Abstract
In Huntington's disease and other polyglutamine (polyQ) disorders, mutant proteins containing a long polyQ stretch are well documented as the trigger of numerous aberrant cellular processes that primarily lead to degeneration and, ultimately, the death of neuronal cells. However, mutant transcripts containing expanded CAG repeats may also be toxic and contribute to cellular dysfunction. The exact nature and importance of RNA toxicity in polyQ diseases are only beginning to be recognized, and the first insights have mainly resulted from studies using simple model systems. In this review, we briefly present the basic mechanisms of protein toxicity in polyQ disorders and RNA toxicity in myotonic dystrophy type 1 and discuss recent results suggesting that the pathogenesis of polyQ diseases may also be mediated by mutant transcripts. This review is focused on the experimental systems used thus far to demonstrate RNA toxicity in polyQ disorders and the design of new systems that will be more relevant to the human disease situation and capable of separating RNA toxicity from protein toxicity.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Nine human hereditary neurodegenerative disorders, including Huntington's disease (HD) and several spinocerebellar ataxias (SCAs), are caused by the abnormal expansion of CAG repeats located in the translated sequences of different genes [1] (Fig. 1). For example, HD is triggered by the expression of at least 36 CAG repeats located in exon 1 of the HTT gene, which encodes the large and multifunctional huntingtin protein (Htt) [2]. SCA1 is caused by at least 39 CAG repeats present in exon 8 of the ATXN1 gene, which codes for the ataxin-1 protein involved in RNA metabolism and transcriptional regulation [3], and SCA3 (also known as Machado–Joseph disease) is induced by at least 60 CAG repeats that occur in exon 10 of the ATXN3 gene encoding a protein with deubiquitinating activity [4]. The CAG repeats are translated into polyglutamine tracts in protein products, and the entire group of resulting disorders is known as the polyglutamine (polyQ) diseases. Each polyQ disorder primarily affects a different population of neurons, although the causative genes are widely expressed in the central nervous system and peripheral tissues. The expanded repeats are thought to exert their pathogenic effects at the protein level, mainly through a gain of toxic function by the mutant protein [1]. For comparison, myotonic dystrophy type 1 (DM1), which is another example of a triplet repeat expansion disease (TRED), is caused by 50–3,000 copies of CTG tandem repeats located in the 3′ untranslated sequence of the dystrophia myotonica protein kinase gene (DMPK) [5]. In DM1, the gain of toxic function by mutant transcript containing CUG repeats is proposed to explain its pathogenesis [6, 7].
Comparison of the lengths of the CAG repeat tracts that occur in polyQ disease-related transcripts and the CUG repeats in the transcript responsible for DM1. The normal repeat range is marked in green, and the mutant repeat range is marked in red and specified. The repeat range marked in gray refers to unidentified or undefined tracts. The starting threshold for DM1 mutation (50 repeats) is denoted by a hatched line
The pathogenic number of CAG repeats in genes implicated in polyQ diseases is, in most cases, lower than the number of CUG repeats in the DMPK gene of DM1 patients, and CAG expansions typically span a much narrower range of repeat lengths (Fig. 1). Nevertheless, a considerable fraction of HD and SCA patients harbor mutant CAG repeats of a length corresponding to the lower range of CUG repeat lengths found in DM1. The expression levels of individual polyQ disease genes and the DMPK gene differ within and between tissues; however, they all belong to the moderate or low expression category [8, 9]. The structural features of the CAG repeats are similar to the CUG repeats, forming hairpin structures in transcripts when the repeat tracts are long enough [10–16]. The repeats of normal length form small and semistable hairpins, and mutant repeats form long hairpins that are more stable [17]. According to the results of crystallographic studies, the CAG repeat duplexes and CUG repeat duplexes show a high degree of similarity [18, 19], suggesting that the stem portion of these hairpins may share some protein-binding properties [20, 21].
In this review, we first discuss the main cellular and molecular hallmarks of the mutant protein and mutant RNA gain-of-function mechanisms from studies of polyQ diseases and DM1 pathogenesis. We then present recent findings indicating that transcripts containing expanded CAG repeats might also be toxic and contribute to the pathogenesis of polyQ disorders. Finally, we focus on the experimental models used to demonstrate RNA toxicity in polyQ diseases. We discuss the features of these models and propose generating new cellular and animal models dedicated to elucidating the toxic effects triggered by mutant RNA and protein in polyQ diseases.
Mechanisms of protein toxicity in polyQ diseases and RNA toxicity in DM1
PolyQ toxicity
The protein products of genes that undergo mutations leading to polyQ diseases differ in their cellular functions and cover a wide range of molecular weights [1]. A common feature of these proteins is the presence of a polyQ tract that is expanded as the result of the mutation. However, the exact nature of protein gain-of-function toxicity in polyQ diseases remains a subject of debate [22, 23], and two mechanisms have been considered: the gain of a new toxic function by the mutant protein and the enhancement of the normal protein function to toxic levels. In either case, the key to understanding the details of such pathogenic mechanisms is the identification and characterization of new toxic protein–protein interactions and imbalanced normal interactions. A systematic search for proteins associated with normal and expanded Htt has recently been performed using quantitative proteomic analysis [24]. The data showed that the proteins that preferentially interacted with mutant Htt are enriched for intrinsically disordered proteins and proteins involved in key cellular functions, such as energy production, protein trafficking, RNA modification and translation, mitochondrial functions, cellular stress, and cell death. These results confirmed and expanded earlier findings regarding the cellular processes and pathways that are altered in HD [25–27].
The pathogenesis of polyQ diseases is believed to be initiated by the expanded polyQ tract, which is thought to acquire a β-sheet conformation and cause the mutant protein to misfold [28]. Alternatively, the expanded polyQ stretch may stabilize one of several conformations characteristic of the wild-type protein that may normally exist in equilibrium [22]; such an effect enhances some intermolecular interactions of the mutant protein at the cost of other interactions. In addition to the polyQ tract, the context of other protein domains may also contribute to pathogenesis, as demonstrated in animal models of spinal–bulbar muscular atrophy [29], SCA1 [30], and HD [31]. In several instances (reviewed in [32–34]), single amino acid substitutions in functional domains of relevant proteins resulted in dramatic changes of disease phenotypes pointing to the critical role of protein toxicity in polyQ diseases.
The cellular hallmarks of polyQ diseases are nuclear and cytoplasmic amyloid-like aggregates formed by mutant proteins, their proteolytic fragments [35, 36], or by fragments translated from aberrantly spliced shorter mRNAs [37] that sequester other proteins, predominantly those containing unstructured regions [24, 28, 38]. The formation of amyloid-like deposits is also observed in Parkinson and Alzheimer diseases, with which the protein toxicity of polyQ diseases is often compared. However, in efforts aimed at demonstrating RNA toxicity in polyQ diseases, reference to DM1 and other RNA-triggered TREDs, e.g., SCA8 [39] and HDL2 (Huntington's disease-like 2) [40, 41], is both appropriate and justified [20, 21, 42–44].
CUG repeat toxicity
DM1 is the prototype of TREDs caused by toxic RNA [6, 7]. Terms such as “toxic RNA” and “RNA toxicity” are used to reflect the fact that the disease-causing mutation is passed from the gene to the transcript only, while the protein structure is unaffected by the mutation. The hallmarks of cells expressing expanded CUG repeats are nuclear RNA foci [45], which are toxic to cells because they sequester important cellular proteins, reducing their normal, functional levels (reviewed in [46, 47]). One such protein trapped in mutant RNA foci is muscleblind-like (MBNL1) alternative splicing factor [48]; together with the concomitant upregulation of the antagonistic splicing factor CUG-BP1 [49], MBNL1 deficiency results in the misregulation of the developmentally regulated alternative splicing of numerous MBNL1-regulated genes. The compromised functions of some of these genes have been linked to the clinical symptoms of DM1, e.g., the altered splicing of insulin receptor in insulin resistance [50], chloride channel 1 in myotonia [51], sarcoplasmic/endoplasmic reticulum Ca2+ ATPase 1 and 2 in muscle wasting [52], cardiac troponin T in cardiac abnormalities [53], and Tau (MAPT) in cognitive deficits [54]. The binding of the multifunctional helicases p68 and p72 to mutant CUG repeats facilitates the binding of MBNL1 to the repeats [55] and likely links the DM1 pathogenesis pathway to the nuclear step of microRNA biogenesis, a process in which these helicases are involved as cofactors of the ribonuclease Drosha complex [56]. Direct evidence was provided for the crosstalk between the DM1 pathogenesis pathway and the cytoplasmic ribonuclease Dicer step of microRNA biogenesis, in which the MBNL1 protein is also implicated [57]. It was demonstrated that MBNL1 sequestered by mutant CUG repeats cannot effectively function in its newly recognized role as a pre-miRNA cleavage regulator, resulting in the compromised expression and function of some miRNAs, including miR-1, an effect linked to DM1-related heart defects. The expanded CUG repeats were also shown to trigger immune responses in DM1 tissue in which the PKR, OAS, TLR3, and RIG I genes were activated [58].
Accumulating evidence for RNA toxicity in polyQ diseases
Although the contribution of RNA toxicity to the pathogenesis of polyQ diseases was previously postulated based on the structural similarities between the CAG and CUG repeats in transcripts [10, 17], the first compelling experimental evidence for such an involvement was provided in 2008 by Bonini and colleagues [59]. Using a Drosophila melanogaster model system, the researchers demonstrated that the expression of both translated and untranslated RNA molecules containing long hairpin-forming CAG repeat tracts causes neurodegeneration, whereas the expression of RNA containing CAA-interrupted CAG repeats resulted in considerably less pronounced neurodegenerative features. CAA triplets also encode glutamine but are unable to form a hairpin structure [11], and their occurrence within CAG repeat tracts [60, 61] destabilizes CAG repeat hairpins [15].
Other studies that were performed in nonhuman model systems provided further evidence for the toxicity of CAG repeats. Hsiao and colleagues showed that in transgenic Caenorhabditis elegans, similar to CUG repeats, CAG repeats of toxic length formed nuclear foci that colocalized with CeMbl, which is the worm homolog of MBNL1 [62]. Worms expressing the toxic CAG repeats showed shortened lifespan and reduced motility rates, which is consistent with the phenotypes of worms expressing toxic CUG repeats. As shown by Pan and colleagues, the expression of 200 CAG repeats from the 3′UTR of EGFP was toxic in transgenic mouse muscle and testis tissues in which the expression of the transgene was directed [63]. Importantly, ribonuclear foci and abnormal phenotypes were observed in the absence of a polyQ product expression.
The formation of CAG repeat RNA foci that colocalize with MBNL1 protein was, for the first time, demonstrated by Cooper and colleagues through the expression of very long exogenous CAG repeats in monkey COSM6 cells [64]. A question that was addressed using human cell lines was whether less CAG repeats could also form nuclear foci and cause alternative splicing aberrations similar to those known for DM1. The results showed that these effects are triggered in both HeLa and neuroblastoma SK-N-MC cells transiently expressing mutant CAG repeats [65]. In addition, we demonstrated that the expanded CAG repeats present in HTT and ATXN3 transcripts also occur within intranuclear MBNL1-positive RNA foci in human HD and SCA3 fibroblasts. Although the focus formation was accompanied by the aberrant alternative splicing of several endogenous MBNL1-dependent transcripts, this only occurred in cells expressing approximately 70 CAG repeats and not a lower number of approximately 45 CAGs [65]. Very recently, Bates and colleagues have demonstrated the CAG repeat length-dependent aberrant splicing of the HTT exon 1 giving rise to short polyadenylated exon 1-intron 1 mRNA which is translated to the toxic HTT exon 1 protein [37]. The aberrant splicing is triggered by the increased binding of the alternative splicing factor SRSF6 to expanded CAG repeats. Thus, it is likely that the pathogenesis of polyQ disorders involves, at least to some extent, the primary elements of the RNA gain-of-function mechanism that is well characterized in DM1. Consistent with the above nuclear effects, a protein that is responsible for the nuclear export of transcripts harboring expanded CAG repeats has been identified by Chan and colleagues as the small nuclear RNA U2-associated factor U2AF65 [66]. This factor binds directly to expanded CAG repeats and forms a complex with the nuclear export receptor NXF1 and its reduction in symptomatic tissues is linked to accumulation of mutant RNA in the nucleus.
An RNAi mechanism has been shown to be involved in pathogenesis in several in vivo and cellular models of polyQ diseases. As demonstrated by Bonini and colleagues [67] and Richards and colleagues [44] in two independent studies, the co-expression of exogenous expanded CAG and CUG repeats in fly cells resulted in a neurodegenerative phenotype. The complementary repeats likely formed a double-stranded RNA that was cleaved by Dicer-2 into 21 nt CAG/CUG repeat siRNAs, which were active in silencing in an Ago2-dependent process. Margolis and colleagues [68] discovered a natural antisense transcript, HTTAS, at the HD repeat locus that contains the CUG repeat tract. This transcript was shown to regulate sense HTT transcript expression in a process dependent on the repeat length and partly dependent on Dicer and the RISC pathway. The involvement of an RNAi mechanism was also demonstrated by Marti and colleagues in human cell lines expressing mutant CAG repeats flanked by the HTT exon 1 sequence [69]. The mutant CAG repeats, both translated and untranslated, gave rise to a toxic small RNA species (sCAG) in a Dicer-dependent manner and caused a downstream silencing effect in an Ago2-dependent manner. Again, the cytotoxic effects were triggered only by CAG repeats and not by CAA repeats. The involvement of the miRNA biogenesis pathway in the pathogenesis of polyQ diseases is only circumstantial, based on the ability of the MBNL1 [16, 64, 65] and p68 [55] proteins to bind to both CUG repeats and, though less efficiently, CAG repeats.
The recent study by Chan and colleagues revealed that CAG repeat expansion induces nucleolar stress in polyglutamine diseases [70], demonstrating that the expanded CAG repeat RNA interacts directly with nucleolin and that this interaction triggers a number of aberrant downstream cellular processes, eventually leading to cell apoptosis. These downstream effects include hypermethylation of the upstream control element of the rRNA promoter and the inhibition of rRNA transcription. The reduced level of rRNA results in the accumulation of free ribosomal proteins, and the interaction of these proteins with MDM2 E3 ubiquitin ligase causes mitochondrial p53 accumulation. The interaction between p53 and the antiapoptotic protein Bcl-xL causes the oligomerization of the proapoptotic protein Bak at the mitochondrial membrane and allows for cytochrome c release to the cytosol, an event that activates the caspase cascade and induces apoptosis [70]. Collectively, several mechanisms of RNA toxicity have been revealed that may likely participate in the pathogenesis of polyQ diseases; however, the entire scale of toxic RNA contribution to the disorders and the range of cellular processes and pathways altered by mutant transcripts remain unknown.
Experimental models used to trace RNA toxicity in polyQ diseases
Various genetic models have been used to demonstrate the first examples of CAG repeat RNA toxicity described in the previous section. The characteristics of these model systems and the main results obtained from their application are compiled in Table 1. It is apparent that this type of RNA toxicity has thus far been modeled in lower invertebrates, nematode worms, and fruit flies in addition to mammalian systems, including mice and monkey and human cell lines. However, a common issue in modeling human diseases in lower organisms is the relevance of the results obtained with simpler model systems of human disease. Indeed, relevance is hard to prove by merely a sufficient similarity of the genetic pathways operating in model organisms and humans. For polyQ diseases and TREDs in general, this task is more difficult, as these late-onset diseases only occur naturally in humans and not in shorter living animals in which the experimental induction of similar effects requires stronger triggers, i.e., longer repeat tracts and in some cases higher expression levels. In most studies, the expression of transgene was determined but the level not referred to the expression of endogene (homologue of human polyQ gene) or any other gene. Most authors related the expression of a single construct to the expression of other transgene variants. In case of mouse models R6/2, BACHD, and YAC128, the expression level of transgene did not differ much from that of homologous endogene [71, 72]. In case of constructs tested in mammalian systems, containing CMV promoters, the expression level of transgene was reported as very high [64, 65, 73]. The worm and fly systems satisfactorily fulfilled the requirement of genetic similarity, and both systems were capable of demonstrating some molecular and phenotypic hallmarks of RNA toxicity. Although more relevant and potentially more informative than the fly systems, transgenic mouse models have less frequently been used, and human cellular models have been used for this purpose only twice.
The genetic systems shown in Table 1 differ considerably with regard to their molecular design. Most models are transgenic and express exogenous expanded repeats that have the potential to trigger pathogenesis by a variety of mechanisms involving aberrant interactions with host proteins. Of importance in these systems is the intragenic localization of repeated sequences. In some experimental systems, the repeats are located within the open reading frame (ORF) and are typically transcribed and translated; the expressed mutant repeats are present within the sequence context of either the full-length human disease transcript or only a fragment of this transcript. Alternatively, the repeats are placed within the 3′UTR sequence of a heterologous gene and are only transcribed. The genes hosting mutant repeats are usually a marker gene, such as GFP or DsRed. In addition, the length and purity of the CAG repeat tracts are important factors in the design of a model system; the repeated sequence typically falls into the upper range of repeat lengths found in human subjects suffering from the disease and is often modified to contain CAA repeats.
The ideal model of a polyQ disease would be one that closely recapitulates the human disease with regard to most of its aspects [36]. Thus, human cell lines are well suited to study the details of the molecular mechanisms that are implicated in triggering pathogenesis. The rapid development of induced pluripotent stem (iPS) cell technology enables somatic cells to be transformed into iPS cells and then differentiated to neuronal cell lines [74]. Moreover, the advent of gene-editing technologies has begun to enable the precise engineering of human disease-related endogens [75]. Both technologies may provide highly informative cellular models for future studies. The expression of mutant exogenous full-length cDNA using the well-established technologies of transgenesis may also be considered as an option for the construction of in vitro models of polyQ diseases. A good transgenic mouse model of polyQ disease should express the CAG repeat mutation within the sequence context of the entire human gene, with sufficient upstream and downstream regulatory sequences to ensure the proper spatial and temporal expression of the human gene at natural levels [72, 76]. In general, model systems expressing a full-length human mRNA, despite showing milder disease phenotypes, are better suited for studies aimed at discovering new mechanisms of RNA toxicity than systems expressing only shorter or longer mRNA fragments. In the full-length RNA models, the mutant transcript has a chance to go through all steps of mRNA cellular journey from its “birth” in the nucleus to “death” in the cytoplasm; therefore, all steps altered by the mutation may be identified.
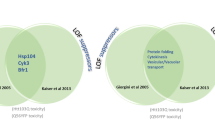
A serious difficulty in identifying the involvement of RNA toxicity in polyQ diseases is the overlap with the polyQ toxicity in models expressing CAG repeats from the ORF, and placing the repeat in the untranslated sequence of a heterologous gene is frequently used to overcome this difficulty and trace the effects of mutant RNA. However, preventing transcript translation by the mutation of the START codon in a polyQ disease gene or its cDNA resolves this problem while preserving the natural sequence context and expression level for the repeats. Based on the assumption that the non-AUG-initiated translation of repeated sequences [77] is not a very common phenomenon, the START codon mutation may indeed be considered a better solution. By mutating the START codon also in constructs expressing CAA repeats or CAA-interrupted CAG repeats, the four model types depicted in Fig. 2 may be generated. With these in vitro and in vivo models critical experiments using both transcriptome-wide and more focused candidate approaches have to be done to obtain clear-cut answers regarding the role of mutant transcripts in the pathogenesis of polyQ diseases.
Four types of models that can be used to separate RNA toxicity and protein toxicity in the investigation of the pathogenesis of CAG repeat diseases. The depicted models express one of two types of repeated sequences, either CAG or CAA. Both sequences code for glutamine, but only the CAG repeat forms a hairpin structure in transcripts. In these types of models, both repeats are expressed in their translated or untranslated forms, depending on the absence or presence of an AUG START codon mutation. Potentially toxic entities, an expanded CAG repeat-containing transcript and a polyQ-containing protein, are marked in red; a transcript containing expanded CAA repeats, presumed to be nontoxic, is marked in blue
Concluding remarks
Research on RNA toxicity in polyQ diseases is still in its infancy. At this stage, there is a marked disproportion between the ability of a model organism to closely reproduce human disease and the frequency with which the model system has been used in the search for RNA toxicity hallmarks. Nevertheless, most of the genetic models provided in Table 1 have succeeded in demonstrating the first examples of RNA toxicity caused by expanded CAG repeats. Future studies along this line will require the more extensive use of dedicated mouse models and the next generation of human cellular models.
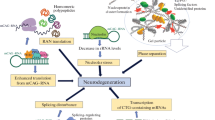
The studies performed thus far have revealed the involvement of several cellular processes and pathways in CAG repeat RNA toxicity. These altered processes include aberrant alternative splicing, transcript nuclear transport and export, RNA interference, and nucleolar stress resulting in apoptosis. All these processes are triggered by aberrant interactions between cellular proteins and mutant RNA. The proteins that participate in direct or indirect interactions with mutant CAG repeat transcripts need to be identified in further studies, and their role in pathogenesis needs to be elucidated. Such efforts may help to determine the contribution of RNA toxicity to the overall toxicity caused by both toxic factors: the protein and the RNA. This endeavor may also provide the basis to answer the question of whether the RNA and protein toxicities interact with each other to enhance pathogenesis or whether these toxic processes are parallel and independent. Another issue to be clarified is whether the RNA and protein toxicities require the same or different repeat length thresholds to become operative. The latter alternative may likely be the case, and a higher number of repeats may be required to initiate the RNA-mediated toxic processes.
Finally, depending on the scale and repeat length thresholds for the various pathogenic mechanisms triggered by mutant RNA, these factors may impact the causative therapeutic approaches to combat polyQ diseases to a higher or lower extent. Should RNA toxicity be proven to contribute significantly to disease development and progression, both the inhibition of mutant protein synthesis and the blockage or degradation of mutant transcript will be required.
References
Orr HT, Zoghbi HY (2007) Trinucleotide repeat disorders. Annu Rev Neurosci 30:575–621
MacDonald ME, Ambrose CM, Duyao MP (1993) A novel gene containing a trinucleotide repeat that is expanded and unstable on Huntington's disease chromosomes. The Huntington's Disease Collaborative Research Group. Cell 72:971–983
Orr HT, Chung MY, Banfi S, Kwiatkowski TJ Jr, Servadio A, Beaudet AL, McCall AE, Duvick LA, Ranum LP, Zoghbi HY (1993) Expansion of an unstable trinucleotide CAG repeat in spinocerebellar ataxia type 1. Nat Genet 4:221–226
Kawaguchi Y, Okamoto T, Taniwaki M, Aizawa M, Inoue M, Katayama S, Kawakami H, Nakamura S, Nishimura M, Akiguchi I et al (1994) CAG expansions in a novel gene for Machado-Joseph disease at chromosome 14q32.1. Nat Genet 8:221–228
Brook JD, McCurrach ME, Harley HG, Buckler AJ, Church D, Aburatani H, Hunter K, Stanton VP, Thirion JP, Hudson T et al (1992) Molecular basis of myotonic dystrophy: expansion of a trinucleotide (CTG) repeat at the 3′ end of a transcript encoding a protein kinase family member. Cell 69:385
Ranum LP, Cooper TA (2006) RNA-mediated neuromuscular disorders. Annu Rev Neurosci 29:259–277
Wheeler TM, Thornton CA (2007) Myotonic dystrophy: RNA-mediated muscle disease. Curr Opin Neurol 20:572–576
Jasinska A, Kozlowski P, Krzyzosiak WJ (2008) Expression characteristics of triplet repeat-containing RNAs and triplet repeat-interacting proteins in human tissues. Acta Biochimica Polonica 55:1–8
Nestor CE, Monckton DG (2011) Correlation of inter-locus polyglutamine toxicity with CAG*CTG triplet repeat expandability and flanking genomic DNA GC content. PLoS One 6:e28260
Sobczak K, de Mezer M, Michlewski G, Krol J, Krzyzosiak WJ (2003) RNA structure of trinucleotide repeats associated with human neurological diseases. Nucleic Acids Res 31:5469–5482
Sobczak K, Michlewski G, de Mezer M, Kierzek E, Krol J, Olejniczak M, Kierzek R, Krzyzosiak WJ (2010) Structural diversity of triplet repeat RNAs. J Biol Chem 285:12755–12764
de Mezer M, Wojciechowska M, Napierala M, Sobczak K, Krzyzosiak WJ (2011) Mutant CAG repeats of Huntingtin transcript fold into hairpins, form nuclear foci and are targets for RNA interference. Nucleic Acids Res 39:3852–3863
Michlewski G, Krzyzosiak WJ (2004) Molecular architecture of CAG repeats in human disease related transcripts. J Mol Biol 340:665–679
Napierala M, Krzyzosiak WJ (1997) CUG repeats present in myotonin kinase RNA form metastable “slippery” hairpins. J Biol Chem 272:31079–31085
Sobczak K, Krzyzosiak WJ (2004) Imperfect CAG repeats form diverse structures in SCA1 transcripts. J Biol Chem 279:41563–41572
Yuan Y, Compton SA, Sobczak K, Stenberg MG, Thornton CA, Griffith JD, Swanson MS (2007) Muscleblind-like 1 interacts with RNA hairpins in splicing target and pathogenic RNAs. Nucleic Acids Res 35:5474–5486
Jasinska A, Michlewski G, de Mezer M, Sobczak K, Kozlowski P, Napierala M, Krzyzosiak WJ (2003) Structures of trinucleotide repeats in human transcripts and their functional implications. Nucleic Acids Res 31:5463–5468
Kiliszek A, Kierzek R, Krzyzosiak WJ, Rypniewski W (2010) Atomic resolution structure of CAG RNA repeats: structural insights and implications for the trinucleotide repeat expansion diseases. Nucleic Acids Res 38:8370–8376
Kiliszek A, Kierzek R, Krzyzosiak WJ, Rypniewski W (2009) Structural insights into CUG repeats containing the ‘stretched U-U wobble’: implications for myotonic dystrophy. Nucleic Acids Res 37:4149–4156
Galka-Marciniak P, Urbanek MO, Krzyzosiak WJ (2012) Triplet repeats in transcripts: structural insights into RNA toxicity. Biol Chem 393:1299–1315
Krzyzosiak WJ, Sobczak K, Wojciechowska M, Fiszer A, Mykowska A, Kozlowski P (2012) Triplet repeat RNA structure and its role as pathogenic agent and therapeutic target. Nucleic Acids Res 40:11–26
Kratter IH, Finkbeiner S (2010) PolyQ disease: too many Qs, too much function? Neuron 67:897–899
Orr HT (2012) Polyglutamine neurodegeneration: expanded glutamines enhance native functions. Curr Opin Genet Dev 22:251–255
Ratovitski T, Chighladze E, Arbez N, Boronina T, Herbrich S, Cole RN, Ross CA (2012) Huntingtin protein interactions altered by polyglutamine expansion as determined by quantitative proteomic analysis. Cell Cycle 11:2006–2021
Olzscha H, Schermann SM, Woerner AC, Pinkert S, Hecht MH, Tartaglia GG, Vendruscolo M, Hayer-Hartl M, Hartl FU, Vabulas RM (2011) Amyloid-like aggregates sequester numerous metastable proteins with essential cellular functions. Cell 144:67–78
Li SH, Li XJ (2004) Huntingtin-protein interactions and the pathogenesis of Huntington's disease. Trends Genet 20:146–154
Boutell JM, Thomas P, Neal JW, Weston VJ, Duce J, Harper PS, Jones AL (1999) Aberrant interactions of transcriptional repressor proteins with the Huntington's disease gene product, huntingtin. Hum Mol Genet 8:1647–1655
Takahashi T, Katada S, Onodera O (2010) Polyglutamine diseases: where does toxicity come from? what is toxicity? where are we going? J Mol Cell Biol 2:180–191
Nedelsky NB, Pennuto M, Smith RB, Palazzolo I, Moore J, Nie Z, Neale G, Taylor JP (2010) Native functions of the androgen receptor are essential to pathogenesis in a Drosophila model of spinobulbar muscular atrophy. Neuron 67:936–952
Lim J, Crespo-Barreto J, Jafar-Nejad P, Bowman AB, Richman R, Hill DE, Orr HT, Zoghbi HY (2008) Opposing effects of polyglutamine expansion on native protein complexes contribute to SCA1. Nature 452:713–718
Keryer G, Pineda JR, Liot G, Kim J, Dietrich P, Benstaali C, Smith K, Cordelieres FP, Spassky N, Ferrante RJ et al (2011) Ciliogenesis is regulated by a huntingtin-HAP1-PCM1 pathway and is altered in Huntington disease. J Clin Invest 121:4372–4382
Pennuto M, Palazzolo I, Poletti A (2009) Post-translational modifications of expanded polyglutamine proteins: impact on neurotoxicity. Hum Mol Genet 18:R40–R47
Orr HT (2012) SCA1-phosphorylation, a regulator of Ataxin-1 function and pathogenesis. Prog Neurobiol 99:179–185
Katsuno M, Tanaka F, Adachi H, Banno H, Suzuki K, Watanabe H, Sobue G (2012) Pathogenesis and therapy of spinal and bulbar muscular atrophy (SBMA). Prog Neurobiol 99:246–256
DiFiglia M, Sapp E, Chase KO, Davies SW, Bates GP, Vonsattel JP, Aronin N (1997) Aggregation of huntingtin in neuronal intranuclear inclusions and dystrophic neurites in brain. Science 277:1990–1993
Ross CA, Tabrizi SJ (2011) Huntington's disease: from molecular pathogenesis to clinical treatment. Lancet Neurol 10:83–98
Sathasivam K, Neueder A, Gipson TA, Landles C, Benjamin AC, Bondulich MK, Smith DL, Faull RL, Roos RA, Howland D et al (2013) Aberrant splicing of HTT generates the pathogenic exon 1 protein in Huntington disease. Proc Natl Acad Sci U S A 110:2366–2370
Jones L, Hughes A (2011) Pathogenic mechanisms in Huntington's disease. Int Rev Neurobiol 98:373–418
Daughters RS, Tuttle DL, Gao W, Ikeda Y, Moseley ML, Ebner TJ, Swanson MS, Ranum LP (2009) RNA gain-of-function in spinocerebellar ataxia type 8. PLoS Genet 5:e1000600
Rudnicki DD, Holmes SE, Lin MW, Thornton CA, Ross CA, Margolis RL (2007) Huntington's disease-like 2 is associated with CUG repeat-containing RNA foci. Ann Neurol 61:272–282
Margolis RL, O'Hearn E, Rosenblatt A, Willour V, Holmes SE, Franz ML, Callahan C, Hwang HS, Troncoso JC, Ross CA (2001) A disorder similar to Huntington's disease is associated with a novel CAG repeat expansion. Ann Neurol 50:373–380
Wojciechowska M, Krzyzosiak WJ (2011) CAG repeat RNA as an auxiliary toxic agent in polyglutamine disorders. RNA Biol 8:565–571
Li LB, Bonini NM (2010) Roles of trinucleotide-repeat RNA in neurological disease and degeneration. Trends Neurosci 33:292–298
Lawlor KT, O'Keefe LV, Samaraweera SE, van Eyk CL, McLeod CJ, Maloney CA, Dang TH, Suter CM, Richards RI (2011) Double-stranded RNA is pathogenic in Drosophila models of expanded repeat neurodegenerative diseases. Hum Mol Genet 20:3757–3768
Taneja KL, McCurrach M, Schalling M, Housman D, Singer RH (1995) Foci of trinucleotide repeat transcripts in nuclei of myotonic dystrophy cells and tissues. J Cell Biol 128:995–1002
Echeverria GV, Cooper TA (2012) RNA-binding proteins in microsatellite expansion disorders: mediators of RNA toxicity. Brain Res 1462:100–111
Wojciechowska M, Krzyzosiak WJ (2011) Cellular toxicity of expanded RNA repeats: focus on RNA foci. Hum Mol Genet 20:3811–3821
Miller JW, Urbinati CR, Teng-Umnuay P, Stenberg MG, Byrne BJ, Thornton CA, Swanson MS (2000) Recruitment of human muscleblind proteins to (CUG)(n) expansions associated with myotonic dystrophy. EMBO J 19:4439–4448
Timchenko NA, Cai ZJ, Welm AL, Reddy S, Ashizawa T, Timchenko LT (2001) RNA CUG repeats sequester CUGBP1 and alter protein levels and activity of CUGBP1. J Biol Chem 276:7820–7826
Savkur RS, Philips AV, Cooper TA (2001) Aberrant regulation of insulin receptor alternative splicing is associated with insulin resistance in myotonic dystrophy. Nat Genet 29:40–47
Charlet BN, Savkur RS, Singh G, Philips AV, Grice EA, Cooper TA (2002) Loss of the muscle-specific chloride channel in type 1 myotonic dystrophy due to misregulated alternative splicing. Mol Cell 10:45–53
Kimura T, Nakamori M, Lueck JD, Pouliquin P, Aoike F, Fujimura H, Dirksen RT, Takahashi MP, Dulhunty AF, Sakoda S (2005) Altered mRNA splicing of the skeletal muscle ryanodine receptor and sarcoplasmic/endoplasmic reticulum Ca2+-ATPase in myotonic dystrophy type 1. Hum Mol Genet 14:2189–2200
Philips AV, Timchenko LT, Cooper TA (1998) Disruption of splicing regulated by a CUG-binding protein in myotonic dystrophy. Science 280:737–741
Jiang H, Mankodi A, Swanson MS, Moxley RT, Thornton CA (2004) Myotonic dystrophy type 1 is associated with nuclear foci of mutant RNA, sequestration of muscleblind proteins and deregulated alternative splicing in neurons. Hum Mol Genet 13:3079–3088
Laurent FX, Sureau A, Klein AF, Trouslard F, Gasnier E, Furling D, Marie J (2012) New function for the RNA helicase p68/DDX5 as a modifier of MBNL1 activity on expanded CUG repeats. Nucleic Acids Res 40:3159–3171
Fukuda T, Yamagata K, Fujiyama S, Matsumoto T, Koshida I, Yoshimura K, Mihara M, Naitou M, Endoh H, Nakamura T et al (2007) DEAD-box RNA helicase subunits of the Drosha complex are required for processing of rRNA and a subset of microRNAs. Nat Cell Biol 9:604–611
Rau F, Freyermuth F, Fugier C, Villemin JP, Fischer MC, Jost B, Dembele D, Gourdon G, Nicole A, Duboc D et al (2011) Misregulation of miR-1 processing is associated with heart defects in myotonic dystrophy. Nat Struct Mol Biol 18:840–845
Rhodes JD, Lott MC, Russell SL, Moulton V, Sanderson J, Wormstone IM, Broadway DC (2012) Activation of the innate immune response and interferon signalling in myotonic dystrophy type 1 and type 2 cataracts. Hum Mol Genet 21:852–862
Li LB, Yu Z, Teng X, Bonini NM (2008) RNA toxicity is a component of ataxin-3 degeneration in Drosophila. Nature 453:1107–1111
Sobczak K, Krzyzosiak WJ (2004) Patterns of CAG repeat interruptions in SCA1 and SCA2 genes in relation to repeat instability. Hum Mutat 24:236–247
Imbert G, Saudou F, Yvert G, Devys D, Trottier Y, Garnier JM, Weber C, Mandel JL, Cancel G, Abbas N et al (1996) Cloning of the gene for spinocerebellar ataxia 2 reveals a locus with high sensitivity to expanded CAG/glutamine repeats. Nat Genet 14:285–291
Wang LC, Chen KY, Pan H, Wu CC, Chen PH, Liao YT, Li C, Huang ML, Hsiao KM (2011) Muscleblind participates in RNA toxicity of expanded CAG and CUG repeats in Caenorhabditis elegans. Cell Mol Life Sci 68:1255–1267
Hsu RJ, Hsiao KM, Lin MJ, Li CY, Wang LC, Chen LK, Pan H (2011) Long tract of untranslated CAG repeats is deleterious in transgenic mice. PLoS One 6:e16417
Ho TH, Savkur RS, Poulos MG, Mancini MA, Swanson MS, Cooper TA (2005) Colocalization of muscleblind with RNA foci is separable from mis-regulation of alternative splicing in myotonic dystrophy. J Cell Sci 118:2923–2933
Mykowska A, Sobczak K, Wojciechowska M, Kozlowski P, Krzyzosiak WJ (2011) CAG repeats mimic CUG repeats in the misregulation of alternative splicing. Nucleic Acids Res 39:8938–8951
Tsoi H, Lau CK, Lau KF, Chan HY (2011) Perturbation of U2AF65/NXF1-mediated RNA nuclear export enhances RNA toxicity in polyQ diseases. Hum Mol Genet 20:3787–3797
Yu Z, Teng X, Bonini NM (2011) Triplet repeat-derived siRNAs enhance RNA-mediated toxicity in a Drosophila model for myotonic dystrophy. PLoS Genet 7:e1001340
Chung DW, Rudnicki DD, Yu L, Margolis RL (2011) A natural antisense transcript at the Huntington's disease repeat locus regulates HTT expression. Hum Mol Genet 20:3467–3477
Banez-Coronel M, Porta S, Kagerbauer B, Mateu-Huertas E, Pantano L, Ferrer I, Guzman M, Estivill X, Marti E (2012) A pathogenic mechanism in Huntington's disease involves small CAG-repeated RNAs with neurotoxic activity. PLoS Genet 8:e1002481
Tsoi H, Lau TC, Tsang SY, Lau KF, Chan HY (2012) CAG expansion induces nucleolar stress in polyglutamine diseases. Proc Natl Acad Sci U S A 109:13428–13433
Mangiarini L, Sathasivam K, Seller M, Cozens B, Harper A, Hetherington C, Lawton M, Trottier Y, Lehrach H, Davies SW et al (1996) Exon 1 of the HD gene with an expanded CAG repeat is sufficient to cause a progressive neurological phenotype in transgenic mice. Cell 87:493–506
Pouladi MA, Stanek LM, Xie Y, Franciosi S, Southwell AL, Deng Y, Butland S, Zhang W, Cheng SH, Shihabuddin LS et al (2012) Marked differences in neurochemistry and aggregates despite similar behavioural and neuropathological features of Huntington disease in the full-length BACHD and YAC128 mice. Hum Mol Genet 21:2219–2232
Goldberg YP, Kalchman MA, Metzler M, Nasir J, Zeisler J, Graham R, Koide HB, O'Kusky J, Sharp AH, Ross CA et al (1996) Absence of disease phenotype and intergenerational stability of the CAG repeat in transgenic mice expressing the human Huntington disease transcript. Hum Mol Genet 5:177–185
Consortium, H.i (2012) Induced pluripotent stem cells from patients with Huntington's disease show CAG-repeat-expansion-associated phenotypes. Cell Stem Cell 11:264–278
Joung JK, Sander JD (2013) TALENs: a widely applicable technology for targeted genome editing. Nat Rev Mol Cell Biol 14:49–55
Figiel M, Szlachcic WJ, Switonski PM, Gabka A, Krzyzosiak WJ (2012) Mouse models of polyglutamine diseases: review and data table. Part I. Mol Neurobiol 46:393–429
Zu T, Gibbens B, Doty NS, Gomes-Pereira M, Huguet A, Stone MD, Margolis J, Peterson M, Markowski TW, Ingram MA et al (2011) Non-ATG-initiated translation directed by microsatellite expansions. Proc Natl Acad Sci U S A 108:260–265
McLeod CJ, O'Keefe LV, Richards RI (2005) The pathogenic agent in Drosophila models of ‘polyglutamine’ diseases. Hum Mol Genet 14:1041–1048
Shieh SY, Bonini NM (2011) Genes and pathways affected by CAG-repeat RNA-based toxicity in Drosophila. Hum Mol Genet 20:4810–4821
Lawlor KT, O'Keefe LV, Samaraweera SE, van Eyk CL, Richards RI (2012) Ubiquitous expression of CUG or CAG trinucleotide repeat RNA causes common morphological defects in a Drosophila model of RNA-mediated pathology. PLoS One 7:e38516
Acknowledgments
This work was supported by Ministry of Science and Higher Education (N N301569340), National Science Centre (DEC-2012/06/A/NZ1/00094) and European Regional Development Fund within Innovative Economy Programme (POIG.01.03.01-30-098/08). The authors would like to thank lab members for discussions and comments on the manuscript.
Conflict of interest
The authors declare that they have no conflict of interests.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Fiszer, A., Krzyzosiak, W.J. RNA toxicity in polyglutamine disorders: concepts, models, and progress of research. J Mol Med 91, 683–691 (2013). https://doi.org/10.1007/s00109-013-1016-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00109-013-1016-2