Abstract
Purpose
Pituitary macroadenoma consistency can influence the ease of lesion removal during surgery, especially when using a transsphenoidal approach. Unfortunately, it is not assessable on standard qualitative MRI. Radiomic texture analysis could help in extracting mineable quantitative tissue characteristics. We aimed to assess the accuracy of texture analysis combined with machine learning in the preoperative evaluation of pituitary macroadenoma consistency in patients undergoing endoscopic endonasal surgery.
Methods
Data of 89 patients (68 soft and 21 fibrous macroadenomas) who underwent MRI and transsphenoidal surgery at our institution were retrospectively reviewed. After manual segmentation, radiomic texture features were extracted from original and filtered MR images. Feature stability analysis and a multistep feature selection were performed. After oversampling to balance the classes, 80% of the data was used for hyperparameter tuning via stratified 5-fold cross-validation, while a 20% hold-out set was employed for its final testing, using an Extra Trees ensemble meta-algorithm. The reference standard was based on surgical findings.
Results
A total of 1118 texture features were extracted, of which 741 were stable. After removal of low variance (n = 4) and highly intercorrelated (n = 625) parameters, recursive feature elimination identified a subset of 14 features. After hyperparameter tuning, the Extra Trees classifier obtained an accuracy of 93%, sensitivity of 100%, and specificity of 87%. The area under the receiver operating characteristic and precision-recall curves was 0.99.
Conclusion
Preoperative T2-weighted MRI texture analysis and machine learning could predict pituitary macroadenoma consistency.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Pituitary adenomas are frequent tumors of the pituitary gland. Although most pituitary macroadenomas have a soft consistency, some are rather fibrous and therefore more challenging to remove by transsphenoidal adenomectomy. Indeed, tumor consistency has been reported as one of the principal determinants of transsphenoidal surgery success rate [1]. For this reason, the ability to preoperatively assess adenoma consistency could improve surgical planning and reduce complication rate and risk of residual tumor presence [2].
Radiomics, consisting of conversion of images into mineable data and subsequent analysis for decision support, has been gaining attention in recent years [3]. In particular, texture analysis is a post-processing technique allowing for quantitative description of pixel gray-level heterogeneity. More recently, texture analysis-derived features have been used in association with data mining and machine learning algorithms, aiding in the interpretation of a large amount of information produced. Machine learning (ML) is the branch of artificial intelligence including algorithms capable of modeling themselves and improving in accuracy by analyzing datasets, without prior explicit programming [4]. It leads to the creation of predictive models that are able, among other tasks, to solve classification problems. The usefulness of the radiomic approach is being assessed in different fields of radiology [5,6,7,8,9].
Our aim was to assess the accuracy of a ML model trained on radiomic data mined from MRI exams to predict pituitary macroadenoma surgical consistency prior to an endoscopic endonasal procedure.
Methods and materials
Patient population
This retrospective study was conducted in accordance with the 1964 Helsinki Declaration and its later amendments. The local Institutional Review Board gave its approval and waived the need for informed consent. We reviewed all patients referred to our institution for endoscopic endonasal pituitary adenoma removal (January 2013–December 2017). Those with history of previous treatment for pituitary adenoma (radiation or medical therapy) at the time of MRI, lesions smaller than 10 mm, extensively necrotic or hemorrhagic areas, or significant artifacts on the images used for the analysis were excluded.
Consistency assessment
All patients were operated on by two neurosurgeons with over 10 years of experience in a third level referral center in the field of pituitary surgery [10]. Tumor consistency, classified as soft or fibrous, was assessed in blinded double-check by the two surgeons according to the lesions’ inner surgical features. In detail, adenomas easily removable with conventional maneuvers of curettage and suction were defined as soft. More resistant ones, difficult to remove and thus requiring more complex maneuvers such as extracapsular dissection, were classified as fibrous [11,12,13,14]. Surgical features of soft and fibrous pituitary macroadenomas are depicted in online Video 1 and 2, respectively.
(MOV 17822 kb)
(MOV 25323 kb)
Image acquisition
All patients underwent MRI exams either on a 1.5 (Gyroscan Intera, Philips, Eindhoven, the Netherlands) or 3 T MR scanner (Magnetom Trio, Siemens Medical Solutions, Erlangen, Germany). The imaging protocol always included a coronal T2-weighted (T2-w) Turbo Spin Echo sequence whose detailed parameters are reported in supplementary Table 1.
Handcrafted radiomics
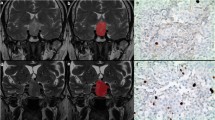
Adenomas are manually annotated by a neuroradiologist (8 years of experience) by placing a 2D polygonal region of interest (ROI) on the coronal slice of maximum lesion extension on a freely available segmentation software (ITKSnap v3.8.0) (Fig. 1). Two other readers (both > 5 years’ experience) also performed lesion segmentation on all patients, blinded to the first neuroradiologist’s ROI placement, to perform radiomic feature stability testing.
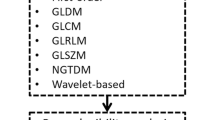
A freely available, well-established, and open-source Python software platform was used for image pre-processing and radiomic parameter extraction (Pyradiomics, v2.2.0). First of all, images and ROIs were resampled to a 2 × 2 × 2 mm isotropic voxel, as required for further pre-processing (i.e., correct use of image filters). All voxel intensity values were also normalized by subtracting the mean intensity and dividing by the standard deviation and discretized by using a fixed bin width (=3). Filtered images were also employed for feature extraction in addition to the pre-processed original T2-w ones. In particular, a Laplacian of Gaussian filter, with sigma values ranging from 2.0 (most fine texture) to 3.5 (most coarse) in 0.5 increments, and all available combinations of wavelet decomposition high- and low-pass filtering in the x, y, and z dimensions were applied. While 2D ROIs were drawn, we still chose to employ a three-dimensional wavelet decomposition as after resampling the software will detect an ROI z-axis value > 1. This is not an issue for the analysis as we excluded shape features, and the following feature selection steps will remove all redundant parameters that could have been extracted from similar wavelet decomposition-derived images.
Data mining and machine learning
Initial assessment and processing of the extracted data were performed on Python in particular using the numpy, pandas, and scikit-learn packages. First of all, the intraclass correlation coefficient (ICC) was calculated for each parameter as extracted using ROIs from the three readers. A two-way, absolute agreement and single rater ICC was employed, and only features with values ≥ 0.75 were considered stable. Non-informative, low variance (variance ≤ 0.1) features were also excluded from the dataset. Then, a pairwise correlation matrix was calculated for these in order to remove all features with an intercorrelation ≥ 0.8. As we expect an unbalanced dataset due to the relative rarity of fibrous adenomas compared with soft ones, the Synthetic Minority Oversampling Technique (SMOTE) was employed [15]. Then, 80% of the data was used for hyperparameter tuning via stratified 5-fold cross-validation, while a 20% hold-out set was employed for its testing on unseen data. In detail, the following steps were exclusively performed on the first set. A normalization scaler was calculated to remove biases due to feature scale and was later applied to the hold-out test set. Finally, recursive feature elimination (RFE), employing a logistic regression algorithm and stratified 5-fold cross-validation, was used to select the better performing feature subset.
The resulting data was used to train an ensemble learning meta-algorithm, the Extra Trees Classifier (ET). These often demonstrate good performance on radiomic medical image data [16]. Its performance for consistency prediction was finally assessed on the test set.
Accuracy metrics were obtained using the scikit-learn package and further analyzed on the R software (R for Unix/Linux, version 3.4.4, the R Foundation for Statistical Computing, 2014). In particular, DeLong’s test (pROC package) was used to obtain 95% confidence intervals (95%CI) of the area under the receiver operating characteristic curve (AUC) and the confusion matrix function (caret package) those of the classifier’s accuracy and compare its performance to the no information rate.
The described radiomics workflow pipeline is illustrated in Fig. 2.
Results
According to selection’s criteria, 89 patients were included in this study; 51 were males and 38 females, with mean age 52.17 ± 14.64 years (range 16–80). Average lesion size was 25 ± 8 mm (range 8–46 mm). The pituitary lesions were classified as soft in 68 patients and fibrous in the remaining 21. In detail, 19 soft (8 ACTH, 7 GH, 4 PRL) and 6 fibrous (2 ACTH, 2 PRL, 1 GH, 1 TSH) were functioning (25/89, 28% in total). In none of the cases, there was discordance among the neurosurgeons in lesion classification. Patient population clinical data are presented in Table 1.
A total of 1118 texture features were extracted, including first- and higher-order texture features from the original and filtered images. The correlation cluster map of the extracted features is shown in Fig. 3. Their detailed description is available in the online Pyradiomics documentation (https://pyradiomics.readthedocs.io/en/latest/features.html). After feature stability analysis, 741 were retained for the subsequent steps. Of these, 4 had low variance, while 625 were highly intercorrelated. RFE then identified a 14-feature subset as most accurate (Fig. 4; feature list is available in supplementary material).
The ET model obtained an overall accuracy, in terms of correctly classified lesions, of 86% (± 10%) in the training set cross-validation. The classifier tuned parameters are reported in the supplementary materials. In the test set, the accuracy was of 93% (95%CI = 77–99%), sensitivity of 100%, and specificity of 87%. The AUC is of 0.99 (95%CI = 0.97–1.00) (Fig. 5), equal to the area under the precision-recall curve (0.99), often used in binary ML classifications (fig AUC). The classifier was significantly better (p = 8e−6) than the no information rate. The confusion matrix and detailed accuracy metrics are shown in Tables 2 and 3.
Discussion
Preoperative assessment of pituitary macroadenoma consistency is useful for planning surgical approach and reducing residuals and recurrence’s rate. For this reason, several studies have investigated the correlation between preoperative MRI features and tumor hardness. In particular, there are conflicting studies on the value of the relative signal on T2-weighted MRI and the macroadenoma consistency, with some works demonstrating a positive correlation between low signal and hardness [17,18,19,20] and other concluding that relative signal intensity values do not correlate [21,22,23,24]. Indeed besides collagen amount, which mainly correlates with the hardness, other factors such as intratumoral hematoma, amyloid, iron, calcification, or protein-rich fluid may affect the T2 signal intensity [25].
Diffusion-weighted imaging ability to predict tumor consistency also showed divergent results, both indicating a significant correlation [1, 26] and not [21, 27]. Furthermore, the lower spatial resolution and the presence of susceptibility artefacts in the sellar region related to bone and sinus pneumatization limit the use of this technique. Finally, in two studies by Romano et al. and Yamamoto et al., contrast-enhanced MRI showed a strong correlation for tumor consistency [2, 28]. Perfusion imaging parameters have also been investigated as possible biomarkers of pituitary macroadenoma consistency, but no added value was found compared with precontrast T1-weighted images [29]. A more interesting advanced technique in this setting is represented by MR elastography. Pituitary adenoma stiffness was found to correlate with their consistency and, if it became widely available, could offer additional data to mine with a radiomic approach [30, 31].
Regarding texture analysis, there are only three studies exploring this issue, to the best of our knowledge. In the first, Rui et al. explored the value of MRI texture analysis in assessing pituitary macroadenoma consistency, obtaining good accuracy values [32]. However, this study was conducted using contrast-enhanced 3D-SPACE images and without a ML approach. Fan and colleagues explored this issue in acromegalic patients using ML for radiomic feature selection prior to building a nomogram obtaining an AUC of 0.81 [33]. Zeynalova et al. also performed an analysis on ML preoperative evaluation of pituitary macroadenoma consistency [34]. Their study presented some similarities with our own. They also used Pyradiomics for feature extraction from bidimensional ROIs, although they utilized different sigma settings (2, 4, and 6 mm) for the LoG filter and obtained a total of 162 parameters. The lower number of features is probably due to their exclusive focus on first-order histogram-derived ones. These are more reproducible but convey less information on tissue texture compared with higher-order parameters. As their in-plane resolution was higher (0.5 × 0.8 mm), they were able to use a 1 × 1 mm resampling size compared with our 2 × 2 mm. It is interesting to note the use of a very narrow bin width value of 0.06, as the number of bins should not exceed 128, following the developer recommendations. They also performed a feature robustness assessment with our same ICC threshold, while their intercorrelation threshold was lower (0.7 vs 0.8). After data dimensionality reduction, they identified 6 informative features using the Weka data mining platform and a wrapper-based selector. In our study, the entire analysis was conducted using the scikit-learn Python package. Some other major differences are represented by the use of cross-validation, without further assessment on a separate test set. Their reported accuracy is 72.5%, with an AUC of 0.71. Therefore, our algorithm presents a clearly superior performance. This could be in part explained by their use of a multilayer perceptron neural network, which may not be the best suited algorithm for a small dataset obtained from 55 patients. Finally, Zenyalova and colleagues also used collagen amount within the tumor on histopathological examination for their reference standard. As consistency information is mainly useful for surgical strategy planning, we believe that intraoperative consistency assessment represents a more practical and useful reference standard as the final recipient of the information should be a neurosurgeon. As the two neurosurgeons involved in our study never had disagreements, we also found this assessment to be reproducible.
By analyzing our confusion matrix, it can be seen that the mistakes made by the classifier were 2 cases of soft lesions identified as fibrous. Given the clinical setting of our investigation, this kind of error is somewhat more acceptable than a false negative, as it would be more auspicable to sometimes overestimate the difficulty of a surgery rather than the opposite.
In our study, we chose to employ an ET ML algorithm. This belongs to the decision tree ensemble methods, in particular constituted by a large number of highly randomized decision trees which are fitted on data subsamples. Each of these outputs a prediction, and a majority vote determines the final outcome. Ensemble learning is based on the assumption that a decision by committee made by a large number of weak classifiers will perform better than a single algorithm. A sufficient diversification of the random trees included in the ET is guaranteed by random sampling, with replacement, of patients from the training dataset (bootstrap aggregation or bagging) and of their available features (n = 3 in our case). This in turn ensures low correlation of each tree, improving the ET’s overall performance [16]. As the dataset lesion classes were imbalanced, SMOTE was employed. This is a known solution to address this issue and has demonstrated its value in the setting of medical imaging radiomic ML analysis [15, 35,36,37].
We have chosen a handcrafted radiomics approach rather than a fully automated deep learning one as this gave us better control on the initial data analysis and following ML model construction. Both approaches have been object of discussion in current literature as they possess peculiar merits and limitations. It is our belief that a handcrafted analysis is more appropriate for relatively smaller datasets as it allows greater involvement of radiologists and better understanding of the whole pipeline. Only when extremely large datasets will become available in medical imaging, as in other fields, the less time-consuming completely neural network-based approach will be a practical necessity. Until then, the value of greater involvement of the radiologist and finer quality control of patient or lesion data outweigh the larger amount of time needed to extract medical imaging radiomics. Furthermore, medicine and especially treatments are evolving in the direction of precision, patient-tailored therapies. Contrary to the current desire in radiology to aggregate as many patients as possible to train ML algorithms, this determines a need to work with ever smaller patient subgroups within each pathological entity. Therefore, a future with space for both deep learning software to apply on large populations and engineered approaches for more specific tasks can be envisioned.
Our study has some limitations which have to be acknowledged. As is often the case for ML, future studies on larger populations are necessary to confirm and possibly expand our results. The need for oversampling given the unbalanced nature of the classes further highlights this necessity but was expected given epidemiological data. Only T2-weighted images were used, without investigating the added value of other sequences. However, obtaining valuable data without contrast agent administration and streamlining the pipeline to incorporate a single MRI sequence could also represent an added value. Furthermore, considering previous works, T2-weighted MRI alone proved effective to provide data concerning proliferative index [38], secretory activity [39], and response prediction to somatostatin analogues in patients with acromegaly and GH secreting pituitary macroadenoma [40, 41].
Conclusion
The ML model trained on radiomic data extracted from T2-weighted MRI demonstrated a high accuracy in the classification of soft and fibrous pituitary macroadenomas. Therefore, this tool could prove valuable in the pre-surgical planning of these patients if further developed and validated on larger datasets.
Abbreviations
- ML:
-
Machine learning
- ROI:
-
Region of interest
- ICC:
-
Intraclass correlation coefficient
- SMOTE:
-
Synthetic Minority Oversampling Technique
- RFE:
-
Recursive feature elimination
- ET:
-
Extra Trees Classifier
- AUC:
-
Area under the receiver operating characteristic curve
References
Alimohamadi M, Sanjari R, Mortazavi A, Shirani M, Moradi Tabriz H, Hadizadeh Kharazi H, Amirjamshidi A (2014) Predictive value of diffusion-weighted MRI for tumor consistency and resection rate of nonfunctional pituitary macroadenomas. Acta Neurochir 156:2245–2252. https://doi.org/10.1007/s00701-014-2259-6
Romano A, Coppola V, Lombardi M, Lavorato L, di Stefano D, Caroli E, Rossi Espagnet MC, Tavanti F, Minniti G, Trillò G, Bozzao A (2017) Predictive role of dynamic contrast enhanced T1-weighted MR sequences in pre-surgical evaluation of macroadenomas consistency. Pituitary 20:201–209. https://doi.org/10.1007/s11102-016-0760-z
Gillies RJ, Kinahan PE, Hricak H (2016) Radiomics: images are more than pictures, they are data. Radiology 278:563–577. https://doi.org/10.1148/radiol.2015151169
Senders JT, Zaki MM, Karhade AV, Chang B, Gormley WB, Broekman ML, Smith TR, Arnaout O (2018) An introduction and overview of machine learning in neurosurgical care. Acta Neurochir 160:29–38. https://doi.org/10.1007/s00701-017-3385-8
Cuocolo R, Stanzione A, Ponsiglione A, Romeo V, Verde F, Creta M, la Rocca R, Longo N, Pace L, Imbriaco M (2019) Clinically significant prostate cancer detection on MRI: a radiomic shape features study. Eur J Radiol 116:144–149. https://doi.org/10.1016/j.ejrad.2019.05.006
Cuocolo R, Cipullo MB, Stanzione A, Ugga L, Romeo V, Radice L, Brunetti A, Imbriaco M (2019) Machine learning applications in prostate cancer magnetic resonance imaging. Eur Radiol Exp 3:35. https://doi.org/10.1186/s41747-019-0109-2
Imbriaco M, Cuocolo R (2018) Does texture analysis of MR images of breast tumors help predict response to treatment? Radiology 286:421–423. https://doi.org/10.1148/radiol.2017172454
Mokrane F-Z, Lu L, Vavasseur A, Otal P, Peron JM, Luk L, Yang H, Ammari S, Saenger Y, Rousseau H, Zhao B, Schwartz LH, Dercle L (2020) Radiomics machine-learning signature for diagnosis of hepatocellular carcinoma in cirrhotic patients with indeterminate liver nodules. Eur Radiol 30:558–570. https://doi.org/10.1007/s00330-019-06347-w
Lotan E, Jain R, Razavian N, Fatterpekar GM, Lui YW (2019) State of the art: machine learning applications in Glioma imaging. Am J Roentgenol 212:26–37. https://doi.org/10.2214/AJR.18.20218
Solari D, Zenga F, Angileri FF, Barbanera A, Berlucchi S, Bernucci C, Carapella C, Catapano D, Catapano G, Cavallo LM, D'Arrigo C, de Angelis M, Denaro L, Desogus N, Ferroli P, Fontanella MM, Galzio RJ, Gianfreda CD, Iacoangeli M, Lauretti L, Locatelli D, Locatelli M, Luglietto D, Mazzatenta D, Menniti A, Milani D, Nasi MT, Romano A, Ruggeri AG, Saladino A, Santonocito O, Schwarz A, Skrap M, Stefini R, Volpin L, Wembagher GC, Zoia C, Zona G, Cappabianca P (2019) A survey on pituitary surgery in Italy. World Neurosurg 123:e440–e449. https://doi.org/10.1016/j.wneu.2018.11.186
Cappabianca P, Cavallo LM, de Divitiis O, Solari D, Esposito F, Colao A (2008) Endoscopic pituitary surgery. Pituitary 11:385–390. https://doi.org/10.1007/s11102-008-0087-5
Cappabianca P, Cavallo LM, Solari D, Stagno V, Esposito F, de Angelis M (2014) Endoscopic endonasal surgery for pituitary adenomas. World Neurosurg 82:S3–S11. https://doi.org/10.1016/j.wneu.2014.07.019
Solari D, Pivonello R, Caggiano C, Guadagno E, Chiaramonte C, Miccoli G, Cavallo LM, del Basso de Caro M, Colao A, Cappabianca P (2019) Pituitary adenomas: what are the key features? What are the current treatments? Where is the future taking us? World Neurosurg 127:695–709. https://doi.org/10.1016/j.wneu.2019.03.049
Di Maio S, Cavallo LM, Esposito F et al (2011) Extended endoscopic endonasal approach for selected pituitary adenomas: early experience. J Neurosurg 114:345–353. https://doi.org/10.3171/2010.9.JNS10262
Romeo V, Ricciardi C, Cuocolo R, Stanzione A, Verde F, Sarno L, Improta G, Mainenti PP, D'Armiento M, Brunetti A, Maurea S (2019) Machine learning analysis of MRI-derived texture features to predict placenta accreta spectrum in patients with placenta previa. Magn Reson Imaging 64:71–76. https://doi.org/10.1016/j.mri.2019.05.017
Breiman L (2001) Random forests. Mach Learn 45:5–32. https://doi.org/10.1023/A:1010933404324
Snow RB, Johnson CE, Morgello S, Lavyne MH, Patterson RH Jr (1990) Is magnetic resonance imaging useful in guiding the operative approach to large pituitary tumors? Neurosurgery 26:801–803. https://doi.org/10.1227/00006123-199005000-00011
Iuchi T, Saeki N, Tanaka M, Sunami K, Yamaura A (1998) MRI prediction of fibrous pituitary adenomas. Acta Neurochir 140:779–786. https://doi.org/10.1007/s007010050179
NAGANUMA H, SATOH E, NUKUI H (2002) Technical considerations of transsphenoidal removal of fibrous pituitary adenomas and evaluation of collagen content and subtype in the adenomas. Neurol Med Chir (Tokyo) 42:202–213. https://doi.org/10.2176/nmc.42.202
Smith K, Leever J, Chamoun R (2015) Prediction of consistency of pituitary adenomas by magnetic resonance imaging. J Neurol Surg Part B Skull Base 76:340–343. https://doi.org/10.1055/s-0035-1549005
Suzuki C, Maeda M, Hori K, Kozuka Y, Sakuma H, Taki W, Takeda K (2007) Apparent diffusion coefficient of pituitary macroadenoma evaluated with line-scan diffusion-weighted imaging. J Neuroradiol 34:228–235. https://doi.org/10.1016/j.neurad.2007.06.007
Chakrabortty S, Oi S, Yamaguchi M et al (1993) Growth hormone producing pituitary adenomas: MR characteristics and pre- and postoperative evaluation. Neurol Med Chir (Tokyo) 33:81–85. https://doi.org/10.2176/nmc.33.81
Bahuleyan B, Raghuram L, Rajshekhar V, Chacko AG (2006) To assess the ability of MRI to predict consistency of pituitary macroadenomas. Br J Neurosurg 20:324–326. https://doi.org/10.1080/02688690601000717
Thotakura AK, Patibandla MR, Panigrahi MK, Mahadevan A (2017) Is it really possible to predict the consistency of a pituitary adenoma preoperatively? Neurochirurgie 63:453–457. https://doi.org/10.1016/j.neuchi.2017.06.003
Hagiwara A, Inoue Y, Wakasa K, Haba T, Tashiro T, Miyamoto T (2003) Comparison of growth hormone–producing and non–growth hormone–producing pituitary adenomas: imaging characteristics and pathologic correlation. Radiology 228:533–538. https://doi.org/10.1148/radiol.2282020695
Pierallini A, Caramia F, Falcone C, Tinelli E, Paonessa A, Ciddio AB, Fiorelli M, Bianco F, Natalizi S, Ferrante L, Bozzao L (2006) Pituitary macroadenomas: preoperative evaluation of consistency with diffusion-weighted MR imaging—initial experience. Radiology 239:223–231. https://doi.org/10.1148/radiol.2383042204
Boxerman JL, Rogg JM, Donahue JE, Machan JT, Goldman MA, Doberstein CE (2010) Preoperative MRI evaluation of pituitary macroadenoma: imaging features predictive of successful transsphenoidal surgery. Am J Roentgenol 195:720–728. https://doi.org/10.2214/AJR.09.4128
Yamamoto J, Kakeda S, Shimajiri S, Takahashi M, Watanabe K, Kai Y, Moriya J, Korogi Y, Nishizawa S (2014) Tumor consistency of pituitary macroadenomas: predictive analysis on the basis of imaging features with contrast-enhanced 3D FIESTA at 3T. Am J Neuroradiol 35:297–303. https://doi.org/10.3174/ajnr.A3667
Ma Z, He W, Zhao Y, Yuan J, Zhang Q, Wu Y, Chen H, Yao Z, Li S, Wang Y (2016) Predictive value of PWI for blood supply and T1-spin echo MRI for consistency of pituitary adenoma. Neuroradiology 58:51–57. https://doi.org/10.1007/s00234-015-1591-8
Sakai N, Takehara Y, Yamashita S, Ohishi N, Kawaji H, Sameshima T, Baba S, Sakahara H, Namba H (2016) Shear stiffness of 4 common intracranial tumors measured using mr elastography: comparison with intraoperative consistency grading. Am J Neuroradiol 37:1851–1859. https://doi.org/10.3174/ajnr.A4832
Hughes JD, Fattahi N, Van Gompel J et al (2016) Magnetic resonance elastography detects tumoral consistency in pituitary macroadenomas. Pituitary. 19:286–292. https://doi.org/10.1007/s11102-016-0706-5
Rui W, Wu Y, Ma Z, Wang Y, Wang Y, Xu X, Zhang J, Yao Z (2019) MR textural analysis on contrast enhanced 3D-SPACE images in assessment of consistency of pituitary macroadenoma. Eur J Radiol 110:219–224. https://doi.org/10.1016/j.ejrad.2018.12.002
Fan Y, Hua M, Mou A, Wu M, Liu X, Bao X, Wang R, Feng M (2019) Preoperative noninvasive radiomics approach predicts tumor consistency in patients with acromegaly: development and multicenter prospective validation. Front Endocrinol (Lausanne) 10. https://doi.org/10.3389/fendo.2019.00403
Zeynalova A, Kocak B, Durmaz ES, Comunoglu N, Ozcan K, Ozcan G, Turk O, Tanriover N, Kocer N, Kizilkilic O, Islak C (2019) Preoperative evaluation of tumour consistency in pituitary macroadenomas: a machine learning-based histogram analysis on conventional T2-weighted MRI. Neuroradiology 61:767–774. https://doi.org/10.1007/s00234-019-02211-2
Antonelli M, Johnston EW, Dikaios N, Cheung KK, Sidhu HS, Appayya MB, Giganti F, Simmons LAM, Freeman A, Allen C, Ahmed HU, Atkinson D, Ourselin S, Punwani S (2019) Machine learning classifiers can predict Gleason pattern 4 prostate cancer with greater accuracy than experienced radiologists. Eur Radiol 29:4754–4764. https://doi.org/10.1007/s00330-019-06244-2
Fehr D, Veeraraghavan H, Wibmer A, Gondo T, Matsumoto K, Vargas HA, Sala E, Hricak H, Deasy JO (2015) Automatic classification of prostate cancer Gleason scores from multiparametric magnetic resonance images. Proc Natl Acad Sci 112:E6265–E6273. https://doi.org/10.1073/pnas.1505935112
Zhang C, Bi J, Soda P (2017) Feature selection and resampling in class imbalance learning: which comes first? An empirical study in the biological domain. In: 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM). IEEE, pp 933–938
Ugga L, Cuocolo R, Solari D, Guadagno E, D’Amico A, Somma T, Cappabianca P, del Basso de Caro ML, Cavallo LM, Brunetti A (2019) Prediction of high proliferative index in pituitary macroadenomas using MRI-based radiomics and machine learning. Neuroradiology 61:1365–1373. https://doi.org/10.1007/s00234-019-02266-1
Sanei Taheri M, Kimia F, Mehrnahad M, Saligheh Rad H, Haghighatkhah H, Moradi A, Kazerooni AF, Alviri M, Absalan A (2019) Accuracy of diffusion-weighted imaging-magnetic resonance in differentiating functional from non-functional pituitary macro-adenoma and classification of tumor consistency. Neuroradiol J 32:74–85. https://doi.org/10.1177/1971400918809825
Heck A, Emblem KE, Casar-Borota O, Bollerslev J, Ringstad G (2016) Quantitative analyses of T2-weighted MRI as a potential marker for response to somatostatin analogs in newly diagnosed acromegaly. Endocrine 52:333–343. https://doi.org/10.1007/s12020-015-0766-8
Kocak B, Durmaz ES, Kadioglu P, Polat Korkmaz O, Comunoglu N, Tanriover N, Kocer N, Islak C, Kizilkilic O (2019) Predicting response to somatostatin analogues in acromegaly: machine learning-based high-dimensional quantitative texture analysis on T2-weighted MRI. Eur Radiol 29:2731–2739. https://doi.org/10.1007/s00330-018-5876-2
Acknowledgements
Open access funding provided by Università degli Studi di Napoli Federico II within the CRUI-CARE Agreement.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical approval
All procedures performed in the studies involving human participants were in accordance with the ethical standards of the institutional and/or national research committee and with the 1964 Helsinki Declaration and its later amendments or comparable ethical standards.
Informed consent
Requirement for informed consent was waived by the local IRB.
Additional information
Publisher’s note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
ESM 1
(DOCX 15 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Cuocolo, R., Ugga, L., Solari, D. et al. Prediction of pituitary adenoma surgical consistency: radiomic data mining and machine learning on T2-weighted MRI. Neuroradiology 62, 1649–1656 (2020). https://doi.org/10.1007/s00234-020-02502-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00234-020-02502-z