F. Comparative approach to the temporo-spatial organization of the tumor microenvironment. Front. Oncol. 9, 1185 (2019).
Gong, L. et al. Comprehensive single-cell sequencing reveals the stromal dynamics and tumor-specific characteristics in the microenvironment of nasopharyngeal carcinoma. Nat. Commun. 12, 1540 (2021).
Balducci, L., Goetz-Parten, D. & Steinman, M. A. Polypharmacy and the management of the older cancer patient. Ann. Oncol. 24(Suppl 7), vii36-40 (2013).
Neagu, M. Metabolic traits in cutaneous melanoma. Front. Oncol. 10, 851 (2020).
Robinson, M. D., McCarthy, D. J. & Smyth, G. K. edgeR: A bioconductor package for differential expression analysis of digital gene expression data. Bioinformatics 26, 139–140 (2010).
Kanehisa, M. & Goto, S. KEGG: Kyoto encyclopedia of genes and genomes. Nucleic Acids Res. 28, 27–30 (2000).
Newman, A. M. et al. Robust enumeration of cell subsets from tissue expression profiles. Nat. Methods 12, 453–457 (2015).
Zhang, L. et al. Genomic analysis of nasopharyngeal carcinoma reveals TME-based subtypes. Mol. Cancer Res. 15, 1722–1732 (2017).
Chen, Y. et al. DNA damage repair status predicts opposite clinical prognosis immunotherapy and non-immunotherapy in hepatocellular carcinoma. Front. Immunol. 12, 676922 (2021).
Zuo, Y. et al. Inhibition of heat shock protein 90 by 17-AAG reduces inflammation via P2X7 receptor/NLRP3 inflammasome pathway and increases neurogenesis after subarachnoid hemorrhage in mice. Front. Mol. Neurosci. 11, 401 (2018).
Funding
The present study was supported by the Hunan Key Laborator Cultivation Base of the Research and Development of Novel Pharmaceutical Preparations (No.2016TP1029); The Hunan Provincial Key Laboratory of Fundamental and Clinical Research on Functional Nucleic Acid; Foundation of Hunan Educational Committee,No.19C0199; Natural Science Foundation of China (No.82104309), Natural Science Foundation of Hunan Province (No.2018JJ3569, No.2020JJ5625, No.2021JJ40639), Outstanding Youth Project of Hunan Provincial Education Department (No.18B538, No.20B075), and the Scientific Research Fund of the Heath and Family Planning Commission of Hunan Province (No. C20180139). ESI Discipline Special Project of Changsha Medical University(No.22B532).
Author information
Authors and Affiliations
Contributions
J.Z. designed the experiments and wrote the draft manuscripts. T.G., S.T. and G.L. performed the experiments. L.Z., L.W. and M.B. analyzed the data. W.Z. and B.H. prepared the figures. Z.G. designed the experiments and revised the paper, and all authors agree to be accountable for the content of the work. All authors reviewed the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zhou, J., Guo, T., Zhou, L. et al. The ferroptosis signature predicts the prognosis and immune microenvironment of nasopharyngeal carcinoma. Sci Rep 13, 1861 (2023). https://doi.org/10.1038/s41598-023-28897-2
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-28897-2
- Springer Nature Limited
This article is cited by
-
Peripheral hemoglobin to albumin ratio predicts prognosis in patients with nasopharyngeal carcinoma underwent concurrent chemoradiotherapy
BMC Cancer (2024)
-
Study of some graph theoretical parameters for the structures of anticancer drugs
Scientific Reports (2024)
-
Increased V-ATPase activity can lead to chemo-resistance in oral squamous cell carcinoma via autophagy induction: new insights
Medical Oncology (2024)
-
Unveiling innovative therapeutic strategies and future trajectories on stimuli-responsive drug delivery systems for targeted treatment of breast carcinoma
Naunyn-Schmiedeberg's Archives of Pharmacology (2024)