Abstract
Programming cell signaling during T-cell activation represents a simple strategy for improving the potency of therapeutic T-cell products. Stim-R technology (Lyell Immunopharma) is a customizable, degradable synthetic cell biomimetic that emulates physiologic, cell-like presentation of signal molecules to control T-cell activation. A breadth of Stim-R formulations with different anti-CD3/anti-CD28 (αCD3/αCD28) antibody densities and stoichiometries were screened for their effects on multiple metrics of T-cell function. We identified an optimized formulation that produced receptor tyrosine kinase-like orphan receptor 1 (ROR1)-targeted chimeric antigen receptor (CAR) T cells with enhanced persistence and polyfunctionality in vitro, as assessed in repeat-stimulation assays, compared with a benchmark product generated using a conventional T-cell–activating reagent. In transcriptomic analyses, CAR T cells activated with Stim-R technology showed downregulation of exhaustion-associated gene sets and retained a unique subset of stem-like cells with effector-associated gene signatures following repeated exposure to tumor cells. Compared with the benchmark product, CAR T cells activated using the optimized Stim-R technology formulation exhibited higher peak expansion, prolonged persistence, and improved tumor control in a solid tumor xenograft model. Enhancing T-cell products with Stim-R technology during T-cell activation may help improve therapeutic efficacy against solid tumors.
Similar content being viewed by others
Introduction
Solid tumors account for more than 90% of cancers1. While adoptive cell transfer (ACT) using chimeric antigen receptor (CAR) T cells has achieved considerable success in hematologic cancers2,3,4, positive outcomes in solid tumors have been limited5,6,7. A major challenge in the treatment of solid tumors is the tumor microenvironment, which suppresses CAR T-cell effector function, limits persistence, and induces T-cell exhaustion8,9,10,11. More potent T cells with increased persistence and reduced susceptibility to exhaustion need to be engineered to overcome these therapeutic hurdles11,12,13,14. To date, efforts to improve CAR T-cell potency have primarily focused on enhancing T-cell function through genetic engineering to modulate cell-intrinsic signaling15. However, T-cell function is also impacted by cell-extrinsic signals received before administration, such as those that mediate T-cell activation16,17,18,19,20,21. Hence, modulation of signaling parameters during T-cell activation represents an opportunity to direct T-cell function and create more potent T-cell products.
T-cell activation is a formative event that directs the fate, function, and durability of mature T cells18,19,20,21. In the body, this process is mediated by antigen-presenting cells (APCs), which provide three signals to T cells in highly coordinated spatiotemporal patterns: T-cell receptor (TCR) stimulation (signal 1), costimulation (signal 2), and paracrine cytokine signaling (signal 3)22,23,24. The strength and duration of these signals influence transcriptional and epigenetic changes that ultimately direct T-cell proliferation, memory differentiation, and effector activity18,19,20,21. Similarly, T-cell activation during ex vivo CAR T-cell production impacts the phenotypic and functional quality of the final ACT product and thereby its activity and persistence in vivo20,25,26,27,28,29,30. The precise optimization of signal presentation during T-cell activation therefore has the potential to generate CAR T cells with enhanced potency and persistence, which could improve efficacy against solid tumors. To date, conventional reagents used for activation were primarily developed to optimize expansion during the manufacturing of early ACT products; however, these activation reagents do not recapitulate physiologic signal presentation or support fine-tuning of signal presentation parameters26.
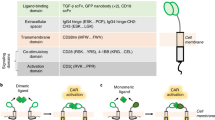
Stim-R technology (Lyell Immunopharma, San Francisco, CA, USA) is a customizable, degradable synthetic cell biomimetic comprised of lipid-coated silica micro-rods that emulate the physiologic, cell-like presentation of signal molecules to control T-cell properties (Fig. 1)25,26,30. The technology incorporates a synthetic lipid membrane to which surface-bound signals, such as agonist antibodies targeting CD3 (anti-CD3 [αCD3]; signal 1: TCR stimulus) and CD28 (anti-CD28 [αCD28]; signal 2: costimulation), can be attached to mimic natural ligand presentation on APC membranes. In addition, soluble signals, such as cytokines, may be loaded into the lipid-coated silica rods and released over time in a controlled manner to emulate paracrine cytokine delivery (signal 3). Because Stim-R can be readily formulated to present surface-bound signals in precise densities and stoichiometries, the technology allows for the rapid tuning of T-cell signal presentation parameters to generate T-cell products with desirable phenotypic and functional properties. Prior work utilizing Stim-R technology with a model CD19-targeted CAR provided proof of concept that CAR T-cell products with different phenotypes and acute cytotoxic function can be generated by varying the total stimulation strength during T-cell activation26.
Stim-R technology is a customizable, degradable, synthetic cell biomimetic that enables the physiologic presentation of signal molecules to control T-cell properties. (A) Stim-R technology is composed of lipid-coated silica micro-rods. (B) Surface-bound signals, such as agonist antibodies targeting CD3 (αCD3; signal 1) and CD28 (αCD28; signal 2), can be attached to the synthetic lipid membrane surface to emulate natural antigen presentation. (C) The lipid-coated silica micro-rods can be loaded with soluble signals (e.g., IL-2, IL-7, and IL-15), which are subject to controlled release over time to mimic paracrine cytokine signaling (signal 3). (D) The fabricated Stim-R material is used for ex vivo activation of T cells during T cell therapy production. (E) Stim-R technology facilitates the physiologic presentation of both surface and soluble signals and supports precise tuning of T-cell signal presentation parameters, such as signal combinations, densities, and stoichiometries, to generate T-cell products with desirable phenotypic and functional properties.
Here, we sought to produce CAR T-cell products with enhanced potency by optimizing signaling parameters during T-cell activation using Stim-R technology (Fig. 2). CAR T-cell products were activated using either Stim-R technology variants or a conventional T-cell–activating benchmark reagent and functionally evaluated to assess persistence and polyfunctionality. We identified an optimized Stim-R formulation that generated CAR T-cell products with enhanced in vitro and in vivo function compared to CAR T cells generated using the benchmark reagent. We also analyzed transcriptomic changes in CAR T cells to understand the effects of Stim-R on stemness and exhaustion. Optimization of cell signaling parameters during T-cell activation using Stim-R technology can be applied to a broad range of T-cell products and potentially improve therapeutic outcomes.
An optimized Stim-R formulation was identified using a suite of in vitro assessments, and subsequently mechanistically interrogated and validated in vivo using a solid tumor xenograft model. Stim-R technology was leveraged to fabricate an array of formulation variants in which signal stoichiometry and cumulative signal intensity were precisely and independently varied. Test variants were used to produce a corresponding array of ROR1-targeted CAR T-cell products, which were then functionally characterized in non-normalized repeat-stimulation assays in combination with ICS to assess polyfunctionality. Based on the trends observed, new arrays of Stim-R formulations were designed and prepared for further optimization in an iterated process. This iterated screening approach identified an optimized Stim-R formulation that generated CAR T-cell products with enhanced function. For benchmarking, ROR1-targeted CAR T cells were also activated in an analogous process using a conventional T-cell–activating reagent. To compare the functionality of these products on a per-cell basis, normalized repeat-stimulation assays were conducted to assess tumor cell killing, T-cell proliferation, and cytokine production. In addition, bulk and single-cell transcriptomic analyses were performed to probe the mechanisms underlying improved CAR T-cell potency following activation using Stim-R technology. The improved function of the CAR T-cell product generated using the optimized Stim-R formulation was validated in vivo in a xenograft solid tumor (lung cancer) model assessing antitumor activity, PK, and survival.
Results
Screening αCD3/αCD28 stoichiometry and signal density in vitro enabled the identification of an optimized Stim-R formulation
To identify a Stim-R formulation that generates a CAR T-cell product with optimal function, we fabricated an array of test formulations in which two attributes of the lipid membrane-bound signals were independently varied: signal stoichiometry and cumulative signal intensity. Signal stoichiometry refers to the relative strength of the αCD3 (TCR) and αCD28 (costimulation) signals (Fig. 3A), while cumulative signal intensity refers to the total strength of these two signals combined (Fig. 3B). The Stim-R array was loaded with interleukin (IL)-2, IL-7 and IL-15 to support T-cell growth and used to generate T-cell products expressing a clinically relevant receptor tyrosine kinase-like orphan receptor 1 (ROR1)-targeted CAR31,32.
Using Stim-R technology to optimize αCD3 and αCD28 signaling during T-cell activation generates CAR T-cell products with enhanced polyfunctionality and cytotoxicity. (A) Signal stoichiometry is the relative density of the αCD28 (costimulation) signal to the αCD3 (TCR) signal on the Stim-R technology lipid coating. (B) Cumulative signal intensity is the total density (mol %) of both the αCD3 and αCD28 signals on the Stim-R technology lipid coating. (C) Cytotoxicity in the non-normalized repeat-stimulation assay (left) and polyfunctionality (right) of CAR T cells activated with Stim-R variants with signal stoichiometry (αCD28:αCD3) ranging between 0.1 and 5.0 at a constant αCD3 intensity of 0.25%. Data represent the mean (± SEM) of products from 1 to 7 independent donors. (D) Cytotoxicity in the non-normalized repeat-stimulation assay (left) and polyfunctionality (right) of CAR T cells activated with Stim-R variants with cumulative signal intensity ranging between 0.1 and 1.0 at a constant αCD28:αCD3 ratio of 1.0. Data represent the mean (± SEM) of products from 3 to 6 independent donors. (E) CAR T-cell products activated using the optimized Stim-R formulation or the benchmark reagent were assessed using a non-normalized repeat-stimulation assay to measure persistent tumor cell clearance. Each panel shows the mean (± SD) of two technical replicates using products from an independent donor. (F) ICS to measure the percentage of polyfunctional (IL-2 + IFN-γ + ; left) and nonresponsive (IL-2–IFN-γ–; right) T cells. Data are shown for individual donors (n = 5), with black lines connecting data for products derived from the same donor. Paired two-tailed t-test, *p = 0.0187 (t = 3.822; df = 4), **p = 0.0095 (t = 4.667; df = 4).
T cells activated using the Stim-R test formulations were screened using a ‘repeat-stimulation’ assay that was designed to measure their ability to proliferate durably in culture in response to repeated stimulation with ROR1 + H1975 tumor cells, while maintaining antitumor cytotoxic function. Following an initial stimulation at an effector-to-target (E:T) ratio of 1:5, restimulations were performed by transferring 25% of the cell culture supernatant to a fresh plate of target cells, and cell killing was monitored in real-time after each stimulation. Since the E:T ratio is not controlled beyond the first stimulation, this assay is referred to as a ‘non-normalized’ repeat-stimulation assay. This assay may identify T-cell products with a capacity for prolonged persistence. In addition, polyfunctionality was assessed based on intracellular coexpression of the cytokines interferon-γ (IFN-γ) and IL-2 following exposure to tumor cells. We selected these assays because both prolonged persistence and polyfunctionality have been associated with improved therapeutic outcomes of CAR T-cell therapies33,34,35.
In the Stim-R formulation test array designed to optimize signal stoichiometry, an array of αCD28:αCD3 ratios ranging between 0.1 and 5.0 was tested, while maintaining a constant αCD3 density of 0.25% (mol%). An αCD28:αCD3 ratio of 1.0 demonstrated the greatest cytotoxicity in the non-normalized repeat-stimulation assay and the highest polyfunctionality (Fig. 3C). Subsequently, we maintained the αCD28:αCD3 ratio at 1.0 and screened Stim-R formulations with a cumulative signal intensity between 0.1 and 1.0% and observed optimal T-cell cytotoxicity and polyfunctionality at 0.5% (Fig. 3D). Based on these data, the Stim-R formulation with signal stoichiometry of 1.0 and cumulative signal intensity of 0.5% was designated as the optimized Stim-R formulation.
We next compared CAR T cells activated using the optimized Stim-R formulation to CAR T cells produced in an analogous process using a benchmark T-cell activation reagent (benchmark). The T cell TransAct reagent was used as the benchmark due to its wide use in clinical CAR T cell manufacturing. The CAR T-cell products generated using either Stim-R technology or the benchmark reagent had comparable total expansion, transduction efficiency, and proportion of stem-like cells at harvest (Fig. S1), as well as the percentage of CD4 + or CD8 + CAR T cells at the time of cryopreservation (Fig. S2). However, across three independent healthy donors, CAR T cells produced using the optimized Stim-R formulation versus the benchmark reagent showed more persistent cytotoxic function in the non-normalized repeat-stimulation assay (Fig. 3E), with significantly more polyfunctional T cells (IL-2 + IFN-γ + ; p = 0.0187) and significantly fewer nonresponsive T cells following tumor cell stimulation (IL-2–IFN-γ–; p = 0.0095) (Fig. 3F). In addition to cytokine supplementation to the cell culture media, the same set of cytokines was also loaded into Stim-R. Since the benchmark reagent does not allow for cytokine loading, we investigated whether the cytokine loading impacted the functional activities of Stim-R CAR T cells. We compared the optimized Stim-R formulation containing loaded cytokines to an analogous formulation that has the same αCD3/αCD28 density and stoichiometry, but that does not contain loaded cytokines. ROR1 CAR T cells were produced using these Stim-R variants or the benchmark reagent. In all conditions, the same concentration of cytokines was also supplemented into the media. T cell function was evaluated using the non-normalized repeat-stimulation assay. Across all donors tested, ROR1 CAR T cells produced using the Stim-R variants exhibited similar cytotoxic function, with both Stim-R products exhibiting improved cytotoxic function compared to the benchmark product (Fig. S3). These observations support that the improved function of the Stim-R ROR1 CAR T cells compared to the benchmark was not due to differences in cytokine loading or supplementation. In all subsequent work, the optimized Stim-R formulation containing loaded cytokines was used.
CAR T cells activated using Stim-R technology showed enhanced potency, expansion, and cytokine production in response to repeated tumor cell killing in vitro
The non-normalized repeat-stimulation assay is readily scalable to facilitate the screening of Stim-R formulation arrays. However, because the E:T ratio is not reset at every stimulation in this assay format, it does not permit the assessment of T-cell function on a per-cell basis. Therefore, to assess CAR T-cell functionality on a per-cell basis, we conducted a ‘normalized’ repeat-stimulation assay, in which the E:T ratio is reset to the initial ratio (1:5) at each restimulation (Fig. 4A).
CAR T cells activated using Stim-R technology exhibit enhanced in vitro function in normalized repeat-stimulation assays. (A) In the normalized repeat-stimulation assay, CAR T cells were exposed to repeated stimulations with ROR1 + H1975 cancer cells. The E:T ratio was reset to 1:5 at each restimulation to enable comparison of CAR T-cell product functionality on a per-cell basis. (B) Target clearance with CAR T cells activated using the optimized Stim-R formulation or the benchmark reagent in the normalized repeat-stimulation assay. Each panel shows the mean (± SD) of two technical replicates using products from an independent donor. (C) Expansion of CAR T cells activated using the optimized Stim-R formulation or the benchmark reagent before each restimulation in the normalized repeat-stimulation assay. Data are shown for individual donors (n = 3), with solid lines connecting the mean expansion at each stimulation. Stimulation 1 is shown in the graph to highlight the trend. Two-way ANOVA was performed using only data from stimulations 2–4 (p = 0.0031; F(2, 8) = 12.98 for interaction) with Šidák’s multiple comparisons test, *p = 0.0225 (t = 9.093; df = 2.296). (D) Concentration of IL-2 (left) and IFN-γ (right) in the culture supernatant 24 h after each stimulation. Data are shown for individual donors (n = 3) with black lines connecting data for products derived from the same donor.
In the normalized assay, CAR T cells activated using the optimized Stim-R formulation showed more persistent cytotoxic function compared with those produced using the benchmark reagent, across three independent donors (Fig. 4B). The percentages of CD4 + and CD8 + CAR T cells remained similar between the Stim-R and the benchmark groups throughout the assay, suggesting that the improved function of the Stim-R CAR T cells was not due to differences in the relative composition of CD4 + and CD8 + CAR T cells (Fig. S4). Furthermore, CAR T cells showed more sustained and increased expansion after each of two rounds of restimulation with tumor cells when activated using Stim-R technology compared with the benchmark reagent, with later peak T-cell expansion (Stimulation 3 versus 2, respectively), and a significantly higher peak expansion level (p = 0.0225) (Fig. 4C). These data are consistent with our findings utilizing the non-normalized assay and suggest that CAR T cells produced with Stim-R technology have a more sustained and higher proliferative potential than those produced with the benchmark reagent in response to repeated stimulation with tumor cells in vitro.
Compared with the benchmark product, CAR T cells activated with Stim-R technology showed comparable IFN-γ and IL-2 secretion at the first stimulation, but trends were observed for higher secretion of both cytokines at subsequent stimulations (Fig. 4d). These increases were most pronounced at the third stimulation, which was the timepoint at which both products inhibited tumor cell growth in the assay but the greatest difference in cytotoxicity was observed. Thus, improved proliferation and maintenance of cytokine production may account for the greater cytotoxicity of CAR T cells produced with Stim-R technology compared with those produced with the benchmark reagent.
CAR T cells activated using Stim-R technology retained a population of stem-like cells with effector-associated gene signatures and showed downregulation of exhaustion-associated gene sets
In order to interrogate the mechanisms underlying the increased potency of Stim-R CAR T cells, bulk RNA-Seq and single-cell CITE-Seq (which captures both RNA and protein expression) were performed on CD8 + CAR T cells produced using the optimized Stim-R formulation or the benchmark reagent at the end of the third stimulation with tumor cells (Day 10) in the normalized repeat-stimulation assay. Bulk RNA-Seq gene set enrichment analysis (GSEA) showed upregulation of both stemness-associated and effector-associated gene sets in cells produced using Stim-R technology compared with the benchmark reagent (Fig. 5A).
CD8 + CAR T cells activated with Stim-R technology retained a unique subset of stem-like cells with effector-associated gene signatures and displayed downregulation of exhaustion-associated gene sets. (A) Bulk RNA-Seq GSEA comparing gene set enrichment in CD8 + CAR T cells activated using Stim-R technology versus the benchmark reagent. The bar chart shows the negative loge of the FDR-adjusted p values, color-coded according to NES. (B) UMAP visualization of single-cell RNA-Seq data from CD8 + T cells activated using Stim-R technology (left, red plot) or the benchmark reagent (left, blue plot), and the composite (right). Putative stem-like clusters for cells activated using Stim-R technology (Cluster 1) or the benchmark reagent (Cluster 4) are indicated along with the range of proportion of cells within each cluster across three independent donors. (C) Single-cell RNA-Seq data showing expression of stemness- and effector-associated markers in CD8 + CAR T cells. The left and middle plots show expression level for stemness-associated markers (TCF7 RNA and CD27 protein) and effector-associated genes (GNLY and CCL5 RNA) on the composite UMAP visualization, with Cluster 1 indicated. The right graphs depict the distribution of TCF7 RNA and CD27 protein expression level in Clusters 1 and 4. Wilcoxon test, ****p < 0.0001. (D) Single-cell RNA-Seq data showing expression of exhaustion-associated genes in CD8 + CAR T cells. The left and middle plots show the TTE gene set module score (left) and TIGIT protein expression (middle) on the composite UMAP visualization. The right graph indicates the proportions of Stim-R technology- or benchmark-activated CD8 + cells within exhaustion-associated Clusters 8 and 14 in three individual donors, with black lines connecting data for the same donor. One-sided t-test, p = 0.067 (t = − 2.46; df = 2). (E) Bulk RNA-Seq GSEA plot for the TTE gene set. The green line indicates the running enrichment score as the algorithm walks through the gene list ranked according to differential expression in CD8 + T cells of the two products. Black lines indicate the positions of TTE genes within the ranking.
Based on the transcriptional profile, unsupervised clustering analysis of single-cell CITE-Seq data identified 15 cell clusters. Two clusters exhibiting enrichment for literature-based stemness markers were identified among cells activated using Stim-R technology (Cluster 1) or the benchmark reagent (Cluster 4) (Fig. 5B). The proportion of these clusters was comparable between conditions (Stim-R technology Cluster 1, 14–19%; benchmark Cluster 4, 10–16%). However, the expression of T-cell stemness markers TCF7 (based on RNA) and CD27 (based on protein) was significantly higher in Cluster 1 of Stim-R technology-activated cells compared with Cluster 4 of benchmark reagent-activated cells (both p < 0.0001). Interestingly, Cluster 1 of Stim-R technology-activated cells was also characterized by increased expression of the effector-associated genes GNLY and CCL5 (Fig. 5C). RNA expression levels of both GNLY and CCL5 were significantly up-regulated in Cluster 1 compared to Cluster 4 in differential expression analysis (CCL5: log2 fold-change = 0.98, false discovery rate [FDR]-adjusted p = 3 × 10–189; GNLY: log2 fold-change = 0.96, FDR-adjusted p = 2 × 10–181). These data suggest that CAR T cells produced with Stim-R technology retain a unique population of stem-like cells that have concomitant expression of effector-associated gene signatures, which may support their enhanced function.
To evaluate exhaustion, we identified two clusters that exhibited enriched module scores for the T-cell terminal exhaustion (TTE) gene set 36 and high T-cell immunoglobulin and ITIM domain (TIGIT) protein expression (Clusters 8 and 14) (Fig. 5D). We observed a trend in which the proportion of cells in these exhaustion-associated clusters was lower in CD8 + T cells activated with Stim-R technology compared with the benchmark reagent across the three donors. However, this difference did not reach statistical significance (p = 0.067). Consistent with this trend, CD8 + T cells activated with Stim-R technology also showed significant downregulation of the TTE gene set versus the benchmark in bulk RNA-Seq GSEA (normalized expression score [NES] = − 1.72; FDR-adjusted p = 0.005) (Fig. 5E). These data suggest that Stim-R technology-activated CAR T cells may show reduced exhaustion after exposure to tumor cells compared to benchmark reagent-activated CAR T cells, which could further support their enhanced function.
CAR T cells activated using Stim-R technology exhibited higher peak expansion, prolonged persistence, and improved tumor control in vivo
Finally, we compared the in vivo antitumor activity and pharmacokinetics (PK) of ROR1 CAR T cells activated using Stim-R technology versus the benchmark reagent in the ROR1 + H1975 non-small cell lung cancer (NSCLC) xenograft model (Fig. 6A)37,38. In one of two donors tested, CAR T cells produced using Stim-R technology showed improved control of established tumors in vivo compared with benchmark CAR T cells (Fig. 6B), and significantly prolonged overall survival of mice (p < 0.0001) (Fig. 6C). In a second donor, differences were not discernible between CAR T cells produced using Stim-R technology and benchmark CAR T cells because both products mediated similarly robust anti-tumor activity at the dose levels tested (Fig. S5).
CAR T cells activated using Stim-R technology showed improved expansion, persistence, and antitumor activity in a solid tumor xenograft model. (A) 5 × 106 ROR1 + H1975 cells were inoculated subcutaneously in NSG MHC I/II dKO mice. After 14 days, mice were infused with 2 × 106 CAR T cells (assessed as CD3 + EGFRopt + T cells) activated using Stim-R technology, the benchmark reagent, or an equivalent total number of mock T cells activated using Stim-R technology. Tumor volume was measured at least twice weekly for up to 60 days. Peripheral blood was collected for PK analysis once per week for up to five weeks. (B) Tumor volume over time in mice bearing ROR1 + H1975 xenografts after infusion of CAR T cells activated using Stim-R technology (left), Stim-R technology with mock transduction (middle), or the benchmark reagent (right) (n = 10 per group). (C) Kaplan–Meier curves showing overall survival in mice bearing ROR1 + H1975 xenografts after infusion of CAR T cells activated using Stim-R technology, Stim-R technology with mock transduction, or the benchmark reagent (n = 10 per group). Mantel–Cox test, ****p < 0.0001. (D) PK analyses showing the blood concentration of CAR T cells (assessed as CD3 + EGFRopt + cells) over time (left) and blood concentration 24 h after infusion, at peak, and cumulative levels calculated as area under the curve (AUC; right) (n = 10 per group). AUC was calculated using PK data up to day 28. Only mice for which PK data could be collected for all timepoints were included in the analysis (n = 8 for the benchmark group, n = 10 for the Stim-R group). Data represent the mean (± SEM) of log-transformed data. Data points from individual mice are overlaid. Data in right graph of d, left were analyzed using a two-tailed Mann–Whitney test ns, not significant (p = 0.0730). Data in right graph of d, middle and right were analyzed using a two-tailed t-test; middle: ****p < 0.0001 (t = 14.68; df = 18); right: ****p < 0.0001 (t = 12.24; df = 16).
PK analyses showed that CAR T cells produced using Stim-R technology reached a 30-fold higher peak concentration and 25-fold higher cumulative level (assessed as area under the time–concentration curve [AUC]) over the duration of the study (28 days) than CAR T cells produced with the benchmark product (Fig. 6D; both p < 0.0001). The increased peak expansion and cumulative levels suggest an improved rate of proliferation since similar levels of engraftment were observed at 24 h for both treatments. The substantially higher peak expansion and prolonged persistence in the peripheral blood of CAR T cells activated using Stim-R technology are consistent with their enhanced and sustained expansion in vitro in the normalized repeat-stimulation assay (Fig. 4C). These data show that T cells activated with Stim-R technology exhibited improved expansion and enhanced antitumor activity, resulting in longer overall survival in vivo compared with T cells activated with benchmark reagent.
Discussion
In the present work, we leveraged Stim-R technology, a synthetic cell biomimetic that emulates the physiologic, cell-like presentation of signal molecules25,26,30 and enables precise signal molecule presentation, to demonstrate that optimizing cell signaling parameters during T-cell activation can enhance CAR T-cell functionality. While a variety of commercial T-cell–activating reagents are available, including those based on microbeads, nanobeads, or multimeric macromolecules25,39,40, many were primarily developed for promoting T-cell expansion rather than for enabling more functional ACT products. Utilizing the Stim-R platform in a broad screening program alongside multiple assays aimed at assessing different functional attributes of the CAR T-cell products, we identified optimal αCD3/αCD28 signal presentation parameters in the context of a specific ROR1-targeted CAR construct and production protocol. The optimized Stim-R formulation generated a more potent T-cell product with enhanced in vitro function and improved in vivo PK and antitumor activity in a solid tumor xenograft model compared with an analogous product generated using a conventional T-cell–activating reagent.
CAR T cells activated using the optimized Stim-R formulation showed improved persistence, as evidenced by significantly increased proliferation in vitro and profoundly improved PK in vivo, alongside greater polyfunctionality and decreased exhaustion. These features have been correlated with the clinical activity of CAR T-cell therapies35,41. For example, CAR T-cell peak expansion and persistence have been associated with both the attainment of clinical response and the durability of benefit in patients with various hematologic malignancies33,34. Furthermore, response to CD19-targeted CAR T-cell therapies has been correlated with pre-infusion product attributes including greater cytokine polyfunctionality35 and reduced T-cell exhaustion as reflected in exhaustion-associated gene signatures42,43.
CD8 + T cells activated with Stim-R technology retained a unique population of stem-like cells that also expressed effector-associated gene signatures. Retention of early memory or stem-like T cells, which have the capacity for self-renewal and are likely important for long-term persistence of adoptive T-cell products, has been correlated with improved clinical response17,28,44. While it is conventionally understood that more differentiated effector T-cell compartments arise from a progenitor stem-like compartment and that the respective attributes of these compartments tend to be mutually exclusive28,29,45,46, our data suggest that activation of CAR T cells using Stim-R technology gives rise to a population with both stem-like and effector-like traits following exposure to antigen stimulation. If these transcriptomic signatures translate to function, the ability of this unique population to persist while actively mediating antitumor cytotoxicity could underlie the improved CAR T-cell function we observed. Further experiments employing multiomics (e.g., epigenetics and metabolomics), lineage tracing, and functional assessments of sorted cell populations are required to understand the origin and behavior of this cell population and to assess its causal relationship to improvements in CAR T-cell potency.
Many ongoing efforts exist to create more potent CAR T cells by genetically engineering T cells to express additional proteins (‘armored’ CAR T cells)12,47, knock out specific genes48,49, or by including small-molecule agents within the T-cell production process50,51. Here, we show that modulating T-cell activation signals with Stim-R technology represents a simple method for improving the potency of CAR T-cell products which could potentially complement these other approaches. Our data demonstrate that changes to the T-cell activation stimulus can generate CAR T-cell products with vastly different functions and unique transcriptomic profiles following exposure to tumor cells. Importantly, this approach for enhancing the potency of T-cell products can be applied to any therapeutic T-cell production process and is highly clinically translatable.
In conclusion, optimizing signal presentation during T-cell activation using Stim-R technology versus a conventional T-cell–activating reagent enabled the production of ROR1-targeted CAR T cells with greater potency. The optimized Stim-R formulation resulted in a CAR T-cell product with sustained cytotoxicity, proliferation, and cytokine production in response to persistent antigen stimulation in vitro and improved PK and antitumor activity in vivo. Enhancement of T-cell products by programming cell signaling during T-cell activation with Stim-R technology is a promising approach to improve CAR T-cell therapeutic efficacy against solid tumors.
Methods
Samples
Primary human CD4 + and CD8 + T cells from healthy donors were purchased from AllCells (Alameda, CA, USA). The H1975 cell line was purchased from ATCC (Manassas, VA, USA).
Chemicals and reagents
Mesoporous silica micro-rods (MSRs) were synthesized by nanoComposix (San Diego, CA, USA) utilizing an optimized process based on a previously described protocol25. Both 16:0–18:1 PC (POPC) and 18:1 Biotinyl Cap PE were purchased from Avanti Polar Lipids (Alabaster, AL, USA). Streptavidin and biotinylated agonistic antibodies for CD3 and CD28 were purchased from BioLegend (San Diego, CA, USA). D-(+)-trehalose dihydrate (trehalose) was purchased from VWR (Radnor, PA, USA). Custom T-cell culture media was purchased from Lonza (Basel, Switzerland) and supplemented with a cocktail of recombinant human cytokines purchased from R&D Systems (Minneapolis, MN, USA). T-cell TransAct was purchased from Miltenyi Biotec and used in studies as a conventional T-cell–activating reagent benchmark. CryoStor® CS10 (BioLife Solutions, Bothell, WA, USA) was purchased from Fisher Scientific (Hampton, NH, USA). Antibodies and reagents for flow cytometry were purchased from BD Biosciences, Thermo Fisher Scientific, BioLegend or Cell Signaling Technology (Danvers, MA, US), as described in Table S1. Fixable viability dye eFluor 780 was purchased from Thermo Fisher Scientific. Fc receptor-blocking solution and cell staining buffer (CSB) were purchased from BioLegend. A transcription factor staining buffer set was purchased from Thermo Fisher Scientific. Cytofix/Cytoperm Fixation/Permeabilization Solution Kit with GolgiPlug was purchased from BD Biosciences (Franklin Lakes, NJ, USA). Antibodies used for sorting and CITE-Seq were purchased from BioLegend, as described in Table S2.
Stim-R technology assembly
A detailed protocol for the assembly of Stim-R technology is available online.25 In brief, liposomes were prepared by dehydrating lipid films, comprising POPC with between 0.1 and 1 mol% 18:1 Biotinyl Cap PE, under nitrogen, rehydrating the lipid films in phosphate buffered saline (PBS), and extruding liposomes through a 100 nm pore size polycarbonate filter using a mini-extruder (Avanti Polar Lipids). Liposomes were stored at 4 °C and used within 24 h. MSRs (nanoComposix) were mixed with a cocktail of IL-2, IL-7, and IL-15 for 1 h at room temperature with gentle mixing every 10 min. Then, liposomes were mixed in a 1:2 w/w (lipid:MSR) ratio for 1 h at room temperature with gentle mixing every 10 min and washed twice with PBS. After washing, the lipid-coated MSRs were blocked using 0.25% w/v bovine serum albumin in PBS for 10 min. Then, streptavidin was added at 30% theoretical saturation of the biotinylated lipid in the respective formulation (based on prior characterization studies30) for 10 min with gentle mixing every 2 min. Biotinylated T-cell–activating cues (αCD3 and αCD28), pre-mixed at αCD28:αCD3 ratios varying between 0.1 and 5.0, were added at 80% molar saturation of the added streptavidin for 40 min with gentle mixing every 10 min, forming Stim-R technology. The Stim-R material was washed twice with PBS. Following the second wash, the Stim-R material was resuspended at 10 mg/ml in PBS containing 10% w/v trehalose, snap frozen in liquid nitrogen, and lyophilized overnight using a Freezone Triad benchtop freeze dryer (Labconco, Kansas City, MO, USA). Lyophilized Stim-R material was stored in a desiccant chamber at 4 °C until used.
CAR T-cell production
Vector production
The lentiviral vector (LVV) was generated using a third-generation vector system. For this, a suspension culture of HEK 293T cells was grown in Freestyle 293 Expression Medium (Thermo Fischer Scientific). The cells were transfected with the necessary helper plasmids expressing gag-pol, rev, and envelope-pseudotype VSV-G, as well as the transfer plasmid containing a lentiviral vector sequence co-expressing a CAR targeting ROR1 and EGFRopt52, a functionally inert cell surface marker for transduced cells. LVV-containing supernatant was harvested, concentrated and purified. Finally, the concentrated LVV was titered and stored at—80 °C until use.
T-cell activation, transduction, and expansion
Primary human CD4 + and CD8 + T cells were mixed 1:1 and seeded in a tissue culture (TC) plate at 1–1.5 × 106 cells/mL in T-cell media. Stim-R material was added to cultures at 200 mg per 1 × 106 cells to activate T cells with Stim-R technology. TransAct suspension was added to cultures at 1% v/v per the manufacturer’s recommendations to activate T cells with TransAct. CAR-encoding LVV was added to cultures at a multiplicity of infection of 7.5. T cells activated with either Stim-R technology or TransAct were expanded for 7 days utilizing a production process based on a proprietary manufacturing protocol (Epi-R manufacturing protocol, Lyell Immunopharma, San Francisco, CA, USA). On day 7, CAR T-cell products were harvested, total live cells were counted, and T-cell products were cryopreserved at ≥ 1 × 107 live cells/mL in CryoStor CS10 using a controlled rate freezer (Thermo Fisher Scientific). Cryopreserved CAR T-cell products were stored in liquid nitrogen until used.
T-cell product phenotyping using flow cytometry
All antibodies and staining reagents are listed in Table S1. Antibodies were titrated and a staining concentration for each antibody was selected based on the optimal staining index.
Cells were washed with CSB and stained with a cocktail containing Fc receptor-blocking solution and CCR7 antibody at 37 °C for 15 min in the dark. Then a cocktail containing the remaining cell surface marker antibodies (CD3, CD4, CD8, EGFR, CD45RA, CD45RO, CD62L, CD27, CD28, CD25, PD1, Lag3, and CD39) and fixable viability dye was added, and staining was allowed to proceed at 4 °C for 20 min in the dark. After surface staining, cells were fixed and permeabilized using a transcription factor staining buffer set (Thermo Fisher Scientific) per the manufacturer’s instructions. Cells were stained with a cocktail containing intracellular marker antibodies (LEF1, TCF1) at 4 °C for 15 min in the dark. Samples were washed and resuspended in CSB for flow cytometry analysis.
Samples were assessed using a Cytek Aurora Spectral Flow Cytometer (Cytek, Fremont, CA, USA) and analyzed using FlowJo version 10.6.1 software (FlowJo, Ashland, OR, USA). The following gating strategy was used to evaluate CAR transduction efficiency: forward scatter/side scatter > single cells > viable cells > CD3 +> EGFRopt+. To assess the memory phenotype of T-cell products, fluorescence minus one controls were used for gating.
Repeat-stimulation assays
Non-normalized repeat-stimulation assay
A total of 2 × 104 ROR1 + H1975 target cells expressing the fluorescent marker IncuCyte® NucLight Rapid Red (NLR; Sartorius, Essen Biosciences, Ann Arbor, MI, USA) were added to 96-well plates and allowed to adhere for ≥ 2 h. At the first stimulation, CAR T cells were added at an E:T ratio of 1:5, and cell killing was monitored in real-time by tracking NLR fluorescence intensity using an IncuCyte Live Cell imager (Sartorius). For all E:T ratio calculations, the number of effector cells was defined as the number of CD3 + EGFRopt + CART cells input into the assay. Cells were restimulated every 3–4 days by transferring 25% of the nonadherent cells from the previous coculture to a new plate with 2 × 104 freshly plated target cells. Restimulations were continued until no CAR T-cell samples in the experiment inhibited tumor cell growth. The killing curves are presented for each round of the restimulation until both conditions have failed to inhibit tumor cell growth.
Normalized repeat-stimulation assay
ROR1+, NLR-expressing H1975 target cells were subjected to an initial stimulation as described for the non-normalized assay. In parallel, CAR T cells were cocultured with target cells at the same E:T ratio (1:5) in separate large vessels (‘large-scale cocultures’) and maintained in the incubator. Restimulations were performed every 3–4 days by resetting the E:T ratio for each coculture to 1:5. Briefly, nonadherent cells from the large-scale cocultures were collected, counted, and assessed via flow cytometry for CAR expression, using EGFR expression as a proxy with the following gating strategy: forward scatter/side scatter > single cells > viable cells > CD3 + > EGFRopt + (Fig. S6a). CAR T cells collected from large-scale cocultures from the previous stimulation were added at an E:T ratio of 1:5 to a new plate with 2 × 104 freshly plated target cells for IncuCyte monitoring, and a new large-scale coculture was set up with the remaining CAR T cells at the same E:T ratio and maintained in the incubator. Target cell killing was monitored in real-time by tracking NLR fluorescence intensity using an IncuCyte Live Cell imager (Sartorius). Restimulations were continued until no CAR T-cell samples in the experiment inhibited tumor cell growth. The killing curves are presented for each round of the restimulation until both conditions have failed to inhibit tumor cell growth. Samples were collected at the end of the third stimulation (Day 10) for transcriptomic analyses.
Intracellular cytokine staining (ICS)
Cytokine production was quantified via ICS, followed by flow cytometry, to assess the production of IFN-γ and IL-2 by CAR T cells in response to ROR1 + target cells. Briefly, 4 × 104 EGFRopt + T cells were co-cultured with 2 × 105 ROR1 + H1975 target cells (1:5 E:T ratio) in 96 well non-TC-treated u-bottom plates. Cells were concentrated by centrifuging at 500 rcf for 1 min and placed in the incubator for 1 h. After 1 h, GolgiPlug (BD Biosciences) was added according to the manufacturer’s recommendations. Samples were mixed and then cells were concentrated by centrifuging at 500 rcf for 1 min. The samples were placed in the incubator for 3.5 h. Following the incubation, cells were collected and washed with CSB. Samples were then stained with a cocktail containing Fc receptor-blocking solution, fixable viability dye, and cell surface marker antibodies (CD45, CD3, CD4, CD8, and EGFR) at 4 °C for 20 min in the dark. After surface staining, cells were fixed and permeabilized with Cytofix/Cytoperm fixation/permeabilization kit (BD Biosciences) per the manufacturer’s instructions. Cells were stained with a cocktail containing intracellular antibodies (IL-2, IFN-γ) at 4 °C for 20 min in the dark. Samples were washed and resuspended in CSB for flow cytometry analysis.
Samples were assessed using a Cytek Aurora Spectral Flow Cytometer and analyzed using FlowJo version 10.6.1 software. The following gating strategy was used to evaluate IL-2 and IFN-γ production: forward scatter/side scatter > single cells > CD45 + lymphocytes > viable cells > EGFRopt +> IL-2, IFN-γ (Fig. S6B).
Soluble cytokine assays
Quantification of secreted IFN-γ and IL-2 in cell culture supernatants was conducted using the V-PLEX Proinflammatory Panel 1 Human Kit (Meso Scale Diagnostics, Rockville, MD, USA) according to the manufacturer’s guidelines, and the plates were read using an MSD Sector S 600 Imager. Cytokine concentrations were calculated using the MSD Workbench software based on the dilution factor of the supernatants and the concentration of the calibrators.
Transcriptomic analyses
Bulk RNA-Seq
Bulk RNA-Seq was performed on live CD45 + EGFR + CD8 + T cells sorted from CAR T cells activated using Stim-R technology or the benchmark reagent collected from three independent donors at the end of the third stimulation (Day 10) in the normalized repeat-stimulation assay. For bulk RNA-Seq, cryopreserved samples were thawed at 37 °C and diluted at least 1:10 by volume with pre-warmed RP10 (RPMI + 10% FBS) media and collected by centrifugation. Sample cell counts and viability were assessed using the Cellaca MX Automated Cell Counter (Nexcelom, Lawrence, MA, USA). Cells were aliquoted for flow staining, followed by Fc receptor blocking using Human TruStain FcX blocking reagent (BioLegend) for 10 min at 4 °C, prior to staining with a cocktail of fluorochrome-conjugated antibodies specific for human CD45, CD8, CD4, EGFR and fixable viability dye eFluor 506 (Table S2) for 30 min at 4 °C. Approximately 3 × 104 live CD45 + EGFR + CD8 + T cells were flow sorted into 100 μL of Lysis Binding Mix (Thermo Fisher Scientific) using the BD FACSAria Fusion cell sorter (BD Biosciences) and stored at − 80 °C until processing. Total RNA was extracted from thawed samples using the MagMAX mirVana RNA kit on the KingFisher Flex System (Thermo Fisher Scientific) and stored at − 80 °C. RNA quality and quantity were assessed using the Agilent RNA 6000 Pico Kit on the 2100 BioAnalyzer (Agilent, Santa Clara, CA, USA). Libraries for mRNA sequencing were prepared with 5 ng of total RNA for cDNA generation using the SMART-Seq v4 Ultra Low Input RNA kit (Takara, Kusatsu, Shiga, Japan) following the manufacturer’s protocol, automated on the Biomek i7 (Beckman Coulter, Brea, CA, USA). The cDNA assessment was performed using the High Sensitivity DNA kit on the 2100 BioAnalyzer (Agilent), and 150 pg of cDNA was used to prepare barcoded libraries using the Nextera XT DNA Library Preparation Kit (Illumina, San Diego, CA, USA) following the manufacturer’s protocol, automated on the Biomek i7 (Beckman Coulter). Barcoded libraries from multiple samples were then pooled in an equimolar ratio, purified, and sequenced using a NovaSeq 6000 (Illumina). Around 2.5 × 107 paired-end 50-base-pair reads were generated per sample.
For bulk RNA-Seq data analysis, Trim_galore (Krueger—Github) was used for adaptor trimming and Spliced Transcripts Alignment to a Reference (STAR)53 was used for alignment to the human GRCh38 reference genome. Gene expression quantification was performed by RNA-Seq by Expectation Maximization (RSEM)54 with default parameters. Differential analysis was performed using the DESeq255 package. GSEA was performed using the fast gene set enrichment analysis (fgsea)56 package in R with a priori-defined gene sets from Molecular Signatures Database (MSigDB, version 7)57 and the literature. The p values yielded by GSEA were corrected for multiple comparisons using FDR adjustment.
Single-cell CITE-Seq
Single-cell CITE-Seq was performed on live CD45 + EGFR + T cells sorted from CAR T cells activated using Stim-R technology or the benchmark reagent collected from three independent donors at the end of the third stimulation (Day 10) in the normalized repeat-stimulation assay. As described for bulk RNA-Seq, cryopreserved samples were prepared for flow sorting. Cells were blocked with Human TruStain FcX blocking reagent (BioLegend) for 10 min at 4 °C, prior to staining with a mix of fixable viability dye eFluor 506; fluorochrome-conjugated antibodies specific for human CD45, CD8, CD4, EGFR; DNA-conjugated Total-Seq C antibodies against cell surface protein markers (Table S2), and unique DNA-barcoded Hashtag antibodies (Table S2) for 30 min at 4 °C. Live CD45 + EGFR + CD8 + T cells and CD45 + EGFR + CD4 + T cells were flow sorted into PBS containing 2% BSA, using the BD FACSAria Fusion cell sorter (BD Biosciences). Sorted cells from samples barcoded with unique hashtag antibodies were pooled together and then counted using the Cellometer K2 Fluorescent Cell Counter (Nexcelom). Approximately 5 × 104 pooled cells were loaded into each channel of the Chromium Next GEM Chip K using a Chromium Next GEM Single Cell 5′ Reagent Kit v2 (10 × Genomics, Pleasanton, CA, USA). After single-cell capture and lysis, cDNA was generated, and DNA conjugated to antibodies bound to cell surface proteins was amplified over 13 cycles according to the manufacturer’s protocol (10 × Genomics). The amplified cDNA from polyadenylated mRNA and the amplified DNA from cell surface protein antibody barcodes were separated by size selection for generating independent barcoded 5′ Gene Expression (GEX) libraries and Antibody derived tag (ADT) libraries, respectively. The GEX and ADT libraries prepared from multiple channels of the Chromium Next GEM Chip K were quantified, pooled, and sequenced together using the NovaSeq 6000 (Illumina). An average sequencing depth of at least 2 × 104 reads per cell for GEX libraries and at least 5 × 103 reads per cell for ADT libraries was obtained.
Single-cell CITE-Seq data was processed using the 10X Cell Ranger software version 6.1.2 (10 × Genomics) with GRCh38 as reference genome and standard parameters. The cell-gene matrix was further processed using Seurat package58 in R. In brief, cells were first filtered using thresholds for quality control metrics (percent mitochondria, nCount_RNA, nFeature_RNA, hashtag empty droplets, and doublets). Then CD8 + T cells were selected by gating on CD8, and CD4 protein expression was measured by CITE-Seq. The CD8 + T cells from the three donors were further analyzed. The CD4 + T cells were not analyzed due to insufficient cell number. The filtered cell-gene matrix was transformed and scaled, including variable feature selection, with the ‘LogNormalize’ pipeline for each donor separately. T cells from the three donors were then integrated using selected features. The FindIntegrationAnchors and IntegrateData functions in Seurat were used to account for donor heterogeneity. Mitochondria, ribosome, TCR, and immunoglobulin complex related genes were excluded from selected features for integration. The effects of cell cycle heterogeneity were corrected by calculating cell cycle phase scores (G2M.Score, S.Score) using the CellCycleScoring function in Seurat and excluding genes correlated with either of the two cell cycle phase scores (with Pearson correlation coefficient greater than 0.25) from selected features for integration. The cell cycle phase scores were further regressed from the data using the ScaleData function. Afterward, Uniform Manifold Approximation and Projection (UMAP) was performed using the RunUMAP function in Seurat to map cells to a two-dimensional space with each dot representing a cell. Cells were subjected to clustering analysis using the FindClusters function in Seurat. CITE-Seq analysis (UMAP, clustering) was performed using RNA expression without protein expression in the T cells.
Solid tumor xenograft model
Animals, housing, and ethical declarations
NSG MHCI/II dKO mice (NOD.Cg-Prkdcscid H2-K1b-tm1Bpe H2-Ab1g 7-em1Mvw H2-D1b-tm1Bpe Il2rgtm1Wjl/SzJ, strain #025216) were purchased from The Jackson Laboratory (Sacramento, CA, USA) and were housed, treated, and maintained at Lyell Immunopharma in an animal facility operated by Explora BioLabs (South San Francisco, CA, USA). All animal procedures were approved and conducted under Explora’s Institutional Animal Care and Use Committee protocol #EB17-010–117. All studies are reported in accordance with ARRIVE guidelines.
Tumor cell implantation and CAR T-cell infusion
Human ROR1 + NSCLC-derived H1975 tumor cells were implanted in the flanks of 8- to 10-week-old, female NSG MHCI/II dKO mice. When tumors reached 125 mm3 in size (range, 100–150 mm3), animals received a single dose of 2 × 106 CAR T cells produced with either Stim-R or a conventional T-cell activation reagent, or a dose of untransduced mock T cells produced with Stim-R that is equivalent to the total T-cell dose of 2 × 106 CAR T cells, by intravenous infusion via the tail vein.
Tumor measurements
Body weight and tumor volume were measured two to three times per week, and clinical observations were performed daily. Tumor volumes and survival curves were visualized using Prism 10.0.3 (GraphPad Software). Mean tumor curves ± standard error of the mean (SEM) were generated for each treatment group from the day of randomization until animals were taken off study for reaching predefined euthanasia criteria of either tumor volume exceeding 2000 mm3, body weight loss greater than 20% from the pre-study value, observation of open weeping tumor ulceration, severe respiratory distress, severely impaired movement, or loss of righting reflex.
Pharmacokinetic analyses
Peripheral blood was collected at 24 h and then once a week for 5 weeks after T cell infusion. In brief, ~ 100 μL of blood was collected from fully anesthetized animals via retro-orbital collection into EDTA-coated tubes. After three rounds of red blood cell lysis using ACK lysis buffer (Gibco), cells were blocked using 10% normal mouse serum (Jackson ImmunoResearch, West Grove, PA, USA) for 10 min at room temperature, followed by staining with an antibody mix (mouse CD45, human CD45, human CD3, human CD4, human CD8, human EGFR, and a ROR1 CAR-specific anti-idiotype [clone 48F6G5E1, Lyell Immunopharma, San Francisco, CA, USA]) in CSB containing 10% Brilliant Plus Buffer for 20 min at room temperature. Cells were washed and stained with LiveDead dye before fixation using BD Cytofix buffer (BD Biosciences). After fixation, cells were washed and resuspended in 50 mL CSB containing 5 × 103 123count eBeads (Thermo Fisher Scientific). Flow cytometry analysis was performed using the ZE5 cytometer (Bio-Rad Laboratories, Hercules, CA, USA), and the results were analyzed using FCS Express 7 (DeNovo Software, Pasadena, CA, USA). Cell counts were calculated using the concentration calculator module and 1 × 105 beads/mL as a reference value for the bead concentration.
Statistical analyses
Statistical analyses were performed using Prism Version 10.0.3 (GraphPad Software, Boston, MA, USA). Data were summarized as means ± SD or SEM. The equality of variance was tested using an F-test. Samples with equal variance were compared using appropriate parametric tests (e.g., paired one- or two-sided Student’s t-test or two-way analysis of variance [ANOVA] with Šidák’s multiple comparisons test), while samples of unequal variance were compared using appropriate nonparametric tests (e.g., Wilcoxon signed rank test or Mann–Whitney test). In the GSEA, p values were corrected for multiple comparisons using FDR adjustment. Survival in the xenograft model was analyzed using the Kaplan–Meier estimator and compared between groups using the Mantel–Cox test. The significance level was p < 0.05. Details of statistical tests are provided in the figure legends.
Data availability
Data from transcriptomic analyses are available in the Gene Expression Omnibus (GEO) repository: single-cell CITE-Seq data is available in GSE269140 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE269140) and bulk RNA-Seq data is available in GSE269134 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE269134). All other data supporting the findings of this study are available within the paper and its supplementary information files.
References
Sung, H. et al. Global Cancer Statistics 2020: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J. Clin. 71, 209–249 (2021).
Neelapu, S. S. et al. Axicabtagene ciloleucel CAR T-cell therapy in refractory large B-Cell lymphoma. N. Engl. J. Med. 377, 2531–2544 (2017).
Maude, S. L. et al. Tisagenlecleucel in children and young adults with B-cell lymphoblastic leukemia. N. Engl. J. Med. 378, 439–448 (2018).
Berdeja, J. G. et al. Ciltacabtagene autoleucel, a B-cell maturation antigen-directed chimeric antigen receptor T-cell therapy in patients with relapsed or refractory multiple myeloma (CARTITUDE-1): A phase 1b/2 open-label study. Lancet 398, 314–324 (2021).
Del Bufalo, F. et al. GD2-CART01 for relapsed or refractory high-risk neuroblastoma. N. Engl. J. Med. 388, 1284–1295 (2023).
Mackensen, A. et al. CLDN6-specific CAR-T cells plus amplifying RNA vaccine in relapsed or refractory solid tumors: The phase 1 BNT211–01 trial. Nat. Med. 29, 2844–2853 (2023).
Guzman, G., Reed, M. R., Bielamowicz, K., Koss, B. & Rodriguez, A. CAR-T therapies in solid tumors: Opportunities and challenges. Curr. Oncol. Rep. 25, 479–489 (2023).
Yamamoto, T. N. et al. T cells genetically engineered to overcome death signaling enhance adoptive cancer immunotherapy. J. Clin. Investig. 129, 1551–1565 (2019).
Philip, M. et al. Chromatin states define tumour-specific T cell dysfunction and reprogramming. Nature 545, 452–456 (2017).
Sade-Feldman, M. et al. Defining T cell states associated with response to checkpoint immunotherapy in melanoma. Cell 175, 998-1013.e1020 (2018).
Hou, A. J., Chen, L. C. & Chen, Y. Y. Navigating CAR-T cells through the solid-tumour microenvironment. Nat. Rev. Drug Discov. 20, 531–550 (2021).
Lynn, R. C. et al. c-Jun overexpression in CAR T cells induces exhaustion resistance. Nature 576, 293–300 (2019).
Eyquem, J. et al. Targeting a CAR to the TRAC locus with CRISPR/Cas9 enhances tumour rejection. Nature 543, 113–117 (2017).
Fraietta, J. A. et al. Disruption of TET2 promotes the therapeutic efficacy of CD19-targeted T cells. Nature 558, 307–312 (2018).
June, C. H., O’Connor, R. S., Kawalekar, O. U., Ghassemi, S. & Milone, M. C. CAR T cell immunotherapy for human cancer. Science 359, 1361–1365 (2018).
Wherry, E. J. et al. Molecular signature of CD8+ T cell exhaustion during chronic viral infection. Immunity 27, 670–684 (2007).
Vodnala, S. K. et al. T cell stemness and dysfunction in tumors are triggered by a common mechanism. Science 363, eaau0135 (2019).
Hwang, J.-R., Byeon, Y., Kim, D. & Park, S.-G. Recent insights of T cell receptor-mediated signaling pathways for T cell activation and development. Exp. Mol. Med. 52, 750–761 (2020).
Kaech, S. M. & Cui, W. Transcriptional control of effector and memory CD8+ T cell differentiation. Nat. Rev. Immunol. 12, 749–761 (2012).
Gett, A. V., Sallusto, F., Lanzavecchia, A. & Geginat, J. T cell fitness determined by signal strength. Nat. Immunol. 4, 355–360 (2003).
Chen, Y., Zander, R., Khatun, A., Schauder, D. M. & Cui, W. Transcriptional and epigenetic regulation of effector and memory CD8 T cell differentiation. Front. Immunol. 9, 2826 (2018).
Bousso, P. T-cell activation by dendritic cells in the lymph node: Lessons from the movies. Nat. Rev. Immunol. 8, 675–684 (2008).
Huppa, J. B. & Davis, M. M. T-cell-antigen recognition and the immunological synapse. Nat. Rev. Immunol. 3, 973–983 (2003).
Sckisel, G. D. et al. Out-of-sequence signal 3 paralyzes primary CD4(+) T-cell-dependent immunity. Immunity 43, 240–250 (2015).
Zhang, D. K. Y., Cheung, A. S. & Mooney, D. J. Activation and expansion of human T cells using artificial antigen-presenting cell scaffolds. Nat. Protoc. 15, 773–798 (2020).
Zhang, D. K. Y. et al. Enhancing CAR-T cell functionality in a patient-specific manner. Nat. Commun. 14, 506 (2023).
Kagoya, Y. et al. Transient stimulation expands superior antitumor T cells for adoptive therapy. JCI Insight 2, e89580 (2017).
Gattinoni, L. et al. Acquisition of full effector function in vitro paradoxically impairs the in vivo antitumor efficacy of adoptively transferred CD8+ T cells. J. Clin. Invest. 115, 1616–1626 (2005).
Cheng, L. E., Ohlén, C., Nelson, B. H. & Greenberg, P. D. Enhanced signaling through the IL-2 receptor in CD8+ T cells regulated by antigen recognition results in preferential proliferation and expansion of responding CD8+ T cells rather than promotion of cell death. Proc. Natl. Acad. Sci. U. S. A. 99, 3001–3006 (2002).
Cheung, A. S., Zhang, D. K. Y., Koshy, S. T. & Mooney, D. J. Scaffolds that mimic antigen-presenting cells enable ex vivo expansion of primary T cells. Nat. Biotechnol. 36, 160–169 (2018).
Zhao, Y. et al. Tyrosine kinase ROR1 as a target for anti-cancer therapies. Front. Oncol. 11, 680834 (2021).
Balakrishnan, A. et al. Analysis of ROR1 protein expression in human cancer and normal tissues. Clin. Cancer Res. 23, 3061–3071 (2017).
Mueller, K. T. et al. Clinical pharmacology of tisagenlecleucel in B-cell acute lymphoblastic leukemia. Clin. Cancer Res. 24, 6175–6184 (2018).
Melenhorst, J. J. et al. Decade-long leukaemia remissions with persistence of CD4+ CAR T cells. Nature 602, 503–509 (2022).
Rossi, J. et al. Preinfusion polyfunctional anti-CD19 chimeric antigen receptor T cells are associated with clinical outcomes in NHL. Blood 132, 804–814 (2018).
Oliveira, G. et al. Phenotype, specificity and avidity of antitumour CD8+ T cells in melanoma. Nature 596, 119–125 (2021).
Liu, M. et al. Targeting PD-L1 in non-small cell lung cancer using CAR T cells. Oncogenesis 9, 72 (2020).
Lee, B. K. et al. Developing ROR1 targeting CAR-T cells against solid tumors in preclinical studies. Cancers (Basel) 14, 3618 (2022).
Wang, X. & Rivière, I. Clinical manufacturing of CAR T cells: Foundation of a promising therapy. Mol. Ther. Oncol. 3, 16015 (2016).
Turtle, C. J. & Riddell, S. R. Artificial antigen-presenting cells for use in adoptive immunotherapy. Cancer J. 16, 374–381 (2010).
Xu, H., Li, N., Wang, G. & Cao, Y. Predictive short/long-term efficacy biomarkers and resistance mechanisms of CD19-directed CAR-T immunotherapy in relapsed/refractory B-cell lymphomas. Front. Immunol. 14, 1110028 (2023).
Fraietta, J. A. et al. Determinants of response and resistance to CD19 chimeric antigen receptor (CAR) T cell therapy of chronic lymphocytic leukemia. Nat. Med. 24, 563–571 (2018).
Deng, Q. et al. Characteristics of anti-CD19 CAR T cell infusion products associated with efficacy and toxicity in patients with large B cell lymphomas. Nat. Med. 26, 1878–1887 (2020).
Krishna, S. et al. Stem-like CD8 T cells mediate response of adoptive cell immunotherapy against human cancer. Science 370, 1328–1334 (2020).
Gattinoni, L., Speiser, D. E., Lichterfeld, M. & Bonini, C. T memory stem cells in health and disease. Nat. Med. 23, 18–27 (2017).
Sun, Q. et al. BCL6 promotes a stem-like CD8(+) T cell program in cancer via antagonizing BLIMP1. Sci. Immunol. 8, eadh1306 (2023).
Narayan, V. et al. PSMA-targeting TGFβ-insensitive armored CAR T cells in metastatic castration-resistant prostate cancer: A phase 1 trial. Nat. Med. 28, 724–734 (2022).
Jung, I. Y. et al. CRISPR/Cas9-mediated knockout of DGK improves antitumor activities of human T cells. Cancer Res. 78, 4692–4703 (2018).
Chen, J. et al. NR4A transcription factors limit CAR T cell function in solid tumours. Nature 567, 530–534 (2019).
Petersen, C. T. et al. Improving T-cell expansion and function for adoptive T-cell therapy using ex vivo treatment with PI3Kδ inhibitors and VIP antagonists. Blood Adv. 2, 210–223 (2018).
Urak, R. et al. Ex vivo Akt inhibition promotes the generation of potent CD19CAR T cells for adoptive immunotherapy. J. Immunother. Cancer 5, 26 (2017).
Shabaneh, T. B. et al. Safety switch optimization enhances antibody-mediated elimination of CAR T cells. Front. Mol. Med. 2, 1026474 (2022).
Dobin, A. et al. STAR: Ultrafast universal RNA-seq aligner. Bioinformatics 29, 15–21 (2013).
Li, B. & Dewey, C. N. RSEM: Accurate transcript quantification from RNA-Seq data with or without a reference genome. BMC Bioinform. 12, 323 (2011).
Love, M. I., Huber, W. & Anders, S. Moderated estimation of fold change and dispersion for RNA-seq data with DESeq2. Genome Biol. 15, 550 (2014).
Korotkevich, G., et al. Fast gene set enrichment analysis. bioRxiv, 060012 (2021).
Subramanian, A. et al. Gene set enrichment analysis: A knowledge-based approach for interpreting genome-wide expression profiles. Proc. Natl. Acad. Sci. U. S. A. 102, 15545–15550 (2005).
Hao, Y. et al. Integrated analysis of multimodal single-cell data. Cell 184, 3573-3587.e3529 (2021).
Acknowledgements
Medical writing assistance was provided by Mark Dyson, DPhil (Berlin, Germany) on behalf of BOLDSCIENCE, and funded by Lyell Immunopharma. We thank Dr. M. Wohlfahrt for coordinating viral vector production and providing associated methods; Lora Zhao, Sheila Lou, Emily Fu-Sum, Stefan Siebert for generating bulk RNA-Seq and single-cell CITE-Seq data; Elizabeth Pedrosa and Andrew Jimena for antibody titrations and flow cytometry cell sorting. Jessica D. Briones, Quinn Walker and Omar A. Ali was an employee of Lyell Immunopharma at the time of study execution.
Author information
Authors and Affiliations
Contributions
A.W.L., O.A.A., and A.S.C. conceptualized the project. A.W.L., J.L., O.A.A., B.B., and A.S.C. designed the experiments. A.W.L., J.D.B., J.L., B.B., Q.W., R.M., and A.S.C. performed the experiments and analyzed the data. H.H., B.B., P.S., S.P., and G.L. provided scientific feedback. A.W.L., O.A.A., and A.S.C. prepared the manuscript with writing assistance from BOLDSCIENCE. All authors read, reviewed, and approved the manuscript.
Corresponding author
Ethics declarations
Competing interests
A.S.C. is an inventor on a patent application related to the Stim-R technology (US patent 11555177B2). All authors were employees of Lyell Immunopharma at the time of study execution, and may additionally hold stock in Lyell Immunopharma, which has licensed intellectual property related to the Stim-R technology.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution-NonCommercial-NoDerivatives 4.0 International License, which permits any non-commercial use, sharing, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if you modified the licensed material. You do not have permission under this licence to share adapted material derived from this article or parts of it. The images or other third party material in this article are included in the article’s Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article’s Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by-nc-nd/4.0/.
About this article
Cite this article
Li, A.W., Briones, J.D., Lu, J. et al. Engineering potent chimeric antigen receptor T cells by programming signaling during T-cell activation. Sci Rep 14, 21331 (2024). https://doi.org/10.1038/s41598-024-72392-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/s41598-024-72392-1
- Springer Nature Limited