Abstract
Whilst most individuals with SARS-CoV-2 infection have relatively mild disease, managed in the community, it was noted early in the pandemic that individuals with cardiovascular risk factors were more likely to experience severe acute disease, requiring hospitalisation. As the pandemic has progressed, increasing concern has also developed over long symptom duration in many individuals after SARS-CoV-2 infection, including among the majority who are managed acutely in the community. Risk factors for long symptom duration, including biological variables, are still poorly defined. Here, we examine post-illness metabolomic profiles, using nuclear magnetic resonance (Nightingale Health Oyj), and gut-microbiome profiles, using shotgun metagenomic sequencing (Illumina Inc), in 2561 community-dwelling participants with SARS-CoV-2. Illness duration ranged from asymptomatic (n = 307) to Post-COVID Syndrome (n = 180), and included participants with prolonged non-COVID-19 illnesses (n = 287). We also assess a pre-established metabolomic biomarker score, previously associated with hospitalisation for both acute pneumonia and severe acute COVID-19 illness, for its association with illness duration. We found an atherogenic-dyslipidaemic metabolic profile, including biomarkers such as fatty acids and cholesterol, was associated with longer duration of illness, both in individuals with and without SARS-CoV-2 infection. Greater values of a pre-existing metabolomic biomarker score also associated with longer duration of illness, regardless of SARS-CoV-2 infection. We found no association between illness duration and gut microbiome profiles in convalescence. This highlights the potential role of cardiometabolic dysfunction in relation to the experience of long duration symptoms after symptoms of acute infection, both COVID-19 as well as other illnesses.
Similar content being viewed by others
Introduction
The devastation caused by the COVID-19 pandemic is unprecedented in recent memory, with > 6.3 million deaths and 543 million cases worldwide in just over two years1. SARS-CoV-2 infection can cause a wide spectrum of illness, even in individuals who do not require acute hospital management; many individuals are asymptomatic (35.1% to 40.5% in meta-analyses2,3) while others report prolonged symptom duration (2.3% to 37.7%4). Colloquially known as “Long COVID”, the National Institute for Health and Care Excellence (NICE), defines two categories: Ongoing Symptomatic COVID (OSC28) for individuals with symptoms lasting 28–83 days (4–12 weeks), and Post-COVID syndrome (PCS84), for individuals with symptoms lasting over 84 days (12 weeks). They should display “signs and symptoms that have developed during or after an infection consistent with COVID-19 … and not explained by an alternative diagnosis”5,6.
The strongest predictors of PCS84 are age (with those aged 35–69 years having highest risk), female sex, and greater severity of acute infection7,8 Whilst vaccination against SARS-CoV-2 reduces the risk and duration of PCS849,10,11, prolonged post-infection symptomatology remains common. The United Kingdom Office for National Statistics report 2 million affected individuals in the United Kingdom by 01 May 2022, and 71% of individuals report that this affects normal daily activities12. The understanding of the pathophysiology and risk factors for these differing phenotypes-from asymptomatic infection to prolonged illness—is still evolving.
Early in the pandemic, it was noted that individuals with cardiovascular disease were at greater risk of severe illness13,14 and ‘Long COVID’15. Metabolomic profiles, particularly lipidomics, can identify risk of cardiovascular disease, with known associations of particular profiles with cardiovascular disease16 and Type 2 diabetes17,18. Such pre-pandemic lipidomics profiles have been associated with risk of hospitalisation for both COVID-19, and pneumonia caused by other pathogens, enabling the generation of an Infectious Diseases risk score (ID score) for hospitalisation due to COVID-1919. Disturbances to the same group of metabolites have also been observed in samples from hospitalised individuals when acutely unwell with COVID-1920,21. What is less clear is whether such metabolomic profiles are associated with disease duration, and/or Long-COVID, with the published studies focusing on specific metabolites in hospitalised individuals22.
However, most cases of COVID-19 are managed in the community rather than hospital. We aimed to assess whether metabolomic profiles differed in community-dwelling individuals with different symptom durations, comparing samples from asymptomatic individuals, to those with short duration, OSC28 and PCS84, approximately 6 months post-infection. This was further extended to include individuals with similar illnesses of prolonged duration, testing negative for SARS-COV-2 infection. We examined metabolites individually and assessed the previously published ID-score.
Metabolism has been related to the gut microbiome composition, with reports that the gut microbiome may separate individuals with PCS84 from healthy controls23,24. Therefore, we further explored whether stool microbiome composition, taken after acute illness and paired with metabolomics, was different between individuals with disease of different symptom duration, with and without previous confirmed SARS-CoV-2 infection. Finally, we tested whether there was any relationship between metabolomic profiles and gut metagenomic composition.
Results
Baseline characteristics of cohort
Of 15 564 individuals invited to the CSSB, 5694 (36.6%) consented and were enrolled. Of these, 4787/5694 individuals (84.1%) returned samples suitable for metabolomic analysis. Participant mean age was 52.5 years (SD 11.8), 78.7% were female, and 94.8% identified as White British, (Table 1 and Supplementary Table 2). The largest group was those with Acute COVID-19 illness (ACI) (n = 1147), lasting 7 days or less, followed by 652 with OSC28, and 307 Asymptomatic participants, our reference group. We included 287 who had a non-COVID-19 illness, of which 48/287 had symptoms over 84 days (Non-COVID-19 illness > 84 days—NC84). 161/287 were confirmed negative for SARS-CoV-2 infection by PCR testing at onset of illness, and all had a negative antibody test at enrolment (Table 1). Baseline characteristics of the groups were broadly similar to the population from which they were recruited and to each other, although the final asymptomatic group was slightly older than the average for the cohort (mean 58.1 years (SD 10.2) vs. 52.7 (SD 11.7) P < 0.001) (Table 1, Supplementary Table 2 + 3). All groups were predominantly female (75–84%, Table 1), with a median BMI of 25.2–26.2 kg/m2 (Tables 1, 2).
Metabolomic analysis
In total, 3718/4787 (77.7%) participants had adequate logging and metabolomic data, of whom 2561/3178 (80.6%) fell into the pre-specified phenotype groups (see Materials and Methods, Table 5).
Our primary analysis, using multinomial regression adjusted for age, sex, and BMI, compared asymptomatic participants to groups with longer symptom duration. We showed 90 of 249 (36%) metabolites differed in participants with OSC28 (28–83 days of COVID-19 symptoms) compared to Asymptomatic SARS-CoV-2-positive individuals (Fig. 1, Supplementary Fig. 1, Supplementary Table 4). 39 of these 90 (43%) also differed in participants with NC28 (Non-COVID-19 illness 28–83 days) compared to Asymptomatic SARS-CoV-2 infection, with the same direction of effect (Fig. 1, Supplementary Fig. 1, Supplementary Table 4).
Relative risk ratio for each illness phenotype, per 1-SD increase in biomarker. Red indicates P ≤ 0.05 after FDR correction. Reference group (OR 1.0): Asymptomatic. ACI: Acute COVID-19 illness. OSC28: Ongoing symptomatic COVID-19 (28–83 days). PCS84: Post COVID-19 syndrome (≥ 84 days). NC28: Non-COVID-19 illness 28–83 days. NC84: Non-COVID-19 illness ≥ 84 days. Fatty Acids: Ratios& Proportions, and Absolute Values. DHA docosahexaenoic acid, SFA saturated fatty acids, MUFA monounsaturated fatty acids, PUFA polyunsaturated fatty acids. Cholesterol, LDL-C: Low density lipoprotein cholesterol, HDL-C: High density Lipoprotein cholesterol, Total-C: Total cholesterol, Total-Tg: Total Triglycerides, VLDL-C: Very low desnity lipoprotein cholesterol.
Amongst the subset of metabolites validated for clinical use (37 of 249 measurements)19, fatty acids differed in asymptomatic cases compared to both positive and negative symptomatic individuals. A higher ratio of polyunsaturated fatty acids (PUFA) compared to monounsaturated fatty acids (MUFA) was associated with a lower odds of prolonged COVID-19 (OR = 0.73, False Discovery Rate adjusted P-value (FDR-P) = 0.01 for OSC28 vs. asymptomatic), and non-COVID-19 illness (OR = 0.68, FDR-P = 0.01 for NC28 vs. asymptomatic). (Fig. 1 and Supplementary Table 5). We also noted an association of absolute MUFA levels with increased length of illness (OR = 1.28 [FDR-P = 0.04] for OSC28 vs. asymptomatic COVID-19). Similarly, in combination, raised triglycerides and VLDL lipids were associated with an increased risk of prolonged illness in both test-positive and test-negative individuals, as was the ratio of triglycerides to phosphoglycerides (Supplementary Table 4 + 5). In contrast, higher HDL lipoprotein levels and larger HDL particles were associated with Asymptomatic cases. Neither amino acids nor glycoprotein acetyls were associated with COVID symptom duration.
In both the clinically validated variables, and the entire metabolomics dataset, only 7/249 (2.8%) variables were significantly different in Long COVID (OSC28 or PCS84 combined) compared to Non-COVID illness (NC28 and NC84 combined) as the reference group (Supplementary Table 6). Of note, HDL-Cholesterol was also raised in Acute COVID-19 Illness (OR 1.24 [FDR P-value 0.005] for ACI vs Non-COVID illness) and further raised in Asymptomatic illness (OR 1.44 [FDR P-value 0.007] for Asymptomatic vs. Non-COVID illness).
The Infectious diseases score
The multi-biomarker infectious diseases score (ID Score) was calculated, whereby higher values were previously associated with hospitalization for COVID-1919. In our cohort, higher values of the ID Score were generally associated with longer duration of symptoms of all illnesses, but notably not Post-COVID Syndrome (Table 2).
Sensitivity analyses adjusting for additional variables
The direction of effect and significance of results were unchanged for most sensitivity analyses, including adjustment for baseline cardiovascular disease and diabetes; however, the effect of the ID Score was marginally stronger after adjusting for Healthy Plant-based diet index (OR = 1.61 with hPDI, OR = 1.53 without hPDI: for OSC28 compared to asymptomatic COVID-19) (Supplementary Fig. 2).
Microbiome demographics
A subset (n = 301) of the metabolomic cohort had corresponding microbiome data (Table 3). The median time between symptom onset and microbiome assessment was 223 (IQR 50 days), with the minimum time between symptom onset and microbiome assessment of 33 days (implications considered further in Discussion).
Quality control of gut microbiome sequence data for these 301 individuals revealed good breadth of coverage of MetaPhlAn markers. In addition, depth of coverage of MetaPhlAn markers was high for most abundant species (~ 3X), with large areas of < 0.5X coverage (Supplementary Fig. 3).
For this subset analysis, individuals were grouped into categories as follows: (1) Asymptomatic: n = 35; (2) ACI: n = 109; (3a) OSC28 n = 52; (3b) Post COVID-19 syndrome (≥ 84 days): n = 20; (4) Negative symptomatic (≥ 28 days, including ≥ 84 days): n = 85 .
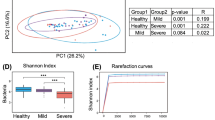
Alpha- diversity analysis
Microbial richness did not differ between groups (Wilcoxon signed-rank test P-value > 0.25 for all comparisons, Supplementary Fig. 4). There were no differences in Alpha-diversity (Simpson or Shannon) between groups, whether they were test positive or negative, symptomatic or asymptomatic, or with long or short symptom duration (Supplementary Fig. 4).
Beta-diversity analysis
Similarly, beta-diversity analysis showed no large-scale shifts in microbial profiles between groups (Supplementary Fig. 4).Ongoing symptomatic COVID-19 and Post COVID-19 syndrome groups were amalgamated and beta-diversity analysis repeated; there remained no significant difference (data not shown).
Microbial differential abundance analysis
A Generalised Linear Model controlled for confounding factors, including age, sex, and BMI identified three species with differences between groups—specifically, Firmicutes bacterium CAG 94, Ruminococcus callidus and Streptococcus vestibularis. Firmicutes bacterium CAG 94 differed between Asymptomatic SARS-CoV-2 infection (ref) and Acute COVID-19 (FDR P-value = 0.03), OSC28 (FDR P-value = 0.01) and PCS84 (FDR P-value = 0.04). Ruminococcus callidus, also a Firmicute, differed between Asymptomatic and Non-COVID-19 illness (≥ 28 days) (FDR P-value = 0.01) (Table 4). Streptococcus vestibularis differed only when comparing OSC28 and non-COVID-19 participants (FDR P-value 0.073).
Correlation between metabolomic and microbiome analysis
Correlations between the metabolites and microbial taxa were assessed using Spearman’s rank coefficient. We found no evidence that microbiome taxa were associated with differences in those metabolites associated with symptom duration in this dataset (Fig. 2). Negatively correlated metabolites and microbial species clustered in the top right-hand corner, with mild positively correlated associations throughout the remainder of the heatmap. A distinctive pattern was unable to be elucidated. The primary species driving the negative correlation were Alistipes finegoldii, an anaerobic, mesophilic, rod-shaped bacterium and Bacteroides cellulosilyticus, a cellulolytic bacterium.
Spearman correlation of microbiome profiles and metabolomic profiles. Single microbial taxa correlated with clinically validated metabolomic data using Spearman’s rank sum non-parametric test. FDR P-values are displayed *P < 0.01. Rows and columns are hierarchically clustered (Euclidean distance). The R package ‘corrplot_0.90' (https://github.com/taiyun/corrplot) was used to compute the variance and the covariance or correlation. The packages ‘pheatmap_1.0.12’ (https://cran.r-project.org/web/packages/pheatmap/index.html) and the ‘cor.mtest' function of ‘corrplot_0.90’ were used to visualise the heatmap and calculate associated P values.
Discussion
Summary of results + results in context
We observed a metabolic profile, particularly in lipid components, that differentiated individuals with longer symptom duration compared with individuals with asymptomatic infection. This profile was evident for individuals with long symptom duration regardless of SARS-CoV-2 test status, compared with asymptomatic infected individuals.
The specific differences identified in association with long-duration illness were small in magnitude and related to atherogenic dyslipidaemia (Fig. 3); in particular, blood fatty acid concentrations, including higher absolute and relative concentrations of MUFA, lower relative concentrations of PUFA, and a lower PUFA/MUFA ratios. In humans, circulating PUFA is derived from dietary sources, and blood levels correlate both to dietary intake and to levels in adipose tissue stores. MUFA, on the other hand, is synthesised in significant quantities in vivo and circulating concentrations (but not dietary intakes) are associated with increased risk of cardiometabolic disease25. Elevated serum MUFA and low serum PUFA have been identified in many studies as associated with ill health, including cardiovascular risk26, metabolic syndrome27, and mortality from infections28. Although our study assayed blood levels up to 9 months after initial COVID illness, these measures, particularly MUFA, are relatively stable over time19,29 and may reflect long-term blood concentrations. Moreover, our results concord with previous work in the UK Biobank, using the same platform, which demonstrated a similar direction of association with the same metabolites with increased risk of acute severe COVID-19 and with pneumonia, using blood samples collected many years beforehand19. Recent studies have reminded clinicians of the increased risk of vascular diagnoses after both COVID-19 and similar respiratory infections30,31,32 and while the risk reduces dramatically after the acute period, there is excess risk which remains many months afterwards. It is possible that our findings may reflect un-detected prior cardiovascular risk in symptomatic COVID-19 cases or vascular changes as a consequence of disease.
Atherogenic-dyslipidaemic biomarkers. Relative risk ratio for each illness phenotype, per 1-SD increase in biomarker. Adjusted for age, sex and body mass index. 95% Confidence intervals displayed with P-values adjusted using Benajmini-Hochberg False Discovery Rate correction. Red indicates FDR corrected P-value ≤ 0.05. ACI: Acute COVID-19 illness, OSC28: Ongoing symptomatic COVID-19 (28–83 days), PCS84: Post COVID-19 syndrome (≥ 84 days), NC28: Non-COVID-19 illness 28–83 days, NC84: Non-COVID-19 illness ≥ 84 days, ApoB_by_ApoA1: Ratio of apolipoprotein B to apolipoprotein A1, HDL_C: High density lipoprotein cholesterol, HDL_L: Total Lipids in high density lipoprotein, LDL_TG: Triglycerides in low density Lipoprotein, L_HDL_P: Concentration of large high density lipoprotein particles, MUFA: Monounsaturated Fatty Acids, PUFA_by_MUFA: Ratio of polyunsaturated fatty acids to monounsaturated fatty acids, Remnant_C: Remnant cholesterol (non-HDL, non-LDL -cholesterol), S_HDL_TG: Cholesterol in small HDL, VLDL_C: Very low density lipoprotein cholesterol, VLDL_L: Total lipids in VLDL, VLDL_TG: Triglycerides in VLDL, VLDL_size: Average diameter for VLDL particles.
Higher levels of VLDL-particles/lipids and TG-enriched lipoproteins were associated with longer illness duration, although again effects were relatively small in absolute terms. Both components have also been associated with ill-health in other (pre-pandemic) studies—in particular, cardiovascular disease, diabetes, renal disease, obesity and depression33,34, although studies did not always adjust for diet35. In our study, adjusting for dietary intakes or co-morbidities did not change the findings. We also demonstrate similar findings to previous work associating peripheral vascular disease with metabolic profiles representing atherogenic dyslipidaemia (Fig. 3)16,36.
We did not find metabolomic profiles specific to Long COVID comparing individuals with long-duration illness who were positive vs. negative for SARS CoV-2 infection. This would suggest that atherogenic-dyslipidaemic metabolic profiles are associated with long symptom duration, regardless of the cause of illness. This is supported by a growing body of literature identifying “conventional” risk factors such as high BMI, diabetes and cardiovascular disease, as risk factors for long COVID, as well as age7,37 all of which are associated with adverse metabolic changes26,34,38. There was no meaningful change in our results with inclusion of self-reported cardiovascular disease and diabetes as co-variates (Supplementary Fig. 2).
There were some unexpected negative findings. Others have associated GlycA with increased mortality39, and it is often increased in other conditions such as diabetes34. It was the biomarker showing the greatest association with hospitalisation for COVID-19 using pre-pandemic UK Biobank samples19. However, we found no association of GlycA with illness duration in our community-based sample. GlycA is considered a marker of inflammation, and our participants were sampled many months after the onset of symptoms and SARS-CoV-2, and were no longer reporting symptoms. It is possible that prolonged symptoms after COVID-19 may not always be related to ongoing inflammation, but previous damage that has not been repaired. There was also no association between any of the amino acid metabolites and length of illness (Supplementary Fig. 1), although some amino acids have previously been associated with hospitalisation for COVID-1919 and were included in the ID Score.
Looking at the ID score, previously associated with severity of acute COVID-1919, we also demonstrated an increased score was associated with increased length of illness. This was replicated for both COVID-19 and those reporting ongoing symptoms without COVID-19. This suggests that there are shared associations which link severe acute illness and prolonged illness. This tallies with our previous observation that participants with a high acute symptom burden early in their illness were at greater risk of Long COVID7. Others have shown perturbations in lipid profiles are also associated with severity of disease, and correlate with changes in immune cell populations and function. In particular, depletion of lipids is often associated with moderate and severe acute disease, compared to mild disease, and associates with shifts in immune cell profiles, including the balance of CD8 and effector T-Cells, and monocyte subsets40. Particular disturbances in natural killer cell (NK-Cell) function and phenotype have also been noted to correlate with severity of acute illness. These further correspond to changes in TNF and INF-α signatures, and FcRγ expression41,42. There are suggestions this may relate to different metabolic profiles of immune cells, metabolic reprogramming, and hepatic injury43,44. Further work would be needed to see if similar correlations with immune function hold true for illness duration, both during the initial phase of the illness, as well as during ongoing, prolonged illness, compared to those who have recovered.
Although 97 metabolites and the ID score were associated with Ongoing Symptomatic COVID-19 illness, only 16 of these were still associated with Post-COVID-19 syndrome. The PCS84 group is much smaller, and may be heterogeneous. Further work is needed to see whether certain subgroups within PCS84 display similar associations to OSC28, whilst other subgroups do not. However, it is of note that those most at risk for PCS84, women in mid-life4,45, are less at risk for some conditions associated with metabolic derangement, such as cardiovascular disease46,47. Our results might suggest that truly persistent symptoms in PCS84 may represent a different type of disease to OSC28, NC28 and NC84, with different risk factors, pathophysiology, and may therefore need different interventions to prevent or treat.
There was little change in the effect of the ID score on length of illness in our sensitivity analyses including pre-morbid cardiovascular disease and diabetes, frailty, lifestyle, IMD, the use of any supplements and the Diet Quality Score, suggesting that our metabolite associations with length of illness are independent of their associations with these variables. Models controlling for whether individuals took omega-3 supplements showed a diminished relationship between ID score and length of illness in non-COVID disease but were unchanged in COVID-19 illness. This finding which might suggest that omega-3 supplementation is a marker explaining this relationship in non-COVID illness only. The addition of the healthy plant-based diet index to the model increased the effect size of the ID score for each length of illness. Further work on hPDI and other correlated variables, such as socio-economic status, might help explore this finding.
We found no large-scale shift in gut microbiome, assessed up to 9 months post symptom onset, in relation to illness duration after SARS CoV-2 infection, and notably, no difference between COVID-19 positive and negative individuals with long-duration symptoms. Analysis of both taxonomic and functional microbial profiles showed increases in relative abundance of some Firmicutes in asymptomatic individuals, for example Firmicutes bacterium CAG:94, an uncharacterized taxon requiring further study to determine its functions. This finding should be interpreted with caution, and we did not find evidence that it related to metabolite alterations. Here, timing of our sampling is highly relevant to the interpretation of our results. Alterations in an individual’s microbiome may occur in the context of acute disease48, whether from infection, dietary changes, medication, usage, and/or immune function; however, an individual’s microbiome regresses over time49 to reflect a stable baseline microbiome50,51. Thus, faecal microbiome samples collected 9 months after the start of illness may represent baseline individual microbiome, which, in our study, did not relate to symptom duration. In any case, our findings would suggest no long-term alterations in microbial profile in individuals who experienced ‘long COVID’. Our results contrast to one small previous study of 106 hospitalised individuals, of whom 76% had ongoing symptoms 6 months after acute SARS-CoV-2 infection. In this study, altered gut microbiome composition was reported in individuals with persistent symptoms in patients with COVID-19 compared to healthy historical pre-pandemic controls52. However, cases were hospitalised for an average of 17 days, received amoxicillin-clavulanic acid among other interventions (including Ribavirin, Interferon and Remdesivir), and changes seen could have been a consequence these treatments, illness severity and/or altered diet in these individuals. Two other studies noted changes in immune-modulating commensal bacteria53,54 potentially specific to COVID-19, but again in hospitalised cohorts, where additional treatments, and illness severity, differ from our community-dwelling participants.
Strengths + Limitations
Our study benefits from large size, with a long duration of prospective symptom reporting, and availability of both metabolite and microbiome data on the same community-based participants. These platforms are well validated, including for clinical use of the metabolomics data. Our participants also reflect the spectrum of COVID cases in the community. As participants were recruited prior to vaccination in the wild-type era, this reduces complexity by variance attributable to virus variant, and type and timing of vaccination55. Infection status was reconfirmed at enrolment, ensuring accuracy of classification by gold standard methodology. During this period, there were also relatively few acute COVID-19 specific treatments available, and none routinely used in UK community-managed individuals allowing our study to reflect the natural history of COVID-19.
We have also been able to conduct sensitivity analyses including variables such as frailty, lifestyle, deprivation, and diet, often not extensively accounted for in metabolomics studies, in addition to more traditional medical co-morbidities. We also have analysed participants who report ongoing symptoms not attributable to SARS-CoV-2, and therefore able to test the specificity of our findings.
Limitations include the cross-sectional nature of the metabolites and microbiome assayed up to 9 months after illness onset. Time of sampling, for both the metabolomics and microbiome analysis is both a strength and a limitation. Post convalescence sampling means that acute perturbations are likely to have resolved, but without pre-pandemic sampling we cannot assess whether changes found were consequential or pre-existing. Longitudinal data from cohorts sampled before and after pandemic are needed to address this issue. We have also considered direct validation of our findings in another cohort, but due to the sui generis method of classification using the ZCS data, other cohorts’ participants would lack the same data to be classified into directly comparable groups.
Conclusion
Metabolic profiles of community cases with asymptomatic COVID-19 were notably different to those with longer illnesses, displaying an atherogenic lipoprotein phenotype, and this difference was apparent regardless of whether the illness was due to COVID-19 or another acute phenomenon. Those with COVID-19 symptoms for ≥ 28 days could not be clearly distinguished from those with non-COVID-19 illnesses of prolonged duration. A biomarker score previously predictive of severe COVID-19 was overall predictive of prolonged illness, although not all individual components were. In contrast, gut microbiome diversity did not differ by length of illness, suggesting no significant gut microbiome dysbiosis post COVID-19 infection.
Further research with longitudinal sampling pre- and post-illness is warranted, to determine if the observed metabolomic associations with longer illness are pre-existing risk factors, or consequential.
Materials and methods
Cohort description
Study participants were volunteers from the COVID Symptom Study Biobank (CSSB, approved by Yorkshire & Humber NHS Research Ethics Committee Ref: 20/YH/0298). Individuals were recruited to the CSSB via the ZOE COVID Study (ZCS)56 using a smartphone-based app developed by Zoe Ltd, King’s College London, the Massachusetts General Hospital, Lund University, and Uppsala University, launched in the UK on 24 March 2020 (approved by the Kings College London Ethics Committee LRS-19/20–18,210). Via the app, participants self-report demographic information, symptoms potentially indicative of COVID-19 disease (both closed/polar questions, and free text), any SARS-CoV-2 testing, and SARS-CoV-2 and influenza vaccinations. Participants can be invited via email to participate in other studies, according to eligibility.
In October 2020, prior to UK vaccination roll-out, 15,564 adult participants from the ZCS were invited to join the CSSB. Invited participants had: (a) a self-reported SARS-CoV-2 test result: A swab test (in this time period RT-PCR) at the start of illness, or a subsequent antibody test, whether positive or negative; and (b) logged at least once every 14 days since start of illness, or since the start of logging if asymptomatic.
Initially, individuals were recruited in four groups based on understanding of Long COVID at that time, and prior to definitions being published: (1) Asymptomatic, with confirmed infection; (2) Short illness (≤ 14 days) after confirmed infection; (3) Long illness (≥ 28 days)5 after confirmed infection; and (4) Long symptom duration (≥ 28 days) but with a negative test for SARS-CoV-2 infection. Invited individuals were matched across these four groups by Euclidian distance for age, sex and BMI9. Participants were invited by email, and consented separately into the CSSB. Participants were sent home sampling kits in November 2020 via post, and returned capillary blood samples for metabolomic analysis. This also enabled antibody testing using an ELISA method57 to confirm prior infection status of all participants, the current standard for retrospective ascertainment58. A subgroup also consented to send in stool samples for analysis of their gut microbiome.
Study participants were subsequently aligned using symptoms ascertained up to sample collection date, and the permissible gap in logging was further tightened to 7 days to increase accuracy of classification. Long-COVID groups were reshaped to match definitions published in November 20205 (see Table 5) corresponding to Ongoing Symptomatic COVID-19 (28–83 days, OSC28) and Post-COVID-19 Syndrome (> 84 days, PCS84). The same duration parameters were applied to those reporting symptoms with the same timeframe parameters around a negative test for SARS-CoV-2, who were presumed to have a non-COVID-19 illness. This yielded six groups for comparison—four SARS-CoV-2 positive groups: Asymptomatic, Acute COVID-19 (≤ 7 days), OSC28, PCS84; and two SARS-CoV-2 negative groups: Non-COVID-19 illness 28–83 days (NC28), Non-COVID-19 illness ≥ 84 days (NC84).
To check that groupings assigned by the recruitment algorithm were clinically accurate, symptom logging maps were scrutinised in a subsample (n = 115) by two researchers (MFÖ, CJS), independently and blind to algorithmic phenotype classification, before analysis. Final categories are detailed in Table 5.
Due to changes in logging stringency criteria, some participants also fell into additional, shorter categories of illness duration, detailed in Supplementary Table 1. These additional phenotypes have been reported in supplementary data tables, but not included in primary analysis as they were not recruited for this purpose, and their classification is less certain.
Metabolomics
Capillary blood samples were obtained between November 2020 and January 2021, when participants had recovered. Samples were returned in plasma collection tubes with initial processing of 20µL used for serology with the remainder frozen. Samples were processed in March/April 2021 by Nightingale Health Oyj (Helsinki, Finland) using high-throughput nuclear magnetic resonance metabolomics, measuring 249 metabolites including lipids, lipoprotein subclasses with lipid concentrations within fourteen subclasses, lipoprotein size, fatty acid composition, and various low-molecular weight metabolites including amino acids, ketone bodies and glycolysis metabolites33. Of these, 37 are certified for clinical diagnostic use and formed the focus of our analysis (referred to herein as “Clinically Validated”)59. Quality control was performed and reported by Nightingale Health. Due to postal transit time, glucose, lactate, and pyruvate could not be assessed and have been excluded from analyses, and creatinine was unavailable. There were no concerns raised with other biomarkers. Metabolites measured using this panel have been associated with the risk of hospitalisation for COVID-19 in the UK Biobank previously19, including 25 of the clinically validated biomarkers used in an Infectious Diseases risk prediction score (ID Score) derived using Lasso regression19.
Gut microbiome
Sample collection and faecal sample processing
Two faecal samples per individual were collected and returned by post: faecal material from both collection tubes were homogenised in a Stomacher® bag, aliquoted out and stored at -80 degrees Celsius. The first 301 samples that would maintain a balance for BMI, age, and sex, were selected to undertake a pilot investigation of microbiome differences.
DNA extraction and sequencing
Genomic DNA (gDNA) was isolated from 1 g faecal sample, using a modified protocol of the MagMax Core Nucleic Acid Purification Kit and MagMax Core Mechanical Lysis Module60. Libraries were prepared using the Illumina DNA Prep (Illumina Inc., San Diego, CA, USA) following the manufacturer’s protocol. Libraries were sequenced (2 × 150 bp reads) using the S4 flow cell on the Illumina NovaSeq 6000 system.
Metagenome quality control and pre-processing
All metagenomes were quality controlled using the pre-processing pipeline (available at https://github.com/SegataLab/preprocessing). Briefly, pre-processing consisted of three main steps: (i) read-level quality control, (ii) removal of host sequence contaminants, and (iii) splitting and sorting of cleaned reads. Read-level quality control removes low-quality reads (quality score < Q20), fragmented short reads (< 75 bp), and reads with ambiguous nucleotides (> 2 Ns), using trim-galore (https://www.bioinformatics.babraham.ac.uk/projects/trim_galore/). Host sequences contaminant DNA were identified using Bowtie 261 with the “–sensitive-local” parameter to remove both the phiX 174 Illumina spike-in and human-associated reads. Splitting and sorting allowed for creation of standard forward, reverse, and unpaired reads output files for each metagenome.
Taxonomic and functional profiling
The metagenomic analysis was performed using the bioBakery 3 suite of tools62. Taxonomic profiling and estimation of species’ relative abundances were performed with MetaPhlAn 3 (v. 3.0.7 with “–stat_q 0.1” parameter)62,63. MetaPhlAn 3 taxonomic profiles were used to compute three alpha diversity measures: (i) the number of species with positive relative abundance in the microbiome (‘Richness’), (ii) the Shannon diversity index, independent of richness, which measures how evenly microbes are distributed64, and (iii) the Simpson diversity index, which accounts for the proportion of species in a sample65. Similarly, species-level relative abundances were used to estimate microbiome dissimilarity between participants (beta diversity) using the Bray–Curtis dissimilarity metric, which accounts for the shared fraction of the microbiome between two individuals and their relative abundance values66. Functional potential profiling of metagenomes was performed with HUMAnN 3 (v. 3.0.0.alpha.3 and UniRef database release 2019_01)62,67 that produced pathway profiles and gene family abundances. We assessed beta diversity by computing a Principal Coordinates Analysis (PCoA)/Metric Multidimensional Scaling (MDS) based on the pairwise Bray–Curtis dissimilarity metric.
Additional covariates
BMI was derived from self-reported weight and height. Other self-reported information (obtained from ZCS app-reported data) included smoking; and co-morbid illness (‘heart disease’, ‘diabetes’ (and type of diabetes), ‘lung disease’ (including asthma), hay fever, eczema, ‘kidney disease’ and current cancer (type, and cancer treatment). Address data was linked to the UK Index of Multiple Deprivation (IMD), with the IMD rank decile used as a categorical variable measuring local area deprivation68,69,70,71. Frailty was assessed using the Prisma-7 scale, with a score > 2 indicating frailty72.
A subset of participants had participated in a dietary assessment during the COVID-19 pandemic, also recruited through the ZCS (published previously73,74). This included detailed information on vitamin supplementation (including omega-3 oils), physical activity, alcohol consumption, dietary habits and a food frequency questionnaire. These data were used to derive a diet quality score73, and a plant-based diet index73, analysed as continuous variables. Both have previously been associated with cardiovascular disease75, Type 2 Diabetes76, a lower risk of COVID-19 illness, and a lower risk of hospitalisation for COVID-19 during the early waves of the pandemic73.
Statistical analysis
The statistical analyses were performed using R software (v. 4.0.5) and Stata (v.17, StataCorp). Baseline characteristics were described by frequency and percentages. Descriptive data on those invited and those enrolled, are presented in Supplementary Table 2 + 3. Metabolites were all log-transformed and standardised (mean 0, standard deviation 1) as per protocol19. To account for 0 values, prior to log transformation, a pseudo-count of 1 was added to all values.
Initial analysis examined association between duration of illness and each metabolite individually, using multinomial logistic regression (all adjusted for age, sex, and BMI). The Asymptomatic group was used as the reference category of the outcome variable. We also performed a secondary analysis, using the non-COVID-19 participants as a reference category. The primary analysis was then extended, assessing association between length of illness and ID score, with asymptomatic as the reference category. The Benjamini–Hochberg False discovery rate method was used to correct P-values for multiple testing77.
To assess potential confounders, we performed eight sensitivity analyses additionally adjusting for: (1) self-reported co-morbidities (cardiovascular disease and diabetes), (2) Frailty, (3) IMD rank decile, (4) lifestyle variables (smoking status, frequency of alcohol consumption, frequency of physical activity), (5) self-reported use of any health supplement, (6) self-reported use of Omega-3 containing supplements, (7) Diet quality score, and (8) Healthy plant-based diet index (hPDI).
For microbiome analyses, differences between the alpha diversity distributions of the groups were assessed using the Wilcoxon rank-sum test within the ‘RClimMAWGEN’ package (P-value ≤ 0.05 considered significant). With a sample size of 300 individuals, we have 79% power at 0.05 significance level, assuming a low effect size of 0.20. PERMANOVA from the ‘adonis2’ function of the ‘vegan’ package, was used to test for differences between groups based on the beta diversity computed from the PCoA/MDS of the pairwise Bray–Curtis dissimilarities. For the microbial differential abundance analysis, we built a generalized linear model, controlling for confounding factors, including age, sex,and BMI. Only species with minimum 20% prevalence were used in this statistical analysis78. P-values were corrected using the Benjamini–Hochberg method77.
Spearman correlation analyses were conducted to associate microbiome profiles of 301 individuals with their metabolome profiles, adjusting for confounding factors (age, sex, BMI). Correlation analyses were conducted using R version 3.6.0. The package ‘corrplot_0.90' was used to compute the variance and the covariance or correlation, ‘pheatmap_1.0.12’ and ‘cor.mtest' were used to visualise the heatmap, calculate associated P values. Hierarchical clustering of both top 39 most abundant microbial species and 39 metabolic profiles was conducted using hclust, implementing the Ward.D2 agglomeration method. The package ‘p.adjust’ was used to perform Benjamini–Hochberg multiple testing correction77.
Ethical approval
The ZOE COVID Study by the King's College London Ethics Committee (Ref: LRS-19/20-18,210) and licensed under the Human Tissue Authority (reference 12,522). All ZCS participants provided informed consent for use of their data for COVID-19 research. The COVID Symptom Study Biobank and related studies, including this study, were approved by the Yorkshire & Humber NHS Research Ethics Committee (Ref: 20/YH/0298). CSSB participants were invited to join from the ZCS user base and provided informed consent to participate in the additional questionnaire and sample collection studies, and for linkage to app-collected data. All research and sample processing has been carried out in line with relevant guidelines including the Declaration of Helsinki.
Data availability
Data collected in the ZCS smartphone application are shared with other health researchers through the UK National Health Service-funded Health Data Research UK (HDRUK) and Secure Anonymised Information Linkage consortium, housed in the UK Secure Research Platform (Swansea, UK). Anonymised data are available to be shared with researchers according to their protocols in the public interest (https://web.www.healthdatagateway.org/dataset/fddcb382-3051-4394-8436-b92295f14259). The code is available in: https://gitlab.com/KCL-BMEIS/covid-zoe/vaccination. Access to data in the CSS Biobank is available to bona fide health researchers on application to the CSS Biobank Management Group. Further details are available online at: https://cssbiobank.com/information-for-researchers including application forms and contact information.
References
WHO. WHO Coronavirus. (2022). Available at: https://www.who.int/health-topics/coronavirus.
Ma, Q. et al. Global Percentage of Asymptomatic SARS-CoV-2 Infections among the Tested Population and Individuals with confirmed COVID-19 diagnosis: A systematic review and meta-analysis. JAMA Netw. Open 4, 1–18 (2021).
Sah, P. et al. Asymptomatic SARS-CoV-2 infection: A systematic review and meta-analysis. Proc. Natl. Acad. Sci. 118, e2109229118 (2021).
Park, C. et al. Short Report on Long COVID. (2021).
NICE (National Institute for Health and Care Excellence), SIGN (Scottish Intercollegiate Guidelines Network) & Royal College of General Practitioners. COVID-19 rapid guideline: Managing the long-term effects of COVID-19. NICE Guidelines (2022).
Soriano, J. B., Murthy, S., Marshall, J. C., Relan, P. & Diaz, J. V. A clinical case definition of post-COVID-19 condition by a Delphi consensus. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(21)00703-9 (2021).
Sudre, C. H. et al. Attributes and predictors of long COVID. Nat. Med. 27, 626–631 (2021).
Thompson, E. J. et al. Long COVID burden and risk factors in 10 UK longitudinal studies and electronic health records. Nat. Commun. 13, 3528 (2022).
Antonelli, M. et al. Risk factors and disease profile of post-vaccination SARS-CoV-2 infection in UK users of the COVID Symptom Study app: A prospective, community-based, nested, case-control study. Lancet Infect. Dis. https://doi.org/10.1016/S1473-3099(21)00460-6 (2021).
Ayoubkhani, D., Bosworth, M. & King, S. Self-reported long COVID after two doses of a coronavirus (COVID-19) vaccine in the UK-Office for National Statistics. 1–5 (2022).
UKHSA. The effectiveness of vaccination against long COVID: A rapid evidence briefing. (2022).
Ayoubkhani, D. & Pawelek, P. Prevalence of ongoing symptoms following coronavirus (COVID-19) infection in the UK: June 2022. Office for National Statistics (2022).
Hu, J. & Wang, Y. The clinical characteristics and risk factors of severe COVID-19. Gerontology 67, 255–266 (2021).
Du, P. et al. A systematic review and meta-analysis of risk factors associated with severity and death in COVID-19 patients. Can. J. Infect. Dis. Med. Microbiol. 2021, 5589 (2021).
Loosen, S. H. et al. Obesity and lipid metabolism disorders determine the risk for development of long COVID syndrome: A cross-sectional study from 50, 402 COVID-19 patients. Infection https://doi.org/10.1007/s15010-022-01784-0 (2022).
Soininen, P., Kangas, A. J., Würtz, P., Suna, T. & Ala-Korpela, M. Quantitative serum nuclear magnetic resonance metabolomics in cardiovascular epidemiology and genetics. Circ. Cardiovasc. Genet. 8, 192–206 (2015).
Wang, T. J. et al. Metabolite profiles and the risk of developing diabetes. Nat. Med. 17, 448–453 (2011).
Yu, D. et al. Plasma metabolomic profiles in association with type 2 diabetes risk and prevalence in Chinese adults. Metabolomics 12, 3 (2016).
Julkunen, H., Cichońska, A., Slagboom, P. E. & Würtz, P. Metabolic biomarker profiling for identification of susceptibility to severe pneumonia and COVID-19 in the general population. Elife 10, 1–15 (2021).
Dierckx, T. et al. The metabolic fingerprint of COVID-19 severity. medRxiv https://doi.org/10.1101/2020.11.09.20228221 (2020).
Ulloque-Badaracco, J. R., Hernandez-Bustamante, E. A., Herrera-Añazco, P. & Benites-Zapata, V. A. Prognostic value of apolipoproteins in COVID-19 patients: A systematic review and meta-analysis. Travel Med. Infect. Dis. 44, 102200 (2021).
Su, Y. et al. Multiple early factors anticipate post-acute COVID-19 sequelae. Cell https://doi.org/10.1016/j.cell.2022.01.014 (2022).
Visconti, A. et al. Interplay between the human gut microbiome and host metabolism. Nat. Commun. 10, 578 (2019).
Menni, C. et al. Serum metabolites reflecting gut microbiome alpha diversity predict type 2 diabetes. Gut Microbes 11, 1632–1642 (2020).
Hodson, L., Skeaff, C. M. & Fielding, B. A. Fatty acid composition of adipose tissue and blood in humans and its use as a biomarker of dietary intake. Prog. Lipid Res. 47, 348–380 (2008).
Würtz, P. et al. Metabolite profiling and cardiovascular event risk: A prospective study of 3 population-based cohorts. Circulation 131, 774–785 (2015).
Julibert, B. & Mateos, A. T. Dietary fat intake and metabolic syndrome in older adults. Nutrients 11, 1901 (2019).
Zhuang, P. et al. Dietary fats in relation to total and cause-specific mortality in a prospective cohort of 521 120 individuals with 16 years of follow-up. Circ. Res. 124, 757–768 (2019).
Nightingale Health. Nightingale Health Metabolic Biomarkers: Phase 1 Release Introduction Biomarker measurements by NMR. (2020).
Bhaskaran, K. et al. Overall and cause-specific hospitalisation and death after COVID-19 hospitalisation in England: A cohort study using linked primary care, secondary care, and death registration data in the OpenSAFELY platform. PLOS Med. 19, e1003871 (2022).
Knight, R. et al. Association of COVID-19 with arterial and venous vascular diseases: A population-wide cohort study of 48 million adults in England and Wales. Circulation 146, 892-906 (2022).
Katsoularis, I., Fonseca-Rodríguez, O., Farrington, P., Lindmark, K. & Fors, C. A. M. Risk of acute myocardial infarction and ischaemic stroke following COVID-19 in Sweden: A self-controlled case series and matched cohort study. Lancet 398, 599–607 (2021).
Würtz, P. et al. Quantitative serum nuclear magnetic resonance metabolomics in large-scale epidemiology: A primer on -omic technologies. Am. J. Epidemiol. 186, 1084–1096 (2017).
Barrios, C. et al. Circulating metabolic biomarkers of renal function in diabetic and non-diabetic populations. Sci. Rep. 8, 1–11 (2018).
Bot, M. et al. Metabolomics profile in depression: A pooled analysis of 230 metabolic markers in 5283 cases with depression and 10,145 controls. Biol. Psychiatry 87, 409–418 (2020).
Aday, A. W. et al. Lipoprotein particle profiles, standard lipids, and peripheral artery disease incidence: Prospective data from the Women’s Health Study. Circulation 138, 2330–2341 (2018).
Thompson, E. J. et al. Long COVID burden and risk factors in 10 UK longitudinal studies and electronic health records Nat. Commun. 13, 3528 (2022).
LeBrasseur, N. K. et al. Identifying biomarkers for biological age: Geroscience and the ICFSR task force. J. Frailty Aging 10, 196–201 (2021).
Deelen, J. et al. A metabolic profile of all-cause mortality risk identified in an observational study of 44,168 individuals. Nat. Commun. 10, 1–8 (2019).
Su, Y. et al. Multi-omics resolves a sharp disease-state shift between mild and moderate COVID-19. Cell 183, 1479-1495.e20 (2020).
Shemesh, A. et al. Diminished cell proliferation promotes natural killer cell adaptive-like phenotype by limiting FcεRIγ expression. J. Exp. Med. 219, 4789 (2022).
Krämer, B. et al. Early IFN-α signatures and persistent dysfunction are distinguishing features of NK cells in severe COVID-19. Immunity 54, 2650-2669.e14 (2021).
Shen, B. et al. Proteomic and metabolomic characterization of COVID-19 patient sera. Cell 182, 59-72.e15 (2020).
Lee, J. W. et al. Integrated analysis of plasma and single immune cells uncovers metabolic changes in individuals with COVID-19. Nat. Biotechnol. 40, 110–120 (2022).
Ayoubkhani, D. Prevalence of ongoing symptoms following coronavirus (COVID-19) infection in the UK: 1 April 2021. Off. Natl. Stat. 5, 1–16 (2021).
Jousilahti, P., Vartiainen, E., Tuomilehto, J. & Puska, P. Sex, age, cardiovascular risk factors, and coronary heart disease. Circulation 99, 1165–1172 (1999).
Mosca, L., Barrett-Connor, E. & Kass, W. N. Sex/gender differences in cardiovascular disease prevention. Circulation 124, 2145–2154 (2011).
Wu, J. et al. Gut microbiota dysbiosis associated with plasma levels of Interferon-γ and viral load in patients with acute hepatitis E infection. J. Med. Virol. 94, 692–702 (2022).
Elvers, K. T. et al. Antibiotic-induced changes in the human gut microbiota for the most commonly prescribed antibiotics in primary care in the UK: A systematic review. BMJ Open 10, e035677 (2020).
Chen, L. et al. The long-term genetic stability and individual specificity of the human gut microbiome. Cell 184, 2302-2315.e12 (2021).
Schloissnig, S. et al. Genomic variation landscape of the human gut microbiome. Nature 493, 45–50 (2013).
Liu, Q. et al. Gut microbiota dynamics in a prospective cohort of patients with post-acute COVID-19 syndrome. Gut 71, 544–552 (2022).
Yeoh, Y. K. et al. Gut microbiota composition reflects disease severity and dysfunctional immune responses in patients with COVID-19. Gut 70, 698–706 (2021).
Zuo, T. et al. Alterations in Gut microbiota of patients with COVID-19 during time of hospitalization. Gastroenterology 159, 944-955.e8 (2020).
Wheeler, S. E. et al. Differential antibody response to mRNA COVID-19 vaccines in healthy subjects. Microbiol. Spectr. 9, 478 (2021).
Menni, C. et al. Real-time tracking of self-reported symptoms to predict potential COVID-19. Nat. Med. 26, 1037–1040 (2020).
Wells, P. M. et al. Estimates of the rate of infection and asymptomatic COVID-19 disease in a population sample from SE England. J. Infect. 81, 931–936 (2020).
Österdahl, M. F. et al. Concordance of B and T cell responses to SARS-CoV-2 infection, irrespective of symptoms suggestive of COVID-19. J. Med. Virol. https://doi.org/10.1002/jmv.28016 (2022).
Nightingale Health. Clinically Validated Biomarkers. (2022). https://research.nightingalehealth.com/clinically-validated-biomarkers.
ABM. MagMAXTM CORE Nucleic Acid Purification Kit. (2022). http://www.abm.com.ge/en/68-Reagents and Consumables/235-MagMAXTM CORE Nucleic Acid Purification Kit. Accessed 1 Mar 2022.
Langmead, B. & Salzberg, S. L. Fast gapped-read alignment with Bowtie 2. Nat. Methods 9, 357–359 (2012).
Beghini, F. et al. Integrating taxonomic, functional, and strain-level profiling of diverse microbial communities with biobakery 3. Elife 10, 1–42 (2021).
Truong, D. T. et al. MetaPhlAn2 for enhanced metagenomic taxonomic profiling. Nat. Methods 12, 902–903 (2015).
Shannon, C. E. A mathematical theory of communication. ACM SIGMOBILE Mob. Comput. Commun. Rev. 5, 3–55 (2001).
Simpson, E. H. Measurement of diversity. Nature 163, 688–688 (1949).
Bray, J. R. & Curtis, J. T. An ordination of the upland forest communities of Southern Wisconsin. Ecol. Monogr. 27, 325–349 (1957).
Franzosa, E. A. et al. Species-level functional profiling of metagenomes and metatranscriptomes. Nat. Methods 15, 962–968 (2018).
Ministry of Housing Communities and Local Government. English Indices of Deprivation 2019. National Satistics: Statistcial Release 2019, (2019).
Northern Ireland Statistics and Research Agency. No Northern Ireland Multiple Deprivation Measure 2017 (NIMDM2017). (2022). https://www.nisra.gov.uk/statistics/deprivation/northern-ireland-multiple-deprivation-measure-2017-nimdm2017. Accessed 05 May 2022.
Scottish Executive. Scottish Index of Multiple Deprivation 2020. (2020). https://www.gov.scot/collections/scottish-index-of-multiple-deprivation-2020/.
Welsh Government. Welsh Index of Multiple Deprivation. (2020). https://gov.wales/welsh-index-multiple-deprivation.
Raîche, M., Hébert, R. & Dubois, M. F. PRISMA-7: A case-finding tool to identify older adults with moderate to severe disabilities. Arch. Gerontol. Geriatr. 47, 9–18 (2008).
Merino, J. et al. Diet quality and risk and severity of COVID-19: A prospective cohort study. Gut https://doi.org/10.1136/gutjnl-2021-325353 (2021).
Mazidii, M. et al. Impact of COVID-19 on health behaviours and body weight: A prospective observational study in a cohort of 1.1 million UK and US individuals. Nat. Food 1, 47 (2021).
Satija, A. et al. Healthful and unhealthful plant-based diets and the risk of coronary heart disease in U.S. adults. J. Am. Coll. Cardiol. 70, 411–422 (2017).
Satija, A. et al. Plant-based dietary patterns and incidence of type 2 diabetes in US men and women: Results from three prospective cohort studies. PLoS Med. 13, 1–18 (2016).
Benjamini, Y. & Hochberg, Y. Controlling the false discovery rate: A practical and powerful approach to multiple testing. J. R. Stat. Soc. Ser. B 57, 289–300 (1995).
Cougoul, A., Bailly, X., Vourc’h, G. & Gasqui, P. Rarity of microbial species: In search of reliable associations. PLoS One 14, e0200458 (2019).
Acknowledgements
We would like to acknowledge support from the Exetera development team (Benjamin Murray, Liyuan Chen, Jie Deng, Eric Kerfoot), the TwinsUK laboratory and administrative teams (Gulsah Akdag, Andy Anastasiou, Andrei Beleanu, Julia Brown, Maria Paz Garcia, Rachel Horsfall, Genevieve Lachance, Ayrun Nessa, Alyce Sheedy, Dovile Vaitkute, Sam Wadge, Sivasubramaniam Rajan Wignarajah, Darioush Yarand). We are also grateful to the CSS Biobank Volunteer Advisory Group for their input on the development of the biobank, and feedback on the significance of this work for affected individuals and the wider public.
Funding
The CSS Biobank is supported by the Chronic Disease Research Foundation (Grant: CDRF 22/2020). We are also supported by the Wellcome EPSRC Centre for Medical Engineering at King's College London (WT 203148/Z/16/Z) and the UK Department of Health via the National Institute for Health Research (NIHR) comprehensive Biomedical Research Centre award to Guy's & St Thomas’ NHS Foundation Trust in partnership with King's College London and King's College Hospital NHS Foundation Trust. Investigators also received support from Medical Research Council (MRC) (Grant codes: MC_PC_20059 (CJS, SO), MR/T005351/1 and MR/V038664/1 (SO), MR/W005611/1 and MR/S023747/1 (MM)), British Heart Foundation (BHF), Alzheimer's Society, European Union, NIHR, COVID-19 Driver Relief Fund (CDRF) and the NIHR-funded BioResource, Clinical Research Facility and Biomedical Research Centre (BRC) based at GSTT NHS Foundation Trust in partnership with KCL. SO was supported by the French government, through the 3IA Côte d'Azur Investments in the Future project managed by the National Research Agency (ANR) with the reference number ANR-19-P3IA-0002. ZOE Limited provided in-kind support for all aspects of building, running and supporting the app and service to all users worldwide. This research was funded in part by the Wellcome Trust [215010/Z/18/Z].
Author information
Authors and Affiliations
Contributions
C.J.S. E.L.D. C.S. S.O. developed the concept for the biobank. C.J.S. developed the current study with input from E.L.D. M.A. C.S. C.J.S. and E.L.D. formulated study recruitment, with M.A., R.W., M.F.Ö. and C.J.S. validating allocation to study groups. M.F.Ö. performed analysis on the metabolomic dataset producing related tables, results and figures, and RW analysed the gut microbiome dataset, producing related tables, results and figures. C.M. and S.E.B. provided additional expertise on the interpretation of metabolomics results, and F.A., A.B.M. and N.S. provided additional support on microbiome analysis. M.F.Ö. and R.W. produced the first draft of the manuscript with oversight from C.J.S. and E.D. All authors have further contributed to the introduction and discussion, and had oversight of the final version of the manuscript. M.F.Ö. and R.W. should be identified as joint first authors.
Corresponding author
Ethics declarations
Competing interests
TDS is a shareholder and cofounder of ZOE Global Ltd, and has received payment for scientific consultancy services to ZOE Global Ltd. JW and CH are employees of ZOE Global Ltd. No other authors have competing interests to declare.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Österdahl, M.F., Whiston, R., Sudre, C.H. et al. Metabolomic and gut microbiome profiles across the spectrum of community-based COVID and non-COVID disease. Sci Rep 13, 10407 (2023). https://doi.org/10.1038/s41598-023-34598-7
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-023-34598-7
- Springer Nature Limited