Abstract
Liver cancer is the 4th most lethal form of cancer with a poor prognosis for patients worldwide. Dysregulation of lipid metabolism is related to FA oxidation alternation which can be modified by peroxisome proliferator-activated receptor-α (PPARα). Therefore, it is important to identify the lipid metabolism-related genes regulated by PPARα in liver cancer. Hub genes related to the lipid metabolism pathway of HCC samples treated with PPARα agonist (WY-14,643) were identified through a weighted gene co-expression network analysis (WGCNA). Gene expression and clinical information were obtained from the Gene Expression Omnibus (GEO) database. The network of top main hub genes was visualized by the Cytoscape software using MCODE and CytoHubba plugins. Finally, the expression and clinical association of each hub gene were evaluated using enrichment analysis, TCGA data, GEPIA, GSCA, and q-PCR. Based on our results, the top 5 co-expressed genes including (CPT2, ACSL1, ACSL3, ACOX1, and SLC27A2) were selected as the main hub genes participating in fatty acid metabolism, fatty acid beta-oxidation, and PPAR signaling pathway. All association of higher ACSL3 expression with lower outcomes and survival rates was detected in HCC patients. Therefore, lipid metabolism-related Hub genes regulated by PPARα are potential biomarkers, and they may offer a therapeutical foundation for targeted therapy directed against the HCC antitumor strategy.
Similar content being viewed by others
Introduction
Liver cancer is one of the 4th most leading cancer with more than 1 million new cases in 2020 worldwide1. Hepatocellular carcinoma (HCC) includes more than 80% of liver cancer cases associated with hepatitis virus infection and other liver diseases such as NAFLD2. Early detection of HCC patients and the 5-year survival rate remained challenging despite the recent progression in HCC treatments3,4. Therefore, it is essential to discover the molecular mechanism of genes related to HCC development and heterogenicity5. Cancer is characterized by dysregulation of fatty acid (FA) metabolism, among other regulatory factors, fatty acid sensors called peroxisome proliferator-activated receptors (PPARs) with 3 different isoforms control metabolism throughout the body6,7.
PPARs involve the aberrant activation of different signaling pathways, which changes the expression and activity of lipid-metabolizing enzymes8,9. As a result, PPARs are the subject of intense research attention for cancer therapy10. Weighted gene co-expression network analysis (WGCNA) is an in-silico approach to examine the expression of regulatory genes in a complicated network11,12.In the current study, we evaluated the differentially Expressed Genes (DEGs) network and their targets which are connected with lipid metabolism in HCC samples treated with PPARα agonist in human HCC samples using the WGCNA approach. The biological role of the DEGs was also evaluated by GO and KEGG pathway analysis. Based on our results most of the genes were enriched in PPAR signaling pathways, fatty acid transport, fatty acid degradation, and lipid metabolism. Next, via Cytoscape software top 10 genes with higher degree levels were detected and their association with clinical outcomes was analyzed. Ultimately ACSL3 was nominated based on its significant relation with liver cancer stage, grade, and survival rates in tumor samples compared to normal samples which can be considered as a new marker to develop a new and effective antitumor strategy against HCC pathology.
Material and methods
Microarray data analysis and DEGs
To evaluate the effect of the PPARα agonist on HCC lipid metabolism. HCC dataset was extracted from the Gene Expression Omnibus (GEO) (http://www.ncbi.nlm.nih.gov/geo): GSE17251 [Affymetrix Human Genome U133 Plus 2.0 Array]. The overall total of 24 samples (12 WY-14,643 treated and 12 untreated) was evaluated during our study and the microarray dataset was normalized by normalized quantile’s function using “limma package” (version 3.48.3) in R software (version 4.3.1)13, (https://www.bioconductor.org/packages/release/bioc/html/limma.html) Table 1.
Weighted correlation network analysis (WGCNA) and modules hub genes identification
WGCNA, is a systematic biological technique, for designing a scale-free network based on gene expression profiles. Via using the R software (version 4.3.1) and the “WGCNA” R package (version 1.72–1) http://www.genetics.ucla.edu/labs/horvath/CoexpressionNetwork/Rpackages/WGCNA, we estimated the genes with a strong correlation with the clinical variable. The expression matrices were transformed into matrices and the matrices were then transformed into adjacency matrices, where the soft-threshold (power value) was applied to highlight significant connections between genes based on the adjacency matrix. The adjacency matrix was then converted into a topological overlap measure (TOM) to show the strength of the gene-to-gene connection14.
Gene ontology (GO) and KEGG pathway enrichment analysis of hub genes
The enrichment analysis of the hub genes was carried out using the “clusterProfiler R package” (version 4.4.4) (https://github.com/GuangchuangYu/enrichment4GTEx_clusterProfiler) to visualize the biological process (BP), and KEGG pathway of hub genes in HCC to better understand their mechanism15.
Identification PPI network
Using the STRING database (https://string-db.org/), the protein–protein interaction (PPI) network of the genes with the highest score was built. Next, Cytoscape softwarehttp://apps.cytoscape.org/apps/stringapp (version 3.7.1) plugins (MCODE)( ftp://ftp.mshri.on.ca/pub/BIND/Tools/MCODE) and Cytohubba (http://hub.iis.sinica.edu.tw/cytohubba) were used to choose the cluster for functional analysis and the Vann diagram (https://bioinformatics.psb.ugent.be/webtools/Venn)was used to candidate the overlapping hub genes16,17.
Validation of hub genes
Genetic changes in the final hub genes in HCC patients were evaluated by evaluating the data from the TCGA database. Lipid metabolism-Related Genes were extracted from the Molecular Signatures Databasev7.5.1 (MSigDB). The final hub gene mRNA expression patterns were evaluated by UALCAN databases UALCAN (http://ualcan.path.uab.edu/index.html), GSCA (https://guolab.wchscu.cn/GSCA/) and GEPIA2 (http://gepia2.cancer-pku.cn/#analysis). The immunohistochemistry (IHC) was visualized by the Human Protein Atlas (HPA) (https://www.proteinatlas.org), and the survival rate was evaluated by the KM plot database (https://kmplot.com/analysis). Gene Set Enrichment Analysis (GSEA) was carried out using R software (version 4.3.1). The networkanalyst.ca was used to identify the TF-miRNA network of hub genes and finally, an association of hub genes and immune cell infiltration was evaluated by CIBERSORT analysis18.
Cell culture and real-time PCR
HepG2, HUH7, and normal liver cell lines (L-02) were obtained from the Chinese Academy of Sciences Cell Bank in Shanghai, China. The cells were regularly cultured in Dulbecco's modified Eagle's medium (DMEM) supplemented with 10% fetal bovine serum, at 37 C in a 5% CO2 atmosphere with 10 U/ml penicillin and 10 g/ml streptomycin. The total RNA of the cells was extracted using an RNA extraction kit (Monadbiotech, China). Using spectrophotometric assays at 260 and 280 nm, the RNA concentration was found. Next, reverse transcriptase (Monadbiotech, China) was used to create the cDNA. The levels of gene expression were ascertained through real-time quantitative PCR using Applied Biosystems 7500 real-time PCR apparatus (ABI 7500, USA) with real-time fluorescence detection. The oligonucleotide primers were as follows: One minute of initial denaturation was conducted at 95 °C, which was followed by ten seconds at 95 °C, sixty seconds at 66 °C, and 40 cycles at 62 °C. The 2−△△Ct method was employed to analyze the PCR data. Primer sequences used are listed below in Table 2.
Western blot analysis
To determine protein levels, the total proteins of liver cells were extracted using RIPA agents and quantified with a BCA protein assay kit (GPL Bio, China). Next, samples were separated by a 10% sodium dodecyl sulfate–polyacrylamide gel electrophoresis and transferred to a polyvinylidene difluoride filter membrane. Next, the membranes were soaked in antibodies against GAPDH, and ACSL3 (Abcam, China) followed by three times wash with 1 × TBST for 5 min and incubated for 1h with secondary antibody and finally visualized by Gel-Pro Analyzer 4.0 software (Media Cybernetics, Inc, Rockville, MD) (Supplementary Information).
Gene expression and therapy response correlation analysis
We conducted a ROC analysis of ACSL3 due to its strong intramodular connectivity and possible significance for the etiology of liver cancer, which were identified as central within their respective co-expression modules from WGCNA. To examine the relationship between gene expression and chemotherapy response the ACSL3 hub gene was chosen. We quantitatively evaluated the predictive power regarding the patient response to standard chemotherapeutic treatments using the ROC Plotter (https://www.rocplot.org/). This approach sought to find possible biomarkers for evaluating the effectiveness of liver cancer treatments.
Statistical analysis
All statistical analyses were performed in R software (version 4.3.1) and a P-value < 0.05 was considered statistically significant.
Results
Screening of DEGS
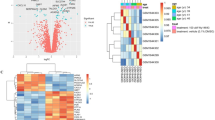
A total of 24 samples from the GEO dataset (GSE17251) were used for identifying the DEGs using the R version (4.3.2). Overall, 96 DEGs were upregulated and 10 were downregulated with the fold change threshold of 0.5 in the treated group compared to the untreated group (Fig. 1A). In addition, the top 20 upregulated genes related to lipid metabolism in HCC are represented in a heatmap plot (Fig. 1B).
Weighted correlation network analysis (WGCNA) of DEGS
WGCNA analysis was carried out to identify the gene clusters related to lipid metabolism. Lipid metabolism-Related Genes were extracted from the Molecular Signatures Databasev7.5.1 (MSigDB). The soft threshold value for the dataset was chosen with a cutoff R2 value of 0.85. A total of eighteen co-expressed gene modules were identified. Next different color was assigned to each module. The grey60 module has the fewest genes with 68 genes, while the turquoise module has been detected to be the largest module as well Table 3. (Fig. 2A–D).
Identification of PPARα agonist (WY-14643) gene modules in GSE17251 using WGCNA analysis. (A) Hierarchical clustering of samples to detect the outlier samples. (B) Hierarchical clustering of modules and heatmap of traits. (C) Scale independence and mean connectivity. (D) Dendrogram of GSE17251 dataset according to (1-TOM) matrix.
The WGCNA network construction to identify PPARα treatment-related module
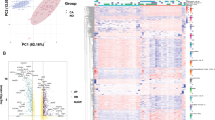
All candidate genes from datasets related to PPARα treatment were subjected to WGCNA analysis to determine whether or not the potential gene modules were connected to PPARα treatment. Among all the distinct modules that have been identified (Table 3), two modules (brown and pink modules) had a significant negative correlation with PPARα treatment based on the heatmap plot of the adjacencies (Fig. 3A–C).
Detecting the PPARα agonist associated modules with WGCNA. (A) Module-trait heatmap to identify the correlation between the gene module and the PPARα agonist treatment in the GSE17251 dataset, while each module contains the corresponding correlation coefficient and P-value. (B) The adjacency heatmap of the trait for each module. (C) Scatter plot of module pink and brown with negative correlation with PPARα agonist treatment.
KEGG pathway and GO enrichment analysis of candidate hub genes
Overlapping genes among the pink and brown modules and the DEGS (P-value < 0.05) were detected by the Vann diagram. Based on our GO enrichment and KEGG pathway results overlapping hub genes were primarily associated with PPAR signaling, Fatty acid metabolism, uptake, transport, and degradation (Fig. 4A–D).
Top PPI network clusters and validation of final hub genes expression levels
The PPI network of the overlapping genes was detected by the STRING online tool and the result was inserted into Cytoscape software version (3.7.1)'s MCODE plugin to find the hub clusters. Next, the Cytohubba plugin was used to detect the top 10 hub genes with the highest maximal clique centrality (MCC) (Fig. 5A). The expression levels of each hub gene were validated using expression data using GEPIA2 and UALCAN database. According to the expression plot, the expression of five genes out of 10 genes was significantly meaningful in HCC samples (P Value < 00.01) (Fig. 5B–G). The biological roles of selected hub genes were examined using a functional enrichment analysis, GSEA. Based on our results, selected DEGs are significantly related to alcohol liver disease, and PI3k-AKT signaling (Fig. 5H). The Go enrichment analysis of lipid metabolism-related hub genes indicates that the selected hub genes are mainly enriched in fatty acid metabolism, degradation, and transport (Fig. 5I).
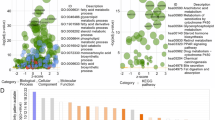
Validation of candidate hub genes. (A) Top 10 hub genes by Cytohubba plugin. (B) Expression plot of top 10 hub genes through GEPIA2 database (C–G). Box plot expression of 5 hub genes in HCC with meaningful P Value < 00.01. (H) GESA analysis of the hub genes using cluster Profiler package (I) GO enrichment analysis of lipid metabolism-related Hub genes.
Hub genes validation and TF-miRNA regulatory network evaluation
The q-PCR result of each hub gene was by the TCGA data indicating the over-expression of ACSL3 and downregulation of (ACSL1, ACOX1, CPT2, and SLC27A2) in HepG2 cells compared to normal control samples (Fig. 6A). Network Analyst database provided evidence of the relationship between transcription factors (TFs), miRNAs, and final hub genes (Fig. 6B). Information on transcription factors and gene targets obtained from ENCODE ChIP-seq data. This algorithm uses only the peak intensity signal < 500 and the predicted regulatory potential score < 1 (BETA Minus). Next, cytoscape 3.8 was used to plot the network after the nodes were filtered according to their degree.
Validation of hub gene expression and visualization of TF-miRNA network for each hub gene. (A) The qRT-PCR evaluation of hub genes mRNA levels in HEPG2 cells compared to the normal control cell line. (B) Diagram of core miRNA and TF regulatory network of hub genes filtered by degree, where red and orange represent genes; yellow represents predicted miRNAs; light-orange represents TF. The data represent the means ± SD. **P < 0.01, *** < 0.001 vs. control.
The significant association of ACSL3 with HCC progression
The function of each hub gene on clinical outcomes was detected, however, only ACSL3 was significantly associated with different pathways activity, worst clinical outcomes including stages (I-IV), and lower Overall survival rate (OS) in HCC patients (HR = 1.61, P = 0.011) (Fig. 7A–C). Moreover, the protein levels of immunohistochemistry (IHC) staining obtained from the Human Protein Atlas (HPA) database showed that the expression of the ACSL3 was significantly higher in tumor tissues than in normal tissues (Fig. 7D), which was consistent with that at the transcriptional level. The higher expression of ACSL3 was also positively associated with immune markers such as T cells, NKT cells, CD4, and monocytes and negatively associated with NK, TH2, and CD4 naive in liver cancer patients (Fig. 7E). ACSL3 expression is also associated with different biological pathways (Fig. 7F). The correlation of ACSL3 over-expression and drug sensitivity also indicates that ACSL3 is positively associated with different anti-cancer drugs such as navitoclax (Fig. 7G).
Higher ACSL3 expression is associated with poor clinical outcomes. (A) The over-expression of ACSL3 was related to poor OS. (B) KM plot of ACSL3 in liver cancer among 364 patients from km plot database. (C) Association of ASCL3 with poor stage in LIHC tumor. (D) The IHC staining of ACSL3 levels in normal human tissues compared to HCC tissues using the human protein atlas database. (E–G) Association of ACSL3 expression with immune markers, biological pathways, and drug sensitivity.
ACSL3 is a potential target for chemotherapy
We analyzed the ACSL3 for immune checkpoint inhibitors (ICIs) chemotherapy in cancers. Since, ROC analysis is used in diagnostic test evaluation, the ROC plotters of the ACSL3 with strong correlation to resistance especially chemotherapy in all samples (n = 167) were evaluated as shown in (Fig. 8A,B). Based on the ROC analysis, we find that ACSL3 has relatively high AUC and low P-value with anti-PD1 (P = 3.8 e − 3, AUC = 0.744) and anti-PD-L1 therapy (P = 3.5e − 2, AUC = 0.589) which can suggest ACSL3 as a potential target of any chemotherapy. The qRT-PCR results and western blot results of ACSL3 expression in HUH7 and HepG2 cell lines also indicated a higher ACSL3 level in compared to the LO2 cell line (Fig. 8C,D).
Discussion
Hepatocellular carcinoma (HCC) is considered the 6th most prevalent cancer with a high morbidity rate globally1. It is necessary to rapidly identify useful diagnostic biomarkers for screening patients to overcome the difficulty of early identification of HCC and to greatly improve the prognosis3. One known indicator of cancer hallmark is the deregulation of lipid metabolism6. Recent research has established a robust connection between dysregulated lipid metabolism and the progression of HCC. Variations in lipid composition can impact carcinogenic pathways and mechanisms19. PPARs are among the Key hepatic transcriptional regulator enzymes involved in lipid metabolism10. Lipids can bind directly to transcription factors as cofactors, which allows them to control lipid metabolism20. Peroxisome proliferator-activated receptor-α (PPARα), a highly expressed transcriptional factor for lipid metabolism in the liver highly regulates the hepatic steatosis21. PPARα activity regulates fatty acid homeostasis and inflammation22. PPARα expression in the human liver is similar to mice21,22. However, the exact role of PPARα whether it inhibits or accelerates cancer growth in cancer is not fully detected22. Several studies have shown the inhibitory effect of PPARα on cancer, while the excessive expression of PPARα can also increase lipid storage in the liver, promote steatosis, and enhance HCC initiation via decreasing FAO levels10,23,24. It has been demonstrated that Wy14643, a PPARα agonist, increases the formation of HCC by promoting PPARα expression levels and causing fatty liver25. According to a recent study, PPARα activation increases fatty acid transport protein (FATP) and fatty acid translocase (FAT) to promote hepatocyte proliferation in vivo19,26. However, in a different study, PPARα agonists decrease tumor growth and chemo-resistance in colon cancer by specifically inhibiting the glucose metabolic pathway through Glut1 transcription activity27. Thus, this study aimed to obtain molecular insights into the function of PPARα agonists in various cancers and to identify genes impacted by PPARα agonist treatment across samples since it has been suggested that lipid metabolism may be a target for cancer diagnosis, prognosis, and treatment and lipid metabolism alternation have been demonstrated to be intriguing biomarkers of the prognosis of cancer in a variety of tumor types. Therefore, by utilizing the WGCNA method, we were able to create a gene co-expression network that may be used to identify and anticipate significant genes and signaling pathways linked to PPARα28.
In this study, the GSE17251 mRNA-seq data was downloaded and examined. Then, the trait heatmap for each module was plotted, and the pink and brown modules were chosen based on the correlation coefficients, which indicated their strong negative association with PPARα agonists treatment in HCC samples following hub gene screening and validation using Cytoscape and the CytoHubba plugin. Next, the top 10 genes with higher degrees potentially regulating lipid metabolism, uptake, transport, and PPAR signaling were identified, while the expression of only five genes—CPT2, ACSL1, ACSL3, ACOX1, and SLC27A2 were significantly meaningful among HCC tumor samples compared to control samples. Finally, acyl-CoA synthetase 3 (ACSL3) was selected based on its great association with the lower survival rate (OS), worst clinical outcomes, and drug sensitivity in LIHC tumors. We also discovered a significant difference in the expression of the biomarker ACSL3 between liver cancer and normal liver tissues using the HPA database. This difference was consistent with the results of our western blot analysis.
In addition, ACSL3 was previously candidate as an ideal marker for various tumors since, the higher expression of ACSL3 correlates with a poor prognosis in different cancers such as NSCLC29, and the ROC curve and AUC of ACSL3 based on response to immunotherapy agents PD-1 (P = 3.8 e − 3, AUC = 0.744) and PD-L1 (P = 3.5e − 2, AUC = 0.589) indicated its potential to be considered as a diagnostic biomarker.
As the main factor causing lipid droplet accumulation in the liver, ACSL3 is a crucial regulatory enzyme that facilitates the esterification of fatty acids with Coenzyme A30. When ACSL3 is pharmacologically or genetically inhibited, tumor cells demonstrate significant cytotoxicity, suggesting that ACSL3 is an essential liver cancer cell viability component30. ACSL3 plays a major role in the regulation of fatty acid metabolism and lipid accumulation. ACSL3 alternation affects several pathological processes such as increasing steatosis, lipid metabolism, and cellular inflammation30,31. It was indicated that ACSL3 controls Clear cell renal cell carcinoma (ccRCC) lipid metabolism and lipid droplet formation30. Consequently, by reducing ACSL3 in human liver tissues, WY-14643 treatment can be regarded as an optimal treatment based on our findings. However, the present study is not without limitations, and given the limitations of our current studies such as the lack of biological experiments, only using the TCGA and GSE17251 databases, and lack of in vivo validation of hub genes to directly evaluate the candidate hub genes, and the small sample size that would allow us to confirm the precise mechanisms of WY-14643 treatment in regulating the HCC lipid metabolism. More research is required to prove its regulatory function in cancer treatment, particularly its function on intracellular lipids to develop better-targeted therapies for liver cancer32.
Conclusion
In conclusion, by using the WGCNA method, we discovered the top 10 co-expressed hub genes associated with lipid metabolism in HCC samples treated with PPARα agonist (WY-14,643) that may serve as diagnostic biomarkers for liver cancer. Of these, ACSL3 was chosen because of its role in lipid metabolism and strong correlation with worse prognosis and the development of liver cancer. However, more studies are needed to evaluate the role of PPARs in lipid metabolism in the molecular mechanism and pathology of HCC.
Data availability
Data will be provided from the corresponding author upon the reasonable request.
References
Siegel, R. L., Miller, K. D. & Jemal, A. Cancer statistics, 2020. CA Cancer J. Clin. 70, 7–30 (2020).
Asafo-Agyei, K. O. & Samant, H. Hepatocellular carcinoma. StatPearls (2021).
Devarbhavi, H. et al. Global burden of liver disease: 2023 update. J. Hepatol. 79, 516–537 (2023).
Brown, Z. J. et al. Management of hepatocellular carcinoma: A review. JAMA Surg. 158, 410–420 (2023).
Hao, X. et al. Targeting immune cells in the tumor microenvironment of HCC: New opportunities and challenges. Front. Cell Dev. Biol. 9 (2021).
Paul, B., Lewinska, M. & Andersen, J. B. Lipid alterations in chronic liver disease and liver cancer. JHEP Rep. 4 (2022).
Corton, J. C., Peters, J. M. & Klaunig, J. E. The PPARα-dependent rodent liver tumor response is not relevant to humans: Addressing misconceptions. Arch. Toxicol. 92, 83–119 (2018).
Peters, J. M., Cattley, R. C. & Gonzalez, F. J. Role of PPAR alpha in the mechanism of action of the nongenotoxic carcinogen and peroxisome proliferator Wy-14,643. Carcinogenesis 18, 2029–2033 (1997).
Saidi, S. A., Holland, C. M., Charnock-Jones, D. S. & Smith, S. K. In vitro and in vivo effects of the PPAR-alpha agonists fenofibrate and retinoic acid in endometrial cancer. Mol. Cancer 5 (2006).
Kimura, O., Kondo, Y. & Shimosegawa, T. PPAR Could contribute to the pathogenesis of hepatocellular carcinoma. PPAR Res. 2012 (2012).
Hartke, J., Johnson, M. & Ghabril, M. The diagnosis and treatment of hepatocellular carcinoma. Semin. Diagn. Pathol. 34, 153–159 (2017).
Zeng, J., Lai, C., Luo, J. & Li, L. Functional investigation and two-sample Mendelian randomization study of neuropathic pain hub genes obtained by WGCNA analysis. Front. Neurosci. 17 (2023).
Ritchie, M. E. et al. Limma powers differential expression analyses for RNA-sequencing and microarray studies. Nucleic Acids Res. 43, e47 (2015).
Li, J. et al. Application of weighted gene co-expression network analysis for data from paired design. Sci. Rep. 8, 1–8 (2018).
Barrett, T. et al. NCBI GEO: Mining tens of millions of expression profiles—Database and tools update. Nucleic Acids Res. 35, D760 (2007).
Shannon, P. et al. Cytoscape: A software environment for integrated models of biomolecular interaction networks. Genome Res. 13, 2498 (2003).
Bindea, G. et al. ClueGO: a Cytoscape plug-in to decipher functionally grouped gene ontology and pathway annotation networks. Bioinformatics 25, 1091 (2009).
Chen, B., Khodadoust, M. S., Liu, C. L., Newman, A. M. & Alizadeh, A. A. Profiling tumor infiltrating immune cells with CIBERSORT. Methods Mol. Biol. 1711, 243–259 (2018).
Meng, F. G., Zhang, X. N., Liu, S. X., Wang, Y. R. & Zeng, T. Roles of peroxisome proliferator-activated receptor α in the pathogenesis of ethanol-induced liver disease. Chem. Biol. Interact. 327, 109176 (2020).
Cheng, H. S. et al. Exploration and development of PPAR modulators in health and disease: An update of clinical evidence. Int. J. Mol. Sci. 20 (2019).
Tan, Y. et al. PPAR-α modulators as current and potential cancer treatments. Front. Oncol. 11, 599995 (2021).
Han, L., Shen, W. J., Bittner, S., Kraemer, F. B. & Azhar, S. PPARs: Regulators of metabolism and as therapeutic targets in cardiovascular disease. Part II: PPAR-β/δ and PPAR-γ. Future Cardiol. 13, 279–296 (2017).
Chung, K. W. et al. Impairment of PPAR α and the fatty acid oxidation pathway aggravates renal fibrosis during aging. J. Am. Soc. Nephrol. 29, 1223–1237 (2018).
Zhang, N. et al. Peroxisome proliferator-activated receptor alpha inhibits hepatocarcinogenesis by mediating the NF-κB signaling pathway. Oncotarget 5, 8330–8340 (2014).
Wang, R. et al. WY-14643 attenuates lipid deposition via activation of the PPARα/CPT1A axis by targeting Gly335 to inhibit cell proliferation and migration in ccRCC. Lipids Health Dis. 21, 1–18 (2022).
Yang, R. et al. WY-14643, a selective agonist of peroxisome proliferator-activated receptor-α, ameliorates lipopolysaccharide-induced depressive-like behaviors by preventing neuroinflammation and oxide-nitrosative stress in mice. Pharmacol. Biochem. Behav. 153, 97–104 (2017).
Gou, Q. et al. PPARα agonist alleviates tumor growth and chemo-resistance associated with the inhibition of glucose metabolic pathway. Eur. J. Pharmacol. 863 (2019).
Langfelder, P. & Horvath, S. WGCNA: An R package for weighted correlation network analysis. BMC Bioinf. 9, 1–13 (2008).
Ansari, I. ul H. et al. Characterization of acyl-CoA synthetase isoforms in pancreatic beta cells: Gene silencing shows the participation of ACSL3 and ACSL4 in insulin secretion. Arch. Biochem. Biophys. 618, 32–43 (2017).
Klasson, T. D. et al. ACSL3 regulates lipid droplet biogenesis and ferroptosis sensitivity in clear cell renal cell carcinoma. Cancer Metab. 10 (2022).
Quan, J., Bode, A. M. & Luo, X. ACSL family: The regulatory mechanisms and therapeutic implications in cancer. Eur. J. Pharmacol. 909 (2021).
Wang, R. et al. WY-14643 attenuates lipid deposition via activation of the PPARα/CPT1A axis by targeting Gly335 to inhibit cell proliferation and migration in ccRCC. Lipids Health Dis. 21 (2022).
Author information
Authors and Affiliations
Contributions
M.A.M. performed the bioinformatic analysis M.A.M write the article draft revised the draft. and X.C supervised the study. All the authors read the submitted version and approved it.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
AmeliMojarad, M., AmeliMojarad, M. & Cui, X. Discovering the lipid metabolism-related hub genes of HCC-treated samples with PPARα agonist through weighted correlation network analysis. Sci Rep 14, 19591 (2024). https://doi.org/10.1038/s41598-024-69998-w
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-69998-w
- Springer Nature Limited