Abstract
Key message The stripe rust resistance gene Yr34 was transferred to polyploid wheat chromosome 5AL from T. monococcum and has been used for over two centuries.
Wheat stripe (or yellow) rust, caused by Puccinia striiformis f. sp. tritici (Pst), is currently among the most damaging fungal diseases of wheat worldwide. In this study, we report that the stripe rust resistance gene Yr34 (synonym Yr48) is located within a distal segment of the cultivated Triticum monococcum subsp. monococcum chromosome 5AmL translocated to chromosome 5AL in polyploid wheat. The diploid wheat species Triticum monococcum (genome AmAm) is closely related to T. urartu (donor of the A genome to polyploid wheat) and has good levels of resistance against the stripe rust pathogen. When present in hexaploid wheat, the T. monococcum Yr34 resistance gene confers a moderate level of resistance against virulent Pst races present in California and the virulent Chinese race CYR34. In a survey of 1,442 common wheat genotypes, we identified 5AmL translocations of fourteen different lengths in 17.5% of the accessions, with higher frequencies in Europe than in other continents. The old European wheat variety “Mediterranean” was identified as a putative source of this translocation, suggesting that Yr34 has been used for over 200 years. Finally, we designed diagnostic CAPS and sequenced-based markers that will be useful to accelerate the deployment of Yr34 in wheat breeding programs to improve resistance to this devastating pathogen.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Wheat is a major staple food crop and provides about 20% of calories and proteins for the human population. Although over 750 million tons of wheat are harvested annually from approximately 220 million hectares globally (FAOSTAT), further increases in wheat production are needed to feed a growing human population. One way to increase wheat productivity is to reduce yield losses due to pathogens. Puccinia striiformis f. sp. tritici (Pst), the causal agent of wheat stripe rust (or yellow rust), is currently one of the most devastating fungal diseases threatening global wheat production. This pathogen became an increasing problem after the year 2000, when more virulent and aggressive strains of Pst with increased tolerance to higher temperatures emerged and spread throughout the world (Chen 2005; Hovmøller et al. 2015; Milus et al. 2009).
While effective fungicides against Pst are available, they are expensive and pose some health and environmental risks if not properly used. The deployment of resistance genes remains the most practical and sustainable approach to control this disease. So far, over 80 stripe rust resistance genes (Yr1–Yr83) have received official designations (Li et al. 2020), but most of them are not effective against the virulent post-2000 Pst races. Therefore, the search for new sources of resistance and the development of molecular markers for the effective deployment of these resistance genes is a valuable research objective.
Stripe rust resistance genes Yr34 and Yr48, discovered in hexaploid wheat lines WAWHT2046 in Australia and PI 610750 in the USA, respectively, have been shown to confer partial adult plant resistance against these post-2000 Pst races. Yr34 was initially mapped on the long arm of chromosome 5A, 12.2 cM distal to the awn inhibitor locus B1 (Bariana et al. 2006). Yr48 was also mapped to chromosome 5AL, but based on their different positions relative to the common marker gwm291 it was initially concluded that Yr34 and Yr48 were different genes (Lowe et al. 2011). However, a more recent study of Yr34 identified an error in the original map, and re-mapped this gene to the same chromosome region as Yr48. A large allelism test (600 F2 plants) failed to detect variation for the Pst response, suggesting that the two genes are either allelic or are tightly linked. Since both Yr34 and Yr48 conferred similar seedling responses to pre-2000 and post-2000 Pst races, it was concluded that they are the same gene, which was designated as Yr34 based on the priority of this name (Qureshi et al. 2018).
Both the Yr34 and Yr48 mapping populations showed suppression of recombination in the distal region of chromosome 5AL (Lan et al. 2017; Lowe et al. 2011; Qureshi et al. 2018), which is characteristic of alien introgressions, but that can also be caused by inverted chromosome segments. In addition, the Yr48 chromosome region showed a slight segregation distortion favoring the markers linked to the resistance allele (67% vs expected 50%) (Lan et al. 2017). Although segregation distortion can occur in both alien segments and segments from the same species, they are particularly frequent in the former. Examples of segregation distortion of alien introgressions carrying resistance genes include Lr53/Yr35 in Triticum dicoccoides (Marais et al. 2005b), Lr54/Yr37 in Aegilops kotschyi (Marais et al. 2005a), Lr19 in Agropyron elongatum (Prins and Marais 1999) and QYrtb.pau-5A in Triticum monococcum (Chhuneja et al. 2008). Based on these observations, we hypothesized that Yr34 may be located within an alien introgression.
To characterize the 5AL distal region carrying the Yr34 and Yr48 genes and the presence of a potential alien introgression, we took advantage of a previously developed wheat exome capture platform (Krasileva et al. 2017) and the recent releases of reference genome sequences for multiple Triticum aestivum varieties (Appels et al. 2018; Walkowiak et al. 2020).
The objectives of this study were to test the hypothesis that the lack of recombination in the distal region of chromosome 5AL including Yr34 was the result of a chromosome translocation from a wheat relative and to characterize the distribution of this translocation in the wheat germplasm. We also aimed to identify some historic recombination events that reduced the Yr34 introgressed region to minimize linkage drag, and to develop molecular markers to facilitate the deployment of this resistance gene in wheat breeding programs.
Materials and methods
Plant materials
As a source of Yr48, we used wheat accession PI 610750, which is a synthetic hexaploid wheat developed by the International Maize and Wheat Improvement Center (CIMMYT) in Mexico. Since this accession has multiple Pst resistance genes, we selected RIL143 from the cross UC1110 × PI 610750 that carries only the 5AL resistance gene (Lowe et al. 2011). A population of 46 F2 plants from the cross RIL143 × Avocet-S was used to confirm the linkage of Yr48 with the resistance to Chinese Pst race CYR34. As a source of Yr34, we used the advanced breeding line WAWHT2046 from Australia, that expressed good level of resistance to the Australian 134 E16A + Pst pathotype (Bariana et al. 2006). We also included the common wheat variety Mediterranean (CItr 11587, CItr 3332 and CItr 5303) that was present in many of the pedigrees identified as carriers of the 5AmL introgression.
For T. monococcum, we generated exome capture data for lines DV92 and G3116, which were the parental lines used in the construction of the first genetic map for this species (Dubcovsky et al. 1996). For the exome capture, we used the NimbleGen assay described in Krasileva et al. (2017) and we deposited the data in the T3/Wheat database (https://triticeaetoolbox.org/wheat/). In addition, we obtained another 31 T. monococcum accessions from the US Department of Agriculture National Small Grains Collection (USDA-NSGC, https://npgsweb.ars-grin.gov/gringlobal/search) that were used to trace the origin of the T. monococcum chromosome segment introgressed into bread wheat.
Stripe rust assays
Yr34 and Yr48 have remained effective to all of the virulent Pst isolates present in California since first tested in 2009. To test if Yr34 and Yr48 confer resistance to Chinese Pst races, plants carring Yr34 or Yr48 and F2 individuals from the cross RIL143 × Avocet-S were challenged with the virulent Pst race CYR34 identified in 2008 (also named V26) (Liu et al. 2010). WAWHT2046, RIL143, Avocet-S and F2 plants from the mapping population were grown in controlled walk-in growth chambers at 24 °C during the day and 22 °C during the night. At the jointing stage, plants were inoculated with fresh urediniospores of race CYR34 mixed with talcum powder at a 1:20 ratio using the shaking off method (Ma et al. 2016; Wang et al. 2020b). Wheat leaves were uniformly dusted with this mixture of urediniospores and talc. The inoculated plants were kept in a dark dew chamber set at 10 °C for ~ 24 h and then moved back to the same walk-in growth chamber set at 18 °C during the day and 15 °C during the night. Infection types were recorded ~ 20 days after inoculation using a 0–4 scale (Liu 1988). For each wheat accession, five fully infected leaves were photographed. Sporulation area was calculated using the image analysis software ASSESS version 2.0 from the American Phytopathology Society as reported previously (Lamari 2008).
Sequences and SNPs
Single nucleotide polymorphisms (SNPs) of T. monococcum accessions DV92 and G3116 were obtained from exome capture data. Exome-capture data of tetraploid wheat accessions Zavitan, D447-DW1, Kronos, Svevo, Gredho, and 280–1-Yr15, and hexaploid wheat accessions PI 610750 (CAP2), Billings, Inayama, Altamo, LCD_Star, PI 70613, CO960293, W7984, Berkut, MN98550-5, McNeal, CItr 7635, UC1036, Dayn, Platte, SS_MVP57, UI_Platinum, C0940610, Overley, RioBlanco, Cheyenne, Duster, Hank, 16REG01643, TAM112, TAM111, RAC875, SY_Capstone, Dharwar_Dry, IDO444, 16REG01644, 26R61, Lyman, Reeder, Opata, Excalibur, LA95135, AGS2000, RSI5, Choteau, 2045A, Vida, Bakahtawar94, CCW3A37, KS05HW14-3, CCW3A49, MN99394-1, PBW343 and UC1419, were obtained from the T3/Wheat database (https://triticeaetoolbox.org/wheat/). The published reference genomes of hexaploid wheat Chinese Spring (Appels et al. 2018) and another 10 wheat varieties in the Wheat Pan Genome project (Walkowiak et al. 2020) were included in our analyses. The SNPs identified by exome capture data from two different wheat panels were used to study the distribution of the Yr34 translocation. The first wheat panel consists of 982 hexaploid genotypes (http://wheatgenomics.plantpath.ksu.edu/1000EC/) (He et al. 2019). The second panel includes 460 hexaploid accessions (Pont et al. 2019). A total of 1,442 hexaploid wheat genotypes were obained with exome sequencing data, and they were separated in four historical groups (every 30 years): group I, before 1930; group II, from 1931 to 1960; group III, from 1961 to 1990; and group IV, after 1991.
Marker development and SNP validation
Genome-specific primers were designed with software Primer3 (https://bioinfo.ut.ee/primer3-0.4.0/primer3/) to amplify gene regions carrying putative T. monococcum-specific SNPs. Techniques and procedures for developing cleaved amplified polymorphic sequence (CAPS) markers were reported previously (Konieczny and Ausubel 1993). NEBcutter V2.0 (http://www.labtools.us/nebcutter-v2-0/) was used to detect restriction sites including the targeted SNPs. PCR reactions were performed in a Veriti 96-Well Fast Thermal Cycler (Applied Biosystems) with an initial denaturation step of 94 °C for 5 min, followed by 40 cycles of 94 °C for 30 s, 50–65 °C for 30 s, and 72 °C for 1 min, with a final extension step at 72 °C for 7 min. The PCR products were visualized in 1–2.5% agarose gels stained with ethidium bromide. The PCR amplification products with the right sizes were then sequenced to confirm the present of 5AmL-specific SNPs. Restriction enzymes from New England BioLabs Inc. were used to digest the amplified products.
Eleven pairs of genome-specific primers, including pku5410F3R3, pku5414F4R4, pku5429F2R2, pku5451F5R5, pku5473F2R2, pku5488F4R4, pku5497F2R2, pku5507F1R1, pku5508F1R1, pku5575F5R5 and pku5585F1R1 (Table 1), were developed from 11 different genes distributed along the translocated T. monococcum segment in RIL143 to detect polymorphic sites among 32 cultivated T. monococcum accessions.
Candidate genes and expression analysis
Candidate genes for our target region were identified from the genome sequences of two European winter wheat varieties ArinaLrFor and SY Mattis included within the 10 + Wheat Genomes Project, and that were the most similar to the lines carrying Yr34 in the distal region of chromosome 5AL. Sequences were obtained from the database of the Institute of Plant Genetics and Crop Plant Research (IPK) (https://webblast.ipk-gatersleben.de/wheat_ten_genomes/viroblast.php). The expression levels for the candidate genes were obtained from the Wheat Expression Browser (expVIP, http://www.wheat-expression.com/) (Borrill et al. 2016).
Results
Assessment of stripe rust responses
WAWHT2046 and RIL143 exhibited a moderate level of resistance to the virulent Chinese race CYR34, whereas the control line Avocet-S was susceptible (Fig. 1a). To quantify the amount of disease present on the leaves, disease measurement were performed on five fully infected leaves of each line using software ASSESS v2. The average percentage of the leaf area covered by Pst pustules was significantly lower (P < 0.001) in WAWHT2046 (24%, ranging from 15 to 34%) and RIL143 (29%, ranging from 18 to 39%) than in Avocet-S (84%, ranging from 72 to 95%). Chlorotic/necrotic responses are marked with arrows on the leaves of WAWHT2046 and RIL143 in Fig. 1b.
Since RIL143 carries only one of the resistant alleles (Yr48) segregating in the UC1110 × PI 610750 RIL population, we hypothesized that its resistance to Pst race CYR34 was conferred by this allele. This was confirmed by phenotyping 46 F2 plants derived from the cross RIL143 × Avocet-S with CYR34, and genotyping them with marker cfa2149, which is completely linked to Yr48 (Lowe et al. 2011). All ten plants homozygous for the RIL143 allele were resistant, whereas all 12 plants homozygous for the Avocet-S allele were susceptible, confirming that RIL143 resistance to race CYR34 is linked to the Yr48 region.
T. monococcum segments introgressed into polyploid wheat
To explore the origin of the Yr34 segment, we compared the SNPs from the exome capture of Yr48 donor line PI 610750 with that of 48 other hexaploid wheat accessions, 6 tetraploid lines, and two diploid T. monococcum lines (DV92 and G3116) generated as part of the USDA-NIFA funded WheatCAP project. This SNP dataset is available in the T3/Wheat database (https://triticeaetoolbox.org/wheat/genotyping/display_genotype.php?trial_code=2017_WheatCAP_UCD). To our surprise, we found that the distal region of chromosome arm 5AL in PI 610750 had a large number of rare polymorphisms that were shared with T. monococcum accessions DV92 and G3116 and the common wheat variety Billings. These results suggested, for the first time, that the distal region of 5AL carrying Yr34 could have originated in T. monococcum. To explore this region in more detail, we focused on the SNPs that were present in the two T. monococcum accessions, but were absent in all other accessions of polyploid wheat (except PI 610750 and Billings), and that are referred hereafter as T. monococcum-specific SNPs.
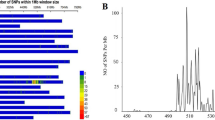
In the 5AL region starting from 685.4 Mb to the end of the chromosome (based on CS RefSeq v1.0 coordinates), we identified 1,047 T. monococcum-specific SNPs (Table S1). To visualize the distribution of these SNPs in the distal 24.3 Mb of chromosome arm 5AL, we represented the T. monococcum-specific SNPs in blue and other SNPs in grey (Fig. 2). This figure shows that the T. monococcum segment in PI 610750 was approximately 15 Mb long, extending from 694.8 Mb to the end of the chromosome and sharing 1,019 T. monococcum-specific SNPs with DV92 and G3116. The T. monococcum segment in Billings was approximately 8 Mb shorter (~ 7 Mb long), and extended from 702.9 Mb to the end of the chromosome sharing 569 T. monococcum-specific SNPs with DV92 and G3116 (Fig. 2).
Distribution of 1,047 T. monococcum-specific SNPs on the distal 24.3 Mb of chromosome 5AL (685.4 Mb to end). 1, DV92; 2, G3116; 3, PI 610,750 (Yr48); 4, Billings; 5, ArinaLrFor; 6, SY Mattis; 7–59, tetraploid and hexaploid wheat accessions from exome-capture sequencing (Table S1). This figure was produced using the Integrative Genomics Viewer (IGV) software version 2.8.9 (Robinson et al. 2011). Vertical lines in blue represent T. monococcum-specific SNPs whereas lines in light grey are normal wheat SNPs or not-polymorphic sites with Chinese Spring. Coordinates were based on CS RefSeq v1.0
To test if the T. monococcum translocation was present in other sequenced T. aestivum accessions, we performed BLASTN searches using the sequences flanking the target SNPs. We found that only the two European winter wheat varieties ‘ArinaLrFor’ and ‘SY Mattis’ have the distal 5AL T. monococcum translocation among the ten T. aestivum accessions assembled as pseudomolecules in the Wheat Pan Genome project (Walkowiak et al. 2020). These two varieties share the 569 T. monococcum-specific SNPs identified in Billings (Table S1) indicating that they have the same translocated segment. Using the genomic sequences of ArinaLrFor and SY Mattis, we were able to estimate more precisely the size of the T. monococcum introgression in these two varieties, which was approximately 9.5 Mb, and extended from 700.7 Mb to the end of the chromosome (710.1 Mb) in ArinaLrFor and from 693.1 Mb to the end of the chromosome (702.6 Mb) in SY Mattis.
Since the T. monococcum segment in PI 610750 was approximately 8 Mb longer than in ArinaLrFor and SY Mattis (Fig. 2), we adjusted the estimate of its length from 15 Mb to 17.5 Mb (distal 9.5 Mb in ArinaLrFor + proximal 8 Mb estimate based on Fig. 2). To define better the translocation breakpoint in PI 610750, we developed 21 A/Am-genome specific primers across the 17.5 Mb introgressed T. monococcum chromosome segment (Table 1) and used them to test DV92, RIL143, ArinaLrFor and WAWHT2046 via Sanger sequencing (Table S2). The physical positions of these markers are presented in Fig. 3. Using these markers, we determined that the translocation point in RIL143 occurred between markers pku5380F3R3 and pku5381F2R2 (Fig. 3b, 694.8 and 695.0 Mb in CS, respectively). Using additional SNP polymorphisms, we determined that the border of the translocation in Billings, ArinaLrFor and SY Mattis was between markers pku5488F4R4 and pkuS5A7761F2R2 (Fig. 3c, 702.8 and 702.9 Mb in CS, 700.7 and 700.8 Mb in ArinaLrFor).
T. monococcum chromosome segments introgressed into common wheat. a Chromosome 5A; b The length of the introgressed T. monococcum segment in RIL143 (grey) is estimated to be 15 Mb based on CS coordinates, but can be adjusted to 17.5 Mb based on ArinaLrFor coordinates; c The introgressed T. monococcum segment in ArinaLrFor (grey) was originally estimated to be 7 Mb based on CS coordinates and was adjusted to 9.5 Mb based on ArinaLrFor coordinates. Coordinates in (b) and (c) are based on CS RefSeq v1.0 and ArinaLrFor, respectively. The dotted horizontal grey lines indicate the recombination events between T. monococcum and T. aestivum chromosomes
We then explored the presence of the T. monococcum translocation in WAWHT2046, which is the original line where Yr34 was discovered. PCR markers in the region that differentiates RIL143 and ArinaLrFor (pku5414F4R4: 698.2 Mb, pku5429F2R2: 698.6 Mb and pku5488F4R4: 702.8 Mb, CS RefSeq v.1 coordinates) showed the T. monococcum allele in RIL143 and DV92 but not in WAWHT2046, ArinaLrFor or the wheat control Avocet-S (Fig. 4a). By contrast, PCR markers in the common T. monococcum distal segment (pku5542F1R1: 706.2 Mb, pku5575F5R5: 708.4 Mb and pku5585F1R1: 709.2 Mb) showed the T. monococcum allele in RIL143, WAWHT2046, ArinaLrFor and DV92, but not in Avocet-S (Fig. 4b). These results confirmed that WAWHT2046 carries the same T. monococcum translocation as ArinaLrFor. We further confirmed that the borders of the translocation were identical using flanking markers pku5488F4R4 and pkuS5A7761F2R2 described above, and that all the tested SNPs starting from position 702.9 Mb were identical in ArinaLrFor and WAWHT2046 (Table S2).
CAPS markers used to characterize the T. monococcum segment present in WAWHT2046 (Yr34). a CAPS markers pku5414F4R4 (698.2 Mb, XhoI), pku5429F2R2 (698.6 Mb, PvuII) and pku5488F4R4 (702.8 Mb, ApoI) in the region that differentiates RIL143 and ArinaLrFor; b CAPS markers pku5542F1R1 (706.2 Mb, Hpy188III), pku5575F5R5 (708.4 Mb, AcuI) and pku5585F1R1 (709.2 Mb, BanII) in the common T. monococcum segment. 1, Avocet-S (wheat control); 2, RIL143 (Yr48, long introgression control); 3, WAWHT2046 (Yr34); 4, ArinaLrFor (short introgression control); 5, DV92 (T. monococcum control); M, markers. Coordinates are based on CS RefSeq v1.0. Arrowheads represent wheat bands and arrows represent T. monococcum bands
Distribution of T. monococcum introgressions in hexaploid wheat
In order to determine the frequency and the distribution of the T. monococcum translocated segments in wheat genotypes, 1,442 hexaploid wheat accessions with exome sequencing data derived from the 1,000 wheat exomes project (includes 982 hexaploid wheat genotypes) (He et al. 2019) and the 500 exomes project (includes 460 hexaploid wheat genotypes) (Pont et al. 2019), were used for SNP calling and comparative analysis. Our survey revealed that 105 accessions out of 982 (10.7%) in the first panel and 147 accessions out of 460 (32%) in the second panel possess T. monococcum-wheat translocations on the distal region of chromosome 5AL (Table S3 and S4). Among these 252 accessions plus the 5 lines mentioned before carrying the T. monococcum-wheat translocation (PI 610750, Billings, ArinaLrFor, SY Mattis and WAWHT2046), we identified T. monococcum segments of 14 different lengths, which were designated as L1 to L14 (Table S3 and S4).
The length of the introgressed T. monococcum segments and the number of lines carrying each introgression are indicated at the bottom of Fig. 5, and a complete list of accessions is available in Table S5. Lines carrying translocations L5 to L11 include the complete L2 region present in WAWHT2046 and, therefore, are likely carriers of the Yr34 resistance gene. By contrast, translocations L3, L4, L13 and L14 include only part of the L2 translocation, so we currently do not know if they carry the Yr34 resistance gene.
Estimated lengths of the L1-L14 T. monococcum introgressions. L1 is the longest (and likely original translocation) and all others were generated by distal or proximal recombination events with chromosome arm 5AL. The first row of numbers below the figure indicates the lengths of the T. monococcum segment estimated using a combination of coordinates of the 9.5 Mb T. monococcum segment in the ArinaLrFor genome and the CS RefSeq v1.0 coordinates for the rest of the T. monococcum segment not present in ArinaLrFor. The second row of numbers indicated the number of wheat accessions carrying each introgression. The T. monococcum introgression is indicated in gray and the T. aestivum segment in white
Using the data from the two exome capture projects (He et al. 2019; Pont et al. 2019), we were able to find a small amount of SNPs among the different introgressions. Since most of these SNPs were frequent in the T. aestivum accessions without the T. monococcum introgressions, we interpreted them as conversion events. Supplemental Table S6 summarizes the 11 most likely conversion events based on the presence of at least two adjacent SNPs separated by less than 4 kb and all identical to T. aestivum alleles. The raw SNPs data used in this analysis are presented in Tables S3 and S4, which also include 23 additional single SNPs. These single SNPs were all frequent in hexaploid lines without the T. monococcum introgressions, and could also be conversion events (not included in Table S6).
The previous results suggest that the T. monococcum introgressions of different lengths may have originated by recombination events from a single T. monococcum introgression. This hypothesis is also supported by shared borders among several of the accessions. For example, the L3 introgression shared its proximal border with L1 (between SNPs S5A_694759680 and S5A_694966923, Table S3). This was further confirmed by a more precise mapping of the L1 and L3 proximal border to a 0.2 Mb region between markers pku5380F3R3 and pku5381F2R2 in accessions PI 619381 and PI 619379. These results suggest that L3 likely originated from L1 by a distal recombination event with 5AL. This distal border of L3 is shared by L4, which also shares a proximal border with L5 suggesting a possible origin of L4 from recombination between L3 and L5. Similarly, the L13 introgression shares the proximal border with L2 and the distal border with L3 and L4, so it could have originated from recombination between L2 and either L3 or L4. L14 shares the proximal border with L13 and could have originated by a distal recombination event with 5AL in L13. More precise mapping of the shared borders will be required to validate these hypotheses. Finally, all other T. monococcum introgressions share the most distal T. monococcum SNPs and are likely terminal introgressions derived by proximal recombination events between 5AL and L1 or other lines with longer distal T. monococcum introgressions.
We detected the 5AmL translocations of different lengths in 50 countries covering all continents where wheat is grown (Table S7), especially in European countries, suggesting a wide distribution. The overall frequency of the 5AmL translocation segments in the present panel of hexaploid wheat genotypes was 17.5% (252/1442), but the proportion in different continents varied significantly (Fig. S1 and Table S7). More specifically, the translocation was detected in 34.4% wheat accessions from Europe, 8.1% accessions from North America, 8.8% accessions from Asia, 2.9% accessions from Oceania, 4.8% accessions from Africa, and 2.1% accessions from South America (Fig. S2 and Table S7).
We then compared the frequencies of the translocation within four historical groups (every 30 years). In the first wheat panel (He et al. 2019), we found that the translocation was rare (1.6%) in varieties released before 1930 (Group I), but its frequency increased sharply (57.1%) in the modern varieties released after 1990 (Group IV). Likewise, in the second wheat panel (Pont et al. 2019), the frequency of the translocation increased from 11.1% in Group I to 46.3% in the more recent varieties of Group IV. In summary, this analysis revealed rapid increases in the frequency of 5AmL.5AL translocation in hexaploid wheat varieties (Table S8).
Tracing the origin of the RIL143/Billings translocation
To trace the origin of the translocation, we performed pedigree analysis to determinate the relationship among wheat accessions carrying the translocations using the wheat pedigree database (http://www.wheatpedigree.net/). Among the 135 lines carrying the 5AmL translocations for which we obtained pedigree information, we found that 103 shared a common parental line named “LV-Mediterranean” (or its derivatives) in their pedigree (Table S9). We found no information about LV-Mediterranean, but we found that its derivative “Mediterranean”, was a late-sown variety introduced into the U.S. from Europe in the year 1819 (Olmstead and Rhode 2002).
We identified three accessions of Mediterranean in the USDA-NSGC (CItr 3332, CItr 5303 and CItr 11587, received in 1912, 1913 and 1933, respectively) and characterized them with markers pku5380F3R3 (694.8 Mb), pku5381F2R2 (695.0 Mb), pku5473F1R1 (701.1 Mb), pku5497F2R2 (703.4 Mb), pku5542F1R1 (706.2 Mb) and pku5585F5R5 (709.2 Mb) (Table 1). These markers showed that CItr 3332 does not carry the T. monococcum translocation, and that both CItr 5303 and CItr 11587 carry a T. monococcum translocation with a common proximal border and different distal borders. The proximal border was located between 694.8 and 695.0 Mb using flanking markers pku5380F3R3 and pku5381F2R2, in the same position as L1 and L3. The 5AmL alleles extended to 709.2 Mb for line CItr 5303, but only to 703.4 Mb for line CItr 11587, with 5AL alleles for markers between 706.2 Mb and 709.2 Mb. These results suggest that CItr 5303 carries the same L1 translocation as PI 610750, whereas CItr 11587 is likely to carry the L3 translocation (Fig. 5). These results suggest that Mediterranean (or LV-Mediterranean) could be the origin of the T. monococcum translocation or at least of its introduction in North America.
To further characterize the T. monococcum introgression in Mediterranean, we determined the alleles present in CItr 5303 for SNPs detected between the L1 translocation in PI 610750 and the L2 translocation in ArinaLrFor. A comparison of the PI 610750 exome capture data with the genomic sequence available for ArinaLrFor (Walkowiak et al. 2020) revealed 13 polymorphisms between L1 and L2. Six of these SNPs appear to be also the result of conversion events in PI 610750 based on the presence of the same SNPs in several of the sequenced T. aestivum genomes (Table S10) and their absence in Mediterranean. The other seven polymorphisms (including one 34-bp deletion and six SNPs, Table S10) were not present in any of the 10 sequenced T. aestivum pseudomolecules, including ArinaLrFor and SY Mattis, nor in the variety Mediterranean (accession CItr 5303). These results suggest that these polymorphisms originated in PI 610750 after the introgression of the T. monococcum segment in T. aestivum.
Identification of the closest source of the T. monococcum segment
To investigate the origin of the T. monococcum segment and to explore the source of the seven polymorphisms between PI 610750 and Mediterranean that we were not able to find in T. aestivum, we sequenced the regions including these SNPs and several other regions in a set of 32 accessions of cultivated T. monococcum subsp. monococcum.
We focused on the cultivated accessions of T. monococcum because a comparison of the exome capture data from the cultivated accession DV92 and the wild T. monococcum subsp. aegilopoides accession G3116, revealed that the L1 introgression shared 249 out of 301 SNPs (82.7%) with DV92 and only 52 (17.3%) with G3116 (Table S11). The numbers were similar for the 9.5-Mb distal region, where the L2 translocation shared 89.5% SNPs (137/153) with DV92 and 10.5% (16/153) with G3116 (Table S11). These results indicate that the translocated segment originated from the cultivated T. monococcum subsp. monococcum.
We evaluated the relationships among 32 T. monococcum subsp. monococcum accessions by Sanger sequencing of 11 different gene regions across the L1 introgression (Table S12). Since our objective was to find the closest T. monococcum accession to the original translocation, we eliminated all the PI 610750 SNPs that were not in Mediterranean CItr 5303 (Table S10). We failed to detect polymorphisms among the 32 T. monococcum accessions and PI 610750 in the 3,300 bp amplified with primers for four regions (pku5410F3R3, pku5414F4R4, pku5507F1R1 and pku5508F1R1). For the other 7 regions, we sequenced 6,135 bp that revealed 67 polymorphic sites (Table S12). A Neighbor-Joining tree based on these polymorphisms (Fig. 6) showed that PI 610750 was located within a cluster that included multiple European accessions (Table S13). The two closest T. monococcum accessions to PI 610750 were PI 289605 and PI 428158, which were both collected in the United Kingdom. The T. monococcum accession PI 289605 showed only 4 SNPs with the L1 introgression in hexaploid wheat (99.958% identical, excluding the PI 610750 unique SNPs not present in Mediterranean), suggesting that this accession is closely related to the one that was the source of the 5AmL.5AL translocation.
SNP-based phylogenetic analysis. Sequences were aligned with Muscle as implemented in software Mega version 7. Phylogenetic tree was produced based on 67 polymorphisms identified among 9,435 bp obtained by Sanger sequencing from 11 loci. The evolutionary history was inferred using the Neighbor-Joining method. All ambiguous positions were removed for each sequence pair (pairwise deletion option). Interactive Tree Of Life (iTOL) version 5 was used to visualize the tree (https://itol.embl.de/). PI 610750 (Yr48) is highlighted in bold. Putative conversion polymorphisms listed in Tables S6 and S10 that were not present in the L1 introgression in Mediterranean were excluded in the comparisons between PI 610750 and the T. monococcum accessions
Among the seven polymorphisms present in PI 610750 and not in the T. aestivum without the T. monococcum introgression (Table S10), two (RefSeq v1.0 coordinates, 703,182,334 and 707,059,015 bp) were also absent in the 32 accessions of T. monococcum. This result suggests that these two SNPs may have originated either by mutations in PI 610750 or by conversion from T. aestivum accessions with different haplotypes than the ones included in our study. Interestingly, the other 5 polymorphisms that were not present in any of the sequenced T. aestivum genomes (one 34-bp deletion: 705,376,608–705,376,641 bp and SNPs 705,375,944, 705,376,462, 705,408,362 and 705,408,374 bp, Table S10) were found in a group of four T. monococcum accession from Armenia, Azerbaijan and Germany (PI 326317, PI 418582, PI 349049 and PI 355524, Table S13). This result suggests the intriguing possibility of recombination with a different T. monococcum accession, but more extensive surveys and sequencing will be required to test this hypothesis.
The vernalization locus VRN2 is included in the T. monococcum introgression region present in L1 but not in L2 (CS RefSeq v1.0 coordinates, 698.2 Mb, Fig. 3b). This locus includes linked genes ZCCT1 and ZCCT2, and both genes are not functional in the A genome of polyploid wheat (Distelfeld et al. 2009). The T. monococcum VRN2 alleles for a spring growth habit have either a deletion of both ZCCT1 and ZCCT2 that can be identified with primers Vrn2F3R3, Zcct2F6R6 and R3C1N3/RACEC1N1 (Table 1) or non-functional copies in both genes characterized by an arginine (R) to tryptophan (W) mutation at position 35 of the CCT domain in the ZCCT1 protein (henceforth RW mutation) that can be detected with CAPS marker R3C1N3/RACEC1N1 (Table 1) (Yan et al. 2004). Analysis of PI 610750 and Mediterranean showed that both accessions carry the ZCCT1 and ZCCT2 deletion. We then screened a collection of 32 cultivated T. monococcum accessions, enriched in the presence of the ZCCT1 and ZCCT2 deletion based on a previous survey (Yan et al. 2004). Analysis with Vrn2F3R3, Zcct2F6R6 and R3C1N3/RACEC1N1 identified 9 accessions where the functional VRN2 alleles were present, 4 carrying the RW mutation and 19 carrying the deletion of both genes (Table S13). Interestingly, among the T. monococcum accessions within the cluster of European varieties including PI 610750, only accession PI 591871 from Georgia showed the VRN2 deletion. Since this was not the closest accession to PI 610750, the origin of the VRN2 deletion in the introgressed L1 segment remains an open question.
Candidate genes for Yr34
Yr34 was initially mapped to the distal region of chromosome 5AL in WAWHT2046 (Qureshi et al. 2018), and we show here that this region is included in the 9.5-Mb introgression from T. monococcum (L2). Based on these results, we concluded that the Yr34 gene is located within the L2 translocation. Since ArinaLrFor shares the same L2 translocation as WAWHT2046, we used the ArinaLrFor genomic sequence (https://webblast.ipk-gatersleben.de/wheat_ten_genomes/viroblast.php) to obtain a list of 134 annotated genes in the candidate region (TraesARI5A01G579500- TraesARI5A01G592800, Table S14). The functional annotation of these genes using Pfam or BLASTN/BLASTX searches in GenBank did not reveal any typical NBS-LRR resistance genes but detected six genes annotated as putative RECEPTOR-LIKE PROTEIN KINASES (RLKs, TraesARI5A01G582700, TraesARI5A01G584100, TraesARI5A01G586200, TraesARI5A01G589400, TraesARI5A01G591100 and TraesARI5A01G591700).
We then analyzed the expression levels of the candidate genes using published RNAseq studies compiled in the wheat expVIP database (http://www.wheat-expression.com/). Among the 134 genes annotated in the candidate gene region in the ArinaLrFor genome, we found that 53 were expressed in wheat leaves infected with Pst, which included four of the six annotated RLKs genes (TraesARI5A01G582700, TraesARI5A01G586200, TraesARI5A01G589400 and TraesARI5A01G591700). We have prioritized these four genes for further functional characterization.
Discussion
Diploid wheat T. monococcum is a good source of resistance genes
Diploid wheat T. monococcum (2n = 2x = 14, AmAm) is closely related but a different species from T. urartu (2n = 2x = 14 = AA) (Johnson and Dhaliwal 1976), which is the donor of the A genome of polyploid wheat (Dvorak et al. 1988). Previous studies have shown that the chromosome 1A of bread wheat and 1Am of T. monococcum recombine poorly in the presence of the Pairing homeologous1 (Ph1b) gene, but that normal recombination can be restored through the use of the ph1b mutation (Dubcovsky et al. 1995). However, in the presence of the wild type Ph1b the reduction in recombination is not the same for all T. monococcum chromosomes, and some recombination was observed between the distal region of chromosomes 5Am and 5A in a wild type hexaploid wheat background (Luo et al. 2000). This result agrees with the discovery of T. monococcum translocation of 14 different lengths in this study (Fig. 5), which suggests multiple 5Am x 5A recombination events during the long breeding history of this introgression.
The ability of the T. monococcum chromosomes to recombine with the A-genome chromosomes (particularly in the ph1b background) has fueled the interest of breeders in the identification of resistance genes in this diploid species and its transfer to the commercial polyploid wheat species. Successful isolation and transfer of resistance genes from T. monococcum to hexaploid wheat include the stem rust resistance genes Sr21 (Chen et al. 2015, 2018b), Sr22 (Steuernagel et al. 2016), Sr35 (Saintenac et al. 2013), SrTm4 (Briggs et al. 2015), Sr60 and SrTm5 (Chen et al. 2018a, 2020); the leaf rust resistance genes Lr63 (Kolmer et al. 2010) and LrTM16 (Sodkiewicz et al. 2008) and the powdery mildew resistance genes Pm1b (Hsam et al. 1998), Pm4d (Schmolke et al. 2012) and Pm25 (Shi et al. 1998).
Although T. monococcum shows good adult plant resistance against Pst (Chhuneja et al. 2008), only two stripe rust resistance QTLs, QYrtm.pau-2A and QYrtb.pau-5A, have been mapped from this species so far (Chhuneja et al. 2008). QYrtb.pau-5A was identified in T. monococcum subsp. aegilopoides accession pau5088 and was mapped on chromosome arm 5AmL flanked by simple sequence repeat (SSR) markers barc151 and cfd12. Using the sequences of these two SSR markers, we determined the physical location of QYrtb.pau-5A in the reference genome of ArinaLrFor was from 557.7 Mb to 561.9 Mb. Since Yr34 was located distal to marker pku5488F4R4 (700.7 Mb), we concluded that QYrtb.pau-5A and Yr34 are likely two different genes.
Stripe rust resistance genes Yr34 and Yr48 were previously suggested to be the same gene on the basis of an allelism test and similar responses to different Pst races (Qureshi et al. 2018). Although the limited recombination within the T. monococcum 5AmL chromosome segment limits the value of the Yr48 (L1) × Yr34 (L2) allelism test, the absence of susceptible plants suggests that both Yr48 and Yr34 are located within the shorter 9.5 Mb segment (L2). This result supports (but does not prove) the suggestion that Yr34 and Yr48 are the same gene. Varieties Billings, ArinaLrFor and SY Mattis carry the same L2 segment as WAWHT2046, suggesting that they also carry the Yr34/Yr48 resistance gene. However, since the Yr34/Yr48 causal gene has not been identified yet, we cannot rule out the possibility that these varieties carry a non-functional copy of this gene.
The presence of this alien T. monococcum translocation likely explains the segregation distortion and the suppression of recombination observed in the chromosome region carrying Yr34 and Yr48 (Lan et al. 2017; Lowe et al. 2011).
The regions of suppressed recombination were not identical for Yr34 and Yr48. In the Yr34 study (Qureshi et al. 2018), the authors reported recombination between Yr34 and the awn inhibitor gene B1 located at 698.5 Mb in CS RefSeq v1.0 (DeWitt et al. 2020). By contrast, the region of suppressed recombination for Yr48 extended to VRN2 at 698.2 Mb (Fig. 3b) (Lowe et al. 2011). This difference in recombination is supported in this study by the finding that the T. monococcum introgressions have different lengths in the donor of Yr48 (L1 in PI 610750) and the donor of Yr34 (L2 in WAWHT2046). The B1 and VRN2 loci are outside the translocation in WAWHT2046 and within the translocation in PI 610750.
The 5AmL translocation has been used across a wide spatial and temporal range
We detected the 5AmL translocation in accessions from 50 countries, which suggests that it has been used in wheat breeding worldwide. However, the frequency of this translocation is not uniform across continents, ranging from less than 5% in South America, Africa and Oceania to 34.4% in Europe (Fig. S2 and Table S7). This data suggests that either the translocation has an older breeding history in Europe or it has been under stronger positive selection in Europe than in other regions. Although it is possible that the presence of stripe rust resistance gene Yr34 contributed to the increased frequency of this segment, we cannot rule out the possibility that other favorable genes within this T. monococcum translocation favored its selection.
This wide geographic distribution of the T. monococcum introgression also suggests that it has a long history. Indeed, the screening of two large and diverse panels of wheat accessions with exome capture data (He et al. 2019; Pont et al. 2019) revealed the presence of the translocation in 11 wheat varieties released before 1931 (Table S5). Pedigree analysis of these varieties found that a wheat variety named LV-Mediterranean (or its derivatives) was frequent in the pedigrees of the varieties carrying the T. monococcum translocation. We confirmed the presence of the longest L1 translocation in Mediterranean accession CItr 5303 and the reduced L3 translocation in another Mediterranean accession (CItr 11587). It should be pointed out that these accessions were collected in different places of the USA nearly 100 years after its introduction in the US from Italy in 1819 under the name “Mediterranean” (Olmstead and Rhode 2002). Mediterranean was a very popular variety due to its better resistance to Hessian fly and rust than other varieties. Nearly 100 years after its introduction, Mediterranean occupied 2,770,000 acres and, in 1924, it was still grown on 600,000 acres (Ball 1930). Mediterranean’s long history and wide area of cultivation likely explain the heterogeneity of the Mediterranean samples maintained in the USDA-NSGC.
The comparison of the longest L1 introgression in Mediterranean and PI 610750 provides additional evidence of the ancestral origin of the T. monococcum segment in Mediterranean. In Mediterranean, we were not able to find the putative conversion events observed in PI 610750. The pedigree of PI 610750 suggests that the L1 segment from Mediterranean passed through at least 11 crosses to reach PI 610750. If we assume an average of three generations of self-pollination before fixation, this would imply that the L1 segment from PI 610750 passed though > 30 meiosis, providing multiple opportunities for conversion events. By contrast, if the L1 from Mediterranean CItr 5303 was never crossed and was never in heterozygous state, it had no opportunities for conversion events.
Taken together, these results suggest that this T. monococcum translocation may have provided Pst resistance for over 200 years and that it may represent one of the oldest alien introgressions in hexaploid wheat.
Source of the T. monococcum introgression
Although we have established with high level of confidence that the introgressed segment originated from a cultivated T. monococcum subsp. monococcum and not from the related wild T. monococcum subsp. aegilopoides, we have not identified the exact accession of diploid wheat where this segment originated.
The closest T. monococcum subsp. monococcum accessions to the L1 segment are from Europe, which is consistent with the origin of Mediterranean in Italy. Two accessions from the UK are particularly close to the L1 introgression, pointing to the UK as a potential origin of the introgression. A more extensive survey of T. monococcum accessions and the sequencing of a larger number of loci will be necessary to provide a more conclusive answer to this question.
Most of the SNPs detected between L1 and L2 or Mediterranean are likely conversion events from T. aestivum chromosome 5A, since the same alleles were found in multiple hexaploid accessions. However, we found a 34-bp deletion linked to 4 SNPs that were not found in any of the sequenced genomes of T. aestivum but were detected instead in a group of four T. monococcum accessions from the Caucasus and Germany. If we assume that Mediterranean and PI 610750 have the same L1 translocation based on their shared proximal border (with a 0.2 Mb resolution), then the absence of these five polymorphisms in Mediterranean would indicate that they represent an introgression or conversion that occurred after the introgression of the T. monococcum segment in hexaploid wheat. We speculate that these polymorphisms may have originated from additional crosses with T. monococcum. Since wheat was grown extensively during the Roman Empire and after in an area that overlaps with T. monococcum geographical distribution, it would be interesting to investigate if this T. monococcum introgression has actually a much longer history.
Candidate genes for Yr34
Most of the cloned disease resistance genes in wheat encode intracellular coiled-coil nucleotide-binding leucine-rich repeat (NLR) proteins (Chen et al. 2018a; Marchal et al. 2018; Saintenac et al. 2013; Wang et al. 2020a; Zhang et al. 2019, 2017), which recognize pathogen effectors and activate effector-triggered immunity (Jones and Dang 2006). However, we did not detect any typical NBS-NLR genes within the 9.5 Mb translocation in the genomes of ArinaLrFor or SY Mattis, which suggests that Yr34 likely belong to a different class of resistance genes.
In the L2 candidate region we identified six RLKs, four of which are expressed in wheat leaves infected with Pst. RLKs have been frequently associated with disease resistance in different plant species (Brueggeman et al. 2002; Hurni et al. 2015; Martin et al. 1993; Song et al. 1995; Wang et al. 1996). Two of the cloned stripe rust resistance genes, Yr36 (Fu et al. 2009) and Yr15 (Klymiuk et al. 2018) encode proteins with kinase domains, and similar to Yr34, provide broad spectrum resistance against Pst and have remained effective for many years. We have prioritized these RLKs for functional characterization to test if they are the causal genes for Yr34.
Conclusions and practical implications
Yr34 confers intermediate levels of resistance against virulent Pst races in a wide range of regions, including China (the current study), the United States (Lowe et al. 2011), Mexico (Lan et al. 2017), and Australia (Qureshi et al. 2018), indicating broad-spectrum resistance to different Pst races. In addition to its broad resistance, the fact that Yr34 has remained effective for more than 100 years after its extensive deployment in commercial agriculture suggests that this gene provides durable resistance to Pst. However, since Yr34 only provides partial resistance, it needs to be combined with other Pst resistance genes to confer economically useful levels of resistance.
The broad resistance spectrum and durability of Yr34 makes this gene a desirable target for introgression into modern wheat varieties. For this purpose, we recommend the utilization of the shorter 9.5 Mb from ArinaLrFor, WAWHT2046, Billings or SY Mattis. The L2 translocation provides similar levels of resistance as the L1 translocation, minimizes potential linkage drag, and reduces the chromosome area with limited recombination associated with the T. monococcum segment. Until Yr34 is mapped more precisely, it is better to use a combination of a proximal marker (pkuS5A7761F2R2, currently a Sanger sequencing marker) and a distal (pku5585F1R1, Fig. 4) marker to confirm that the L2 segment was transferred without recombination. These CAPS and sequenced-based markers represent a useful tool to accelerate the deployment of this broad spectrum and durable resistance gene in modern wheat breeding programs.
References
Appels R, Eversole K, Feuillet C, Keller B, Rogers J, Stein N, Pozniak CJ, Choulet F, Distelfeld A, Poland J (2018) Shifting the limits in wheat research and breeding using a fully annotated reference genome. Science 361:eaar7191
Ball CR (1930) The history of American wheat improvement. Agr hist 4:48–71
Bariana HS, Parry N, Barclay IR, Loughman R, McLean RJ, Shankar M, Wilson RE, Willey NJ, Francki M (2006) Identification and characterization of stripe rust resistance gene Yr34 in common wheat. Theor Appl Genet 112:1143–1148
Borrill P, Ramirez-Gonzalez R, Uauy C (2016) expVIP: a customizable RNA-seq data analysis and visualization platform. Plant physiol 170:2172–2186
Briggs J, Chen S, Zhang W, Nelson S, Dubcovsky J, Rouse MN (2015) Mapping of SrTm4, a recessive stem rust resistance gene from diploid wheat effective to Ug99. Phytopathology 105:1347–1354
Brueggeman R, Rostoks N, Kudrna D, Kilian A, Han F, Chen J, Druka A, Steffenson B, Kleinhofs A (2002) The barley stem rust-resistance gene Rpg1 is a novel disease-resistance gene with homology to receptor kinases. Proc Natl Acad Sci USA 99:9328–9333
Chen X (2005) Epidemiology and control of stripe rust [Puccinia striiformis f. sp tritici] on wheat. Can J of Plant Pathol 27:314–337
Chen S, Rouse MN, Zhang W, Jin Y, Akhunov E, Wei Y, Dubcovsky J (2015) Fine mapping and characterization of Sr21, a temperature-sensitive diploid wheat resistance gene effective against the Puccinia graminis f. sp. tritici Ug99 race group. Theor Appl Genet 128:645–656
Chen S, Guo Y, Briggs J, Dubach F, Chao S, Zhang W, Rouse MN, Dubcovsky J (2018a) Mapping and characterization of wheat stem rust resistance genes SrTm5 and Sr60 from Triticum monococcum. Theor Appl Genet 131:625–635
Chen S, Zhang W, Bolus S, Rouse MN, Dubcovsky J (2018b) Identification and characterization of wheat stem rust resistance gene Sr21 effective against the Ug99 race group at high temperature. PLoS Genet 14:e1007287
Chen S, Rouse MN, Zhang W, Zhang X, Guo Y, Briggs J, Dubcovsky J (2020) Wheat gene Sr60 encodes a protein with two putative kinase domains that confers resistance to stem rust. New Phytol 225:948–959
Chhuneja P, Kaur S, Garg T, Ghai M, Kaur S, Prashar M, Bains NS, Goel RK, Keller B, Dhaliwal HS, Singh K (2008) Mapping of adult plant stripe rust resistance genes in diploid A genome wheat species and their transfer to bread wheat. Theor Appl Genet 116:313–324
DeWitt N, Guedira M, Lauer E, Sarinelli M, Tyagi P, Fu D, Hao Q, Murphy JP, Marshall D, Akhunova A (2020) Sequence-based mapping identifies a candidate transcription repressor underlying awn suppression at the B1 locus in wheat. New Phytol 225:326–339
Distelfeld A, Tranquilli G, Li C, Yan L, Dubcovsky J (2009) Genetic and molecular characterization of the VRN2 loci in tetraploid wheat. Plant Physiol 149:245–257
Dubcovsky J, Luo M, Dvorak J (1995) Differentiation between homoeologous chromosomes 1A of wheat and 1Am of Triticum monococcum and its recognition by the wheat Ph1 locus. Proc Natl Acad Sci USA 92:6645–6649
Dubcovsky J, Luo M-C, Zhong G-Y, Bransteitter R, Desai A, Kilian A, Kleinhofs A, Dvořák J (1996) Genetic map of diploid wheat, Triticum monococcum L., and its comparison with maps of Hordeum vulgare L. Genetics 143:983–999
Dvorak J, McGuire PE, Cassidy B (1988) Apparent sources of the A genomes of wheats inferred from polymorphism in abundance and restriction fragment length of repeated nucleotide sequences. Genome 30:680–689
Fu D, Uauy C, Distelfeld A, Blechl A, Epstein L, Chen X, Sela H, Fahima T, Dubcovsky J (2009) A novel kinase-START gene confers temperature-dependent resistance to wheat stripe rust. Science 323:1357–1360
He F, Pasam R, Shi F, Kant S, Keeble-Gagnere G, Kay P, Forrest K, Fritz A, Hucl P, Wiebe K (2019) Exome sequencing highlights the role of wild-relative introgression in shaping the adaptive landscape of the wheat genome. Nat Genet 51:896–904
Hovmøller MS, Walter S, Bayles RA, Hubbard A, Flath K, Sommerfeldt N, Leconte M, Czembor P, Rodriguez-Algaba J, Thach T, Hansen JG, Lassen P, Justesen AF, Ali S, de Vallavieille-Pope C (2015) Replacement of the European wheat yellow rust population by new races from the centre of diversity in the near-Himalayan region. Plant Pathol 65:402–411
Hsam SLK, Huang X, Ernst F, Hartl L, Zeller F (1998) Chromosomal location of genes for resistance to powdery mildew in common wheat (Triticum aestivum L. em Thell.). 5. Alleles at the Pm1 locus. Theor Appl Genet 96:1129–1134
Hurni S, Scheuermann D, Krattinger SG, Kessel B, Wicker T, Herren G, Fitze MN, Breen J, Presterl T, Ouzunova M, Keller B (2015) The maize disease resistance gene Htn1 against northern corn leaf blight encodes a wall-associated receptor-like kinase. Proc Natl Acad Sci USA 112:8780–8785
Johnson B, Dhaliwal H (1976) Reproductive isolation of Triticum boeoticum and Triticum urartu and the origin of the tetraploid wheats. Am J Bot 63:1088–1094
Jones JDG, Dang JL (2006) The plant immune system. Nature 444:323–329
Klymiuk V, Yaniv E, Huang L, Raats D, Fatiukha A, Chen S, Feng L, Frenkel Z, Krugman T, Lidzbarsky G, Chang W, Jaaskelainen MJ, Schudoma C, Paulin L, Laine P, Bariana H, Sela H, Saleem K, Sorensen CK, Hovmoller MS, Distelfeld A, Chalhoub B, Dubcovsky J, Korol AB, Schulman AH, Fahima T (2018) Cloning of the wheat Yr15 resistance gene sheds light on the plant tandem kinase-pseudokinase family. Nat Commun 9:3735
Kolmer JA, Anderson JA, Flor JM (2010) Chromosome location, linkage with simple sequence repeat markers, and leaf rust resistance conditioned by gene Lr63 in wheat. Crop Sci 50:2392–2395
Konieczny A, Ausubel FM (1993) A procedure for mapping Arabidopsis mutations using co-dominant ecotype-specific PCR-based markers. Plant J 4:403–410
Krasileva KV, Vasquez-Gross H, Howell T, Bailey P, Paraiso F, Clissold L, Simmonds J, Ramirez-Gonzalez RH, Wang X, Borrill P, Fosker C, Ayling S, Phillips A, Uauy C, Dubcovsky J (2017) Uncovering hidden variation in polyploid wheat. Proc Natl Acad of Sci USA 114:E913–E921
Lamari L (2008) ASSESS 2.0: image analysis software for plant disease quantification. 2008. St Paul, MN: Amer Phytopathological Society
Lan C, Hale IL, Herrera-Foessel SA, Basnet BR, Randhawa MS, Huerta-Espino J, Dubcovsky J, Singh RP (2017) Characterization and mapping of leaf rust and stripe rust resistance loci in hexaploid wheat lines UC1110 and PI610750 under Mexican environments. Front Plant Sci 8:1450
Li J, Dundas I, Dong C, Li G, Trethowan R, Yang Z, Hoxha S, Zhang P (2020) Identification and characterization of a new stripe rust resistance gene Yr83 on rye chromosome 6R in wheat. Theor Appl Genet 133:1095–1107
Liu X (1988) Study on the yellow rust resistance to common wheat (T. aetivum). Plant Prot 15:33–39
Liu TG, Peng YL, Chen WQ, Zhang ZY (2010) First detection of virulence in Puccinia striiformis f. sp. tritici in China to resistance genes Yr24 (=Yr26) present in wheat cultivar Chuanmai 42. Plant Dis 94:1163–1163
Lowe I, Jankuloski L, Chao S, Chen X, See D, Dubcovsky J (2011) Mapping and validation of QTL which confer partial resistance to broadly virulent post-2000 North American races of stripe rust in hexaploid wheat. Theor Appl Genet 123:143–157
Luo M-C, Yang Z-L, Kota RS, Dvořák J (2000) Recombination of chromosomes 3Am and 5Am of Triticum monococcum with homeologous chromosomes 3A and 5A of wheat: The distribution of recombination across chromosomes. Genetics 154:1301–1308
Ma L-J, Qiao J-x, Kong X-y, Wang J-j, Xu X-m, Hu X-p (2016) An improved method for RNA extraction from urediniospores of and wheat leaves infected by an obligate fungal pathogen, Puccinia striiformis f. sp. tritici. J Integr Agr 15:1293–1303
Marais GF, Mccallum B, Snyman JE, Pretorius ZA, Marais AS (2005a) Leaf rust and stripe rust resistance genes Lr54 and Yr37 transferred to wheat from Aegilops kotschyi. Plant Breeding 124:538–541
Marais GF, Pretorius ZA, Wellings CR, Mccallum B, Marais AS (2005b) Leaf rust and stripe rust resistance genes transferred to common wheat from Triticum dicoccoides. Euphytica 143:115–123
Marchal C, Zhang J, Zhang P, Fenwick P, Steuernagel B, Adamski NM, Boyd L, McIntosh R, Wulff BBH, Berry S, Lagudah E, Uauy C (2018) BED-domain-containing immune receptors confer diverse resistance spectra to yellow rust. Nat Plants 4:662–668
Martin GB, Brommonschenkel SH, Chunwongse J, Frary A, Ganal MW, Spivey R, Wu T, Earle ED, Tanksley SD (1993) Map-based cloning of a protein kinase gene conferring disease resistance in tomato. Science 262:1432–1436
Milus EA, Kristensen K, Hovmøller MS (2009) Evidence for increased aggressiveness in a recent widespread strain of Puccinia striiformis f. sp. tritici causing stripe rust of wheat. Phytopathology 99:89–94
Olmstead AL, Rhode PW (2002) The red queen and the hard reds: productivity growth in American wheat, 1800–1940. J Econ Hist 62:929–966
Pont C, Leroy T, Seidel M, Tondelli A, Duchemin W, Armisen D, Lang D, Bustos-Korts D, Goué N, Balfourier F (2019) Tracing the ancestry of modern bread wheats. Nat Genet 51:905–911
Prins R, Marais GF (1999) A genetic study of the gametocidal effect of the Lr19 translocation of common wheat. S Afr J Plant Soil 16:10–14
Qureshi N, Bariana HS, Zhang P, McIntosh R, Bansal UK, Wong D, Hayden MJ, Dubcovsky J, Shankar M (2018) Genetic relationship of stripe rust resistance genes Yr34 and Yr48 in wheat and identification of linked KASP markers. Plant Dis 102:413–420
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G, Mesirov JP (2011) Integrative genomics viewer. Nat Biotechnol 29:24–26
Saintenac C, Zhang W, Salcedo A, Rouse MN, Trick HN, Akhunov E, Dubcovsky J (2013) Identifcation of wheat gene Sr35 that confers resistance to Ug99 stem rust race group. Science 341:783–786
Schmolke M, Mohler V, Hartl L, Zeller FJ, Hsam SLK (2012) A new powdery mildew resistance allele at the Pm4 wheat locus transferred from einkorn (Triticum monococcum). Mol Breeding 29:449–456
Shi AN, Leath S, Murphy JP (1998) A major gene for powdery mildew resistance transferred to common wheat from wild einkorn wheat. Phytopathology 88:144–147
Sodkiewicz W, Strzembicka A, Apolinarska B (2008) Chromosomal location in triticale of leaf rust resistance genes introduced from Triticum monococcum. Plant Breeding 127:364–367
Song WY, Wang GL, Chen LL, Kim HS, Pi LY, Holsten T, Gardner J, Wang B, Zhai WX, Zhu LH, Fauquet C, Ronald P (1995) A receptor kinase-like protein encoded by the rice disease resistance gene, Xa21. Science 270:1804–1806
Steuernagel B, Periyannan SK, Hernandez-Pinzon I, Witek K, Rouse MN, Yu G, Hatta A, Ayliffe M, Bariana H, Jones JD, Lagudah ES, Wulff BB (2016) Rapid cloning of disease-resistance genes in plants using mutagenesis and sequence capture. Nat Biotechnol 34:652–655
Walkowiak S, Gao LL, Monat C, Haberer G, Kassa MT, Brinton J, Ramirez-Gonzalez RH, Kolodziej MC, Delorean E, Thambugala D, Klymiuk V, Byrns B, Gundlach H, Bandi V, Siri JN, Nilsen K, Aquino C, Himmelbach A, Copetti D, Ban T, Venturini L, Bevan M, Clavijo B, Koo DH, Ens J, Wiebe K, N’Diaye A, Fritz AK, Gutwin C, Fiebig A, Fosker C, Fu BX, Accinelli GG, Gardner KA, Fradgley N, Gutierrez-Gonzalez J, Halstead-Nussloch G, Hatakeyama M, Koh CS, Deek J, Costamagna AC, Fobert P, Heavens D, Kanamori H, Kawaura K, Kobayashi F, Krasileva K, Kuo T, McKenzie N, Murata K, Nabeka Y, Paape T, Padmarasu S, Percival-Alwyn L, Kagale S, Scholz U, Sese J, Juliana P, Singh R, Shimizu-Inatsugi R, Swarbreck D, Cockram J, Budak H, Tameshige T, Tanaka T, Tsuji H, Wright J, Wu JZ, Steuernagel B, Small I, Cloutier S, Keeble-Gagnere G, Muehlbauer G, Tibbets J, Nasuda S, Melonek J, Hucl PJ, Sharpe AG, Clark M, Legg E, Bharti A, Langridge P, Hall A, Uauy C, Mascher M, Krattinger SG, Handa H, Shimizu KK, Distelfeld A, Chalmers K, Keller B, Mayer KFX, Poland J, Stein N, McCartney CA, Spannagl M, Wicker T, Pozniak CJ (2020) Multiple wheat genomes reveal global variation in modern breeding. Nature 588:277–283
Wang X, Zafian P, Choudhary M, Lawton M (1996) The PR5K receptor protein kinase from Arabidopsis thaliana is structurally related to a family of plant defense proteins. Proc Natl Acad Sci USA 93:2598–2602
Wang Y, Cao Q, Zhang J, Wang S, Chen C, Wang C, Zhang H, Wang Y, Ji W (2020) Cytogenetic analysis and molecular marker development for a new wheat–Thinopyrum ponticum 1Js (1D) disomic substitution line with resistance to stripe rust and powdery mildew. Front Plant Sci 11:1282
Wang H, Zou S, Li Y, Lin F, Tang D (2020) An ankyrin-repeat and WRKY-domain-containing immune receptor confers stripe rust resistance in wheat. Nat commun 11:1–11
Yan L, Loukoianov A, Blechl A, Tranquilli G, Ramakrishna W, SanMiguel P, Bennetzen JL, Echenique V, Dubcovsky J (2004) The wheat VRN2 gene is a flowering repressor down-regulated by vernalization. Science 303:1640–1644
Zhang W, Chen S, Abate Z, Nirmala J, Rouse MN, Dubcovsky J (2017) Identification and characterization of Sr13, a tetraploid wheat gene that confers resistance to the Ug99 stem rust race group. Proc Natl Acad Sci USA 114:E9483-9492
Zhang C, Huang L, Zhang H, Hao Q, Lyu B, Wang M, Epstein L, Liu M, Kou C, Qi J, Chen F, Li M, Gao G, Ni F, Zhang L, Hao M, Wang J, Chen X, Luo MC, Zheng Y, Wu J, Liu D, Fu D (2019) An ancestral NB-LRR with duplicated 3’UTRs confers stripe rust resistance in wheat and barley. Nat Commun 10:4023
Acknowledgments
Work at JD laboratory was supported by the Howard Hughes Medical Institute and by the Agriculture and Food Research Initiative Competitive Grant 2017-67007-25939 (WheatCAP) from the USDA National Institute of Food and Agriculture (NIFA). Work at SC laboratory was supported by Peking University Institute of Advanced Agricultural Sciences project 2019-0401-SC-01.
Author information
Authors and Affiliations
Contributions
SC, JH and TS performed most of the experimental work. SC analyzed the data and wrote the first version of the manuscript. JH performed the initial sequence analysis. LH, HnaL and JL contributed primers development. HyuL and SB contributed the phenotyping experiments. CZ performed the genotyping of Mediterranean. JD supervised the project and generated the final version of the manuscript. All authors revised the manuscript and provided suggestions.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Additional information
Communicated by Hermann Buerstmayr.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chen, S., Hegarty, J., Shen, T. et al. Stripe rust resistance gene Yr34 (synonym Yr48) is located within a distal translocation of Triticum monococcum chromosome 5AmL into common wheat. Theor Appl Genet 134, 2197–2211 (2021). https://doi.org/10.1007/s00122-021-03816-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00122-021-03816-z