Abstract
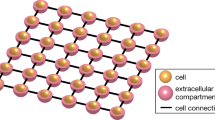
Excitable cells are of vital importance in biology, and mathematical models have contributed significantly to understand their basic mechanisms. However, classical models of excitable cells are based on severe assumptions that may limit the accuracy of the simulation results. Here, we derive a more detailed approach to modeling that has recently been applied to study the electrical properties of both neurons and cardiomyocytes. The model is derived from first principles and opens up possibilities for studying detailed properties of excitable cells.We refer to the model as the EMI model because both the extracellular space (E), the cell membrane (M) and the intracellular space (I) are explicitly represented in the model, in contrast to classical spatial models of excitable cells. Later chapters of the present text will focus on numerical methods and software for solving the model. Also, in the next chapter, the model will be extended to account for ionic concentrations in the intracellular and extracellular spaces.
Chapter PDF
Similar content being viewed by others
References
Agudelo-Toro A (2012) Numerical simulations on the biophysical foundations of the neuronal extracellular space. PhD thesis, Niedersächsische Staats-und Universitätsbibliothek Göttingen
Anastassiou CA, Perin R, Markram H, Koch C (2011) Ephaptic coupling of cortical neurons. Nature neuroscience 14(2):217–223
Buccino A, Kuchta M, Jæger KH, Ness T, Berthet P, Mardal KA, Cauwenberghs G, Tveito A (2019) How does the presence of neural probes affect extracellular potentials? Journal of Neural Engineering 16:026030
Cuellar AA, Lloyd CM, Nielsen PF, Bullivant DP, Nickerson DP, Hunter PJ (2003) An overview of CellML 1.1, a biological model description language. Simulation 79(12):740–747
Einevoll GT (2006) Mathematical modeling of neural activity. In: Dynamics of Complex Interconnected Systems: Networks and Bioprocesses, Springer, pp 127–145
Einevoll GT, Kayser C, Logothetis NK, Panzeri S (2013) Modelling and analysis of local field potentials for studying the function of cortical circuits. Nature Reviews Neuroscience 14(11):770
Ellingsrud AJ, Daversin-Catty C, Rognes ME (2020) A cell-based model for ionic electrodiffusion in excitable tissue. In: Tveito A, Mardal KA, Rognes ME (eds) Modeling excitable tissue - The EMI framework, Simula Springer Notes in Computing, SpringerNature
Franzone PC, Pavarino LF, Scacchi S (2014) Mathematical cardiac electrophysiology, vol 13. Springer
Griffiths DJ (1989) Introduction to electrodynamics, 2nd edn. Prentice Hall
Henriquez AP, Vogel R, Muller-Borer BJ, Henriquez CS, Weingart R, Cascio WE (2001) Influence of dynamic gap junction resistance on impulse propagation in ventricular myocardium: a computer simulation study. Biophysical Journal 81(4):2112–2121
Henriquez CS, Ying W (2009) The bidomain model of cardiac tissue: from microscale to macroscale. In: Cardiac Bioelectric Therapy, Springer, pp 401–421
Herz AV, Gollisch T, Machens CK, Jaeger D (2006) Modeling single-neuron dynamics and computations: a balance of detail and abstraction. Science 314(5796):80–85
Hille B (2001) Ion channels of excitable membranes, vol 507. Sinauer Sunderland, MA 14. Hodgkin AL, Huxley AF (1952) A quantitative description of membrane current and its application to conduction and excitation in nerve. The Journal of Physiology 117(4):500–544
Holt GR (1998) A critical reexamination of some assumptions and implications of cable theory in neurobiology. PhD thesis, California Institute of Technology
Holt GR, Koch C (1999) Electrical interactions via the extracellular potential near cell bodies. Journal of computational neuroscience 6(2):169–184
Jæger KH (2019) Cell-based mathematical models of small collections of excitable cells. PhD thesis, University of Oslo
Jæger KH, Edwards AG, McCulloch A, Tveito A (2019) Properties of cardiac conduction in a cell-based computational model. PLoS computational biology 15(5):e1007042
Jæger KH, Hustad KG, Cai X, Tveito A (2020) Operator splitting and finite difference schemes for solving the emi model. In: Tveito A, Mardal KA, Rognes ME (eds) Modeling excitable tissue - The EMI framework, Simula Springer Notes in Computing, SpringerNature
Keener J, Sneyd J (2010) Mathematical physiology. Springer Science & Business Media
Kucera JP, Rohr S, Kleber AG (2017) Microstructure, cell-to-cell coupling, and ion currents as determinants of electrical propagation and arrhythmogenesis. Circulation: Arrhythmia and Electrophysiology 10(9):e004665
Kuchta M, Mardal KA (2020) Iterative solvers for cell-based emi models. In: Tveito A, Mardal KA, Rognes ME (eds) Modeling excitable tissue - The EMI framework, Simula Springer Notes in Computing, SpringerNature, pp 0–100
Kuchta M, Mardal KA, Rognes ME (2020) Solving the emi equations using finite element methods. In: Tveito A, Mardal KA, Rognes ME (eds) Modeling excitable tissue - The EMI framework, Simula Springer Notes in Computing, SpringerNature
Lin J, Keener JP (2010) Modeling electrical activity of myocardial cells incorporating the effects of ephaptic coupling. Proceedings of theNationalAcademy of Sciences 107(49):20935–20940
Noble D (1962) A modification of the Hodgkin–Huxley equations applicable to purkinje fibre action and pacemaker potentials. The Journal of Physiology 160(2):317–352
Qu Z, Hu G, Garfinkel A, Weiss JN (2014) Nonlinear and stochastic dynamics in the heart. Physics Reports 543(2):61–162
Rudy Y (2012) From genes and molecules to organs and organisms: heart. Comprehensive Biophysics pp 268–327
Spach MS, Heidlage JF, Barr RC, Dolber PC (2004) Cell size and communication: role in structural and electrical development and remodeling of the heart. Heart Rhythm 1(4):500–515
SperelakisN, McConnellK(2002) Electric field interactions between closely abutting excitable cells. IEEE Engineering in Medicine and Biology Magazine 21(1):77–89
Sterratt D, Graham B, Gillies A,Willshaw D (2011) Principles of computational modelling in neuroscience. Cambridge University Press
Stinstra JG, Roberts SF, Pormann JB, MacLeod RS, Henriquez CS (2006) A model of 3D propagation in discrete cardiac tissue. In: Computers in Cardiology, 2006, IEEE, pp 41–44
Stinstra JG, Henriquez CS, MacLeod RS (2009) Comparison of microscopic and bidomain models of anisotropic conduction. In: Computers in Cardiology, 2009, IEEE, pp 657–660
Trayanova NA (2011) Whole-heart modeling: applications to cardiac electrophysiology and electromechanics. Circulation Research 108(1):113–128
Tveito A, Jæger KH, Lines GT, Paszkowski Ł, Sundnes J, Edwards AG, Mäki-Marttunen T, Halnes G, Einevoll GT (2017) An evaluation of the accuracy of classical models for computing the membrane potential and extracellular potential for neurons. Frontiers in Computational Neuroscience 11:27
Weinberg SH (2017) Ephaptic coupling rescues conduction failure in weakly coupled cardiac tissue with voltage-gated gap junctions. Chaos: An Interdisciplinary Journal of Nonlinear Science 27(9):093908
Author information
Authors and Affiliations
Corresponding author
Editor information
Editors and Affiliations
Rights and permissions
Open Access This chapter is licensed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license and indicate if changes were made.
The images or other third party material in this chapter are included in the chapter's Creative Commons license, unless indicated otherwise in a credit line to the material. If material is not included in the chapter's Creative Commons license and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder.
Copyright information
© 2021 The Author(s)
About this chapter
Cite this chapter
Jæger, K.H., Tveito, A. (2021). Derivation of a Cell-Based Mathematical Model of Excitable Cells. In: Tveito, A., Mardal, KA., Rognes, M.E. (eds) Modeling Excitable Tissue. Simula SpringerBriefs on Computing(), vol 7. Springer, Cham. https://doi.org/10.1007/978-3-030-61157-6_1
Download citation
DOI: https://doi.org/10.1007/978-3-030-61157-6_1
Published:
Publisher Name: Springer, Cham
Print ISBN: 978-3-030-61156-9
Online ISBN: 978-3-030-61157-6
eBook Packages: Mathematics and StatisticsMathematics and Statistics (R0)