Abstract
Septoria tritici blotch (STB), caused by the ascomycete Mycosphaerella graminicola, is one of the most devastating foliar diseases of wheat. We screened five synthetic hexaploid wheats (SHs), 13 wheat varieties that represent the differential set of cultivars and two susceptible checks with a global set of 20 isolates and discovered exceptionally broad STB resistance in SHs. Subsequent development and analyses of recombinant inbred lines (RILs) from a cross between the SH M3 and the highly susceptible bread wheat cv. Kulm revealed two novel resistance loci on chromosomes 3D and 5A. The 3D resistance was expressed in the seedling and adult plant stages, and it controlled necrosis (N) and pycnidia (P) development as well as the latency periods of these parameters. This locus, which is closely linked to the microsatellite marker Xgwm494, was tentatively designated Stb16q and explained from 41 to 71% of the phenotypic variation at seedling stage and 28–31% in mature plants. The resistance locus on chromosome 5A was specifically expressed in the adult plant stage, associated with SSR marker Xhbg247, explained 12–32% of the variation in disease, was designated Stb17, and is the first unambiguously identified and named QTL for adult plant resistance to M. graminicola. Our results confirm that common wheat progenitors might be a rich source of new Stb resistance genes/QTLs that can be deployed in commercial breeding programs.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Since early history, wheat (Triticum aestivum L., 2n = 6× = 42, AABBDD) was a main source of food and feed. It is the oldest and has been the most widely grown crop since 10,000–8,000 bc (Heun et al. 1997; Luo et al. 2007; Nesbitt and Samuel 1998). Due to its importance and increasing demand, it is a key commodity to eradicate global hunger not only by ensuring sufficient production to feed a world population that will grow by 50% and reach 9 billion by 2050, but also by guaranteeing access to food (FAO 2010). Still, in 2010 annual bread wheat production is projected to decline and diseases play a significant role in such reductions (USDA 2010). In Western Europe, which is among the largest wheat production areas, septoria tritici blotch (STB) caused by the ascomycete Mycosphaerella graminicola (Fuckel) J. Schröt is the most recurrent and important wheat disease. STB is also a major disease in the Americas, Central and West Asia, and particularly on durum wheat in North Africa. STB can cause yield losses that typically range from 10–15%, but under conducive weather conditions, losses can easily exceed 50%, particularly in low-input agriculture where disease management is frequently suboptimal (Duveiller et al. 2007; Eyal 1999; King et al. 1983).
In intensive wheat production areas, disease management is often accomplished by fungicide applications and the deployment of resistant wheat cultivars (Goodwin 2007; Lehoczki-Krsjak et al. 2010). STB is the major target of the agrochemical industry that has Western Europe as its prime market (Jorgensen 2008), but fungicide applications are not always timely, environmentally sound or economically viable (Paveley et al. 1997). Under conditions favorable for disease, 2–12 fungicide applications are required to control STB (Burke and Dunne 2008), and the costs easily reach approximately 150 Euro per hectare (Beest et al. 2009). Most importantly, fungicide efficacy towards STB is hampered by the development of fungicide resistant strains of the pathogen (Fraaije et al. 2005; Mavroeidi and Shaw 2005; Stergiopoulos et al. 2003). Therefore, host resistance is an important component of effective disease management strategies for commercial wheat production.
To date, 15 major resistance genes, Stb1–Stb15, have been identified and characterized, but compared to yellow rust, leaf rust, stem rust and powdery mildew—with 73, 89, 61 and 95 mapped resistance genes, respectively—this number is limited. Moreover, the majority of these genes have narrow spectra of specificity towards M. graminicola isolates that represent current field populations in major wheat producing areas, and this limits their use (Arraiano and Brown 2006; Chartrain et al. 2005b). Furthermore, M. graminicola is a heterothallic filamentous fungus with multiple sexual cycles during the growing season that defines its complex genetic population structure and influences disease management (Chen and McDonald 1996; Kema et al. 1996c; McDonald et al. 1996). The wheat–M. graminicola pathosystem complies with the gene-for-gene hypothesis where a pathogen effector interacts with a host target (Brading et al. 2002). Hence, the selection pressure that new Stb genes may exert on natural M. graminicola populations calls for responsible deployment strategies and a continuous effort to unveil key genes that control this disease (Cowger et al. 2000; Linde et al. 2002; Zhang et al. 2001).
Modern wheat improvement programs and wheat domestication processes resulted in narrow diversity of wheat germplasm (Christiansen et al. 2002; Raman et al. 2010). For this reason, wild wheat progenitors have been considered potential sources for the recovery of genetic diversity (Dreisigacker et al. 2008; Ortiz et al. 2008; Warburton et al. 2006; Zhang et al. 2006). The production of synthetic hexaploid (SH) wheats goes back to the 1940s but is recently considered a strategic approach to exploit germplasm of wild wheat progenitors in commercial breeding programs (Mizuno et al. 2010; van Ginkel and Ogbonnaya 2007; Warburton et al. 2006; Xie and Nevo 2008; Yang et al. 2009). SHs are produced by crossing tetraploid wheat (T. turgidum L., 2n = 4× = 28, A and B genomes) with diploid goatgrass (Aegilops tauschii Coss., 2n = 2× = 14, DD genomes) followed by chromosome doubling of the F1 hybrid. The resulting synthesized hexaploids provide a rich source of genetic variation and can be readily hybridized with elite bread wheat cultivars and germplasm. Breeders have exploited these sources for resistance to a wide range of biotic and abiotic stresses (Adhikari et al. 2003; Arraiano et al. 2001b; Assefa and Fehrmann 1998, 2000, 2004; Berzonsky et al. 2004; Cakmak et al. 1999; Genc and McDonald 2004; Gororo et al. 2001; Konik-Rose et al. 2009; Lage et al. 2003, 2004; Lage and Trethowan 2008; Mujeeb-Kazi et al. 2001a, b; Sotelo et al. 2009; Xu et al. 2004, 2006). Here, we further investigate the potential of SHs and derived breeding lines as sources of resistance to M. graminicola in commercial resistance breeding programs.
Materials and methods
Mycosphaerella graminicola isolates, plant materials and experimental design
A global panel of 20 M. graminicola isolates (Table 1) was assembled and used to characterize the response of mapped Stb genes and compare their resistance spectrum with uncharacterized resistance to STB in SHs and derived breeding lines. A set of 20 wheat accessions comprising 13 M. graminicola differential cultivars, five SHs, and the susceptible checks cv. Taichung 29 and the hard red spring wheat cv. Kulm (Table 2), was tested in a triplicate seedling experiment.
F1 and F2 plants, and an F6:7 population of recombinant inbred lines (RILs) developed by single-seed descent were produced from a cross between the SH M3 and cv. Kulm. M3 (W-7976) was developed at CIMMYT by A. Mujeeb-Kazi and has the pedigree Cando/R143//Mexi’S’/3/Ae. tauschii (C122), whereas cv. Kulm was developed at North Dakota State University, Fargo, ND. The hexaploid wheat cv. Chinese Spring (CS) and CS chromosome 5A deletion lines 5AS-1, 5AS-3, 5AL-10, 5AL-12, and 5AL-17 (Endo and Gill 1996) were used to locate chromosome 5A markers to deletion bins.
The various wheat accessions were grown in VQB 7 × 7 × 8 cm TEKU® plastic pots with ten linearly sown seeds per pot. RILs were planted in 5.5 × 5 cm round Jiffy® pots with three seeds per pot using a steam-sterilized peat/sand mixture. All plants were grown in a controlled greenhouse compartment with a 16/8 h light/dark cycle supplemented with son-T Agro 400 W lamps (Hortilux, Boca Raton, Florida, USA). Pre-inoculation temperature and relative humidity (RH) were 18/16°C (day/night rhythm) and 70% RH, while post-inoculation temperature and RH were 22/21°C and ≥85% RH, respectively. Plants were grown in an alpha lattice experimental design with pots as experimental units that were randomly arranged for each isolate-replication combination on separate parallel tables in the greenhouse compartment.
Inoculation procedures
Pre-cultures of each isolate (Table 1) were prepared in an autoclaved 100 ml Erlenmeyer flask containing 50 ml yeast-glucose (YG) liquid medium (30 g glucose, 10 g yeast per liter demineralized water). The flasks were inoculated using a small piece of frozen isolate mycelium maintained at −80°C and were placed in an incubated rotary shaker (Innova 4430, New Brunswick Scientific, USA) set at 125 rpm and 18°C for 5–6 days. These pre-cultures were then used to inoculate three 250 ml Erlenmeyer flasks containing 100 ml YG media per isolate that were incubated under the aforementioned conditions to provide enough inoculum for the seedling inoculation assays at growth stage (GS) 11 (Zadoks et al. 1974). The inoculum concentration was adjusted to 107 spores/ml in a total volume of 40 ml for a set of 18 plastic pots or 24 Jiffy® pots and was supplemented with two drops of Tween 20 (MERCK®, Nottingham, UK). The screening of the 20 wheat accessions as seedlings was conducted using the collection of 20 isolates (Table 1). Seedlings of the entire RIL population were initially tested with M. graminicola isolates IPO92004, IPO92034, IPO94218 and IPO88018, and the results of these pre-screening experiments were used to select the most appropriate isolates (IPO94218 and IPO88018) for the second and third replications that were also used to screen F1 and F2 seedlings.
Adult plant screening of the RILs and parents was carried out in a greenhouse experiment with three replications using M. graminicola isolate IPO88018 (0.6 × 106 spores/ml) at GS 47–58.
Data collection and analysis
Wheat germplasm
Disease severity was evaluated 21 days after inoculation by estimating the percentage necrosis (N) and pycnidia (P) on the inoculated first leaves (GS 11–12) (Zadoks et al. 1974) in the seedling assays. Data were transformed to the logit scale for statistical analysis using residual maximum likelihood (REML) variance component analysis (Genstat 13th edition, VSN International Ltd, Hemel Hempstead, UK). Significant differences were determined using the least significant difference (LSD) of back-transformed N and P values. Logit transformed data analysis resulted in minor changes between observed and processed data to cope with zero scores of N or P.
RILs
A total of 96 RILs were evaluated in the pre-screening (first replication) and 103 RILs in the second and third replications. Disease severity on the seedlings was evaluated 23 days post inoculation (dpi) by scoring N and P on the primary leaves. Latency periods (NLP and PLP: days between inoculation and first N and P appearance) were also determined in the second and third replications of the seedling assays. Adult plant responses—total leaf area covered with sporulating STB lesions—were scored on the flag leaves (F) or the second leaf layer (F-1) at 21 and 28 dpi. Bartlett’s χ2 test was employed to evaluate the homogeneity of replication error variances and calculated using the Excel formula option. Data homogeneous across replications were subsequently averaged and used for QTL analysis (Chu et al. 2010; Friesen et al. 2009).
Molecular mapping in the RIL population
DNA was extracted from M3, cv. Kulm and the RILs as described in Faris et al. (2000). A total of 609 microsatellite (simple sequence repeat; SSR) primer pairs were tested on M3 and cv. Kulm to reveal polymorphisms. The microsatellite primers were derived from the following sets: GWM (Roder et al. 1998), WMC (Somers et al. 2004), HBG, HBD, HBE (Torada et al. 2006), CFA, CFD (Sourdille et al. 2004), BARC (Song et al. 2005), and FCP (Faris et al. 2010; Reddy et al. 2008; Zhang et al. 2009). Methods for PCR, polyacrylamide gel electrophoresis, and fragment visualization were as described in Lu et al. (2006). Primer pairs revealing polymorphism between M3 and cv. Kulm were subsequently used to genotype the 103 RILs.
A total of 284 of the 609 (47%) primer sets revealed polymorphisms and detected 349 marker loci (1.2 loci per primer set). Linkage analysis of the 349 loci was conducted using Mapmaker (Lander et al. 1987) for Macintosh and the Kosambi mapping function (Kosambi 1944) as described in Liu et al. (2005).
QTL analysis
Linkage maps consisting of 296 markers giving the most complete genome coverage were used to detect genomic regions associated with phenotypic means. Composite interval mapping (CIM) was performed using the computer program QGene (Joehanes and Nelson 2008). A permutation test with 1,000 permutations was conducted to determine that a critical LOD threshold of 4.7 in this population yields an experiment-wise significance level of 0.05.
Genotype to phenotype discrepancy
Analysis of the allelic marker segregation and concurrent phenotypic data of the RILs enabled us to study genotype to phenotype discrepancies with respect to STB resistance. We used all observed disease parameters (N, P, NLP and PLP) and distributed the RILs in statistically significantly different (χ 21:1 ) groups. Subsequently, marker segregation was superimposed on these data to determine sliding windows of lower to upper limits of the aforementioned disease parameters for each isolate to determine the threshold values for segregation analyses. At a later stage 11 individual RILs (KM7, KM8, KM14, KM15, KM20, KM21, KM32, KM41, KM63, KM73 and KM88) were screened with the entire panel of M. graminicola isolates to confirm broad efficacy of the identified resistance loci.
Results
Wheat germplasm screen
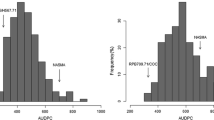
All control inoculations resulted in excellent disease development enabling precise phenotyping of wheat germplasm and the Kulm/M3 RIL population. None of the differential cultivars was completely resistant to the global M. graminicola panel, whereas all SHs, including M3, were widely resistant to the entire set of isolates (Table 3). The number of identified Stb genes in each differential cultivar (Table 2) positively correlated with broader efficacy (r = 0.75, P < 0.01; N = 13, df = 11) indicating that accumulation of Stb genes is a valid resistance breeding strategy. In contrast, the SHs showed a significantly different pattern for they were resistant to all M. graminicola isolates (Fig. 1; Table 3). We therefore focused further analyses on the cv. Kulm/M3 RIL population. The parental lines differed significantly for N (values for cv. Kulm and M3 ranging from 2.2 to 91.8 and 1.1 to 6.8, respectively) and P (values for cv. Kulm and M3 ranging from 0 to 37.5 and 0, respectively) over the 20 isolates (Fig. 1). This enabled the selection of isolates IPO94218, IPO92004, IPO88018 and IPO92034 for further analysis.
Mapping
The 349 microsatellite markers were assembled into linkage groups representing the 21 hexaploid wheat chromosomes and spanned a genetic distance of 2,465 cM. Only chromosomes 3D and 5A were associated with STB resistance and these will be shown here, details of map construction and analysis will be published elsewhere. The genetic map of chromosome 3D in the cv. Kulm/M3 RIL population consisted of 27 markers spanning a genetic distance of 67.9 cM and included a cluster of 18 co-segregating markers near the distal end of the long arm (Fig. 3). Comparison with the 3D deletion-based physical map indicated that this suppressed recombination occurred across much of the long arm of chromosome 3D (Fig. 4). Closer evaluation of the 3D marker profiles indicated that most were codominant, and hence, there was no indication of a large deletion on chromosome 3D in either M3 or cv. Kulm.
The linkage map of chromosome 5A consisted of 13 markers spanning 125.4 cM (Fig. 3). Of the markers mapped to 5A in the cv. Kulm/M3 population, only Xbarc180, Xcfa2250, Xbarc141, Xgwm617, Xgwm595, and Xgwm291 were previously located on the deletion-based physical map (Sourdille et al. 2004). Therefore, we tested markers Xhbd160, Xhbg247, Xhbg219, Xbarc232, Xhbd150, and Xwmc524 on the 5A deletion lines to determine their locations on the physical map. Comparison of the cv. Kulm/M3 5A genetic map with the 5A physical map indicated that the genetic linkage map of 5A developed in the cv. Kulm/M3 population accounted for most of the chromosome (Fig. 4).
Phenotyping and QTL analyses
RIL screening
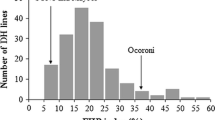
We produced 103 cv. Kulm/M3 RILs and 96 were inoculated with M. graminicola isolates IPO94218, IPO92004, IPO88018 and IPO92034 in the first replication (Fig. 2). The results of this experiment indicated that segregation ratios of P fit 1:1 ratios for M. graminicola isolates IPO92004, IPO88018 and IPO92034, suggesting segregation of a single genetic factor. The result with IPO94218, however, indicated that more genes could be involved. We, therefore, continued analyses for the second and third replications with M. graminicola isolates IPO88018 and IPO94218. Ranking of the RILs for N and P showed highly significant correlations for N and P, indicating that the same genetic factor(s) could control resistance to these isolates (Table 4).
QTL analyses
Seedling resistance
QTL analysis using CIM indicated that, for both M. graminicola isolates IPO88018 and IPO94218, markers located on the long arm of chromosome 3D were significantly associated with N, P, NLP, and PLP in seedlings (Table 5; Fig. 3). The QTLs peaked at position 58.0 cM between SSR loci Xwmc494 and Xbarc125 for each trait (Figs. 3, 4), and resistance effects were derived from M3. LOD values were highly significant ranging from 11.7 to 22.3 for the phenotypes caused by isolate IPO94218 and 19.0–27.0 for those caused by isolate IPO88018 (Fig. 3; Table 5). The QTL explained from 41 to 64% of the phenotypic variation for the disease caused by isolate IPO94218, and 58–71% of the variation for disease caused by isolate IPO88018.
Comparison of the Chinese Spring chromosome 3D and 5A deletion-based physical maps with the 3D and 5A genetic linkage maps developed in the cv. Kulm/M3 population. Deletion breakpoints are indicated to the left of the physical maps and bin-located markers are shown along the right. On the linkage maps, cM distances are shown along the left and markers along the right. The QTL regions associated with STB resistance are indicated by the red lines
Adult plant resistance
QTL analysis of adult plant reactions to M. graminicola isolate IPO88018 indicated that the resistance locus on 3DL identified at the seedling stage, was also significantly associated with resistance at both the 21 and 28 dpi readings (Fig. 3; Tables 6, 7). The QTL peaked at the same cM position as for the seedling data for both isolates and had LOD values of 7.2 and 8.4 for the 21 and 28 dpi readings, respectively. The locus explained 28% of the variation in STB at 21 dpi, which increased to 31% at 28 dpi. In addition to the resistance locus on 3DL, an additional QTL associated with adult plant resistance derived from M3 was identified on the long arm of chromosome 5A (Fig. 3). The 5AL QTL had a LOD value of 3 and explained 12% of the variation at 21 dpi, but had stronger effects at 28 dpi with an LOD of 8.9, explaining 32% of the variation (Table 6). The 5AL QTL was flanked by SSR loci Xgwm617 and Xhbg247, and it peaked approximately 3.1 cM proximal to Xhbg247 (Fig. 3). Comparisons between the genetic and physical maps indicated that this QTL was located in the deletion bin defined by the breakpoints in deletion lines 5AL-10 and 5AL-17, which is in the distal half of 5AL (Figs. 4, 5). Comparative RIL (KM7, KM20, KM41 and KM73) genotyping/phenotyping showed that the presence of the 5AL locus in KM41 specifically incited resistance to isolate IPO88018 in adults plants, but KM41 was susceptible in the seedling stage similar to KM73 that lacks the 3DL as well as the 5AL QTL (Table 7).
Segregation for N, P, NLP and PLP in the cv. Kulm/M3 RIL population inoculated with M. graminicola isolates IPO88018 and IPO94218 overlaid with allelic segregation of the Xwmc494 SSR marker which is linked to Stb16q. ‘M’ and ‘K’ indicate parental bin-values. Blue and purple triangles indicate average values of RILS with ‘M’ and ‘K’ alleles, respectively. The vertical dashed line is the χ 21:1 validated threshold position between resistant and susceptible RILs
F1 and F2 screening
Four F1 plants were inoculated with M. graminicola isolate IPO88018 and showed only minor tip leaf necrosis and no pycnidia formation at 21 dpi (data not shown). Thirty-two and 28 F2 plants were then inoculated with M. graminicola isolates IPO88018 and IPO94218, respectively. Segregation ratios (resistant:susceptible) for N and P did not significantly differ from the expected 3:1 (Table 8), suggesting the inheritance of a single dominant gene.
Genotyping versus phenotyping discrepancies
Analyses of the phenotypic and genotypic data indicated that lines with the Xwmc494 allele from M3 had P values that ranged from 0 to 5 and N values from 0 to 30, with averages over both isolates of 1 and 15, respectively. On the contrary, RILs carrying the cv. Kulm allele for Xwmc494 had values that ranged from 8 to 70 P and 37–100 N, and averaged over both isolates of 30 and 80, respectively.
Despite the indications for a single locus inheritance, the observed recombination suppression on chromosome 3D could also mask several genes at the 3D locus. We, therefore, tested RILs KM7, KM8, KM14, KM15, KM20, KM21, KM32, KM41, KM63, KM73 and KM88 with the entire panel of isolates (Table 1) to confirm either broad susceptibility or resistance with the presence of the Kulm or M3 alleles of the flanking Xwmc494 and Xbarc125 SSR loci, respectively (Table 9). The phenotypes of the majority of RILs was as expected either broadly resistant (KM7 and KM20 with M3 alleles of the flanking markers) or susceptible (KM15, KM41, KM21, KM63 and KM73 with M3 alleles of the flanking markers), although KM63 was unexpectedly resistant to isolate IPO86068. This could have been an incidental escape as Kulm itself also showed some variation compared to the earlier screen (Table 3). However, RILs KM8, KM32, KM88 and KM14, which also have the M3 alleles of the flanking markers, clearly differed from the parental phenotypes by showing specificity to isolate panel.
Discussion
Here we report two new STB resistance genes that were derived from the SH wheat line M3. Segregation and QTL analyses as well as genetic and physical mapping suggested that a single locus on chromosome 3D derived from M3 conferred resistance to all STB disease parameters in the seedling stage in the cv. Kulm/M3 RIL population. Although (1) no additional QTLs were significantly associated with any of the seedling phenotypes caused by either isolate in genome-wide scans, (2) none of the known Stb genes were mapped on chromosome 3D and, (3) the 3D QTL was highly significant and explained a large portion of the phenotypic variation, we cannot unequivocally conclude on single gene inheritance due to the substantial recombination suppression along the long arm of chromosome 3D, which is not due to a large deletion. However, it is possible that a large inversion exists in 3D of one of the parents, which could be the cause of the extreme suppression of recombination on 3DL. Due to the highly suppressed recombination along chromosome arm 3DL, comparison with the physical map of 3D yielded little additional information. We therefore propose to tentatively designate this locus Stb16q in accordance with a suggestion in a recent community wide discussion on Stb nomenclature to add ‘q’ for cases where presented data do not unequivocally show that a single gene underlies a detected QTL. Indeed additional inoculation studies with 11 selected RILs using the full isolate panel showed that RILs KM8, KM32, KM88 and KM14 showed a differential pattern that cannot be explained by a single gene. Nevertheless, the results indicate that Stb16q lies on the long arm of chromosome 3D and that it was derived from the Ae. tauschii accession C122, which was the donor of the D-genome chromosomes in M3. Zwart et al. (2010) reported a SH derived QTL with multiple unrelated functions including STB resistance on chromosome 3D, but the LOD scores were relatively low and STB resistance was only tested with a single non-characterized M. graminicola isolate. Our study showed that Stb16q had an unusually broad efficacy in the seedling stage as shown by the resistance to the global panel of isolates, and is also expressed in adult plants.
In addition we determined a QTL on chromosome 5AL that does not confer resistance to STB in seedlings, but specifically in adult plants. None of the previously characterized Stb genes was mapped on chromosome 5A (Arraiano et al. 2007; Chartrain et al. 2009; Goodwin 2007) and we, therefore, conclude that this QTL represents a novel gene for STB resistance. At 28 dpi is showed a highly significant LOD that explained a substantial percentage (32%) of the observed STB variation compared with the 21 dpi observations. This is not surprising since disease development in adult plants usually takes slightly longer, particularly under greenhouse conditions. A recent community wide discussion on Stb nomenclature suggested that Stb genes should at least explain 50% of the observed STB variation before a number could be assigned to such a locus. We currently consider this as an unjustified criterion for gene designation as we recently showed that the percentage of explained variation strongly depends on the applied isolates (Tabib Ghaffary et al. 2011). Hence, we designate the 5AL locus as Stb17, which originated from the tetraploid durum wheat line used in the development of M3. Previous experiments conducted to compare seedling and adult plant STB resistance suggested the occurrence of specific seedling resistance loci, but no specific adult plant resistance genes were identified (Kema and van Silfhout 1997). All previously reported Stb genes are effective in the seedling and adult plant stage as indicated in the present study with Stb16q. Evidently, the unequivocal identification of specific adult plant resistance loci can only come from comparative seedling/adult plant mapping studies using the same populations. The majority of analyses has been performed in either of both stage and cannot conclude on the occurrence of specific adult plant resistance (Adhikari et al. 2003, 2004a, c; Chartrain et al. 2005a, b, c, 2009; Risser et al. 2011). Three studies used seedling and adult plant data in mapping analyses. Arraiano et al. (2001b) mapped Stb5 in seedling and adult plant experiments. In the study of Simon et al. (2004) none of the scored parameters resulted in a significant QTL except for AUDPC (QStb.ipk-7B) that mapped on the same position as Stb8, which is also expressed in seedlings (Table 3). Finally, Simon et al. (2010) mapped resistance in seedlings and adult plants on chromosome 7D, but only interval mapping suggested that the adult plant locus was apart from Stb4 or Stb5 that map on the same chromosome. Moreover, the adult plant data only included limited necrosis scores, no LOD values of the identified QTLs were provided and none of them were named. We, therefore, claim that Stb17 is the first gene for adult plant resistance to M. graminicola since the 5A QTL is specifically expressed in adult plants after inoculation with isolate IPO88018 and chromosome 5 has not yet been associated with resistance (the efficacy of Stb17 to a wider set of isolates has to be determined and we cannot exclude that it might be expressed in seedlings with other isolates). This complies with what is classically described as adult plant resistance, which is very common to other cereal diseases such as the rusts and has been associated with temperature sensitivity and other abiotic environmental factors (McIntosh et al. 1995), and we adopt that interpretation here.
Interestingly, the response of M3 to the global panel of M. graminicola isolates was very similar to those of the other tested SHs. The broad resistance spectrum of Stb16q might be due to the apparent dichotomy of host specificity in the wheat-M. graminicola pathosystem. Kema et al. (1996a, b) summarized and extended these observations and showed that M. graminicola isolates are in general either pathogenic on bread wheat or durum wheat. Recently, Wittenberg et al. (2009) and Ware Sarrah (2006) showed that genetic recombination during sexual reproduction in M. graminicola easily results in progeny with altered cultivar and host specificity. However, tetraploid wheats are in general resistant to M. graminicola isolates derived from bread wheat and vice versa. This was confirmed in the current experiments because neither of the durum wheat-derived isolates IPO86022 and IPO95052 were virulent on any of the tested bread wheat accessions including the susceptible parent cv. Kulm and the susceptible check cv. Taichung 29. Therefore, a SH is expected to be resistant to such bread wheat derived M. graminicola isolates unless the D genome component affects the expression of resistance, which has been shown for rust diseases (Kerber and Green 1980, Kema et al. 1995). Assefa and Fehrmann (1998) also documented broad-spectrum resistance to M. graminicola (99% of 194 accessions) in seven Aegilops species, while only 8, 11, 16 and 24% of this collection was resistant to stem rust, leaf rust, eyespot and powdery mildew, respectively. Similar broad spectrum resistance was observed in phenotypic screens of the diploid wheat T. monococcum, which led to the identification of the resistance locus TmStb1 and the linked microsatellite locus Xbarc174 on chromosome 7Am (Jing et al. 2008). Because SHs effectively combine the genomes of tetraploid and diploid wheat progenitors and relatives (Mujeeb-Kazi et al. 1996; Yang et al. 2009), they may carry a reservoir of novel genes for resistance to M. graminicola. Despite the value of the genes that we discovered, exposure to M. graminicola populations may potentially enable the fungus to adapt and circumvent them (Wittenberg et al. 2009; Ware Sarrah 2006; McDonald and Linde 2002a, b; Linde et al. 2002; Zhan et al. 2007). Hence, their commercial deployment should take these observations into consideration to maximize their efficacy under practical conditions.
To date, there has been no report of mapping host QTLs associated with life strategy parameters such as latency period and the lesion development rate of M. graminicola. Here, we characterized classical (N and P) and new parameters (NLP, PLP) to investigate whether a major STB resistance gene also controls underlying pathogenicity factors, which is relevant, as resistance to STB is characterized by the absence of the hypersensitive response (HR) (Kema et al. 1996d). Interestingly, all the analyzed parameters mapped to the Stb16q locus. In the absence of the HR, resistance is achieved by reducing the development of fungal biomass, which may occur by reducing infection rates. Such partial, or ‘horizontal’, resistance has been observed in some cereal rust interactions (Aghnoum and Niks 2010; Marcel et al. 2008). One of the best-known ‘slow rusting’ loci is the Lr34/Yr18/Pm38 complex (Singh et al. 2007), which confers partial resistance to stripe rust, leaf rust and powdery mildew. Molecular cloning of the Lr34/Yr18/Pm38 locus indicated that it is a unique functional ABC transporter (Krattinger et al. 2009; Lagudah et al. 2009). On the contrary, genes that confer complete, or ‘vertical’, resistance to pathogens with biotrophic lifestyles and susceptibility to necrotrophic pathogens usually harbor NBS and LRR domains (Bent and Mackey 2007; Jones and Dangl 2006; McDowell and Simon 2006; Lorang et al. 2007; Nagy and Bennetzen 2008; Faris et al. 2010). Tsn1, a gene controlling sensitivity to a host-selective toxin produced by the necrotrophic fungal pathogens Stagonospora nodorum and Pyrenophora tritici-repentis has resistance gene-like features including protein kinase and NBS-LRR domains (Faris et al. 2010). Interestingly, S. nodorum, P. tritici-repentis and M. graminicola are close relatives and belong to the Dothideomycete class of fungi. However, nothing is currently known about the molecular characteristics of Stb resistance genes. Therefore, the wide efficacy of the Stb16q locus and the abovementioned findings call not only for further deciphering and understanding of the resistance mechanism exerted by these new genes for resistance to STB, but also for unveiling their molecular structure.
Because the M. graminicola-wheat pathosystem is characterized by the absence of an HR, resistance and susceptibility are currently usually expressed on a quantitative scale. However, symptom expression is strongly affected by environmental fluctuations and hence repeatability of experiments might be low (Arraiano et al. 2001a; Bearchell et al. 2005; Czembor et al. 2010; Kema et al. 1996a). Early reports determined an arbitrary threshold of resistance and susceptibility by using a 0–5 scoring scale (Rosielle 1972) that was more qualitative than quantitative. Later, applications of complex statistics were used to turn qualitative data into qualitative determinants (Eyal and Levy 1987; Eyal et al. 1985; Yechilevich-Auster et al. 1983). Eventually, Kema et al. (1996a, b) used quantitative data in cluster analyses based on interaction components of analyses of variance to group isolates and cultivars with similar responses and hypothesized that N and P were controlled by different genetic factors in the fungal genome. This was later corroborated by formal fungal genetics (Kema et al. 2000, 2002; Wittenberg et al. 2009; Ware Sarrah 2006). Adhikari et al. (2003, 2004a, b, c) used a modified 0–5 scale, which considered pycnidia percentage and density, for the mapping of several Stb genes, but phenotypic classifications were not matched with allelic segregations of the associated markers. A detached leaf assessment method also has been established for the characterization and mapping of some Stb genes (Arraiano et al. 2001a; Chartrain et al. 2005a, c, 2009). Essentially, all these phenotyping assays address the phenotyping versus genotyping problem (Dowell et al. 2010). Here we had the opportunity to study phenotype/genotype variation in more detail using the allelic information of all RILs along with all observed disease assessment parameters. As Stb16q controls all the observed disease parameters for a global panel of unrelated M. graminicola isolates, the phenotypes of RILs with alternative parental alleles at the Xwmc494 locus are of interest. RILs with the Xwmc494 allele of M3 showed 0–5 P and 0–30 N values, whereas RILs with cv. Kulm allele showed 8–70 P and 37–100 N values. We do not know the origin of these sliding disease parameter windows, but we cannot exclude phenotyping errors due to environmental fluctuations, despite the accordance of all replications. We can exclude genotyping errors and recombination events between the Xwmc494 marker and Stb16q as possible sources of error because the results indicate significant recombination suppression in this region evidenced by the fact that 18 SSR markers that co-segregated at a single locus on the genetic map were distributed across 3DL on the deletion-based physical map. However, unknown genetic modifiers could also play an important role in genotype to phenotype variation in wheat. What counts, however, is that despite the presence of Stb16q, resistant plants may develop up to 5% P and 30% N, which is close to the lowest values for plants lacking Stb16q, which had values as low as 8% P and 37% N. The application is that the distinguishing threshold between resistance and susceptibility in a given population should not be taken arbitrarily, as indicated previously (Adhikari et al. 2003, 2004b; Chartrain et al. 2005b), and ought to be based on appropriate genotype versus phenotype analyses.
In conclusion, the present results show that Stb16q and Stb17 are valuable new resistance loci that can be easily deployed in national and international marker-assisted resistance breeding programs. However, M. graminicola is classified as a high to moderate risk pathogen due to its multiple asexual and sexual cycles per year and its effective spore dissemination mechanism (McDonald and Linde 2002a, b), which enabled the fungus to circumvent Stb genes deployed in commercial wheat (Linde et al. 2002; Wittenberg et al. 2009; Zhan et al. 2007). We, therefore, discourage using Stb16q or Stb17 as single genes, but rather suggest pyramiding strategies with other STB resistance genes in order to maximize their commercial life span.
References
Adhikari TB, Anderson JM, Goodwin SB (2003) Identification and molecular mapping of a gene in wheat conferring resistance to Mycosphaerella graminicola. Phytopathology 93:1158–1164
Adhikari TB, Cavaletto JR, Dubcovsky J, Gieco JO, Schlatter AR, Goodwin SB (2004a) Molecular mapping of the Stb4 gene for resistance to septoria tritici blotch in wheat. Phytopathology 94:1198–1206
Adhikari TB, Wallwork H, Goodwin SB (2004b) Microsatellite markers linked to the Stb2 and Stb3 genes for resistance to septoria tritici blotch in wheat. Crop Sci 44:1403–1411
Adhikari TB, Yang X, Cavaletto JR, Hu X, Buechley G, Ohm HW, Shaner G, Goodwin SB (2004c) Molecular mapping of Stb1, a potentially durable gene for resistance to septoria tritici blotch in wheat. Theor Appl Genet 109:944–953
Aghnoum R, Niks RE (2010) Specificity and levels of nonhost resistance to nonadapted Blumeria graminis forms in barley. New Phytol 185:275–284
Arraiano LS, Brown JKM (2006) Identification of isolate-specific and partial resistance to septoria tritici blotch in 238 European wheat cultivars and breeding lines. Plant Pathol 55:726–738
Arraiano LS, Brading PA, Brown JKM (2001a) A detached seedling leaf technique to study resistance to Mycosphaerella graminicola (anamorph Septoria tritici) in wheat. Plant Pathol 50:339–346
Arraiano LS, Worland AJ, Ellerbrook C, Brown JKM (2001b) Chromosomal location of a gene for resistance to septoria tritici blotch (Mycosphaerella graminicola) in the hexaploid wheat ‘Synthetic 6×’. Theor Appl Genet 103:758–764
Arraiano LS, Chartrain L, Bossolini E, Slatter HN, Keller B, Brown JKM (2007) A gene in European wheat cultivars for resistance to an African isolate of Mycosphaerella graminicola. Plant Pathol 56:73–78
Assefa S, Fehrmann H (1998) Resistance in Aegilops species against leaf rust, stem rust, septoria tritici blotch, eyespot and powdery mildew of wheat. Zeitschrift Fur Pflanzenkrankheiten Und Pflanzenschutz-Journal of Plant Diseases and Protection 105:624–631
Assefa S, Fehrmann H (2000) Resistance to wheat leaf rust in Aegilops tauschii Coss. and inheritance of resistance in hexaploid wheat. Genet Resour Crop Evol 47:135–140
Assefa S, Fehrmann H (2004) Evaluation of Aegilops tauschii Coss. for resistance to wheat stem rust and inheritance of resistance genes in hexaploid wheat. Genet Resour Crop Evol 51:663–669
Bearchell S, Fraaije B, Shaw M, Fitt B (2005) Wheat archive links long-term fungal pathogen population dynamics to air pollution. Proc Natl Acad Sci USA 102:5438
Beest DET, Shaw MW, Paveley ND, van den Bosch F (2009) Evaluation of a predictive model for Mycosphaerella graminicola for economic and environmental benefits. Plant Pathol 58:1001–1009
Bent AF, Mackey D (2007) Elicitors, effectors, and R genes: the new paradigm and a lifetime supply of questions. Annu Rev Phytopathol 45:399–436
Berzonsky WA, Hartel KD, Kianian SF, Leach GD (2004) Registration of four synthetic hexaploid wheat germplasm lines with resistance to fusarium head blight. Crop Sci 44:1500–1501
Brading PA, Verstappen ECP, Kema GHJ, Brown JKM (2002) A gene-for-gene relationship between wheat and Mycosphaerella graminicola, the septoria tritici blotch pathogen. Phytopathology 92:439–445
Burke JJ, Dunne B (2008) Investigating the effectiveness of the Thies Clima “Septoria Timer” to schedule fungicide applications to control Mycosphaerella graminicola on winter wheat in Ireland. Crop Prot 27:710–718
Cakmak I, Cakmak O, Eker S, Ozdemir A, Watanabe N, Braun HJ (1999) Expression of high zinc efficiency of Aegilops tauschii and Triticum monococcum in synthetic hexaploid wheats. Plant Soil 215:203–209
Chartrain L, Berry ST, Brown JKM (2005a) Resistance of wheat line Kavkaz-K4500 L.6.A.4 to septoria tritici blotch controlled by isolate-specific resistance genes. Phytopathology 95:664–671
Chartrain L, Brading PA, Brown JKM (2005b) Presence of the Stb6 gene for resistance to septoria tritici blotch (Mycosphaerella graminicola) in cultivars used in wheat-breeding programmes worldwide. Plant Pathol 54:134–143
Chartrain L, Joaquim P, Berry ST, Arraiano LS, Azanza F, Brown JKM (2005c) Genetics of resistance to septoria tritici blotch in the Portuguese wheat breeding line TE 9111. Theor Appl Genet 110:1138–1144
Chartrain L, Sourdille P, Bernard M, Brown JKM (2009) Identification and location of Stb9, a gene for resistance to septoria tritici blotch in wheat cultivars Courtot and Tonic. Plant Pathol 58:547–555
Chen RS, McDonald BA (1996) Sexual reproduction plays a major role in the genetic structure of populations of the fungus Mycosphaerella graminicola. Genetics 142:1119–1127
Christiansen MJ, Andersen SB, Ortiz R (2002) Diversity changes in an intensively bred wheat germplasm during the 20th century. Mol Breed 9:1–11
Chu CG, Faris JD, Xu SS, Friesen TL (2010) Genetic analysis of disease susceptibility contributed by the compatible Tsn1-SnToxA and Snn1-SnTox1 interactions in the wheat-Stagonospora nodorum pathosystem. Theor Appl Genet 120:1451–1459
Cowger C, Hoffer ME, Mundt CC (2000) Specific adaptation by Mycosphaerella graminicola to a resistant wheat cultivar. Plant Pathol 49:445–451
Czembor P, Radecka Janusik M, Ma kowski D (2010) Virulence spectrum of Mycosphaerella graminicola isolates on wheat genotypes carrying known resistance genes to septoria tritici blotch. J Phytopathol. doi:10.1111/j.1439-0434.2010.01734.x
Dowell R, Ryan O, Jansen A, Cheung D, Agarwala S, Danford T, Bernstein D, Rolfe P, Heisler L, Chin B (2010) Genotype to phenotype: a complex problem. Science 328:469
Dreisigacker S, Kishii M, Lage J, Warburton M (2008) Use of synthetic hexaploid wheat to increase diversity for CIMMYT bread wheat improvement. Aust J Agric Res 59:413–420
Duveiller E, Singh RP, Nicol JM (2007) The challenges of maintaining wheat productivity: pests, diseases, and potential epidemics. Euphytica 157:417–430
Endo T, Gill B (1996) The deletion stocks of common wheat. J Hered 87:295–307
Eyal Z (1999) The septoria tritici and Stagonospora nodorum blotch diseases of wheat. Eur J Plant Pathol 105:629–641
Eyal Z, Levy E (1987) Variations in pathogenicity patterns of Mycosphaerella-graminicola within Triticum spp. in Israel. Euphytica 36:237–250
Eyal Z, Scharen AL, Huffman MD, Prescott JM (1985) Global insights into virulence frequencies of Mycosphaerella graminicola. Phytopathology 75:1456–1462
FAO (2010) http://www.fao.org/wsfs/world-summit/wsfs-challenges/en/
Faris JD, Haen KM, Gill BS (2000) Saturation mapping of a gene-rich recombination hot spot region in wheat. Genetics 154:823–835
Faris J, Zhang Z, Lu H, Lu S, Reddy L, Cloutier S, Fellers J, Meinhardt S, Rasmussen J, Xu S, Oliver R, Simons KJ, Friesen TL (2010) A unique wheat disease resistance-like gene governs effector-triggered susceptibility to necrotrophic pathogens. Proc Natl Acad Sci 107:13544–13549
Fraaije BA, Cools HJ, Fountaine J, Lovell DJ, Motteram J, West JS, Lucas JA (2005) Role of ascospores in further spread of QoI-resistant cytochrome b alleles (G143A) in field populations of Mycosphaerella graminicola. Phytopathology 95:933–941
Friesen TL, Chu CG, Liu ZH, Xu SS, Halley S, Faris JD (2009) Host-selective toxins produced by Stagonospora nodorum confer disease susceptibility in adult wheat plants under field conditions. Theor Appl Genet 118:1489–1497
Genc Y, McDonald GK (2004) The potential of synthetic hexaploid wheats to improve zinc efficiency in modern bread wheat. Plant Soil 262:23–32
Goodwin SB (2007) Back to basics and beyond: increasing the level of resistance to Septoria tritici blotch in wheat. Australas Plant Pathol 36:532–538
Gororo NN, Flood RG, Eastwood RF, Eagles HA (2001) Photoperiod and vernalization responses in Triticum turgidum × T.tauschii synthetic hexaploid wheats. Ann Bot 88:947–952
Heun M, Schafer-Pregl R, Klawan D, Castagna R, Accerbi M, Borghi B, Salamini F (1997) Site of einkorn wheat domestication identified by DNA fingerprinting. Science 278:1312
Jing HC, Lovell D, Gutteridge R, Jenk D, Kornyukhin D, Mitrofanova OP, Kema GHJ, Hammond-Kosack KE (2008) Phenotypic and genetic analysis of the Triticum monococcum–Mycosphaerella graminicola interaction. New Phytol 179:1121–1132
Joehanes R, Nelson JC (2008) QGene 4.0, an extensible Java QTL-analysis platform. Bioinformatics 24:2788–2789
Jones JDG, Dangl JL (2006) The plant immune system. Nature 444:323–329
Jorgensen LN (2008) Resistance situation with fungicides in cereals. Zemdirbyste-Agriculture 95:373–378
Kema GHJ, van Silfhout CH (1997) Genetic variation for virulence and resistance in the wheat Mycosphaerella graminicola pathosystem 3. Comparative seedling and adult plant experiments. Phytopathology 87:266–272
Kema GHJ, Lange W, Van Silfhout CH (1995) Differential suppression of stripe rust resistance in synthetic wheat hexaploids derived from Triticum turgidum subsp. dicoccoides and Aegilops squarrosa. Phytopathology 85:425–429
Kema GHJ, Annone JG, Sayoud R, VanSilfhout CH, Van Ginkel M, de Bree J (1996a) Genetic variation for virulence and resistance in the wheat-Mycosphaerella graminicola pathosystem 1. Interactions between pathogen isolates and host cultivars. Phytopathology 86:200–212
Kema GHJ, Sayoud R, Annone JG, Van Silfhout CH (1996b) Genetic variation for virulence and resistance in the wheat-Mycosphaerella graminicola pathosystem 2. Analysis of interactions between pathogen isolates and host cultivars. Phytopathology 86:213–220
Kema GHJ, Verstappen ECP, Todorova M, Waalwijk C (1996c) Successful crosses and molecular tetrad and progeny analyses demonstrate heterothallism in Mycosphaerella graminicola. Curr Genet 30:251–258
Kema GHJ, Yu DZ, Rijkenberg FHJ, Shaw MW, Baayen RP (1996d) Histology of the pathogenesis of Mycosphaerella graminicola in wheat. Phytopathology 86:777–786
Kema G, Verstappen E, Waalwijk C (2000) Avirulence in the wheat Septoria tritici leaf blotch fungus Mycosphaerella graminicola is controlled by a single locus. Mol Plant Microbe Interact 13:1375–1379
Kema G, Goodwin S, Hamza S, Verstappen E, Cavaletto J, Van der Lee T, de Weerdt M, Bonants P, Waalwijk C (2002) A combined amplified fragment length polymorphism and randomly amplified polymorphism DNA genetic linkage map of Mycosphaerella graminicola, the septoria tritici leaf blotch pathogen of wheat. Genetics 161:1497–1505
Kerber E, Green G (1980) Suppression of stem rust resistance in the hexaploid wheat cv. Canthatch by chromosome 7DL. Can J Bot 58:1347–1350
King JE, Cook RJ, Melville SC (1983) A review of septoria diseases of wheat and barley. Ann Appl Biol 103:345–373
Konik-Rose CM, Rahman S, Appels R, Moss R, McMaster G, Marshall DR, Stoddard FL (2009) Starch characterisation and variability in GBSS loci of synthetic hexaploid wheats and their durum and Aegilops tauschii parents. Euphytica 167:203–216
Kosambi D (1944) The estimation of map distances from recombination values. Ann Eugen 12:172–175
Krattinger SG, Lagudah ES, Spielmeyer W, Singh RP, Huerta-Espino J, McFadden H, Bossolini E, Selter LL, Keller B (2009) A putative ABC transporter confers durable resistance to multiple fungal pathogens in wheat. Science 323:1360–1363
Lage J, Trethowan RM (2008) CIMMYT’s use of synthetic hexaploid wheat in breeding for adaptation to rainfed environments globally. Aust J Agric Res 59:461–469
Lage J, Skovmand B, Andersen SB (2003) Expression and suppression of resistance to greenbug (Homoptera: Aphididae) in synthetic hexaploid wheats derived from Triticum dicoccum × Aegilops tauschii crosses. J Econ Entomol 96:202–206
Lage J, Skovmand B, Andersen SB (2004) Field evaluation of emmer wheat-derived synthetic hexaploid wheat for resistance to Russian wheat aphid (Homoptera: Aphididae). J Econ Entomol 97:1065–1070
Lagudah ES, Krattinger SG, Herrera-Foessel S, Singh RP, Huerta-Espino J, Spielmeyer W, Brown-Guedira G, Selter LL, Keller B (2009) Gene-specific markers for the wheat gene Lr34/Yr18/Pm38 which confers resistance to multiple fungal pathogens. Theor Appl Genet 119:889–898
Lander E, Green P, Abrahamson J, Barlow A, Daly M, Lincoln S, Newburg L (1987) MAPMAKER: an interactive computer package for constructing primary genetic linkage maps of experimental and natural populations. Genomics 1:174–181
Lehoczki-Krsjak S, Szabo-Hever A, Toth B, Kotai C, Bartok T, Varga M, Farady L, Mesterhazy A (2010) Prevention of Fusarium mycotoxin contamination by breeding and fungicide application to wheat. Food Addit Contam Part A Chem Anal Control Expo Risk Assess 27:616–628
Linde CC, Zhan J, McDonald BA (2002) Population structure of Mycosphaerella graminicola: From lesions to continents. Phytopathology 92:946–955
Liu Z, Anderson J, Hu J, Friesen T, Rasmussen J, Faris J (2005) A wheat intervarietal genetic linkage map based on microsatellite and target region amplified polymorphism markers and its utility for detecting quantitative trait loci. TAG Theor Appl Genet 111:782–794
Lorang JM, Sweat TA, Wolpert TJ (2007) Plant disease susceptibility conferred by a ‘resistance’ gene. Proc Natl Acad Sci USA 104:14861–14866
Lu H, Fellers J, Friesen T, Meinhardt S, Faris J (2006) Genomic analysis and marker development for the Tsn1 locus in wheat using bin-mapped ESTs and flanking BAC contigs. TAG Theor Appl Genet 112:1132–1142
Luo M, Yang Z, You F, Kawahara T, Waines J, Dvorak J (2007) The structure of wild and domesticated emmer wheat populations, gene flow between them, and the site of emmer domestication. TAG Theor Appl Genet 114:947–959
Marcel TC, Gorguet B, Ta MT, Kohutova Z, Vels A, Niks RE (2008) Isolate specificity of quantitative trait loci for partial resistance of barley to Puccinia hordei confirmed in mapping populations and near-isogenic lines. New Phytol 177:743–755
Mavroeidi VI, Shaw MW (2005) Sensitivity distributions and cross-resistance patterns of Mycosphaerella graminicola to fluquinconazole, prochloraz and azoxystrobin over a period of 9 years. Crop Prot. 24:259–266
McCartney CA, Brule-Babel AL, Lamari L, Somers DJ (2003) Chromosomal location of a race-specific resistance gene to Mycosphaerella graminicola in the spring wheat ST6. Theor Appl Genet 107:1181–1186
McDonald BA, Linde C (2002a) Pathogen population genetics, evolutionary potential, and durable resistance. Annu Rev Phytopathol 40:349–379
McDonald BA, Linde C (2002b) The population genetics of plant pathogens and breeding strategies for durable resistance. Euphytica 124:163–180
McDonald BA, Mundt CC, Chen RS (1996) The role of selection on the genetic structure of pathogen populations: evidence from field experiments with Mycosphaerella graminicola on wheat. Euphytica 92:73–80
McDowell JM, Simon SA (2006) Recent insights into R gene evolution. Mol Plant Pathol 7:437–448
McIntosh R, Wellings C, Park R (1995) Wheat rusts: an atlas of resistance genes. Kluwer Academic Publishers, Sydney
Mizuno N, Yamasaki M, Matsuoka Y, Kawahara T, Takumi S (2010) Population structure of wild wheat D-genome progenitor Aegilops tauschii Coss.: implications for intraspecific lineage diversification and evolution of common wheat. Mol Ecol 19:999–1013
Mujeeb-Kazi A, Rosas V, Roldan S (1996) Conservation of the genetic variation of Triticum tauschii (Coss.) Schmalh. (Aegilops squarrosa auct. non L.) in synthetic hexaploid wheats (T. turgidum L. s. lat. × T. tauschii; 2n = 6× = 42, AABBDD) and its potential utilization for wheat improvement. Genet Resour Crop Evol 43:129–134
Mujeeb-Kazi A, Cano S, Rosas V, Cortes A, Delgado R (2001a) Registration of five synthetic hexaploid wheat and seven bread wheat lines resistant to wheat spot blotch. Crop Sci 41:1653–1654
Mujeeb-Kazi A, Fuentes-Davila G, Villareal RL, Cortes A, Roasas V, Delgado R (2001b) Registration of 10 synthetic hexaploid wheat and six bread wheat germplasms resistant to karnal bunt. Crop Sci 41:1652–1653
Nagy ED, Bennetzen JL (2008) Pathogen corruption and site-directed recombination at a plant disease resistance gene cluster. Genome Res 18:1918–1923
Nesbitt M, Samuel D (1998) Wheat domestication: archaeobotanical evidence. Science 279:1431
Ortiz R, Braun HJ, Crossa J, Crouch JH, Davenport G, Dixon J, Dreisigacker S, Duveiller E, He ZH, Huerta J, Joshi AK, Kishii M, Kosina P, Manes Y, Mezzalama M, Morgounov A, Murakami J, Nicol J, Ferrara GO, Ortiz-Monasterio JI, Payne TS, Pena RJ, Reynolds MP, Sayre KD, Sharma RC, Singh RP, Wang JK, Warburton M, Wu HX, Iwanaga M (2008) Wheat genetic resources enhancement by the International Maize and Wheat Improvement Center (CIMMYT). Genet Resour Crop Evol 55:1095–1140
Paveley ND, Lockley KD, SylvesterBradley R, Thomas J (1997) Determinants of fungicide spray decisions for wheat. Pest Sci 49:379–388
Raman H, Stodart BJ, Cavanagh C, Mackay M, Morell M, Milgate A, Martin P (2010) Molecular diversity and genetic structure of modern and traditional landrace cultivars of wheat (Triticum aestivum L.). Crop Pasture Sci 61:222–229
Reddy L, Friesen T, Meinhardt S, Chao S, Faris J (2008) Genomic analysis of the Snn1 locus on wheat chromosome arm 1BS and the identification of candidate genes. Plant Genome 1:55
Risser P, Ebmeyer E, Korzun V, Hartl L, Miedaner T (2011) Quantitative-trait loci for adult plant resistance to Mycosphaerella graminicola in two winter wheat populations. Phytopathology. doi: 10.1094/phto-08-10-0203
Roder MS, Korzun V, Wendehake K, Plaschke J, Tixier MH, Leroy P, Ganal MW (1998) A microsatellite map of wheat. Genetics 149:2007–2023
Rosielle AA (1972) Sources of resistance in wheat to speckled leaf blotch caused by septoria-tritici. Euphytica 21:152–161
Simon MR, Ayala FM, Cordo CA, Röder MS, Börner A (2004) Molecular mapping of quantitative trait loci determining resistance to septoria tritici blotch caused by Mycosphaerella graminicola in wheat. Euphytica 138:41–48
Simon MR, Khlestkina EK, Castillo NS, Börner A (2010) Mapping quantitative resistance to septoria tritici blotch in spelt wheat. Eur J Plant Pathol 128:317–324
Singh D, Park RF, McIntosh RA (2007) Characterisation of wheat leaf rust resistance gene Lr34 in Australian wheats using components of resistance and the linked molecular marker csLV34. Aust J Agric Res 58:1106–1114
Somasco OA, Qualset CO, Gilchrist DG (1996) Single-gene resistance to Septoria tritici blotch in the spring wheat cultivar ‘Tadinia’. Plant Breed 115:261–267
Somers DJ, Isaac P, Edwards K (2004) A high-density microsatellite consensus map for bread wheat (Triticum aestivum L.). Theor Appl Genet 109:1105–1114
Song QJ, Shi JR, Singh S, Fickus EW, Costa JM, Lewis J, Gill BS, Ward R, Cregan PB (2005) Development and mapping of microsatellite (SSR) markers in wheat. Theor Appl Genet 110:550–560
Sotelo P, Starkey S, Voothuluru P, Wilde GE, Smith CM (2009) Resistance to Russian wheat aphid biotype 2 in CIMMYT synthetic hexaploid wheat lines. J Econ Entomol 102:1255–1261
Sourdille P, Singh S, Cadalen T, Brown-Guedira G, Gay G, Qi L, Gill B, Dufour P, Murigneux A, Bernard M (2004) Microsatellite-based deletion bin system for the establishment of genetic-physical map relationships in wheat (Triticum aestivum L.). Funct Integr Genomics 4:12–25
Stergiopoulos I, van Nistelrooy JGM, Kema GHJ, De Waard MA (2003) Multiple mechanisms account for variation in base-line sensitivity to azole fungicides in field isolates of Mycosphaerella graminicola. Pest Manag Sci 59:1333–1343
Tabib Ghaffary SM, Robert O, Laurent V, Lonnet P, Margalé E, van der Lee TAJ, Visser RGF, Kema GHJ (2011) Genetic analysis of resistance to septoria tritici blotch in the French winter wheat cultivars Balance and Apache. Theor Appl Genet. doi:10.1007/s00122-011-1623-7
Torada A, Koike M, Mochida K, Ogihara Y (2006) SSR-based linkage map with new markers using an intraspecific population of common wheat. Theor Appl Genet 112:1042–1051
USDA (2010) http://usda.mannlib.cornell.edu/usda/ers/WHS//2010s/2010/WHS-06-14-2010.pdf
van Ginkel M, Ogbonnaya F (2007) Novel genetic diversity from synthetic wheats in breeding cultivars for changing production conditions. Field Crop Res 104:86–94
Warburton ML, Crossa J, Franco J, Kazi M, Trethowan R, Rajaram S, Pfeiffer W, Zhang P, Dreisigacker S, van Ginkel M (2006) Bringing wild relatives back into the family: recovering genetic diversity in CIMMYT improved wheat germplasm. Euphytica 149:289–301
Ware SB (2006) Aspect of sexual reproduction in M. graminicola species on wheat and barley, genetic studies on sepecificity, mapping and fungicide resistance. PhD thesis, Wageningen Universisty, The Netherlands
Wittenberg AHJ, van der Lee TAJ, Ben M’Barek S, Ware SB, Goodwin SB et al (2009) Meiosis drives extraordinary genome plasticity in the haploid fungal plant pathogen Mycosphaerella graminicola. PLoS ONE 4(6):e5863. doi:10.1371/journal.pone.0005863
Xie WL, Nevo E (2008) Wild emmer: genetic resources, gene mapping and potential for wheat improvement. Euphytica 164:603–614
Xu SS, Friesen TL, Mujeeb-Kazi A (2004) Seedling resistance to tan spot and Stagonospora nodorum blotch in synthetic hexaploid wheats. Crop Sci 44:2238–2245
Xu SS, Cai X, Wang T, Harris MO, Friesen TL (2006) Registration of two synthetic hexaploid wheat germplasms resistant to hessian fly. Crop Sci 46:1401–1402
Yang WY, Liu DC, Li J, Zhang LQ, Wei HT, Hu XR, Zheng YL, He ZH, Zou YC (2009) Synthetic hexaploid wheat and its utilization for wheat genetic improvement in China. J Genet Genomics 36:539–546
Yechilevich-Auster M, Levi E, Eyal Z (1983) Assessment of interactions between cultivated and wild wheats and septoria tritici. Phytopathology 73:1077–1083
Zadoks JC, Chang TT, Konzak CF (1974) Decimal code for growth stages of cereals. Weed Res 14:415–421
Zhan J, Mundt CC, McDonald BA (2007) Sexual reproduction facilitates the adaptation of parasites to antagonistic host environments: evidence from empirical study in the wheat-Mycosphaerella graminicola system. Int J Parasitol 37:861–870
Zhang X, Haley SD, Jin Y (2001) Inheritance of Septoria tritici blotch resistance in winter wheat. Crop Sci 41:323–326
Zhang P, Dreisigacker S, Buerkert A, Alkhanjari S, Melchinger AE, Warburton ML (2006) Genetic diversity and relationships of wheat landraces from Oman investigated with SSR markers. Genet Resour Crop Evol 53:1351–1360
Zhang ZC, Friesen TL, Simons KJ, Xu SS, Faris JD (2009) Development, identification, and validation of markers for marker-assisted selection against the Stagonospora nodorum toxin sensitivity genes Tsn1 and Snn2 in wheat. Mol Breed 23:35–49
Zwart R, Thompson J, Milgate A, Bansal U, Williamson P, Raman H, Bariana H (2010) QTL mapping of multiple foliar disease and root-lesion nematode resistances in wheat. Mol Breed 26:107–124
Acknowledgments
This project is financially supported by the Fonds de Soutien à l’Obtention Végétale (FSOV), France and by the USDA-Agricultural Research Service CRIS project 5442-22000-033-00D. S.M. Tabib Ghaffary acknowledges partial financial support from the Agricultural Research and Education Organization (AREO) of Iran. We are grateful to Drs. Maarten van Ginkel (ICARDA), Abbas Keshavarz (Former Head of AREO-SPII) and Mohammad Reza Jalal Kamali (CIMMYT-Iran) for their role in initiating this project. We thank our colleagues Pieter Vereijken and Jacques Withage for statistical support and Els Verstappen for excellent M. graminicola maintenance and greenhouse management.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by B. Keller.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Tabib Ghaffary, S.M., Faris, J.D., Friesen, T.L. et al. New broad-spectrum resistance to septoria tritici blotch derived from synthetic hexaploid wheat. Theor Appl Genet 124, 125–142 (2012). https://doi.org/10.1007/s00122-011-1692-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-011-1692-7