Abstract
The objective of this study was to assess genotype by environment interaction for 21 physiological traits in sugar beet (Beta vulgaris L.) parents and hybrids grown in Rodasht Agricultural Research Station in Iran by the additive main effects and multiplicative interaction model. The study comprised of 51 sugar beet genotypes [10 multigerm pollen parents, four monogerm seed parents and 36 F1 hybrids], evaluated at four environments in a randomized complete block design, with three replicates. The additive main effects and multiplicative interaction analyses revealed significant environment main effects with respect to all observed traits, except extraction coefficient of sugar. The additive main effects and multiplicative interaction stability values ranged from 0.009 (G17 for leaf Ca2+) to 9.698 (G09 for extraction coefficient of sugar). The parental forms 2 7233-P.29 (G38) and C CMS (G49) as well as hybrids 2(6)*C (G27) and 5*C (G33) are recommended for further inclusion in the breeding programs because of their stability and good average values of observed traits.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Sugar beet approximately supplies about 30 percent of the world’s sugar [24]. In the chenopodiaceae family, sugar beet is considered as salt-tolerant plant. This plant with deep root system shows high tolerance to water stress conditions, such as salinity and drought [8]. The previous study was shown that about 50 mM NaCl can stimulate plant growth in sugar beet [32]. Therefore, this plant could be a good resource to explore salt tolerance mechanisms to product salt tolerant hybrid seeds.
Environmental instability causes a variety of genotype reactions during the growing season, which complicates understanding of the physiological traits of sugar beet. The additive main effects and multiplicative interaction (AMMI) model have analyzed the main effects of genotype and environment as well as the genotype-by-environmental interaction (GEI) [7, 34]. The AMMI model as a reliable way combines the analysis of variance and the principal component analysis (PCA) in a particular single analysis to dedicate cultivars to different environments or locations. There is little information about how sugar beet varieties respond to different conditions [14, 17, 18, 31].
Evaluation of genotypes in different years and locations (different environments) is one of the important part of breeding programs. In fact, the assessment of the adaptation abilities of cultivars in mega-environment, is a reliable criterion in the detection of stable or specific cultivars in the area. The most stable cultivars are characterized by a negligible effect of genotype by environment interaction [16]. The specifically adapted genotype is known when a genotype shows superior rank in an area over several years particularly under a restricted set of condition that related to the phenomenon of genotype-by-environment interaction (GEI). By changing the rank of genotypes in different environments (due to GEI), the interpretation of the information obtained becomes difficult and complicated [21]. Thus, an understanding of GEI using multi environmental trails (METs) will leads to achievement of guaranteed superior genotypes across a range of environmental conditions [21, 30, 33].
The additive main effect and multiplicative interaction model (AMMI) is effective method for studying GEI, stability and adaptation analysis and identifying the specifically adapted cultivars or mega-environments [10]. So far, the variability assessment of genotypes for different environments and stability analysis has been carried out for sugar beet to study the interaction of the genotype per environment. In a study on nine sugar beet genotypes in 52 different environments, the value of total variance for environment, genotype and their interaction were shown 80%, 5%, and 3%, respectively [18]. Mostafavi et al. [25] studied the interaction of nine sugar beet cultivars with six regions using GGE biplot and AMMI methods, and (I13*A37.1) * SH-1-HSF.5 genotype was introduced as the best genotype for stability. From the stability analysis of white sugar yield and the adaptation of 36 sugar beet cultivars under 11 examined locations, four mega-environments and high compliance of the sugar yield rankings were distinguished [31].
The objective of this study were: (1) to assess genotype by environment interaction for the 21 physiological traits in 51 sugar beet genotypes across four environments and 2 years grown in Rodasht Agricultural Research Station in Iran by the AMMI model, and (2) to select genotypes combining a high stability with good average values of observed traits.
Materials and methods
Plant materials and experimental conditions
Four CMS lines and 10 populations [S0 = open-pollinated (parental) and S2 = the second generation of selfing] were subjected to crossing in this study using North Carolina mating design II [9]. The description of these parent populations are shown in Table 1. The drought tolerant and salinity tolerant lines displayed in Table 1 were derived from two independent recurrent selection programs.
To carry out the crosses, the pollen parents were divided into two groups each with five populations. Biparental mating and bulk mating schemes were employed for the first and second group, respectively. In biparental mating, 10 plants were randomly selected from each male parent and independently crossed with four CMS lines in a cross section. Thus, 40 crosses (4 × 10) from each male parent and a total of 200 cross combinations is expected to produce from five pollen parents of the first group. In bulk mating, 20 crosses (5 × 4) from five pollen parents of the second group with four CMS lines were obtained. Due to lack of reproductive synchronization between some of the female and male lines, in a number of crosses, the hybrid seed was not formed. In a number of crosses, seed was only sufficiently evaluated for 1 year. Therefore, a total of 51 genotypes including 36 F1 hybrids along with 14 parents and 1 control (Table 2) were grown in two environments saline (EC of soil and water was 8 and 12 dSm−1, respectively) and normal (soil ECe = 4 and irrigation water ECw = 3 dSm−1) field conditions during the 2011 and 2012 growing seasons at Rodasht Agricultural Research Station, Iran (65 km East of Isfahan, 32° 82′ 90'' N and 52° 81′ 00'' E, 1560 m altitude). A complete block design with three replications was used. Each plot contained three 5 m long rows and spaced 0.5 m between rows.
Standard cultural practices were followed for seed-bed preparation, sowing, irrigation and control of weed and pests. Thinning was conducted at 6-leaf stage by hand to settle 17 cm plant distance. The salinity treatment was commenced at 4 weeks after emergence.
Agro-physiological traits
The physiological traits (leaf Ca2+, leaf Na+, leaf K+, Ca2+/ Na+, leaf K+/ Na+, net CO2 assimilation rate, transpiration rate), root related traits (root Na+, root K+, root α-N, root K+/Na+, sugar content (SC), white sugar content (WSC), molasses sugar (MS), extraction coefficient of sugar (ECS), alkaline level content (ALC), dry matter (DM) and yield related traits (root number, root yield, sugar yield, white sugar yield) were evaluated in this study. Measurements of physiological traits were performed at the 5–7-leaf stage which corresponded to about 2 months after applying salinity stress.
Leaf gas exchange parameters (net CO2 assimilation rate (PN) and transpiration rate (E)) were measured using a Li-Cor 6400 gas-exchange portable photosynthesis system (Li-Cor, Lincoln, Nebraska, USA).
The concentration of Na+, K+ and Ca2+ ions in leaf tissue were measured about 3 months after sowing in the both experiments in 2011 and 2012. Samples of dried tissues of the four uppermost leaves were ashed for 6 h at 550 °C, dissolved in 1% (v/v) hydrochloric acid (HCl) and made to volume with distilled water. The leaf Na+ and K+ concentrations were determined by flame photometer (JENWAY Clinical PFP7 model) and Ca2+ concentration was determined by atomic absorption spectrometer (Perkin-Elmer 2380 Atomic Absorption).
The sugar beet root yield (RY) was determined after harvest. Root pulps were quickly frozen and stored at − 26ºC until analysis of quality-related traits.
Sugar beet root quality-related traits [Na+ and K+, α-amino-N and sucrose (sugar) contents] in the saline and non-saline experiments conducted in 2011 and 2012 were determined using an automatic beet laboratory system (Venema, Groningen, the Netherlands) linked to Betalyser system (W. Kernchen, GmbH, Seelze, Germany). Molasses sugar (MS), extraction coefficient of sugar (ECS) and alkaline level content (ALC) were calculated using the following formula [29]:
Roots were oven dried to a constant weight at 80 °C for 48 h to measure dry matter (DM). White sugar content (WSC), sugar yield (SY) and white sugar yield (WSY) were calculated according to the following formulas [29]:
The details of other measurements are as described earlier [2].
Statistical analysis
A two-way fixed effect model was fitted to determine the magnitude of the main effects of variation and their interaction on the 21 physiological traits. Least-squares means were simultaneously produced for the AMMI model. The AMMI model is fitted in two stages. In first stage, the main additive effects of genotypes (G) and environments (E) are fitted by least squares through analysis of variance (ANOVA). In the second stage, genotype-by-environmental interaction GEI, the non-additive residuals will be subjected to singular value decomposition to obtain multiplicative terms referred to as interaction principal components (IPC) scores for genotypes and environments. The traditional AMMI model for fixed effects [13, 26] is given by
where yge is the mean value of observed trait of genotype g in environment e, μ is the grand mean, αg is the genotypic mean deviations, βe is the environmental mean deviations, N is the number of PCA axis retained in the adjusted model, λn is the eigenvalue of the PCA axis n, γgn is the genotype score for PCA axis n, δen is the score eigenvector for PCA axis n, and Qge is the residual, including AMMI noise and pooled experimental error (with expected normal distribution).
The AMMI stability value (ASV) was used to compare the stability of genotypes as described by Purchase et al. [28]:
where SS is the sum of squares, IPCA1 and IPCA2 is the first and the second interaction principal component axes, respectively; and the IPCA1 and IPCA2 scores were the genotypic scores in the AMMI model. Lower ASV score indicate a more stable genotype across environments.
Genotype selection index (GSI) was calculated for each genotype which incorporates both mean 1000-kernel weight and ASV index in single criteria (GSIi) as [12]
where GSIi is genotype selection index for ith genotype, RMi is rank of trait mean (from maximal to minimal for: leaf Ca2+, leaf K+, leaf Ca2+/Na+, leaf K+/Na+, net CO2, root number, root yield, root Na+, sugar yield, white sugar yield, white sugar content, extraction coefficient of sugar, dry matter, and from minimal to maximal for: leaf Na+, transpiration rate, sugar content, root K+, root α-N, root K+/Na+, alkaline level content and molasses sugar) for ith genotype, RASVi is rank for the AMMI stability value for the ith genotype. Finally, total genotype selection index (TGSI) was calculated for each genotype as sum of GSIs for all 21 physiological traits of study.
The relationships between the particular 21 physiological traits were assessed based on Pearson’s correlation coefficients, independently for four environments. All the analyses were conducted using the GenStat v. 18 statistical software package.
Results
The three sources of variation (genotypes, environments and GEI) were statistically significant for: net CO2, transpiration rate, root number, root yield, root α-N, sugar yield and white sugar yield (Table 3). The differences between genotypes explained from 2.92% (for leaf K+/Na+) to 25.21% (for root yield) of the total quantitative trait variation (Table 3). The differences between genotypes were significant for: leaf Ca2+, leaf K+, net CO2, transpiration rate, root number, root yield, sugar content, root K+, root α-N, sugar yield, white sugar yield, white sugar content, extraction coefficient of sugar and molasses sugar (Table 3). The sums of squares for environment main effect were highly significant for all observed traits, except extraction coefficient of sugar and ranged from 1.30% (for white sugar content) to 79.35% (for leaf K+/Na+) of the total physiological trait variation (Table 3). The GEI was statistically significant for ten from 21 observed traits: leaf Na+, net CO2, transpiration rate, root number, root yield, root α-N, alkaline level content, sugar yield, white sugar yield and dry matter (Table 3). The effects of GEI explained form 7.32% (for leaf K+/Na+) to 54.79% (for net CO2) of the total variation.
The first principal component (IPCA 1) was significant for all observed traits except sugar content and white sugar content and accounted from 43.61% (for dry matter) to 63.61% (for extraction coefficient of sugar) of the variation caused by interaction (Table 3), while IPCA 2 was significant for: net CO2, transpiration rate, root number, root yield, molasses sugar and dry matter and 3 accounted from 22.94% (for root Na+) to 47.06% (for root number). Values for the first two principal components were accounted jointly from 78.01% (for transpiration rate) to 99.56% (for leaf Ca2+) of the whole effect it had on the variation of observed trait (Table 3).
Lack of significant change in genotype rank in different environmental and climatic conditions and stability against stresses indicate genotype stability. In this study, climatic conditions were considered as the source of this variation component. The analysis showed that some genotypes have high adaptation; however, most of them have specific adaptability. ASVs revealed variations in observed physiological trait stability among the 51 genotypes were calculated. According to Purchase et al. [28], a stable genotype is defined as one with ASV close to zero. ASV ranged different for particular physiological traits, for: leaf Ca2+: from 0.009 (for G17) to 1.582 (for G20), leaf Na+: from 0.038 (for G50) to 1.105 (for G07), leaf K+: from 0.089 (for G13) to 1.297 (for G01), leaf Ca2+/Na+: from 0.056 (for G27) to 0.815 (for G22), leaf K+/Na+: from 0.073 (for G22) to 1.719 (for G38), net CO2: from 0.100 (for G46) to 3.748 (for G35), transpiration rate: from 0.044 (for G03) to 2.486 (for G11), root number: from 0.106 (for G44) to 4.517 (for G24), root yield: from 0.211 (for G18) to 8.279 (for G11), sugar content: from 0.162 (for G26) to 2.198 (for G31), root Na+: from 0.025 (for G39) to 3.272 (for G30), root K+: from 0.078 (for G03) to 1.178 (for G44), root α-N: from 0.068 (for G37) to 1.821 (for G42), root K+/Na+: from 0.057 (for G34) to 1.435 (for G15), alkaline level content: from 0.078 (for G38) to 3.150 (for G42), sugar yield: from 0.147 (for G20) to 3.659 (for G11), white sugar yield: from 0.080 (for G05) to 2.671 (for G11), white sugar content: from 0.045 (for G14) to 1.856 (for G09), extraction coefficient of sugar: from 0.135 (for G28) to 9.698 (for G09), molasses sugar: from 0.097 (for G10) to 0.932 (for G16), and dry matter: from 0.035 (for G36) to 1.612 (for G31) (Table 4).
The genotype selection index ranged different for particular physiological traits, for: leaf Ca2+: from 18 (for G01) to 98 (for G11), leaf Na+: from 13 (for G31) to 97 (for G40), leaf K+: from 11 (for G13) to 89 (for G19), leaf Ca2+/Na+: from 21 (for G27) to 101 (for G11), leaf K+/Na+: from 3 (for G22) to 93 (for G11), net CO2: from 19 (for G12) to 76 (for G32), transpiration rate: from 11 (for G17) to 101 (for G35), root number: from 19 (for G29) to 101 (for G11), root yield: from 6 (for G44) to 91 (for G17), sugar content: from 9 (for G26) to 93 (for G17), root Na+: from 9 (for G26) to 97 (for G05), root K+: from 9 (for G41) to 101 (for G48), root α-N: from 6 (for G37) to 100 (for G42), root K+/Na+: from 13 (for G06) to 100 (for G15), alkaline level content: from 2 (for G38) to 100 (for G16), sugar yield: from 10 (for G10) to 90 (for G37), white sugar yield: from 9 (for G10) to 88 (for G37), white sugar content: from 7 (for G41) to 102 (for G09), extraction coefficient of sugar: from 11 (for G50) to 102 (for G09), molasses sugar: from 8 (for G03) to 98 (for G48), and dry matter: from 6 (for G36) to 95 (for G31) (Table 4).
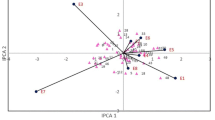
The stability of tested genotypes can be evaluated according to biplot for physiological trait. The most interesting results were obtained for seven traits: net CO2, transpiration rate, root number, root yield, sugar yield, white sugar yield and dry matter. Figure 1 shows distribution of the first two interaction principal components for net CO2. Sugar beet parents and hybrids interacted differently with climate conditions in the observed environments. The genotypes G15, G22, G24, G27, G35 and G45 interacted positively with the S1 environment, but negatively with the N2 and S2 (Fig. 1). The genotypes G03, G05, G17, G21, G42 and G43 interacted positively with the N1 environment, but negatively with the both saline and non-saline experiments in 2012.
The first two interaction principal components for transpiration rate accounted jointly for 78.01% of the variation caused by interaction (Fig. 2). The genotypes G11 and G35 interacted positively with the S1 environment, but negatively with the N1 and S2. The genotypes G21, G28 and G42 interacted positively with the N1 environment. Genotypes G04 and G50 interacted positively with the S2 environment.
Biplot for genotype by environment interaction of transpiration rate in sugar beet (Beta vulgaris L.) parents and hybrids in four environments [saline (S) and non-saline (N) experiments conducted in 2011 and 2012], showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively)
The stability of tested genotypes can be evaluated according to biplot for root number (Fig. 3). The genotypes G24 and G43 interacted positively with the N2 environment, but negatively with the N1 and S1. The genotypes G04, G09, G18, G19 and G32 interacted positively with the S2 environment. Genotypes G01, G03, G06, G17 and G31 interacted positively with the S1 and N1 environments.
For root yield we observed that the genotype G11 interacted positively with the N1 environment, but negatively with the S2 as well as the genotype G35 interacted positively with the S1 environment, but negatively with the N2 (Fig. 4). The first two interaction principal components for this trait accounted jointly for 83.92%% of the variation caused by interaction (Fig. 4, Table 3).
Figures 5, 6 present the biplots of the stability of tested genotypes for sugar yield and white sugar yield, respectively. Only the first interaction principal component, IPCA1, was statistically significant for these traits. The saline and non-saline experiments in both years of study were strongly different (Figs. 5, 6). The genotype G35 interacted positively with the S1 environment; the G03 and G11 interacted positively with the N1; and genotypes G24 and G43 with N2 environment for both traits.
Biplot for genotype by environment interaction of white sugar yield in sugar beet (Beta vulgaris L.) parents and hybrids in four environments [saline (S) and non-saline (N) experiments conducted in 2011 and 2012], showing the effects of primary and secondary components (IPCA 1 and IPCA 2, respectively)
The first two interaction principal components for dry matter accounted jointly for 79.24% of the variation caused by interaction (Fig. 7). The genotypes G15, G17, G20 and G45 interacted positively with the S1 environment, but negatively with the saline environment in 2012. The genotypes G24, G29, G38, G41 and G49 interacted positively with the N1 environment. Genotypes G11, G23 and G35 interacted positively with the S2 environment, but negatively with the saline environment in 2011. The genotypes G16 and G19 interacted positively with the N2 environment (Fig. 7).
The best total genotype selection index calculated for all 21 physiological traits was observed for genotype G38 (TGSI = 794), while the worst—G11 (TGSI = 1464) (Table 4). The parental forms 2 7233-P.29 (G38) and C CMS (G49) as well as hybrids 2(6)*C (G27) and 5*C (G33) are recommended for further inclusion in the breeding programs because of their stability and good average values of observed traits.
The positive statistical significant (α = 0.05) correlation coefficients in all four environments (saline and non-saline experiments conducted in 2011 and 2012) were observed between: leaf Ca2+ and leaf Na+, root number and root yield, root number and sugar yield, root number and white sugar yield, root yield and sugar yield, root yield and white sugar yield, sugar content and white sugar content, sugar content and extraction coefficient of sugar, root Na+ and alkaline level content, root Na+ and molasses sugar, root K+ and root α-N, root K+/Na+ and extraction coefficient of sugar, sugar yield and white sugar yield, white sugar yield and extraction coefficient of sugar as well as white sugar content and extraction coefficient of sugar (Tables 5, 6). Negative correlation coefficients in all four environments were observed between: leaf Na+ and leaf K+/Na+, root Na+ and root α-N, root Na+ and extraction coefficient of sugar, root α-N alkaline level content, root K+/Na+ and molasses sugar as well as extraction coefficient of sugar and molasses sugar (Tables 5, 6).
Discussion
Genotype by environment interaction is one of the unifying challenges facing plant breeders [19, 20]. The quantity and quality traits of sugar beet (Beta vulgaris L.) are determined by genotype and environment that sugar beet varieties usually differ in different environments. To introduce new varieties in the final stages of breeding program, they must be tested in several environments to identify their stability across different environments. If the genotypes ranking do not vary the environment that is a GEI is absent or low, showing general adaption [4]. As an advantage of the change of ranks in different environments (crossover) is the availability of special varieties which are adapted to certain areas stress situations [3]. Thus, GEI is of major importance to improve sugar beet production. The aim of this study was to disclosure the importance of environment on quantity and quality traits of sugar beet genotypes. For that purpose, 21 traits of 53 sugar beet genotypes were investigated in four environments (combination of two locations and 2 years) in field trails.
AMMI allow for a large set of technical interpretations, and they used more commonly to evaluate the genotype–environment interactions. The additive main effects and multiplicative interaction (AMMI) model addresses the limitations of ANOVA and PCA. The AMMI model effectively explains the GEI patterns. The AMMI method used for three primary purposes for stability. The first is that the model involves variance analysis and principal component analysis. Second, AMMI clarifies GEI and summarizes G and E patterns and relationships, and the third use is the accuracy of yield estimates. AMMI model is more suitable and simplifies genotypes' instantaneous choice for stability. The model helps establish the relationship of genotypes, environment and their interaction. The AMMI model as the most popular multiplicative models was initially proposed by Gollob [15] and [22, 23] in the context of fixed effects. AMMI can have several models: AMMI0, which estimates the main additive effect of genotypes and environments and does not include any major axis (IPCA) AMMI1, which combines AMNI0 genotype additive effects with environmental interactions estimated and combines from the first major axis (IPCA 1). AMMI2 and others up to the full model with all IPCA axes. In this paper we used the traditional AMMI model for fixed effects. The traditional AMMI model has been used extensively for many species [1, 5, 11, 26].
Researchers apply the proposed strategy to extract specific cultivars with competitive performance across different environments that can extract more information from GE interaction [27]. Hassani et al. [17] using AMMI model analyzed GEI for 49 sugar beet genotypes in four different geographical locations in 2 years for three morphological traits: root yield, sugar yield and white sugar yield. Ghareeb et al. [14] also analyzed root yield, sugar yield and white sugar yield by AMMI model. Seven sugar beet cultivars were appraised in eight environments: two consecutive seasons and four locations. Their results showed that the AMMI model clarified most of the GEI (85.97%, 83.34% and 86.47%) for root yield, sugar content and, sugar yield, respectively. Hoffmann et al. [18] analyzed GEI of nine genotypes in 52 environments for yield and quality (sugar, K, Na, amino N, total soluble N, betaine, glutamine, invert sugar and raffinose) of sugar beet in Europe. According to their results, the GEI with about 3% was less than the main effect of genotypes (about 80%).
High genotypes stability is linked with the AMMI stability value. AMMI stability value he distance of the coordinates of each genotype from the origin of the bi-plot coordinate diagram of the two principal components of the interaction is based on the scores of the first and second for interaction principal component axis (IPCA) model for each genotype. Genotypes with the lowest ASV values are identified by their shortest projection from the bi-plot origin and considered the most stable. Using GE bi-plot displayed based on the AMMI results, the main effect of the genotype, the environment, and the most significant GEI could be determined. Regarding the AMMI model, the results of the analysis of variance indicated that significant genotype × environment interaction for all considering physiological traits.
References
Abakemal D, Shimelis H, Derera J (2016) Genotype-by-environment interaction and yield stability of quality protein maize hybrids developed from tropical-highland adapted inbred lines. Euphytica 209:757–769. https://doi.org/10.1007/s10681-016-1673-7
Abbasi Z, Majidi MM, Arzani A, Rajabi A, Bocianowski J (2015) Association analysis of morpho-physiological traits with salinity tolerance using SSR markers in sugar beet (Beta vulgaris L.). Euphytica 205:785–797. https://doi.org/10.1007/s10681-015-1408-1
Annicchiarico P (2007) Wide-versus specific-adaptation strategy for lucerne breeding in northern Italy. Theor Appl Genet 114:647–657. https://doi.org/10.1007/s00122-006-0465-1
Baker RJ (1988) Tests for crossover genotype-environmental interactions. Can J Plant Sci 68:405–410. https://doi.org/10.4141/cjps88-051
Bocianowski J, Niemann J, Nowosad K (2019) Genotype-by-environment interaction for seed quality traits in interspecific cross-derived Brassica lines using additive main effects and multiplicative interaction model. Euphytica 215:7. https://doi.org/10.1007/s10681-018-2328-7
Bocianowski J, Nowosad K, Szulc P (2019) Soil tillage methods by years interaction for harvest index of maize (Zea mays L) using additive main effects and multiplicative interaction model. Acta Agric Scand B Soil Plant Sci 69(1):75–81. https://doi.org/10.1080/09064710.2018.1502343
Bocianowski J, Warzecha T, Nowosad K, Bathelt R (2019) Genotype by environment interaction using AMMI model and estimation of additive and epistasis gene effects for 1000-kernel weight in spring barley (Hordeum vulgare L.). J Appl Genet 60(2):127–135. https://doi.org/10.1007/s13353-019-00490-2
Brar NS, Dhillon BS, Saini KS, Sharma PK (2015) Agronomy of sugarbeet cultivation–A review. Agricul Rev 36(3):184–197. https://doi.org/10.5958/0976-0741.2015.00022.7
Comstock RE, Robinson HF (1952) Estimation of average dominance of genes. In: Gowen JW (ed) Heterosis. Iowa State College Press, Ames, pp 494–516
Crossa J, Cornelius PL, Yan W (2001) Biplot of linear-bilinear models for studying crossover genotype × environment interaction. Crop Sci 41:158–163. https://doi.org/10.2135/cropsci2002.6190
Edwards JW (2016) Genotype × environment interaction for plant density response in maize (Zea mays L.). Crop Sci 56:1493–1505. https://doi.org/10.2135/cropsci2015.07.0408
Farshadfar E, Sutka J (2003) Locating QTLs controlling adaptation in wheat using AMMI model. Cereal Res Commun 31:249–256. https://doi.org/10.1007/BF03543351
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761. https://doi.org/10.1007/BF00224240
Ghareeb ZE, Ibrahim HEA, Elsheikh SRE, Bachoash SMI (2014) Genotype× environment interaction for characteristics of some sugar beet genotypes. J Plant Prod 5(5):853–867. https://doi.org/10.21608/jpp.2014.55434
Gollob HF (1968) A statistical model which combines features of factor analytic and analysis of variance techniques. Psychometrika 33:73–115. https://doi.org/10.1007/BF02289676
Götze P, Rücknagel J, Wensch-Dorendorf M, Märländer B, Christen O (2017) Crop rotation effects on yield, technological quality and yield stability of sugar beet after 45 trial years. Eur J Agron 82:50–59. https://doi.org/10.1016/j.eja.2016.10.003
Hassani M, Heidari B, Dadkhodaie A, Stevanato P (2018) Genotype by environment interaction components underlying variations in root, sugar and white sugar yield in sugar beet (Beta vulgaris L.). Euphytica 214(4):79. https://doi.org/10.1007/s10681-018-2160-0
Hoffmann CM, Huijbregts T, van Swaaij N, Jansen R (2009) Impact of different environments in Europe on yield and quality of sugar beet genotypes. Eur J Agron 30(1):17–26. https://doi.org/10.1016/j.eja.2008.06.004
Lin CS, Binns MR (1988) A superiority measure of cultivar performance for cultivar × location data. Canad J Plant Sci 68:193–198. https://doi.org/10.4141/cjps88-018
Lin CS, Binns MR (1994) Concepts and methods for analyzing regional trial data for cultivar and location selection. Plant Breed Rev 12:271–297. https://doi.org/10.1002/9780470650493.ch10
Malosetti M, Ribaut JM, van Eeuwijk FA (2013) The statistical analysis of multi-environment data: modeling genotype-by-environment interaction and its genetic basis. Front Physiol 4(44):1–17. https://doi.org/10.3389/fphys.2013.00044
Mandel J (1969) The partitioning of interactions in analysis of variance. J Res Nat Bureau Standards Series B 73:309–328
Mandel J (1971) A new analysis of variance model for non-additive data. Technometrics 13:1–18
Monteiro F, Frese L, Castro S, Duarte MC, Paulo OS, Loureiro J, Romeiras MM (2018) Genetic and genomic tools to assist sugar beet improvement: the value of the crop wild relatives. Front Plant Sci 9:74. https://doi.org/10.3389/fpls.2018.00074
Mostafavi K, Orazizadeh M, Rajabi A, Ilkaei MN (2018) Stability and adaptability analysis in sugar beet varieties for sugar content using GGE-biplot and AMMI methods. Bulg J Agric Sci 24(1):40–45
Nowosad K, Liersch A, Popławska W, Bocianowski J (2016) Genotype by environment interaction for seed yield in rapeseed (Brassica napus L.) using additive main effects and multiplicative interaction model. Euphytica 208:187–194. https://doi.org/10.1007/s10681-015-1620-z
Nowosad K, Tratwal A, Bocianowski J (2018) Genotype by environment interaction for grain yield in spring barley using additive main effects and multiplicative interaction model. Cereal Res Commun 46(4):729–738. https://doi.org/10.1556/0806.46.2018.046
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. South African J Plant Soil 17:101–107. https://doi.org/10.1080/02571862.2000.10634878
Reinefeld E, Emmerich A, Baumgarten G, Winner C, Beiß U (1974) Zur voraussage des melassezuckers aus rübenanalysen. Zucker 27:2–15
Rodrigues PC, Malosetti M, Gauch HG, van Eeuwijk FA (2014) A weighted AMMI algorithm to study genotype-by-environment interaction and QTL-by-environment interaction. Crop Sci 54(4):1555–1570. https://doi.org/10.2135/cropsci2013.07.0462
Studnicki M, Lenartowicz T, Noras K, Wójcik-Gront E, Wyszyński Z (2019) Assessment of stability and adaptation patterns of white sugar yield from sugar beet cultivars in temperate climate environments. Agronomy 9(7):405. https://doi.org/10.3390/agronomy9070405
Wu GQ, Feng RJ, Liang N, Yuan HJ, Sun WB (2015) Sodium chloride stimulates growth and alleviates sorbitol-induced osmotic stress in sugar beet seedlings. Plant Growth Regul 75:307–316. https://doi.org/10.1007/s10725-014-9954-4
Yan W (2016) Analysis and handling of G×E in a practical breeding program. Crop Sci 56(5):2106–2118. https://doi.org/10.2135/cropsci2015.06.0336
Zobel RW, Wright MJ, Gauch HG (1988) Statistical analysis of yield trial. Agron J 80:388–393. https://doi.org/10.2134/agronj1988.00021962008000030002x
Author information
Authors and Affiliations
Contributions
Conceptualization, ZA and JB; methodology, ZA and JB; software, JB; formal analysis, JB; data curation, ZA and JB; writing—original draft preparation, ZA and JB; writing—review and editing, ZA and JB; visualization, JB. All authors have read and agreed to the published version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
Zahra Abbasi declares that she has no conflict of interest. Jan Bocianowski declares that he has no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Abbasi, Z., Bocianowski, J. Genotype by environment interaction for physiological traits in sugar beet (Beta vulgaris L.) parents and hybrids using additive main effects and multiplicative interaction model. Eur Food Res Technol 247, 3063–3081 (2021). https://doi.org/10.1007/s00217-021-03861-4
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00217-021-03861-4