Abstract
Key message
One hundred and thirty four introgressions from Thinopyrum elongatum have been transferred into a wheat background and were characterised using 263 SNP markers.
Abstract
Species within the genus Thinopyrum have been shown to carry genetic variation for a very wide range of traits including biotic and abiotic stresses and quality. Research has shown that one of the species within this genus, Th. elongatum, has a close relationship with the genomes of wheat making it a highly suitable candidate to expand the gene pool of wheat. Homoeologous recombination, in the absence of the Ph1 gene, has been exploited to transfer an estimated 134 introgressions from Th. elongatum into a hexaploid wheat background. The introgressions were detected and characterised using 263 single nucleotide polymorphism markers from a 35 K Axiom® Wheat-Relative Genotyping Array, spread across seven linkage groups and validated using genomic in situ hybridisation. The genetic map had a total length of 187.8 cM and the average chromosome length was 26.8 cM. Comparative analyses of the genetic map of Th. elongatum and the physical map of hexaploid wheat confirmed previous work that indicated good synteny at the macro-level, although Th. elongatum does not contain the 4A/5A/7B translocation found in wheat.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Modern hexaploid wheat (Triticum aestivum) is a staple food crop that contributes ~ 20% of global daily dietary calories (Reynolds et al. 2012). By 2050, the global population is predicted to exceed 9.7 billion, and global wheat demand is predicted to reach 900 million tonnes (Charmet 2011; FAOSTAT 2018). However, current global wheat production is only 749 million tonnes (achieved in 2016) (FAOSTAT 2017). The annual yield percentage increase, which for the last decade has averaged 0.9% globally, will need to increase to ~ 2.4%. In Europe, however, yield increases have steadily plateaued to just 0.1% (Ray et al. 2013).
Wheat, which evolved circa 8000 to 10,000 years ago, has been through a significant genetic bottle neck due to its monophyletic or diphyletic evolution and subsequent domestication (Shewry 2009). Intensive selection pressure applied by centuries of farming has also eroded the variation within the wheat gene pool, reducing the level of genetic variation available in breeding programmes for the production of elite cultivars. Emerging diseases and climate change are further impacting wheat yields (Lobell et al. 2011; Curtis and Halford 2014; Price et al. 2014). It is thus essential to increase the genetic diversity available for breeders for producing new elite cultivars of wheat that are climate change ready (Dempewolf et al. 2014).
The wild relatives of wheat represent a vast and underutilised source of genetic variation for virtually all agronomic traits of interest. Interspecific crossing with wheat’s wild relatives has repeatedly been shown to successfully transfer traits of interest (e.g. Ayala-Navarrete et al. 2007; Riar et al. 2012; Ceoloni et al. 2017).
Th. elongatum belongs to the genus Thinopyrum, which was segregated from the much larger genus Elytrigia in the late twentieth century, alongside the genera Lophopyrum and Trichopryum (Baum and Johnson 2018). Diploid (2n = 2x = 14:EE), tetraploid (2n = 4x = 28), hexaploid (2n = 6x = 42) and decaploid accessions (2n = 10x = 70) of Th. elongatum have been identified (Chen et al. 2013; Guo et al. 2016; Mao et al. 2010). However, the literature contains considerable confusion in distinguishing between decaploid Th. elongatum and another of the decaploid Thinopyrum species, Th. ponticum. Previously, these two species were frequently placed under the same name, Agropyrum elongatum (Shepherd and Islam 1988; Li et al. 2017). Th. ponticum has been described with three diverging genomic constitutions: EbEbEbEbEbEbEbEbEbEb (Arterburn et al. 2011), JJJJJJJSJSJSJS (Chen et al. 1998) and EeEeEbEbExExStStStSt (Zhang et al. 1996), where the Eb/Ee/Ex/J genome is derived from the closely related genomes Th. bessarabicum/Th. elongatum (diploid) and the St genome from Pseudoroegneria strigosa. Research has shown a range of Thinopyrum species carry genetic variation for a range of agronomically important traits including salinity tolerance (Dvorák et al. 1988; Colmer et al. 2006), perennial growth habit (Lammer et al. 2004), water logging tolerance (Taeb et al. 1993), improved photosynthetic capacity (Reynolds et al. 2001), resistance to a wide range of diseases (Friebe et al. 1996; Zhang et al. 2009; Fu et al. 2012; Li et al. 2017) and improved flour quality (Tanaka et al. 2017). Research has also shown a close relationship between the genomes of wheat and Th. elongatum, suggesting it is a highly suitable candidate to expand the gene pool of wheat (Liu et al. 2007).
This paper describes the identification and characterisation of a series of introgression lines generated between hexaploid wheat and Th. elongatum using genomic in situ hybridisation (GISH) and an Axiom® Wheat-Relative SNP Genotyping Array.
Materials and methods
Generation of introgressions
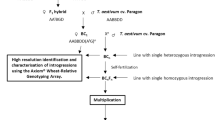
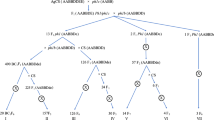
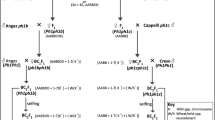
Th. elongatum, accession 401007 (2n = 10x = 70 (+4), seed obtained from the United States Department of Agriculture) was used to pollinate hexaploid wheat ph1/ph1 mutant (cv. Chinese Spring) (Fig. 1). The resulting F1 hybrids were then pollinated using wheat Ph1/Ph1 (cv. Paragon) to produce backcrossed lines (BC1 generation). Both of the wheat genotypes were obtained from the Germplasm Resource Unit (GRU), John Innes Centre. Further rounds of backcrossing using Ph1/Ph1 wheat as the pollen donor produced BC2, BC3, BC4 and BC5 populations with self-fertilised lines also produced at each generation after the BC2 generation (Fig. 1).
An overview of the wheat/Th. elongatum breeding programme showing the production of each generation via crossing (whole arrow) or via self-fertilisation (dashed arrow). Red boxes represent lines produced from self-fertilisation of the previous generation. Blue boxes represent lines produced via crossing
Detection and characterisation of introgressions
Genotyping and genetic map construction
Three hundred and thirty individuals from the BC1 to BC5 backcross generations, 17 Chinese Spring-Th. elongatum addition lines (obtained from the GRU) and three replicates of each parental line were genotyped using the Axiom® Wheat-Relative Genotyping Array as described in King et al. (2017). All SNPs incorporated in the array show polymorphisms between wheat and ten wild relatives and were selected from the Axiom® 820 K array (Wilkinson et al. 2012, 2016; Winfield et al. 2016). Allele calling was carried out as described by King et al. (2017). SNP markers from the categories Poly High Resolution (PHR) and Call Rate Below Threshold (CRBT) were selected for further analysis. Markers were only used in further analysis if polymorphic and co-dominant between the three replicates of wheat and Th. elongatum. Flapjack™ (Milne et al. 2010) was used to visualise the genotypes, and markers were removed from further analysis if they showed no call, were heterozygous for either parent or if they showed no polymorphism between the parents. Remaining markers were then analysed in JoinMap® 4.0 (van Ooijen 2011) using the Haldane map function (Haldane 1919), a LOD score of 20 and a recombination frequency of 0.1. The seven highest ranking linkage groups were assigned to a chromosome group using chromosome locations for each marker described in the Axiom® Wheat HD Genotyping Array (Winfield et al. 2016). Markers showing more than 20% erroneous calls were removed and markers at the same genetic position were ordered by their physical positions on the wheat reference genome, found by performing a BLAST analysis of the marker sequences against the wheat reference sequence RefSeq v1.0 (Alaux et al. 2018; International Wheat Genome Sequencing Consortium [IWGSC] et al. 2018) and obtaining the best BLAST hit from each of the three genomes of wheat, where available. Linkage group data was used to produce a genetic map using MapChart 2.3 (Voorrips 2002) (Fig. 2) and genotypes for individual lines were visualised using Graphical GenoTypes 2.0 (GGT; van Berloo 2008) (Figs. 3, 4).
Genetic linkage map of Th. elongatum showing the 263 Affymetrix SNP markers spread across all seven linkage groups. SNP marker names and calculated cM distances for each group are shown. Map created using MapChart 2.3 (Voorrips 2002)
A progression of GGT genotypes (red represents Th. elongatum chromatin and blue represents wheat chromatin) for a BC3 parent, one of its BC4 offspring and a subsequent BC5 offspring. The GISH image shows a metaphase spread of BC5-26A and shows the introgression detected via the SNP markers (white arrow). The SNP markers enable the introgression to be identified as a linkage group 5E segment
GGT genotypes for a BC3 plant and three BC4 offspring, all confirmed using GISH below their respective genotype and showing the segregation of segments in a family as the breeding programme progresses. In the GGT images, red represents Th. elongatum chromatin and blue represents wheat chromatin. In the GISH images, all introgressions are indicated using a white arrow, whole Th. elongatum chromosomes are indicated using a red arrow
Synteny analysis
Synteny between wheat and Th. elongatum was analysed using the 263 SNP markers selected for the genotyping analysis. The sequences of the mapped markers were used in a BLAST search (e-value cut-off of 1e−05) against the wheat genome IWGSC RefSeq v1.0 (Alaux et al. 2018; IWGSC et al. 2018) to obtain the corresponding physical positions of the top hit in the A, B and D genomes of wheat Supplementary Table S1). The results were visualised using Circos (v. 0.67; Krzywinski et al. 2009) to show synteny between the genetic position in cM for Th. elongatum and the corresponding physical positions on the D genome of wheat (Fig. 5). Some markers showed the same score for the top hit for more than one genome.
Graphical representation made using Circos showing the introgression sizes in panel plants from the wheat/Th. elongatum breeding programme. Each band in each linkage group represents a different individual line. Lines selected may contain whole chromosomes in other linkage groups. A total of 48 plants make up the panel
Cytogenetic analysis
Root tips, collected from germinated seeds, were treated with nitrous oxide gas at a pressure of 10 bar for 2 h, fixed in 90% acetic acid, washed with water and then digested in 20 μl of 1% pectolyase Y23 and 2% cellulose Onozuka R-10 (Yakult Pharmaceutical, Tokyo) enzyme solution at 37 °C for 55 min (adapted from Kato et al. 2004). Samples were crushed in 70% ethanol and the cells collected via centrifugation at 2.5 g for 1 min before being briefly dried and re-suspended on ice in 25 μl of 100% acetic acid. Cell suspensions were dropped onto glass slides (7 μl per slide).
Single colour genomic in situ hybridisation (sc-GISH) was carried out as described in (King et al. 2017 and 2018; Grewal et al. 2018a) using total genomic DNA from Th. elongatum labelled by nick translation with Chroma Tide Alexa Fluor 488-5-dUTP (Invitrogen; C11397). Genomic DNA from T. aestivum (cv. Chinese Spring) was fragmented to 300-500 bp and used as blocking DNA in a ratio of 1:20 probe: blocking DNA.
For multi-colour GISH (mc-GISH) of wheat/Th. elongatum introgression lines, total genomic DNAs from Triticum urartu, Aegilops tauschii and Th. elongatum were labelled by nick translation with Chroma Tide Alexa Fluor 488-5-dUTP (Invitrogen; C11397), Chroma Tide Alexa Fluor 594-5-dUTP (Invitogen; C11400) and Chroma Tide Alexa Fluor 546-14-dUTP (Invitrogen; C11401), respectively. Slides selected from the sc-GISH analysis were then re-probed with labelled DNA from T. urartu (100 ng), Th. elongatum (100 ng), Ae. tauschii (200 ng) and fragmented DNA of Ae. speltoides (4000 ng) as blocking DNA in a ratio of 1:1:2:40.
Multi-colour GISH was also carried out on Th. elongatum accession 401007. Total genomic DNAs from Th. bessarabicum and P. strigosa were labelled by nick translation with Chroma Tide Alexa Fluor 488-5-dUTP and Chroma Tide Alexa Fluor 594-5-dUTP, respectively. Slides were probed with labelled DNA from Th. bessarabicum (100 ng), P. strigosa (100 ng) and genomic DNA of Chinese Spring (4000 ng), fragmented to 300-500 bp, as blocking DNA in a ratio of 1:1:40.
All sc-GISH and mc-GISH slides were counterstained with Vectashield mounting medium containing 4′,6-diamidino-2-phenylindole,dihydrochloride (DAPI) and analysed using a Zeiss Axio Imager.Z2 upright epifluorescence microscope (Carl Zeiss Ltd, Germany) with filters for DAPI and Alexa Fluor 488, Alexa Fluor 594 and Alexa Fluor 546. A Metasystems Coolcube 1 m CCD camera was used to capture images which were analysed using Metafer (automated metaphase image capture) and ISIS (image processing) software (Metasystems GmbH, Germany).
Results
GISH analysis of Thinopyrum elongatum
The decaploid accession used in this study (accession 401007) carried 74 chromosomes (Fig. 6). When the parental Th. elongatum was analysed with mc-GISH, large blocks of the St genome were observed at the centromeres of 32 chromosomes, while the remaining 42 chromosomes either did not carry any DNA from the St genome or a very faint non-centromric St genome fluorescence (Fig. 6).
Production of wheat/Th. elongatum introgression lines
The number of seeds sown, germination success, heads crossed, cross-fertility and seed set for each generation is summarised in Table 1. In total, 29 crosses were made between Th. elongatum and wheat homozygous for ph1 deletion resulting in the production of 222 F1 seed. The lowest germination rate was shown by the F1 seeds (57%) while the highest rate was shown by the BC1 (91%). In contrast, the BC1 showed the lowest cross-fertility (56%) and the lowest number of seeds set per cross (2.5) while the highest fertility was shown by the F1 (88%).
Genotyping and genetic map construction
Initial genotyping analysis identified 1594 polymorphic markers between wheat and Th. elongatum from the 35 K Axiom® Wheat-Relative Genotyping Array that were either PHR or CRBT SNP markers with good cluster resolution. Sample call rate ranged from 83.7 to 99.6% with an average of 98.3% for the 356 samples genotyped (330 backcross lines, 17 addition lines and 9 parental replicates) and the lowest call rates were obtained for the three Th. elongatum samples with an average of 87.3%. A total of 497 erroneous markers were removed after analysis with Flapjack™ and a further 834 markers were removed as they showed unique or inconsistent patterns of segregation. The remaining 263 SNP markers were genetically mapped to seven linkage groups using JoinMap® 4.0 (Table 2) with the highest number of SNP markers mapping to linkage group 1 (26%) and the lowest to linkage group 5 (7%). The genetic linkage map of Th. elongatum (Fig. 2) had a total map length of 187.8 cM and an average chromosome length of 26.8 cM. From the genetic linkage maps, an estimated 134 wheat/Th. elongatum introgressions were generated. Plants from different generations were used for the genotyping, and thus, while the linkage map allowed the identification and characterisation of the introgressed segments and tracking through the backcross generations (Fig. 3), the cM distances need to be treated with considerable caution.
Detection of introgressions using GISH
Lines analysed by sc-GISH confirmed the presence of high numbers of both whole Th. elongatum chromosomes and wheat/Th. elongatum introgressions revealed by genotyping (Fig. 7). Thus, the sc-GISH was also used to validate the introgressions identified by genotyping. In the lines analysed with both genotyping and GISH, the number of whole chromosomes and introgressions detected was the same (Figs. 3 and 4). Mc-GISH showed that recombination had taken place between Th. elongatum and all three genomes of wheat (Fig. 8) (E with A = 13, E with B = 11 and E with D = 4) and that the recombination events were not localised in distal regions of the chromosomes (Fig. 8).
GISH images from lines from different families in the breeding programme. a BC4-118E, b BC4F1-101D, c BC4-183A, d BC4-172C, e BC4F1-90D and f BC3-670C. Th. elongatum chromatin is shown as green and wheat chromatin is shown as blue. All introgressions are indicated using a red arrow and whole Th. elongatum chromosomes are indicated using a white arrow
Mc-GISH results showing 11 interspecific chromosomes with the recombination events at different locations. In all images, the yellow represents Th. elongatum chromatin (E genome). In the A-E grouping, the green represents T. urartu chromatin (A genome), in the B-E grouping, the blue represents Ae. speltoides chromatin (blue) and in the D-E grouping, the red represents Ae. tauschii (D genome)
Synteny
Figure 5 shows the syntenic relationship between the D genome of wheat and the E genome of Th. elongatum. The BLAST results of the SNP markers showed that 89.4%, 92.4% and 94.5% of the markers had a significant BLAST hit on the A, B and D genomes of wheat, respectively, with 25.9%, 41.1% and 39.9% of the markers having an overall top hit on the A, B and D genomes of wheat, respectively (Supplementary Table S1). The D genome was thus selected for syntenic analysis as it was shown to have the highest number of significant BLAST hits for the markers on the Th. elongatum genetic map (Fig. 2).
Discussion
The genus Thinopyrum has the potential to significantly increase the level of genetic variation available for wheat improvement., e.g. salt tolerance (King et al. 1997; Colmer et al. 2006), Fusarium head blight (Oliver et al. 2006; Ceoloni et al. 2017), increased biomass and seed number (Reynolds et al. 2001). Much of the work previously reported on Th. elongatum has been carried out on the diploid (2x = 2n = 14; EE). However, the species contains accessions at different ploidy levels including decaploid (Chen et al. 2013; Guo et al. 2016; Mao et al. 2010). Mc-GISH observations of the Th. elongatum 401007 accession indicate that the genome is composed of DNA sequences derived from at least two genomes, i.e. Th. bessarabicum and P. strigosa, and that considerable recombination/translocation has occurred between the two genomes in the past. It would also suggest that the correct genome composition of the accession under study here (401007) is EEEEEEEStEStEStESt or JJJJJJJStJStJStJSt. This configuration was also found in Th. ponticum by Kruppa and Molnár-Lang (2016) again suggesting a close relationship between Th. elongatum 401007 and Th. ponticum.
In this work, introgressions were generated in the gametes of interspecific wheat/Th. elongatum hybrids which lacked the wild type Ph1 locus, which normally restricts recombination to homologous chromosomes. In addition, the F1 hybrids only carried the haploid chromosome complement of the genomes of wheat and Th. elongatum. Thus, the A, B and D genome chromosomes could only undergo homoeologous recombination, i.e. with homoeologous wheat or Th. elongatum chromosomes. This strategy was employed in a direct attempt to increase the frequency of homoeologous recombination and hence introgression in the gametes of the F1 hybrids.
The germination rate (Table 1) of the F1 interspecific hybrids (57%), although the lowest of the generations in this crossing programme, was higher than that seen in F1 interspecific hybrids between wheat and other wild relatives, e.g. Amblyopyrum muticum (28.6%) and Ae. speltoides (15%) (King et al. 2017, 2018).
The rate of fertility of the F1 hybrids was also surprisingly high at 88% (Table 1). In crossing programmes with other wild relatives, the fertility of the F1 interspecific hybrids has been observed to be considerably lower, e.g. 1.6% for Th. bessarabicum, 16% with Am. muticum, 21% with T. urartu and 29% with Ae. speltoides (Grewal et al. 2018a, b; King et al. 2017, 2018). In our hands, the low fertility of F1 hybrids, although they frequently result in the generation of high numbers of introgressions, remains the rate limiting step with many of the wild relatives of wheat. The lower rate of fertility normally observed in wheat/wild relative F1 hybrids presumably results from the fact that they are essentially haploid for the genomes of wheat and those of the wild relative resulting in substantial chromosome pairing failure at meiosis leading to the generation of unbalanced gametes which as a result are often unviable.
The higher rate of fertility in the wheat/Th. elongatum F1 hybrids might be attributable to the close synteny between the E and Est genomes of Th. elongatum and also between these genomes and the D genome of wheat (Liu et al. 2007; Wang, 2011; Hu et al. 2012), resulting in a level of chromosome pairing during meiosis that reduces the frequency of unbalanced gametes. This could also explain why the F1 interspecific hybrids between wheat and Th. elongatum show male fertility. In contrast, F1 hybrids between wheat and Am muticum (King et al. 2017), Ae. speltoides (King et al. 2018), Th. bessarabicum (Grewal et al. 2018a), T. urartu (Grewal et al. 2018b) and T. timopheevii (Devi et al. 2019) all show complete male sterility. Some of the GISH results obtained here, however, are not fully supportive of the D genome similarity (see below). The fertility rate then dropped in the BC1 generation.
In this work, we have utilised the Axiom® Wheat-Relative Genotyping Array for the characterisation of the wheat/Th. elongatum introgression lines. This array was designed to contain SNPs polymorphic between wheat and all ten wild relatives under study at the Nottingham BBSRC Wheat Research Centre and thus allowed the genotyping of introgression lines from different wild relatives on the same 384 array. However, many other markers have been designed to characterise Th. elongatum introgressions in a wheat background including microsatellites (SSRs), expressed sequence tags (ESTs), PCR-based landmark unique genes (PLUGs), sequence-characterised amplified regions (SCARs), conserved orthologous set (COS) markers and SNPs (Hu et al. 2012; Chen et al. 2015; Dong et al. 2017; Gaál et al. 2018).
The level of interspecific recombination in the gametes of the F1 hybrids was such that it was possible to generate 7 discrete genetic linkage groups of Th. elongatum (Fig. 2), from which it could be estimated that 134 introgressions, covering the whole genome of Th. elongatum, had been generated. (There was no evidence from the genetic mapping or GISH of any further recombination in later generations.) The genetic map also allowed the characterisation and tracking of the introgressions through the backcross generations (Figs. 3, 4). In order to validate the genotyping data obtained, lines were also analysed via GISH. In each case, the presence, number and size of introgressions predicted via genotyping was confirmed via GISH. The production of only 7 linkage groups would be considered an unexpected observation from an allopolyploid, i.e. 2n = 10x = 70 + 4; EEEEEEEStEStEStESt. However, this observation could be explained by the high level of recombination/translocation that has occurred between the two genomes of Th. bessarabicum and P. strigosa in the past. Alternatively, the assembly of only 7 linkage groups may have resulted from the relatively low numbers of Th. elongatum polymorphic SNPs generated that could be used to identify introgressions as compared to similar work undertaken in other species, e.g. (King et al. 2017, 2018; Grewal et al. 2018a, b). The generation of a SNP between a wild relative and wheat requires that a base change is identified in the genome(s) of the species in question relative to the equivalent sequences in the three genomes of wheat. With diploid wild relatives, the identification of polymorphic SNPs is a relatively straight forward process, i.e. a single change in the genome of the wild relative relative to wheat is required. However, a polymorphic SNP between an allopolyploid species and wheat requires that each of the genomes of the wild relative carry the same polymorphism relative to the three genomes of wheat. As a result, the attrition rate for identifying polymorphic SNPs is much greater in allopolyploid wild relatives.
GISH has validated the genotyping work and significant numbers of introgressions have been indentified and characterised (Fig. 7). However, we are not presently able to determine whether we lack SNPs for parts of the genome. If this is the case, then we would not have been able to detect introgressions from these regions of the genome.
Previous results have suggested that the E genome shares the greatest level of homology with the D genome of wheat (Liu et al. 2007; Wang 2011; Hu et al. 2012). This was also suggested by the results of the synteny analysis in this work where the D genome was shown to have the highest number of significant BLASTS hits for the markers on the Th. elongatum genetic map (Fig. 2). It was therefore unexpected to find an almost equal number of recombination events between the E genome and the A and B genomes and the least number with the D genome (Fig. 8). The number of lines looked at with mc-GISH, however, was relatively small and thus more introgressions need to be studied. Much of the previous work has also been done on diploid accessions of Th. elongatum and thus greater clarity is required as to the genome composition of the higher ploidy levels. Good overall synteny is maintained between wheat and Th. elongatum, showing the close relationship of the two genomes. However, Th. elongatum does not carry a reciprocal 4A/5A/7B translocation that is observed in the A and B genomes of wheat (Devos et al. 1995).
Graphical representation made using Circos showing the introgression sizes in panel plants from the wheat/Th. elongatum breeding programme. Each band in each linkage group represents a different individual line. Lines selected may contain whole chromosomes in other linkage groups. A total of 48 plants make up the panel
Most recombination events in wheat occur towards the distal regions of the chromosomes with large blocks in the pericentric regions experiencing only very low levels of recombination (Lukaszewski and Curtis 1993; Akhunov et al. 2003). However, in contrast to wheat, recombination in the interspecific wheat/Th. elongatum hybrids was not restricted to the distal regions of chromosomes, i.e. recombination was observed in both the distal and proximal regions of chromosomes (Fig. 8). It will be interesting to compare the recombination observed in the wheat/Th. elongatum hybrids with hybrids between wheat and other wild relatives, i.e. there is some evidence that mainly distal recombination occurs between wheat and Haynaldia villosa group 4 chromosome (Dai et al. 2020). A direct consequence of the localisation of chiasma to the distal regions of chromosomes in wheat is that genes located in the proximal regions of chromosomes will be inherited as unrecombined blocks, i.e. the generation of new allelic combinations will not occur in the proximal regions of chromosomes. Thus, the development of new allelic combinations for use in developing superior wheat varieties in breeding programmes will be limited to the genes located in the distal regions of chromosomes. Therefore, in order to generate new allelic combinations, a major focus of fundamental research in wheat is to shift recombination to the proximal regions of chromosomes. Thus, the observations described here demonstrate that the use of the wheat wild relative germplasm will provide an important means in manipulating and understanding the fundamental process of the position of recombination in wheat.
At present, work is underway using molecular markers complemented with cytogenetic analyses to select plants with single introgressions that represent the entire genome of Th. elongatum in overlapping segments (Fig. 9). After further rounds of self-fertilisation, a final panel will be composed of lines homozygous for different single introgressions and made available upon request.
References
Akhunov ED, Goodyear AW, Geng S, Qi L-L, Echalier B, Gill BS, Miftahudin Gustafson JP, Lazo G, Chao S, Anderson OD, Linkiewicz AM, Dubcovsky J, La Rota M, Sorrells ME, Zhang D, Nguyen HT, Kalavacharia V, Hossain K, Kianian SF, Peng J, Lapitan NLV, Gonzalez-Hernandez JL, Anderson JA, Choi D-W, Close TJ, Dilbirligi M, Gill KS, Walker-Simmons MK, Steber C, McGuire PE, Qualset CO, Dvorak J (2003) The organization and rate of evolution of wheat genomes are correlated with recombination rates along chromosome arms. Genome Res 13:753–763
Alaux M, Rogers J, Letellier T, Flores R, Alfama F, Pommier C, Mohellibi N, Durand S, Kimmel E, Michotey C, Guerche C, Loaec M, Lainé M, Steinback D, Choulet F, Rimbert H, Leroy P, Guilhot N, Salse J, Feuillet C, Paux E, Eversole K, Adam-Blondon A-F, Quesneville H, International Wheat Genome Sequencing Consortium (2018) Linking the International Wheat Genome Sequencing Consortium bread wheat reference genome sequence to wheat genetic and phenomic data. Genome Biol 19:111
Arterburn M, Kleinhofs A, Murray T, Jones S (2011) Polymorphic nuclear gene sequences indaicate a novel genome donor in the polyploid genus Thinopyrum. Hereditas 148:8–27
Ayala-Navarrete L, Bariana HS, Singh RP, Gibson JM, Mechanicos AA, Larkin PJ (2007) Trigenomic chromosomes by recombination of Thinopyrum intermedium and Th. ponticum translocations in wheat. Theor Appl Genet 116:63–75
Baum BR, Johnson DA (2018) Lophopyrum Á Löve (1980), Thinopyrum Á Löve (1980) and Trichopyrum Á Löve (1986): one, two or three genera? A study based on the nuclear 5S DNA. Genet Resour Crop Ev 65:1–26
Ceoloni C, Forte P, Kuzmanović L, Tundo S, Moscetti I, De Vita P, Virili ME, D’Ovidio R (2017) Cytogenetic mapping of a major locus for resistance to Fusarium headblight and crown rot of wheat on Thinopyrum elongatum 7EL and its pyramiding with valuable genes from a Th. ponticum homoeologous arm onto bread wheat 7DL. Theor Appl Genet 130:2005–2024. https://doi.org/10.1007/s00122-017-2939-8
Charmet G (2011) Wheat domestication: lessons for the future. CR Biol 334:212–220
Chen Q, Conner RL, Laroche A, Thomas JB (1998) Genome analysis of Thinopyrum intermedium and Thinopyrum ponticum using genomic in situ hybridization. Genome 41:580–586
Chen S, Huang Z, Dai Y, Qin Y, Zhang L, Gao Y, Chen J (2013) The development of 7E chromosome-specific molecular markers for Thinopyrum elongatum based on SLAF-seq technology. PLoS ONE 8:e65122
Chen S, Gao Y, Zhu X, Zhang C, Cao W, Fedak G, He Z, Chen X, Chen J (2015) Development of E-chromosome specific molecular markers for Thinopyrum elongatum in a wheat background. Crop Sci 55:2777–2785. https://doi.org/10.2135/cropsci2014.08.0539
Colmer TD, Flowers TJ, Munns R (2006) Use of wild relatives to improve salt tolerance in wheat. J Ex Bot 57:1059–1078
Curtis T, Halford NG (2014) The challenge of increasing wheat yield and the importance of not compromising food safety. Ann Appl Biol 164:354–372. https://doi.org/10.1111/aab.12108
Dai K, Zhao R, Shi M, Xiao J, Yu Z, Wang Z, Yuan C, Sun H, Cao A, Zhang R, Chen P, Li Y, Wang H, Wang X (2020) Dissection and cytological mapping of chromosome arm 4VS by the development of tructural aberration library. Theor Appl Genet 133:217–226. https://doi.org/10.1007/s00122-019-03452-8
Dempewolf H, Eastwood RJ, Guarino L, Khoury CK, Müller JV, Toll J (2014) Adapting agriculture to climate change A global initiative to collect, conserve and use crop wild relatives. Agroecol Sust Food 38:369–377. https://doi.org/10.1080/21683565.2013.870629
Devi U, Grewal S, Yang C, Hubbart-Edwards S, Scholefield D, Ashling S, Burridge A, King IP, King J (2019) Development and characterisation of interspecific hybrid lines with genome-wide introgressions from Triticum timopheevii in a hexaploid wheat background. BMC Plant Bio. https://doi.org/10.1186/s12870-019-1785-z
Devos KM, Dubcovsky J, Dvorák J, Chinoy CN, Gale MD (1995) Structural evolution of wheat chromosomes 4A, 5A and 7B and its impact on recombination. Theor Appl Genet 91:282–288
Dong L, Zhang K, Wang D, Zhao M, Li Y, Rong C, Qin H, Zhang A, Dong Z, Wang D (2017) High-throughput mining of E-genome-specific SNPs for characterizing Thinopyrum elongatum introgressions in common wheat. Mol Ecol Res 17:1318–1329. https://doi.org/10.1111/1755-0998.12659
Dvorák J, Edge M, Ross K (1988) On the evolution of the adaptation of Lophopyrum elongatum to growth in saline environments. Proc Natl Acad Sci USA 85:3805–3809
FAOSTAT (2017). FAO cereal supply and demand brief. http://www.fao.org/worldfoodsituation/csdb/en/. Accessed Dec 2017
FAOSTAT (2018) Predicted annual population, food and agriculture organisation of the United Nations Statistics Division
Friebe B, Jiang J, Raupp WJ, McIntosh RA, Gill BS (1996) Characterization of wheat-alien translocations conferring resistance to disease and pests: current status. Euphytica 91:59–87
Fu S, Zhenling I, Qi B, Guo X, Li J, Liu B, Han F (2012) Molecular cytogenetic characterization of wheat-Thinopyrum elongatum addition, substitution and translocation lines with a novel source of resistance to wheat Fusarium head blight. J Genet Genomics 39:103–110
Gaál E, Valárik M, Molnár I, Farkas A, Linc G (2018) Identification of COS markers for Thinopyrum elongatum chromosomes preliminary revealed high level of macrosyntenic relationship between the wheat and Th. elongatum genomes. PLoS ONE 13:e0208840. https://doi.org/10.1371/journal.pone.0208840
Grewal S, Yang C, Edwards S, Scolefield D, Ashling S, Burridge AJ, King IP, King J (2018a) Characterisation of Thinopyrum bessarabicum chromosomes through genome-wide introgressions into wheat. Theor Appl Genet 131:389–406. https://doi.org/10.1007/s00122-017-3009-y
Grewal S, Hubbart-Edwards S, Yang C, Scholefield D, Ashling S, Burridge A, Wilkinson PA, King IP, King J (2018b) Detection of T. urartu introgressions in wheat and development of a panel of interspecific introgression lines. Front Plant Sci 9:1565. https://doi.org/10.3389/fpls.2018.01565
Guo J, Yu X, Yin H, Liu G, Li A, Wang H, Kong L (2016) Phylogenetic relationships of Thinopyrum and Triticum species revealed by SCoT and CDDP markers. Plant Syst Evol 302:1301–1309
Haldane JBS (1919) The probable errors of calculated linkage values, and the most accurate method of determining gametic from certain zygotic series. J Hered 8:291–297
Hu L-J, Liu C, Zeng Z-X, Li G-R, Song X-J, Yang Z-J (2012) Genomic rearrangement between wheat and Thinopyrum elongatum revealed by mapped functional markers. Genes Genom 34:67–75
International Wheat Genome Sequencing Consortium [IWGSC], Appels R, Eversole K, Feuillet C, Keller B, Rojers J, Stein N et al (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:eaar7191
Kato A, Lamb JC, Birchler JA (2004) Chromosome painting using repetitive DNA sequences as probes for somatic chromsome identification in maize. Proc Natl Acad Sci USA 101:13554–13559
King IP, Forster BP, Law CC, Cant KA, Orford SE, Gorham J, Reader S, Miller TE (1997) Introgression of salt-tolerance genes from Thinopyrum bessarabicum into wheat. New Phytol 137:75–81
King J, Grewal S, Yang CY, Hubbart-Edwards S, Scholefield D, Ashling S, Edwards KJ, Allen AM, Burridge A, Bloor C, Davassi A, da Silva GJ, Chalmers K, King IP (2017) A step change in the transfer of interspecific variation into wheat from Amblyopyrum muticum. Plant Biotechnol J 15:217–226. https://doi.org/10.1111/pbi.12606
King J, Grewal S, Yang CY, Hubbart-Edwards S, Scholefield D, Ashling S, Harper JA, Allen AM, Edwards KJ, Burridge AJ, King IP (2018) Introgression of Aegilops speltoides segments in Triticum aestivum and the effect of the gametocidal genes. Ann Bot 121:229–240. https://doi.org/10.1093/aob/mcx149
Kruppa K, Molnár-Lang M (2016) Simultaneous visualization of different genomes (J, JSt and St) in a Thinopyrum intermedium x Thinopyrum ponticum synthetic hybrid (Poaceae) and in its parental species by multicolour genomic in situ hybridization (mcGISH). Comp Cytogenet 10:283–293
Krzywinski M, Schein J, Birol I, Connors J, Gascoyne R, Horsman D, Jones SJ, Marra MA (2009) Circos: an information aesthetic for comparative genomics. Genome Res 19:1639–1645
Lammer D, Cai X, Arterburn M, Chatelain J, Murray T, Jones S (2004) A single chromosome addition from Thinopyrum elongatum confers a polycarpic, perennial habit to annual wheat. J Exp Bot 55:1715–1720
Li X, Jiang X, Chen X, Song J, Ren C, Xiao Y, Gao X, Ru Z (2017) Molecular cytogenetic identification of a novel wheat-Agropyron elongatum chromosomes translocation line with powdery mildew resistance. PLoS ONE 12(9):e0184462
Liu Z, Li D, Zhang X (2007) Genetic relationships among five basic genomes St, E, A, B and D in triticeae revealed by genomic southern and in situ hybridization. J Integr Plant Biol 49:1080–1086
Lobell D, Schlenker W, Costa-Roberts J (2011) Climate trends and global crop production since 1980. Science 333:616–620
Lukaszewski AJ, Curtis CA (1993) Physical distribution of recombination in B-genome chromosomes of tetraploid wheat. Theor Appl Genet 86:121–127
Mao P, Huang Y, Wang X, Meng L, Mao P, Zhang G (2010) Cytological evaluation and karyotype analysis in plant germplasms of Elytrigia Desv. Agr Sci China 9:553–1560
Milne I, Shaw P, Stephen G, Bayer M, Cardle L, Thomas WTB, Flavell AJ, Marshall D (2010) Flapjack—graphical genotype visualisation. Bioinformatics 26:3133–3134
Oliver RE, Xu SS, Stack RW, Friesen TL, Jin Y, Cai X (2006) Molecular cytogenetic characterization of four partial wheat-Thinopyrum ponticum amphiploids and their reactions to Fusarium head blight, tan spot and Stagonospora nodorum blotch. Theor Appl Genet 112:1473–1479
Price JA, Simmons AR, Rashed A, Workneh F, Rush CM (2014) Winter wheat cultivars with temperature-sensitive resistance to wheat streak mosaic virus do not recover from early-season infections. Plant Dis 84:525–531
Ray DK, Mueller ND, West PC, Foley JA (2013) Yield trends are insufficient to double global crop production by 2050. PLoS ONE 8:e66428
Reynolds M, Calderini DF, Condon AG, Rajaram S (2001) Physiological basis of yield gains in wheat associated with the LR19 translocation from Agropyron elongatum. Euphytica 119:139–144
Reynolds M, Foulkes J, Furbank R, Griffiths S, King J, Murchie E, Parry M, Slafer G (2012) Achieving yield gains in wheat. Plant, Cell Environ 35:1799–1823
Riar AK, Kaur S, Dhaliwal HS, Singh K, Chhuneha P (2012) Introgression of a leaf rust resistance gene from Aegilops caudata to bread wheat. J Genet 91:155–161
Shepherd KW, Islam AKMR (1988) Fourth compendium of wheat-alien chromosome lines. In: Miller TE, Koebner RMD (eds) Proceedings of the 7th international genetic symposium. Bath Press, Bath, pp 1373–1398
Shewry PR (2009) Wheat. J Exp Bot 60:1537–1553
Taeb M, Koebner RM, Forster BP (1993) Genetic variation for waterlogging tolerance in the Triticeae and the chromosomal location of genes conferring waterlogging tolerance in Thinopyrum elongatum. Genome 36:825–830
Tanaka H, Nabeuchi C, Kurogaki M, Garg M, Saito M, Ishikawa G, Nakamura T, Tsujimoto H (2017) A novel compensating wheat-Thinopyrum elongatum Robertsonian translocation line with a positive effect on flour quality. Breed Sci 67:509–517
van Berloo R (2008) GGT 2.0: versatile software for visualization and analysis of genetic data. J Hered 99:232–236
van Ooijen JW (2011) Multipoint maximum likelihood mapping in a full-sib family of an outbreeding species. Genet Res 93:343–349
Voorrips RE (2002) MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93:77–78
Wang RRC (2011) Agropyron and psathyrostachys. In: Kole C (ed) Wild crop relatives: genomic and breeding resources, cereals. Springer, Berlin, pp 1–247
Wilkinson PA, Winfield MO, Barker GLA, Allen AM, Burridge A, Coghill JA, Edwards KJ (2012) CerealsDB 2.0: an integrated resource for plant breeders and scientists. BMC Bioinf 13:219
Wilkinson PA, Winfield MO, Barker GLA, Tyrell S, Bian X, Allen AM, Burridge A, Coghill JA, Waterfall C, Caccamo M, Davey RP, Edwards KJ (2016) CerealsDB 3.0: expansion of resources and data integration. BMC Bioinf 17:256
Winfield MO, Allen AM, Burridge AJ, Barker GLA, Benbow HR, Wilkinson PA, Coghill J, Waterfall C, Davassi A, Scopes G, Pirani A, Webster T, Brew F, Bloor C, King J, West C, Griffiths S, King I, Bentley AR, Edwards KJ (2016) High-density SNP genotyping array for hexaploid wheat and its secondary and tertiary gene pool. Plant Biotechnol J 13:733–742
Zhang X, Dong Y, Wang RRC (1996) Characterisation of genomes and chromosomes in partial amphiploids of the hybrid Triticum aestivum x Thinopyrum ponticum by in situ hybridization, isozyme analysis, and RAPD. Genome 39:1062–1071
Zhang Z, Lin Z, Xin Z (2009) Research progress in BYDV resistance genes derived from wheat and its wild relatives. J Genet Genomics 36:567–573
Acknowledgements
This work was supported by the Biotechnology and Biological Sciences Research Council as part of the Wheat Institute Strategic Programme [grant number BB/J004596/1] and the Designing Future Wheat Programme [grant number BB/P016855/1]. The funding body played no role in the design of the study, collection, analysis and interpretation of data and in writing the manuscript.
Author information
Authors and Affiliations
Contributions
LB, SG, CY SHE, DS, SA, IPK and JK carried out the crossing programme. LB and CY performed the in situ hybridisation experiments. LB, SHE, DS, SA and CY prepared the samples for genotyping and AJB ran the samples on the array. LB and SG analysed the genotyping data and constructed the physical map. SG, AMPA and PW performed the comparative genomic studies. IPK and JK conceived and designed the experiments. LB wrote the manuscript with assistance from JK, IPK and SG. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Conflict of interest
On behalf of all authors, the corresponding author states that there is no conflict of interest.
Additional information
Communicated by Aimin Zhang.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Baker, L., Grewal, S., Yang, Cy. et al. Exploiting the genome of Thinopyrum elongatum to expand the gene pool of hexaploid wheat. Theor Appl Genet 133, 2213–2226 (2020). https://doi.org/10.1007/s00122-020-03591-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-020-03591-3