Abstract
Aims/hypothesis
Despite the current pandemic of metabolic diseases, our understanding of the diverse nature of the development of metabolic alterations in people who eat a high-fat diet (HFD) is still poor. We recently demonstrated a cardio-metabolic adaptation in mice fed an HFD, which was characterised by a specific gut and periodontal microbiota profile. Since the severity of hepatic disease is characterised by specific microRNA (miRNA) signatures and the gut microbiota is a key driver of both hepatic disease and miRNA expression, we analysed the expression of three hepatic miRNA and studied their correlation with hepatic triacylglycerol content and gut microbiota.
Methods
Two cohorts of C57BL/6 4-week-old wild-type (WT) male mice (n = 62 and n = 96) were fed an HFD for 3 months to provide a model of metabolic adaptation. Additionally 8-week-old C57BL/6 mice, either WT or of different genotypes, with diverse gut microbiota (ob/ob, Nod1, Cd14 knockout [Cd14KO] and Nod2) or without gut microbiota (axenic mice) were fed a normal chow diet. Following which, glycaemic index, body weight, blood glucose levels and hepatic triacylglycerol levels were measured. Gut (caecum) microbiota taxa were analysed by pyrosequencing. To analyse hepatic miRNA expression, real-time PCR was performed on total extracted miRNA samples. Data were analysed using two-way ANOVA followed by the Dunnett’s post hoc test, or by the unpaired Student’s t test. A cluster analysis and multivariate analyses were also performed.
Results
Our results demonstrated that the expression of miR-181a, miR-666 and miR-21 in primary murine hepatocytes is controlled by lipopolysaccharide in a dose-dependent manner. Of the gut microbiota, Firmicutes were positively correlated and Proteobacteria and Bacteroides acidifaciens were negatively correlated with liver triacylglycerol levels. Furthermore, the relative abundance of Firmicutes was negatively correlated with hepatic expression of miR-666 and miR-21. In contrast, the relative abundance of B. acidifaciens was positively correlated with miR-21.
Conclusions/interpretation
We propose the involvement of hepatic miRNA, liver triacylglycerols and gut microbiota as a new triad that underlies the molecular mechanisms by which gut microbiota governs hepatic pathophysiology during metabolic adaptation to HFD.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The current pandemic of metabolic diseases, such as obesity and type 2 diabetes, cannot be completely explained by genetic alterations and the growing consumption of a Western diet [1, 2]. Moreover, obesity is not an inevitable consequence of a fat-rich diet, since both people and mice consuming a high-fat diet (HFD) can display the opposite metabolic outcome, suggesting the existence of metabolic adaptations in some individuals [3, 4]. Among the factors that may affect the metabolic processes on an individual basis [5] are the gut microbiota [6], the impact of which on host metabolism has been established [7,8,9]. We previously found that adaptation to obesity in terms of insulin sensitivity was characterised by a specific gut microbiota profile in insulin-resistant vs insulin-sensitive obese individuals [10]. In addition, we showed that divergent gut microbiota profiles characterise the different metabolic phenotypes developed during metabolic adaptation to an HFD in mice [3, 4]. Recently, we reported that the periodontal microbiota profile correlates with cardio-metabolic adaptations to an HFD in mice [11].
Furthermore, along with xenobiotics [12], diet is considered to be the strongest modulator of gut microbiota [13]. Evidence from hepatic transcript profiles in mice has suggested that liver pathophysiology may be affected during metabolic adaptation to HFD in mice [14]. The existence of a gut–liver axis has been previously demonstrated and the liver is the organ in which xenobiotic metabolism occurs, especially with regard to our capacity of responding to gut microbial antigens [15].
Moreover, the alteration of gut microbiota, termed dysbiosis, is an additional causal factor in the development of hepatic steatosis [16], a condition that involves the accumulation of hepatic triacylglycerols, which is a common feature of metabolic disease [17]. Indeed, the different stages of hepatic diseases, including steatosis, hepatitis and hepatocellular carcinoma (HCC), are identifiable by a precise microRNA (miRNA) signature [18]. miRNA are pleiotropic modulators of gene expression [19] that have been shown to be under the control of gut microbiota [20]. Some miRNA, for example miR-181a, miR-666 and miR-21, are specifically involved in the modulation of liver pathophysiology [18, 21].
In the present study, we aimed to elucidate the gut microbiota profiles that are associated with metabolic adaptations to HFD in mice. We also aimed to investigate the associations between specific taxa of gut microbiota and hepatic expression of miR-181a, miR-666 and miR-21 in mouse models of hepatic steatosis. In addition, we explored the link between miRNA expression levels and metabolic parameters, such as glucose tolerance, body weight and fasting blood glucose.
Methods
Animal models and dietary treatment
All animal experimental procedures were approved by the local ethical committee of Rangueil University Hospital (Toulouse, France). All experimenters were blind to group assignment and outcome assessment. No data, samples or animals were excluded from this study.
Animal model for metabolic adaptation to HFD
An initial cohort of 62 and a second cohort of 96 C57BL/6 4-week-old wild-type (WT) male mice (Charles River, L'Arbresle, France) were fed an HFD (∼72% fat [corn oil and lard], 28% protein and <1% carbohydrate; SAFE, Augy, France) for 3 months [4]. Mice were housed in groups (10–11 mice per cage) in a specific pathogen-free controlled environment (inverted 12 h light cycle; lights off at 10:00 hours). Mice were killed by cervical dislocation after a 6 h fast. Tissues were collected and snap frozen in liquid nitrogen.
Axenic, WT, ob/ob, Nod1, Cd14KO and Nod2 mice
Eight-week-old C57BL/6 mice (Charles River) either WT or of different genotypes, with diverse gut microbiota (ob/ob, Nod1, Cd14 knockout [Cd14KO] and Nod2) or without gut microbiota (axenic mice) were fed a normal chow diet. Mice were housed in groups (4–6 mice per cage) in a specific pathogen-free controlled environment (inverted 12 h light cycle; lights off at 10:00 hours). After the mice were killed, the livers were collected and snap frozen in liquid nitrogen and stored at −80°C until analysis.
GTT and hepatic triacylglycerol measurement
After 3 months of HFD, an IPGTT or OGTT were performed. Briefly, for the IPGTT, 6 h fasted mice were injected with glucose (1 g/kg) into the peritoneal cavity, as previously described [22]. An OGTT was performed via oral administration of glucose (1.5 mg/g) following a 6 h fast. Blood glucose levels were measured 30 min before glucose administration and at 0, 15, 30, 60, 90 and 120 min following glucose challenge.
For both IPGTT and OGTT, the glycaemic index was calculated as the sum of the blood glucose values (mmol/l) divided by the total time of the curve in min to present value in mmol/l × min, or additionally multiplied by 1000 to give value in μmol/l × min.
Liver triacylglycerol content was measured by a colorimetric assay using free glycerol and triacylglycerol reagents (Sigma Aldrich, St Louis, MO, USA) and the plate was read using the Multiskan Spectrum plate reader and the SkanIt RE software (both Thermo Labsystems, Beverly, MA, USA).
Taxonomic analysis of gut microbiota by pyrosequencing
Caecum total DNA was extracted as previously described [4]. The whole 16S bacterial ribosomal RNA V2 region was targeted by the 28F-519R primers (designed by Research and Testing Laboratory [www.researchandtesting.com/, accessed 1 September 2016; Lubbock, TX, USA]) and pyrosequenced by the 454 GS FLX+ system (Roche, Branford, CT, USA) at the Research and Testing Laboratory. On average, 3000 sequences were generated per mouse. The minimum number of sequences guaranteed per mouse was 1606 (n = 62).
Preparation of murine primary hepatocytes and lipopolysaccharide stimulation
Hepatocytes were isolated by a non-recirculating collagenase perfusion through the portal vein of anesthetised 8-week-old C57BL/6 WT or Cd14KO male mice fed a normal chow diet. Isolated cells were filtered through a 100 μm pore mesh nylon filter and cultured (2.5 × 106 cells per well) onto 96-well plates in DMEM (BE12-614F; Lonza, Levallois, France) supplemented with 10% (vol./vol.) FCS, 1% (vol./vol.) penicillin/streptomycin and 0.2 nmol/l l-glutamine. After 12 h, the medium was replaced with medium plus industrially purified lipopolysaccharide (LPS; Sigma Aldrich, St Louis, MO, USA) either from the proinflammatory Escherichia coli serotype O55:B5 [22], or the E. coli strain O111:B4, which stimulates human hepatocytes [23]. Two doses of LPS were tested: 10 ng/ml (low dose) and 100 ng/ml (high dose), and cells were stimulated for 6 h. Experiments were performed in quadruplicates (control) or pooled duplicates (LPS).
miRNA-based quantitative PCR
Real-time PCR for miRNA expression was performed on total miRNA extracted from cells or livers using the miRNeasy kit (Qiagen, Courtaboeuf, France). The expression of each miRNA was normalised to U6 small nuclear RNA (snRNA) expression [23]. For in vitro analyses, cells were directly harvested into Qiazol solution, which was provided with the miRNeasy kit. For ex vivo analyses, frozen pieces of liver were put directly into the Qiazol and total miRNA was extracted following the manufacturer’s protocol. Expression values were quantified using the \( {2}^{-\Delta \Delta {\mathrm{C}}_{\mathrm{t}}} \) method [4].
Hepatic microarray analysis
Total RNA was isolated from the right lobe of the liver using TRIzol (Life Technologies, Villebon sur Yvette, France) according to the manufacturer’s protocol. Preparation, labelling and hybridisations of cDNA were performed as per the manufacturer’s protocol. Samples were analysed using an Agilent SurePrint G3 Mouse GE 8 × 60K chip (design 028005; Agilent Technologies, Courtaboeuf, France). The hybridised microarrays were washed and scanned using an Agilent G2505C scanner. Data were extracted from the scanned image using the Agilent Feature Extraction software version 10.10.1.1. All of these steps were performed at the GENOTOUL GeT-TRIX facility at the French National Institute for Agricultural Research (INRA; Toulouse, France).
Statistical analysis
Statistical analyses were performed by two-way ANOVA followed by the Dunnett’s post hoc test, or using the unpaired Student’s t test, using GraphPad Prism version 7.00 for Windows 7 (GraphPad, San Diego, CA, USA). A p value <0.05 was considered significant. Cluster analysis was performed using PermutMatrixEN software (http://download.cnet.com/PermutMatrix/3000-20432_4-75325452.html, accessed 20 June 2016) [24]. Multivariate analyses were performed using the Spearman correlation coefficient and p values were adjusted using the Benjamini–Hochberg correction (available at www.marum.de/Binaries/Binary745/BenjaminiHochberg.xlsx, accessed 12 November 2016). A String analysis was performed to study the network of genes targeted by miR-21; these genes were identified using the software miRTarBase (http://mirtarbase.mbc.nctu.edu.tw/, accessed 20 June 2016) [25].
Results
Hepatic triacylglycerol content correlates with metabolic adaptation to HFD feeding in mice
We aimed to investigate the effect of diabetogenic/non-obesogenic HFD in hepatic steatosis. To do this, hepatic triacylglycerol content was analysed in a murine model of metabolic adaptation to HFD [4, 11]. Glycaemic index (evaluated using IPGTT; Fig. 1a,c), body weight (Fig. 1a,b; trend over 3 months reported in ESM Fig. 1) and liver triacylglycerol content (Fig. 1b,c) were significantly correlated with each other. Liver triacylglycerol content showed high interindividual variation, which is typical of this animal model and suggests that liver triacylglycerol levels are dependent on the metabolic response of each mouse to HFD feeding (Fig. 1). Thus, we further confirm our previous findings from an analysis of hepatic gene expression during metabolic adaptation to HFD in mice, reinforcing the reproducibility of the animal model used in this study [14].
Metabolic diversity in C57BL/6 4-week-old WT male mice (n = 62) fed an HFD for 3 months. Three main metabolic variables are shown: glycaemic index (−30 to 90 min) during an IPGTT [4], body weight and liver triacylglycerols. Correlations between (a) glycaemic index vs body weight, (b) liver triacylglycerols vs body weight and (c) glycaemic index vs liver triacylglycerols
Specific bacterial groups in the gut microbiota are associated with hepatic triacylglycerol content during metabolic adaptation to HFD feeding in mice
We previously demonstrated that a specific gut microbiota profile characterises the mouse model of metabolic adaptation to HFD used in this study [4]. Hence, since the gut–liver axis manages our capacity to sense gut microbes [15], we investigated whether bacterial groups from the gut microbiota may be associated with the diversity of hepatic triacylglycerol content in HFD-fed mice. We sequenced the gut microbiota from all mice and performed a non a priori-based analysis to identify putative correlations between bacterial phyla and metabolic variables of interest. Out of the total 12 phyla identified, three were present in all mice: Firmicutes, Proteobacteria and Bacteroidetes (Table 1). As presented in Fig. 2a, all of the metabolic variables of interest (glycaemic index, body weight and liver triacylglycerol) were positively associated with Firmicutes, and negatively associated with Proteobacteria and Bacteroidetes. We challenged these associations by performing single linear regression analyses and found that Firmicutes showed a significant positive correlation with liver triacylglycerol content (Fig. 2b) and Proteobacteria showed a significant negative correlation with liver triacylglycerols (Fig. 2c). In contrast, single linear regression analysis revealed that Bacteroidetes showed a non-significant correlation with liver triacylglycerol content (Fig. 2d).
Integration data analysis between metabolic and metagenomic variables during metabolic adaptation to an HFD. (a) Heat map between metagenomics data (phylum level) and metabolic variables (glycaemic index, body weight and liver triacylglycerols [TG]). The dendrogram on the left-hand side of the heat map identifies clusters of metabolic and metagenomic variables; the dendrogram above the heat map identifies clusters of mice (the identification numbers are presented below this dendrogram). Cluster analysis was performed using PermutMatrixEN software [24]. Correlation matrices for the overall variables (right-hand side of the heat map) and mice (below the heat map) are also shown. Correlation analyses between liver triacylglycerols and the relative abundance (%) of (b) Firmicutes, (c) Proteobacteria, (d) Bacteroidetes and (e) B. acidifaciens. n = 62
We also tested whether specific bacterial species were associated with liver triacylglycerol levels. Of the 421 bacterial species identified in the gut microbiota, 24 were present in all mice (Table 1). Among those that were significantly correlated with liver triacylglycerol content, Bacteroides acidifaciens (belonging to the Bacteroidetes phylum) showed the most robust negative correlation (Fig. 2e).
In summary, these data identify that Firmicutes, Proteobacteria and B. acidifaciens of the gut microbiota correlate with hepatic triacylglycerol content during metabolic adaptation to HFD feeding in mice.
Bacterial antigens drive hepatic miRNA expression in vitro
Gut microbiota dysbiosis can drive liver disease [26]; miRNA are known modulators of liver pathophysiology [18] and the gut microbiota modulates miRNA expression [20]. Therefore, we tested whether liver triacylglycerol content was associated with both gut microbiota and hepatic miRNA expression in a mouse model of metabolic adaptation to an HFD. First, we tested the capacity of bacterial antigens to modulate hepatic miRNA expression in vitro. The expression of three miRNA involved in the modulation of liver pathophysiology [18, 21], miR-181a, miR-666 and miR-21, was under the dose-dependent control of proinflammatory LPS from E. coli O55:B5 in primary WT murine hepatocytes. Also, 10 ng/ml LPS from the E. coli O111:B4 serotype, a known stimulator of nitric oxide synthase (NOS) in human hepatocytes [27], significantly decreased the expression of miR-181a, whilst treatment with 100 ng/ml showed a trend towards decreased expression of miR-181a and both 10 ng/ml and 100 ng/ml non-significantly decreased miR-21 expression. There was no significant effect of E. coli O111:B4 LPS on miR-666 levels (Fig. 3a). This LPS-induced modulation of miRNA expression was highly specific since it was not observed in primary hepatocytes from Cd14KO mice (Fig. 3b), known not to respond to LPS [22, 28].
Real-time PCR miRNA analysis in 6 h LPS-stimulated primary murine hepatocytes from either WT or Cd14KO mice. miR-181a, miR-666 and miR-21 expression in primary murine hepatocytes from either (a) WT or (b) Cd14KO C57BL/6 male mice stimulated for 6 h with the purified LPS at various doses. The expression of each miRNA was normalised to U6 snRNA gene expression. Data are shown as mean ± SEM. Black bars, control (PBS [vehicle]); white bars, 10 ng/ml E. coli O55:B5 LPS ; dark grey bars, 100 ng/ml E. coli O55:B5 LPS; light grey bars, 10 ng/ml E. coli O111:B4 LPS; white bars with diagonal stripes, 100 ng/ml E. coli O111:B4 LPS. Two mice were used in two independent experiments to prepare primary hepatocytes; 2.5 × 106 cells were plated per well; all experiments were performed in quadruplicates (control) and pooled duplicates (treatment). *p < 0.05, **p < 0.01 vs control; two-way ANOVA and Dunnett’s post hoc test
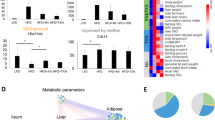
We then analysed the hepatic expression of the above miRNA in animal models of metabolic diseases characterised by gut microbiota dysbiosis, fed a normal chow diet (Fig. 4). First, we analysed the diversity of liver triacylglycerol content between the different models (Fig. 4a) and confirmed that the liver of axenic mice had lower triacylglycerol content than all other models [29]. With regard to miRNA expression, there was less variation in miR-181a expression between models suggesting this miRNA did not undergo metabolic adaptation and that there was no link with liver triacylglycerol levels (Fig. 4b). In contrast, the regulation of miR-666 and, to a larger extent, miR-21 expression was significantly altered depending on the animal model used (Fig. 4c,d).
Analysis of triacylglycerol content and miRNA expression in the livers of 8-week-old C57BL/6 mice as models of gut microbiota dysbiosis: WT axenic, WT conventional (WT), ob/ob, Nod1, Cd14KO and Nod2 mice fed normal chow. (a) Liver triacylglycerol content in each mouse model. Hepatic expression of (b) miR-181a, (c) miR-666 and (d) miR-21. The expression of each miRNA was normalised to U6 snRNA gene expression. Data are mean ± SEM; *p < 0.05, **p < 0.01, ***p < 0.001; Student t test
Together these data show that: (1) in vitro, bacterial antigens control hepatic miRNA expression in a dose-dependent manner; and (2) in vivo, hepatic miRNA expression is associated with liver triacylglycerol content during gut microbiota dysbiosis.
Hepatic miRNA expression is significantly associated with liver triacylglycerol content and with Firmicutes and B. acidifaciens relative abundance during metabolic adaptation to HFD feeding in mice
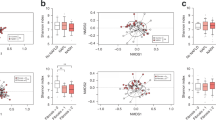
The hepatic expression of miR-181a, miR-666 and miR-21 was studied in our model of metabolic adaptation to HFD. We observed a high diversity of expression for all three miRNA (Fig. 5a–c). Furthermore, miR-181a was not significantly correlated with liver triacylglycerols (Fig. 5d), whereas miR-666 and, to a larger extent, miR-21 showed significant correlations with this variable (Fig. 5e,f). Given that Firmicutes modulation is a marker of dysbiosis during metabolic adaptation (as we have already shown [4, 11]), we analysed whether this phylum may be correlated with hepatic miRNA expression in this model, along with B. acidifaciens. Of note, miR-181a was not significantly correlated with Firmicutes or B. acidifaciens relative abundance (Fig. 5g,j). In contrast, miR-666 was significantly correlated with Firmicutes but not with relative abundance of B. acidifaciens (Fig. 5h,k), whereas miR-21 was significantly correlated with both taxa (Fig. 5i,l).
Hepatic miRNA expression analysis and correlation with liver triacylglycerols and gut microbiota during metabolic adaptation to HFD in mice. A cohort (n = 62) of C57BL/6 4-week-old WT male mice was fed an HFD for 3 months. Expression of (a) miR-181a, (b) miR-666 and (c) miR-21 was analysed. (d–l) Correlations between miRNA expression with (d–f) liver triacylglycerols, (g–i), Firmicutes relative abundance and (j–l) B. acidifaciens relative abundance. For (e, f), (h, i) and (k, l), significance determined by Spearman correlation adjusted according to the Benjamini–Hochberg correction for multiple comparisons (false-discovery rate <0.05)
In summary, hepatic miRNA expression is associated with both variations in liver triacylglycerol content and specific taxa of the gut microbiota during metabolic adaptation to HFD (Fig. 2 and Fig. 5).
Hepatic miR-21-targeted genes and their metabolic relationships
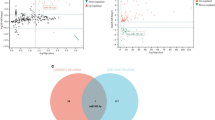
To provide insight on the functional consequences of hepatic miR-21 expression on hepatic metabolic pathways, we performed a String analysis [30] of the network of genes targeted by this miRNA, which were identified by miRTarBase [25]. Notably, some genes targeted by miR-21 are involved in pathways associated with the hepatocyte apoptotic process, non-alcoholic fatty liver disease (NAFLD), insulin signalling and the proinflammatory RIG-I-like receptor signalling pathway (Fig. 6). This pathway regulates the production of proinflammatory cytokines, such as TNF-α and IL-8, in response to microbial antigens (i.e. LPS). Interestingly, we recently found that the RIG-I-like receptor signalling pathway is modulated in the periodontitis-induced periodontal dysbiosis during cardio-metabolic adaptation to HFD [11]. This result suggests an important association between the RIG-I-like receptor signalling microbial pathway and metabolic adaptation to an HFD in mice.
Next, we investigated whether genes targeted by miR-21 may correlate with key metabolic variables: glucose tolerance, body weight and fasting blood glucose. To corroborate our observations in the first HFD-fed murine cohort, the correlation analysis was conducted in a second independent murine cohort (n = 96) of metabolic adaptation to HFD, in which we performed a liver microarray analysis. Importantly, the expression of Fas, Pdcd4, Reck and Rmnd5a was significantly correlated with the glycaemic index evaluated during an OGTT (Fig. 7a–d). Moreover, the expression of Akt1, Fadd, Pten, Reck and Tns1 was significantly correlated with body weight (Fig. 7e–i). Additionally, the expression of Akt1 and Rmnd5a was significantly correlated with fasting blood glucose (Fig. 7j,k).
Correlations of genes targeted by miR-21 with metabolic variables during metabolic adaptation to a HFD in mice. (a–k) Correlation of specified genes with (a–d) glycaemic index (−30 to 120 min) during an OGTT, (e–i) body weight and (j, k) 6 h fasting blood glucose in an independent cohort of C57BL/6 4-week-old WT male mice (n = 96) fed a diabetogenic/non-obesogenic HFD for 3 months
Together these data suggest that the association of gut microbiota with hepatic miR-21 expression may have metabolic consequences via the modulation of genes targeted by miR-21.
Discussion
In this study we report that, beyond the diversity already observed for blood glucose and body weight [3, 4, 11], liver triacylglycerol content is characterised by a high heterogeneity according to the individual response of mice to a diabetogenic/non-obesogenic HFD. Furthermore, liver triacylglycerol content was positively associated with the relative abundance of Firmicutes, and negatively associated with hepatic miR-21 expression and the relative abundance of Proteobacteria and B. acidifaciens.
Our finding that hepatic steatosis follows metabolic diversity on an individual basis, in mice fed a diabetogenic/non-obesogenic HFD, confirms our previous observations where hepatic lipid metabolism in this model, evaluated at the level of gene expression, was shown to be modulated according to the response to an HFD [14]. Therefore, our results reinforce the reproducibility of this animal model.
The observed positive association between hepatic triacylglycerol content and the relative abundance of Firmicutes is in contrast with the results of Henao-Mejia et al [26]. The authors found a reduction of Firmicutes in a model for inflammasome-mediated dysbiosis, regulating the progression of NAFLD and obesity [31]. However, the murine model used in our study is very different from the one used by Henao-Mejia and colleagues; we used C57BL/6 WT mice, whereas these authors used C56BL/6 Nlrp6 KO mice. Therefore, it is likely that the different genetic backgrounds and genotypes of these mouse models account for disparity in the manifestations of dysbiosis, and are also responsible for the observed colitogenic phenotype. Differences in murine models may also explain the discrepancies found in the literature with regard to B. acidifaciens. This bacterium has recently been shown to be associated with liver disease [32], in contrast to our findings. Again, the murine model used in our study is very different from the mouse model used by Xie et al (streptozotocin/HFD-induced non-alcoholic steatohepatitis [NASH]/HCC C57BL/6 J mice) [32]. Thus, it is likely that under two very different dietary conditions, B. acidifaciens may have different associations with hepatic pathophysiology. This explanation is corroborated by another recent study by Yang et al that showed that B. acidifaciens prevents obesity and improves insulin sensitivity in mice [33]. Hence, based on the adaptation of both mice and microbes to a fatty environment, a divergent metabolic phenotype may arise, as we also recently reported with regard to cardio-metabolic adaptation to HFD in mice [11].
From a molecular perspective, miRNA represent promising molecules that may link gut microbiota dysbiosis to metabolic outcomes. Gut microbiota can modulate intestinal miRNA expression [20]. Moreover, a specific miRNA profile defines every stage of hepatic pathophysiology during the progression of disease from NAFLD (characterised by accumulated hepatic triacylglycerols) to NASH, (characterised by inflammation and fibrosis), up to HCC [18]. Thus, taking into account the gut–liver axis with regard to our capacity to sense gut microbes [15] and the fact that gut microbiota dysbiosis can drive NAFLD [25], it is plausible to consider that, in our study microbes, or even their antigens (e.g. LPS), may affect the liver via the modulation of hepatic miRNA expression, as we have shown here in vitro.
Specifically, miR-21 regulates both regeneration [34] and progression of fibrosis [35] in the liver, and long-term (18 weeks) inhibition of miR-21 reduces both body weight and adipocyte size in aged db/db mice [36]. These data are in accordance with the definition of miR-21 as a ‘disease miRNA’ [21]. However, this definition appears to be dependent on both animal model and diet. In fact, in the absence of HFD feeding, hepatic miR-21 expression is not correlated with hepatic triacylglycerols (Fig. 4; r 2 = 0.08, p = 0.1 [correlation not shown]). Notably, there is no clear evidence in the literature as to whether miR-21 may act as a marker of hepatic fat deposition. On one hand, Wu et al recently showed that miR-21 knockdown impairs lipid accumulation [37], which is in contrast to our findings. In contrast, Ahn et al reported that lycopene inhibits hepatic steatosis via the upregulation of miR-21, which is in accordance with our findings [38].
However, during metabolic adaptation to HFD, hepatic miR-21 expression was observed to be negatively associated with liver triacylglycerol content (Fig. 5f). This evidence suggests that during the adaptation to HFD, hepatic miRNA expression follows a different pattern of regulation. We confirmed this interpretation by showing that among the genes targeted by miR-21 are Casp8 and Fadd, which are also part of the microbial RIG-I-like receptor signalling pathway. As previously mentioned, this pathway regulates the production of proinflammatory cytokines (such as TNF-α and IL-8) in response to microbial antigens (i.e. LPS). Importantly, we recently showed that the microbial RIG-I-like receptor signalling pathway is one of the most upregulated pathways during cardio-metabolic adaptation to HFD in mice [11], corroborating its implication in this phenomenon.
In conclusion, we propose a new triad linking gut microbiota, hepatic miRNA expression and liver triacylglycerol content. These findings may help to explain hepatic metabolism adaptation to an HFD in mice.
Abbreviations
- Cd14KO:
-
Cd14 knockout
- HCC:
-
Hepatocellular carcinoma
- HFD:
-
High-fat diet
- LPS:
-
Lipopolysaccharide
- miRNA:
-
MicroRNA
- NAFLD:
-
Non-alcoholic fatty liver disease
- NASH:
-
Non-alcoholic steatohepatitis
- snRNA:
-
Small nuclear RNA
- WT:
-
Wild-type
References
Fretts AM, Follis JL, Nettleton JA et al (2015) Consumption of meat is associated with higher fasting glucose and insulin concentrations regardless of glucose and insulin genetic risk scores: a meta-analysis of 50,345 Caucasians. Am J Clin Nutr 102:1266–1278
Ruchat SM, Elks CE, Loos RJ et al (2009) Association between insulin secretion, insulin sensitivity and type 2 diabetes susceptibility variants identified in genome-wide association studies. Acta Diabetol 46:217–226
Burcelin R, Crivelli V, Dacosta A, Roy-Tirelli A, Thorens B (2002) Heterogeneous metabolic adaptation of C57BL/6J mice to high-fat diet. Am J Physiol Endocrinol Metab 282:E834–842
Serino M, Luche E, Gres S et al (2012) Metabolic adaptation to a high-fat diet is associated with a change in the gut microbiota. Gut 61:543–553
Nicholson JK, Holmes E, Wilson ID (2005) Gut microorganisms, mammalian metabolism and personalized health care. Nat Rev Microbiol 3:431–438
Eckburg PB, Bik EM, Bernstein CN et al (2005) Diversity of the human intestinal microbial flora. Science 308:1635–1638
Backhed F, Ding H, Wang T et al (2004) The gut microbiota as an environmental factor that regulates fat storage. Proc Natl Acad Sci U S A 101:15718–15723
Ley RE, Backhed F, Turnbaugh P, Lozupone CA, Knight RD, Gordon JI (2005) Obesity alters gut microbial ecology. Proc Natl Acad Sci U S A 102:11070–11075
Turnbaugh PJ, Ley RE, Mahowald MA, Magrini V, Mardis ER, Gordon JI (2006) An obesity-associated gut microbiome with increased capacity for energy harvest. Nature 444:1027–1031
Serino M, Fernandez-Real JM, Garcia-Fuentes E et al (2013) The gut microbiota profile is associated with insulin action in humans. Acta Diabetol 50:753–761
Branchereau M, Reichardt F, Loubieres P et al (2016) Periodontal dysbiosis linked to periodontitis is associated with cardio-metabolic adaptation to high-fat diet in mice. Am J Physiol Gastrointest Liver Physiol 310:G1091–G1101
Maurice CF, Haiser HJ, Turnbaugh PJ (2013) Xenobiotics shape the physiology and gene expression of the active human gut microbiome. Cell 152:39–50
Carmody RN, Gerber GK, Luevano JM Jr et al (2015) Diet dominates host genotype in shaping the murine gut microbiota. Cell Host Microbe 17:72–84
de Fourmestraux V, Neubauer H, Poussin C et al (2004) Transcript profiling suggests that differential metabolic adaptation of mice to a high fat diet is associated with changes in liver to muscle lipid fluxes. J Biol Chem 279:50743–50753
Szabo G, Bala S, Petrasek J, Gattu A (2010) Gut-liver axis and sensing microbes. Dig Dis 28:737–744
Dumas ME, Barton RH, Toye A et al (2006) Metabolic profiling reveals a contribution of gut microbiota to fatty liver phenotype in insulin-resistant mice. Proc Natl Acad Sci U S A 103:12511–12516
Collaborators GBDRF, Forouzanfar MH, Alexander L et al (2015) Global, regional, and national comparative risk assessment of 79 behavioural, environmental and occupational, and metabolic risks or clusters of risks in 188 countries, 1990-2013: a systematic analysis for the Global Burden of Disease Study 2013. Lancet 386:2287–2323
Wang XW, Heegaard NH, Orum H (2012) MicroRNAs in liver disease. Gastroenterology 142:1431–1443
Ambros V (2001) MicroRNAs: tiny regulators with great potential. Cell 107:823–826
Dalmasso G, Nguyen HT, Yan Y et al (2011) Microbiota modulate host gene expression via microRNAs. PLoS One 6:e19293
Androsavich JR, Chau BN, Bhat B, Linsley PS, Walter NG (2012) Disease-linked microRNA-21 exhibits drastically reduced mRNA binding and silencing activity in healthy mouse liver. RNA 18:1510–1526
Cani PD, Amar J, Iglesias MA et al (2007) Metabolic endotoxemia initiates obesity and insulin resistance. Diabetes 56:1761–1772
Boon RA, Iekushi K, Lechner S et al (2013) MicroRNA-34a regulates cardiac ageing and function. Nature 495:107–110
Caraux G, Pinloche S (2005) PermutMatrix: a graphical environment to arrange gene expression profiles in optimal linear order. Bioinformatics 21:1280–1281
Hsu SD, Lin FM, Wu WY et al (2011) miRTarBase: a database curates experimentally validated microRNA-target interactions. Nucleic Acids Res 39:D163–169
Henao-Mejia J, Elinav E, Jin C et al (2012) Inflammasome-mediated dysbiosis regulates progression of NAFLD and obesity. Nature 482:179–185
Geller DA, Lowenstein CJ, Shapiro RA et al (1993) Molecular cloning and expression of inducible nitric oxide synthase from human hepatocytes. Proc Natl Acad Sci U S A 90:3491–3495
Luche E, Cousin B, Garidou L et al (2013) Metabolic endotoxemia directly increases the proliferation of adipocyte precursors at the onset of metabolic diseases through a CD14-dependent mechanism. Mol Metab 2:281–291
Velagapudi VR, Hezaveh R, Reigstad CS et al (2010) The gut microbiota modulates host energy and lipid metabolism in mice. J Lipid Res 51:1101–1112
Szklarczyk D, Franceschini A, Wyder S et al (2015) STRING v10: protein-protein interaction networks, integrated over the tree of life. Nucleic Acids Res 43:D447–D452
Elinav E, Strowig T, Kau AL et al (2011) NLRP6 inflammasome regulates colonic microbial ecology and risk for colitis. Cell 145:745–757
Xie G, Wang X, Liu P et al (2016) Distinctly altered gut microbiota in the progression of liver disease. Oncotarget 7:19355–19366
Yang JY, Lee YS, Kim Y et al (2016) Gut commensal Bacteroides acidifaciens prevents obesity and improves insulin sensitivity in mice. Mucosal Immunol doi:10.1038/mi.2016.42
Song G, Sharma AD, Roll GR et al (2010) MicroRNAs control hepatocyte proliferation during liver regeneration. Hepatology 51:1735–1743
Zhang J, Jiao J, Cermelli S et al (2015) miR-21 inhibition reduces liver fibrosis and prevents tumor development by inducing apoptosis of CD24+ progenitor cells. Cancer Res 75:1859–1867
Seeger T, Fischer A, Muhly-Reinholz M, Zeiher AM, Dimmeler S (2014) Long-term inhibition of miR-21 leads to reduction of obesity in db/db mice. Obesity (Silver Spring) 22:2352–2360
Wu H, Ng R, Chen X, Steer CJ, Song G (2016) MicroRNA-21 is a potential link between non-alcoholic fatty liver disease and hepatocellular carcinoma via modulation of the HBP1-p53-Srebp1c pathway. Gut 65:1850–1860
Ahn J, Lee H, Jung CH, Ha T (2012) Lycopene inhibits hepatic steatosis via microRNA-21-induced downregulation of fatty acid-binding protein 7 in mice fed a high-fat diet. Mol Nutr Food Res 56:1665–1674
Acknowledgements
We thank the Société Française d’Hypertension Artérielle and the Société Francophone du Diabète for supporting V. Blasco-Baque; the zootechnie-Rangueil Inserm/UPS US006 CREFRE (Toulouse, France); the GeT-TRIX facility at INRA (Toulouse, France). We also thank S. Dejean from the Institut de Mathématiques de Toulouse (Toulouse, France) for helping with the statistical analysis.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Data availability
The data that support the findings of this study are available from Vaiomer SAS but restrictions apply to the availability of these data, which were used under license for the current study, and so are not publicly available. However, data are available from the authors upon reasonable request and with permission of Vaiomer SAS.
Funding
This work was supported in part by grants from Agence Nationale pour la Recherche (ANR) to RB and collaborators (ANR-Bactimmunodia and LPS diagra), and by the European Commission’s Seventh Framework programme under grant agreement No. 241913 (FLORINASH) to RB.
Duality of interest statement
The authors declare that there is no duality of interest associated with this manuscript.
Author contribution
VB-B, BC, AF, SH made substantial contributions to acquisition of data and reviewed the article critically for important intellectual content; PG, J-FA, MC, CS-K, BG, FT and RB made substantial contributions to the analysis and interpretation of data, and reviewed the article critically for important intellectual content; MS made substantial contributions to conception and design, acquisition, analysis and interpretation of data, as well as in drafting the article. All authors gave final approval of the version to be published. MS is the guarantor of this work.
Electronic supplementary material
Below is the link to the electronic supplementary material.
ESM Fig. 1
(PDF 35 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Blasco-Baque, V., Coupé, B., Fabre, A. et al. Associations between hepatic miRNA expression, liver triacylglycerols and gut microbiota during metabolic adaptation to high-fat diet in mice. Diabetologia 60, 690–700 (2017). https://doi.org/10.1007/s00125-017-4209-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00125-017-4209-3