Abstract
Marine animals such as the southern elephant seal (Mirounga leonina) rely on a productive marine environment and are vulnerable to oceanic changes that can affect their reproduction and survival rates. Davis Base, Antarctica, acts as a moulting site for southern elephant seals that forage in Prydz Bay, but the mitochondrial haplotype diversity and natal source populations of these seals have not been characterized. In this study, we combined genetic and animal tracking data on these moulting seals to identify levels of mitochondrial haplotype diversity, natal source population, and movement behaviours during foraging and haul-out periods. Using partial sequences of the mitochondrial control region, we identified two major breeding mitochondrial lineages of seals at Davis Base. We found that the majority of the seals originated from breeding stocks within the South Atlantic Ocean and South Indian Ocean. One seal was grouped with the Macquarie Island breeding stock (South Pacific Ocean). The Macquarie Island population, unlike the other two stocks, is decreasing in size. Tracking data revealed long-distance foraging activity of the Macquarie Island seal around Crozet Islands. We speculate that changes to the Antarctic marine environment can result in a shift in foraging and movement strategies, which subsequently affects seal population growth rates.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Warmer oceans result in a loss of sea ice, which is likely to affect reproduction and survival rates in species that depend on sea ice for foraging (Bryndum-Buchholz et al. 2019). In other species, loss of sea ice has reduced movement corridors, which has resulted in loss of connectivity pathways for some marine animal populations (Laidre et al. 2018). As global temperatures continue to increase, changes to habitat structure in ice-locked regions are likely to cause shifts in marine animal movements, effective dispersal, foraging behaviours, and population numbers (Hindell et al. 2017, 2020). These changes can lead to population restructuring or even loss of genetic diversity (Laidre et al. 2018; Siegert et al. 2019).
Over a 40-year data record from the late 1970s, the highest peak in Antarctic sea ice was evident in 2014, but large declines occurred in 2017 and 2018, possibly caused by the El Niño Southern Oscillation (Parkinson 2019). Polar marine animals, such as pinnipeds, occupy the upper trophic levels in the Antarctic region and therefore integrate large environmental signals (Bestley et al. 2020). Sea ice is crucial to the survival of these predators, especially those that require sea ice for breeding, foraging, and moulting (Bestley et al. 2020). A recent study used a climate model to project that colonies of the Emperor Penguin (Aptenodytes forsteri) would become quasi-extinct by 2100 because of the reduced availability of sea ice and foraging habitat (Jenouvrier et al. 2021). Similarly, changes in the extent of sea ice have altered the foraging behaviour and survival rates of a number of pinnipeds, such as the southern elephant seal (Mirounga leonina), which has ultimately affected the population dynamics of these species (McMahon and Burton 2005; Bestley et al. 2020; Bester 2021).
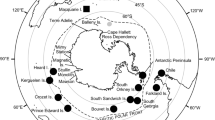
Southern elephant seals (SES) have a circumpolar distribution and haul out twice a year on sub-Antarctic islands to breed and to moult (Le Boeuf and Law 1994). The four main breeding stocks are South Georgia, Kerguelen Islands and Heard Island, Macquarie Island, and Península Valdés in Argentina (McMahon et al. 2017), with smaller breeding colonies on other sub-Antarctic islands (Slade et al. 1998; Hoelzel et al. 2001) (Fig. 1). Population decreases of SES were first documented in the mid-1980s (McMahon et al. 2005a). Population estimates between the 1980s and early 2000s demonstrated stable and increasing population sizes across three of the main breeding stocks and most of the smaller breeding stocks (McMahon et al. 2005a). However, the Macquarie Island breeding stock has continued to decrease and has been listed as vulnerable due to dramatic decreases in population size (McMahon et al. 2005a; van den Hoff and Burton 2007). Initial decreases were likely to have been driven by hunting for the seal’s oil-rich blubber in the early nineteenth century (Hindell and Burton 1988; van den Hoff and Burton 2007), but recent population declines were most likely caused by changes in food supply (McMahon et al. 2005a).
Circumpolar distribution of southern elephant seals around Antarctica. Blue squares represent the four main breeding stocks (South Georgia, Kerguelen Islands and Heard Island, Macquarie Island, and Península Valdés in Argentina). Black circles represent smaller breeding colonies (Marion Island, King George Island, Elephant Island, and Falkland Islands). The red star represents the sample collection site (Davis Base, Antarctica) for this study. Pink dashed outline represents the Polar Front. Solid yellow represents the Kerguelen Plateau. Outline of continents and island locations are from Free Vector Maps
During the 2.5-month period between the breeding and moulting seasons, and the eight-month period between moulting and breeding, SES spend the majority of their time foraging at sea. The timing and location of the seals’ foraging movements depend on energy requirements and the availability of food sources (Goedegebuure et al. 2018; de Kock et al. 2021). Polar research stations such as Casey Station (66°16′57″ S, 110°31′36″ E) and Davis Base (68°34′36″ S, 77°58′03″ E) act as moulting sites for some seal colonies (Rodríguez et al. 2017). Davis Base is an ice-free area that is situated near Vestfold Hills, covering roughly 400 km2 (Australian Antarctic Program 2020). Previous studies indicated that the male seals at Davis Base originated from the Kerguelen Islands, Heard Island, and some of the smaller populations in the Kerguelen stock at Marion and Crozet islands (Tierney 1977; Bester 1988). The seals’ annual visits might be due to the abundance of food along Prydz Bay during the austral summer (Bester 1988). The underlying genetic make-up of this and indeed other moulting aggregations can potentially be identified through genetic analysis. The results of such an analysis would allow comparison with traditional capture-mark-recapture studies that formed the basis of much previous work on population composition at Davis Station.
The genetic structure and genetic diversity of the breeding colonies of SES have been investigated previously using mitochondrial DNA (mtDNA) and microsatellites (Hoelzel et al. 1993; Slade et al. 1998; Fabiani 2002; Chauke 2008; Bogdanowicz et al. 2013). Previous studies have shown lower genetic diversity in the Península Valdés and Macquarie Island breeding stocks in both mtDNA and microsatellites (Slade et al. 1998; Fabiani 2002; Chauke 2008), whereas the study by Hoelzel et al. (1993) investigated only mtDNA. In contrast, genetic diversity was higher within the South Georgia stock (South Georgia, Elephant Island, Sea Lion Island, and Falkland Islands seals), and the Kerguelen and Heard Island stock (Heard Island and Marion Island seals), as seen from mitochondrial studies (Hoelzel et al. 1993) and other studies that investigated genetic diversity in both mtDNA and microsatellites (Slade et al. 1998; Chauke 2008; Bogdanowicz et al. 2013). Evidence of sex-biased dispersal was found in previous studies where male-biased gene flow and male dispersal occurred more than in females (Slade et al. 1998; Fabiani 2002; Chauke 2008; Bogdanowicz et al. 2013; Corrigan et al. 2016). However, there remains a gap in our knowledge regarding the natal origins of seals at moulting sites such as Davis Base, where seals potentially from multiple breeding sites aggregate (Bester 1988).
Southern elephant seals are polygynous and display philopatry, so breeding colonies have high female-to-male ratios (one male beachmaster to a female harem of up to 100 seals) (de Bruyn et al. 2011). Therefore, the reproductive success and dispersal of a single male can affect disproportionately the genetic structure of an entire breeding colony (Fabiani 2002; Chauke 2008). Nuclear DNA markers such as microsatellites are commonly used to infer male gene flow because these markers are inherited by both males and females. However, the natal source population of seal moulting aggregations can be inferred using molecular markers such as mtDNA, which is maternally inherited. Sequences of the mitochondrial control region effectively delineate the major breeding stocks of SES (Hoelzel et al. 1993; Slade et al. 1998; Bogadanowicz et al. 2013).
In this study, we aim to resolve the natal origins of SES that haul out at the moulting site at Davis Base, Antarctica. We analysed a partial sequence of the mitochondrial control region from blood samples and used tracking data from these seals to determine their mtDNA diversity, infer natal source location, and explore movement from Davis Base. We compared these data with previous studies of the genetic structure of surrounding breeding colonies. Our study shows that genetic and physical tracking data provide complementary information on the natal affiliations of the seals and their resource usage around the Kerguelen Plateau.
Materials and methods
Blood sample collection and animal tracking tags
We captured, collected blood samples from, and attached animal tracking devices to 12 male SES (10 subadults and 2 juveniles) at Davis Base in 2016 and 2017. Animal tracking data were collected over one year of the seals’ lifecycle, from the start of their trip (moulting at Davis Base) to returning to moult the following year. After collection, blood from each sampled seal was spotted onto Whatman FTA cards (Stowell et al. 2018) and stored at − 20 °C until required for DNA extraction. The 12 blood samples were then sent to the University of Sydney for the purpose of this study.
DNA library assembly
We prepared seal blood samples using the standard sterile technique, which involved taking a sterilized (Bunsen flame) standard office one-hole punch from the centre of the dried blood sample (Stowell et al. 2018). We extracted and purified the sampled DNA using the Qiagen DNeasy Blood & Tissue Kit following the Spin-Column protocol for blood with non-nucleated erythrocytes. To amplify the mitochondrial control region, we used the primers ancF (5′-GCTGACATTCTACTTAAACT-3′) and mdbR (5′-CAGTATAGAAACCCCCACATGA-3′) (de Bruyn et al. 2009). We ran PCRs for approximately 3 h using the following cycle protocol: 10 min at 95 °C followed by 35 cycles of 30 s at 52 °C, 30 s at 72 °C, 1 min at 72 °C, followed by an additional extension step for 10 min, and then 4 °C for cooldown. After successful PCR, we cleaned up the PCR product following the ExoSAP-IT Express PCR Product Cleanup kit and standard protocol. The cleaned PCR products were stored at 4 °C prior to DNA sequencing by Macrogen (Seoul, South Korea).
We collected a total of 201 additional sequences of the mitochondrial control region from GenBank (Hoelzel et al. 1993; Slade et al. 1998; Fabiani 2002; Chauke 2008; Curtis et al. 2009; Bogdanowicz et al. 2013; Zappes et al. 2017). These comprised 191 sequences from southern elephant seals and 10 sequences from Weddell seals (Leptonychotes weddellii), which we included as an outgroup because they belong to the same subfamily of Phocidae (Slade et al. 1994). The southern elephant seal data downloaded from GenBank had been sampled from the following locations: Marion Island (n = 50), King George Island (n = 23), Macquarie Island (n = 53), Elephant Island (n = 12), Falkland Islands (n = 16), Península Valdés (n = 32), Heard Island (n = 6), and South Georgia (n = 28). We aligned the 12 newly generated sequences and the 201 published sequences using Geneious Prime (Kearse et al. 2012).
Genetic analyses
Uncorrected pairwise distances were calculated from sampled sequences of the mitochondrial control region using MEGA X (Kumar et al. 2018). We used pairwise deletion to account for gaps in sequences. We used DnaSP version 6.12 (Rozas et al. 2017) to calculate the number of segregating sites, number of haplotypes and haplotype diversity, nucleotide diversity and average nucleotide differences, and neutrality tests using Fu’s FS (Fu 1997). Rarefaction was used to correct for unequal sample sizes by comparing haplotype richness between the samples from Davis Base and from all other populations. The sampled haplotype sequences were rarefied to generate the expected haplotype richness using the function rarefy (Hurlbert 1971; Heck et al. 1975) in R package vegan (Oksanen et al. 2020).
To visualize the relationships among mitochondrial haplotypes, we constructed a median-joining haplotype network using the software package POPART (Leigh and Bryant 2015). We assigned each seal sequence to one of nine geographical locations: Marion Island (MR), King George Island (KG), Macquarie Island (MQ), Elephant Island (EI), Falkland Islands (FI), Península Valdéz (PV), Heard Island (HD), South Georgia (SG), and Davis Base. The sequence data collected from GenBank were assigned locations according to where they were collected for the referenced study.
Animal tracking
We immobilized the 12 seals as part of an integrated oceanography and animal behaviour study (McMahon et al. 2021). From each seal, we took morphometric measurements including standard body length, maximum girth, and weight (Field et al. 2002). Each seal was anaesthetised using Zoletil 100, or available combinations of Tiletamine and Zolazepam (McMahon et al. 2000). We then attached identification tags to the hind flippers using Dalton Jumbo Robotags (Wilkinson and Bester 1997). We tracked the 12 seals over a year of their lifecycle from post-moulting to the next moulting haul-out season the following year (Table 1). Conductivity-Temperature-Depth Satellite Data Relay Loggers (CTD-SRDL, Sea Mammal Research Unit, University of St Andrews, UK) were glued to the top of the seal’s head for improved satellite reception and transmission at sea (Horning et al. 2019). Based on a previous study, the tag attachments to the seals were believed to have had no impact on their reproduction or survival patterns (McMahon et al. 2008). Tracking data received from the tags, such as location and diving behaviour, were transmitted via the ARGOS satellite network (Myers et al. 2006; Henderson et al. 2020).
We analysed the tracking data using the R package foieGras (Jonsen and Patterson 2020). We first used the fit_ssm function to fit a continuous-time correlated random walk state-space model (SSM; Jonsen et al. 2020) to the ARGOS satellite-derived locations. This model accounted for well-known measurement errors in the ARGOS locations and predicted locations at regular 12-h time intervals along the seal tracks (as per Jonsen et al. 2019). We then used the fit_mpm function to fit a movement persistence model (Jonsen et al. 2019) to the predicted locations to infer changes in the seals’ movement behaviour, possibly arising in response to stimuli such as changes in prey density or ice concentration, along their estimated tracks. Movement persistence (γt) is the autocorrelation in both speed and direction (scaled from 0 to 1) between successive displacements along a movement pathway. Low γt values represent low speed and/or directionality that are typical of resident or area-restricted searching behaviours, whereas high γt values represent higher speed and/or directionality that are typical of directed travel associated with dispersal or migration. Using the SSM-predicted locations, we also calculated the following track summary statistics: maximum displacement from deployment location; total deployment duration; maximum displacement scaled by deployment duration; and path tortuosity (mean vector of turning angles along each seal’s track).
Results
Population genetic diversity
Nucleotide sequences of the mitochondrial control region (348 bp), sampled from the 12 SES at Davis Base, showed extensive divergence between sample 48_SES and the other 11 samples (the latter were all more closely related to each other, and do not include 48_SES). Analysis of these 12 sequences and 191 published southern elephant seal sequences revealed a close relationship between sample 48_SES and sequences from Macquarie Island seals. The largest pairwise genetic distance (0.065 substitutions site−1) is seen between seals from Macquarie Island and Península Valdés (Table 2). This is expected, given that the two locations are geographically the farthest apart. The pairwise distance between the Davis Base and Macquarie Island seals (0.049 substitutions site−1) is greater than that between the Davis Base seals and other population groups. This confirms the divergence between sample 48_SES and the other 11 Davis Base seals.
We identified a total of 65 haplotypes in the combined data set of 203 mitochondrial sequences (Table 3). From the 12 seals from Davis Base, 11 haplotypes were identified (two individual samples 42_SES and 62_SES shared the same haplotype). A single haplotype was carried by 26 individual seals that were previously sampled from King George Island, Marion Island, Falkland Islands, Heard Island, and one individual from Elephant Island. Six haplotypes from Davis Base seals were shared with those from Marion Island, and three haplotypes with those from the Falkland Islands. The final Davis Base haplotype grouped with the Macquarie Island population as described above. The seals from Davis Base had high haplotype diversity (0.99 ± 0.04; \(n\) = 12) and had higher nucleotide diversity (2.52 ± 0.004; \(n\) = 12) than the seals from the other populations (Macquarie Island, Marion Island, Heard Island, King George Island, and Península Valdés). Haplotype richness and average number of nucleotide differences were elevated in the Davis Base seals compared with all other populations.
Haplotype network
We assigned locations to the 12 sequences from Davis Base seals based on their haplotype grouping (Fig. 2). For the seal sequences from GenBank, we assigned locations based on their sampling information. Of the 12 Davis Base seals, three had unique mitochondrial haplotypes. Two seals had haplotypes that were shared with Marion Island seals, three seals had haplotypes that were shared with seals from breeding colonies in the South Atlantic and South Indian Ocean, three seals had haplotypes shared with those from Marion Island and the South Atlantic Ocean breeding colonies, and one seal shared a haplotype with Macquarie Island seals (see below). From the combined analysis of all population groups and the Davis Base seals, 12 haplotypes were unique to the Macquarie Island stock. One Davis Base seal (48_SES) shared a haplotype with four seals from Macquarie Island. Eleven haplotypes were unique among seals from Marion Island. Out of the 11 Marion Island haplotypes, four haplotypes were shared by seals from the Falkland Islands, three shared by seals from each of Elephant Island, Heard Island, and King George Island, and two shared by seals from South Georgia. Islands in the South Atlantic Ocean region mostly shared haplotypes with one another, with only 7.7% of haplotypes shared with populations on islands in the South Indian Ocean. We found no common haplotypes between Macquarie Island (South Pacific Ocean), the South Atlantic Ocean, and South Indian Ocean islands.
Median-joining network of mitochondrial haplotypes shared between southern elephant seals from Davis Base and all other major populations. Sizes of circles represent the number of individuals per haplotype. Each colour in the pie charts represents a unique population group. The South Georgia breeding stocks are indicated as shades of green (dark blue for Península Valdés). The Kerguelen and Heard Island breeding stocks are indicated as shades of red. The Macquarie Island breeding stock are indicated by yellow. Davis Base samples are indicated as purple. Black dots represent unobserved haplotypes that have one mutational step from adjacent haplotypes. Hatch marks on lines connecting haplotypes indicate mutations. ‘\(n\) = x’ represents the number of Davis Base individuals that carry the specific haplotype
Animal tracking analyses
We analysed track summaries, estimated move persistence, and predicted locations of the 12 seal samples from Davis Base (Table 4). Between 2005 and 2017, we estimated that the 12 tagged seals account for approximately 9.7% of the total moulting population (see Online Resource 1). Track summaries indicated that the average maximum displacement of the seals from Davis Base was 1330 km, with 44_SES travelling the greatest distance (3056 km). The average maximum displacement scaled by duration was found to be 8.1 km day−1, with the greatest being 14.5 km day−1 (62_SES), and the smallest being 1.30 km day−1 (seal 60). The average path tortuosity was found to be 0.078, with 58_SES having the highest path tortuosity (0.188), and 44_SES and 54_SES with the lowest (0.020 and 0.021, respectively).
The sampled seals showed high move persistence during their outbound trips into open waters, and low move persistence when approaching areas with high sea-ice coverage or when returning to breeding and moulting areas (Fig. 3). Predicted locations have indicated visits to Kerguelen Islands by 44_SES, 50_SES, 56_SES, and 62_SES (Fig. 4). Seal 48_SES made visits to Crozet Islands and appeared to spend the majority of its journey around those islands. The predicted locations also indicated that five individuals did not venture out of the Davis Base ice shelf regions. This might be due to seal mortality or detachment of the tags.
Map of sampled seals’ move persistence by measure of path tortuosity. γt values approaching 1.00 (yellow) indicate relatively fast, directed movement. γt values approaching 0.00 (navy) indicate high tortuosity and slow movement. Davis Base is indicated by the purple star. Main breeding islands are represented by purple circles. Ice concentrations (maximum extent of sea ice coverage > 15% during winter) are represented by light grey along the coastline. The Kerguelen Plateau is shown in solid brown
a State-space model predicted locations of all sampled seals. Seal 60_SES failed to converge on predicted locations due to small scales and low contrasts of movement. The blue box indicates a narrowed view of the cluster of seal tracks close to Davis Base as seen in (b). Davis Base is indicated by the purple star. Breeding islands are represented by purple circles. Ice concentrations (maximum extent of sea ice coverage > 15% during winter) are represented by light grey along the coastline. Small grey boxes represent end data points of the individual seal tracks
Discussion
By combining genetic and animal tracking data, we determined the natal locations and at-sea movements of moulting seals at Davis Base, Antarctica. Our analysis of genetic data revealed a mixed sample of likely natal locations from the 12 Davis Base seals, including some of the main breeding stocks (South Georgia, Macquarie Island, and Marion Island) and two of the smaller breeding colonies (King George Island and the Falkland Islands). We also identified two distinct mtDNA lineages, representing three of the four main breeding stocks (Macquarie Island, South Georgia, and Kerguelen and Heard Islands stocks, but excluding Península Valdés). Our tracking data showed that all seals, despite their varied lineages, remained within the vicinity of Crozet Island, Kerguelen Islands, and Heard Island for the duration of the tracking study. Seals that showed short displacements and less move persistence travelled more frequently around areas of high ice concentration. Overall, our integrated genetic and telemetry analyses provided longer-term insights into the seals’ natal affiliations and migration strategies, and additional short-term information on their annual life-cycle movements and foraging ecology.
The genetic data from 12 SES collected at Davis Base suggested high nucleotide diversity and haplotypes shared with elephant seals from other breeding colonies within the South Atlantic Ocean, South Indian Ocean, and South Pacific Ocean. However, given the small sample size, our study might not be representative of the larger population of all male SES at Davis Base. Replication of this study with a larger sample size and analysing nuclear markers, such as microsatellites, to clarify male gene flow and levels of interbreeding would assist in describing more comprehensively the genetic make-up of this moulting population. Our mitochondrial evidence shows that one seal had natal source affinities with the Macquarie Island stock, approximately 4207 km away, whilst the other 11 Davis Base seals showed affinities with the South Georgia stock (South Georgia, Elephant Island, Falkland Island, and King George Island), and the Kerguelen and Heard Island stock (Heard Island and Marion Island).
Animal tracking analyses
The male seals from our study spent more time in areas of high ice concentration and typically involved intense search behaviours exemplified by highly tortuous movements. Similar patterns have been observed previously in male seals that adopted sea-ice foraging strategies rather than pelagic foraging (Labrousse et al. 2017; Rodríguez et al. 2017; Jonsen et al. 2019; Hindell et al. 2021). Moreover, the preferences of foraging locations along the Kerguelen Plateau were shown by seal 50_SES, which had less move persistence near Kerguelen Islands. Five other seal tracks (44_SES, 48_SES, 50_SES, 56_SES, and 62_SES) showed patterns of inter-island movements by outbound trips made towards the Kerguelen Plateau and Crozet Islands, which might indicate pelagic foraging trips away from the ice shelf areas. Similarly, seal 48_SES showed less move persistence and had the second-longest maximum displacement per day around Crozet Islands (which is approximately 6900 km from its natal affinity, Macquarie Island). This suggests a long-distance movement event to gain foraging (and potentially future breeding) advantage away from the seal’s natal location.
SES are philopatric to breeding and foraging sites, so long-distance movements are rare, particularly for a juvenile seal (Hindell and McMahon 2000; Reisinger and Bester 2010). Our findings show that the only two male juvenile seals, out of the 12 seals included in this study, remained close to the ice shelf region to forage close to Davis Base, which was indicated by the reduced move persistence. Seals that remain close to the ice shelf region might have easier access to sea-ice polynyas where their prey (myctophids and ice fish) may be more easily found (Labrousse et al. 2017). However, a previous study found that a juvenile female from Macquarie Island was sighted at Peter 1 Øy, which is approximately 5,200 km to the east (Hindell and McMahon 2000). This finding corresponds with that observed here of 48_SES (Macquarie Island natal affinity), which foraged around Crozet Islands, as discussed above. The parallel finding between this study and previous studies might indicate a lack of resource availability around Macquarie Island where the population has been in decline in recent decades (McMahon et al. 2005a), which could be forcing seals from this location to travel farther to find food.
Changes to foraging site and the consequences on populations
The Kerguelen Plateau is a popular foraging location for male SES. However, there are high levels of predation on SES here (Hindell et al. 2021), which might have caused terminations of seal tracking signals. The Kerguelen Plateau lies within the Antarctic Polar Front boundary, which is an area of elevated productivity for most Antarctic marine species due to the decrease in water temperature down to 2 °C at 200 m, and the distribution of water masses and associated abundance of primary producers (O’Toole et al. 2014; Cristofari et al. 2018). SES will often forage on the Antarctic Continental Shelf and Polar Front, and because they are a deep-diving species, sea-surface temperatures have less of an effect on their diving behaviours (Hindell et al. 1991; O’Toole et al. 2014). Alternatively, previous studies have found that seals have greater foraging opportunities in colder waters, particularly along the Antarctic shelf; temperatures slow down the movements of prey, allowing their capture with less energy expenditure from the seals (Bailleul et al. 2007).
Popular foraging locations around the Kerguelen Plateau, such as Crozet Islands and Kerguelen Islands, have been reported to show either a slight increase or a stabilization in SES population numbers over the last decade (Guinet et al. 2004). The stabilized population numbers might have been caused by the seals’ inter-island movements and foraging between islands in the Kerguelen province, including Marion Island, Crozet Islands, Kerguelen Islands, and Heard Island (Oosthuizen et al. 2011). However, despite showing philopatry to foraging sites, the seals’ foraging strategies and movements might shift based on the changes to oceanic conditions that will influence where resources are available (Bailleul et al. 2007). Moreover, poor foraging success by females has led to a decrease in first-year pup survival and would thus reduce reproductive success in populations such as Macquarie Island (Arnbom et al. 1997; McMahon et al. 2003; Clausius et al. 2017; Mestre et al. 2020). As oceanic conditions continue to change with climate, the entire marine ecosystem will also shift. Therefore, the lack of resources around Macquarie Island is likely to continue causing decreases in population size (McMahon et al. 2005a; Clausius et al. 2017).
Population declines impacted by climate change
Broad climate events such as El Niño can affect the oceanic structure of the Southern Ocean. The Southern Ocean ecosystem relies heavily on phytoplankton and krill abundance for other species’ survival, previously reported for Antarctic whales, King Penguins (Aptenodytes patagonicus), and SES (McIntyre et al. 2014; Cristofari et al. 2018; Bestley et al. 2020; Rogers et al. 2020; Agrelo et al. 2021; Volzke et al. 2021). Over the last four decades, the global SES population has seen dramatic declines in some populations due to the changes in food availability (McMahon et al. 2005a; Volzke et al. 2021). Populations of the four main breeding stocks, South Georgia, Kerguelen Islands and Heard Island, Macquarie Island, and Península Valdés, have all decreased since the 1970s. However, presently all the populations barring the Macquarie population are either stable or increasing (McMahon et al. 2005b; Hindell et al. 2016). A more thorough understanding of seal foraging behaviour, the selection and variations in foraging sites, and how this is expressed and transmitted within a population is central to understanding how foraging site selection affects population growth and ultimately population viability.
In summary, we were able to identify the likely natal locations of the 12 Davis Base seals through genetic data. We combined findings from genetic data with satellite telemetry tracking data and identified that the majority of the seals are spending most of the time foraging along the Kerguelen Plateau. Our data suggest that a seal from Macquarie Island has travelled a long distance, probably to gain a foraging advantage over its conspecifics. Long-range migrations and movements to distant feeding grounds might be one way for seals to maximize foraging efficiency, which may affect population growth rates through changes in survival and reproduction.
Data availability
The data sets generated during and/or analysed during the current study are available from the corresponding author on reasonable request. DNA sequences have been deposited in GenBank, with accession numbers ON571492–ON571503.
References
Agrelo M, Daura-Jorge FG, Rowntree VJ, Sironi M, Hammond PS, Ingram SN, Marón CF, Vilches FO, Seger J, Payne R, Simões-Lopes PC (2021) Ocean warming threatens southern right whale population recovery. Sci Adv 7:2823
Arnbom T, Fedak MA, Boyd IL (1997) Factors affecting maternal expenditure in southern elephant seals during lactation. Ecology 7:471–483
Australian Antarctic Program (2020) Davis Environment. Australian Government: Australian Antarctic Division. https://www.antarctica.gov.au/antarctic-operations/stations/davis/environment/. Accessed 14 Oct 2021
Bailleul F, Charrassin JB, Monestiez P, Roquet F, Biuw M, Guinet C (2007) Successful foraging zones of southern elephant seals from the Kerguelen Islands in relation to oceanographic conditions. Philos Trans R Soc B 362:2169–2181
Bester MN (1988) Marking and monitoring studies of the Kerguelen stock of southern elephant seals Mirounga leonina and their bearing on biological research in the Vestfold Hills. Hydrobiologia 165:269–277
Bester MN (2021) Status of pinnipeds on mid-Atlantic ridge islands, South Atlantic Ocean. Polar Biol 44:865–871
Bestley S, Ropert-Coudert Y, Nash SB, Brooks CM, Cotte C, Dewar M, Friedlaender AS, Jackson JA, Labrousse S, Lowther AD, McMahon CR, Phillips RA, Pistorius P, Puskic PS, Reis AOA, Reisinger RR, Santos M, Tarszisz E, Tixier P, Trathan PN, Wege M, Wienecke B (2020) Marine ecosystem assessment for the Southern Ocean: birds and marine mammals in a changing climate. Front Ecol Evol 8:566936
Bogdanowicz W, Pilot M, Gajewska M, Suchecka E, Golachowski M (2013) Genetic diversity in a moulting colony of southern elephant seals in comparison with breeding colonies. Mar Ecol Progr Ser 478:287–300
Bryndum-Buchholz A, Tittensor DP, Blanchard JL, Cheung WWL, Coll M, Galbraith ED, Jennings S, Maury O, Lotze HK (2019) Twenty-first-century climate change impacts on marine animal biomass and ecosystem structure across ocean basins. Glob Change Biol 25:459–472
Chauke LF (2008) Genetic variation and population structure of southern elephant seals Mirounga leonina from Marion Island. Dissertation, University of Pretoria
Clausius E, McMahon CR, Hindell MA (2017) Five decades on: Use of historical weaning size data reveals that a decrease in maternal foraging success underpins the long-term decline in population of southern elephant seals (Mirounga leonina). PLOS ONE 12:e0173427
Corrigan LJ, Fabiani A, Chauke LF, McMahon CR, de Bruyn M, Bester MN, Bastos A, Campagna C, Muelbert MMC, Hoelzel AR (2016) Population differentiation in the context of Holocene climate change for a migratory marine species, the southern elephant seal. J Evol Biol 29:1667–1679
Cristofari R, Liu X, Bonadonna F, Cherel Y, Pistorius P, Maho YL, Raybaud V, Stenseth NC, Bohec CL, Trucchi E (2018) Climate-driven range shifts of the king penguin in a fragmented ecosystem. Nat Clim Change 8:245–251
Curtis C, Stewart BS, Karl SA (2009) Pleistocene population expansions of Antarctic seals. Mol Ecol 18:112–2121
de Bruyn M, Hall BL, Chauke LF, Baroni C, Koch PL, Hoelzel AR (2009) Rapid response of a marine mammal species to holocene climate and habitat change. PLOS Genet 5:e1000554
de Bruyn PJN, Tosh CA, Bester MN, Cameron EZ, McIntyre T, Wilkinson IS (2011) Sex at sea: alternative mating system in an extremely polygynous mammal. Anim Behav 82:445–451
de Kock L, Oosthuizen WC, Beltran RS, Bester MN, de Bruyn PJN (2021) Determinants of moult haul-out phenology and duration in southern elephant seals. Sci Rep 11:13331
Fabiani A (2002). Molecular ecology of southern elephant seals (Mirounga leonina): mating system and population genetics. Dissertation, Durham University.
Field IC, Bradshaw CJA, McMahon CR, Harrington J, Burton HR (2002) Effects of age, size and condition of elephant seals (Mirounga leonina) on their intravenous anaesthesia with tiletamine and zolazepam. Vet Rec 151:235–240
Fu YX (1997) Statistical tests of neutrality of mutations against population growth, hitchhiking and background selection. Genetics 147:915–925
Goedegebuure M, Melbourne-Thomas J, Corney SP, McMahon CR, Hindell MA (2018) Modelling southern elephant seals (Mirounga leonina) using an individual-based model coupled with a dynamic energy budget. PLOS ONE 13:e0194950
Guinet C, Jouventin P, Weimerskirch H (2004) Recent population change of the southern elephant seal at Îles Crozet and Îles Kerguelen: the end of the decrease? Antarct Sci 11:193–197
Heck KL, van Belle G, Simberloff D (1975) Explicit calculation of the rarefaction diversity measurement and the determination of sufficient sample size. Ecology 56:1459–1461
Henderson AF, McMahon CR, Harcourt R, Guinet C, Picard B, Wotherspoon S, Hindell MA (2020) Inferring variation in southern elephant seal at-sea mortality by modelling tag failure. Front Mar Sci 7:517901
Hindell MA, Burton HR (1988) The history of the elephant seal industry at Macquarie Island and an estimate of pre-sealing numbers. Pap Proc R Soc Tasmania 122:159–176
Hindell MA, Burton HR, Slip DJ (1991) Foraging areas of southern elephant seals, (Mirounga leonina), as inferred from water temperature data.Aust J Mar Freshwater Res 42:115–128
Hindell MA, McMahon CR (2000) Long distance movement of a southern elephant seal (Mirounga leonina) from Macquarie Island to Peter 1 Øy. Mar Mammal Sci 16:504–507
Hindell MA, McMahon CR, Bester MN, Boehme L, Costa D, Fedak MA, Guinet C, Herraiz-Borreguero L, Harcourt RG, Huckstadt L, Kovacs KM, Lydersen C, McIntyre T, Muelbert MMC, Patterson T, Roquet F, Williams G, Charrassin JB (2016) Circumpolar habitat use in the southern elephant seal: implications for foraging success and population trajectories. Ecosphere 7:e01213
Hindell MA, Sumner M, Bestley S, Wotherspoon S, Harcourt R, Lea M-A, Alderman R, McMahon CR (2017) Decadal changes in habitat characteristics influence population trajectories of southern elephant seals. Glob Change Biol 23:5136–5150
Hindell MA, Reisinger RR, Ropert-Coudert Y, Hückstädt LA, Trathan PN et al (2020) Tracking of marine predators to protect Southern Ocean ecosystems. Nature 580:87–92
Hindell MA, McMahon CR, Jonsen I, Harcourt R, Arce F, Guinet C (2021) Inter- and intrasex habitat partitioning in the highly dimorphic southern elephant seal. Ecol Evol 11:1620–1633
Hoelzel AR, Campagna C, Arnbom T (2001) Genetic and morphometric differentiation between island and mainland southern elephant seal populations. Proc Biol Sci 268:325–332
Hoelzel AR, Halley J, O’Brien SJ, Campagna C, Arnbom T, Le Boeuf B, Ralls K, Dover GA (1993) Elephant seal genetic variation and the use of simulation models to investigate historical population bottlenecks. J Hered 84:443–449
Horning M, Andrews RD, Bishop AM, Boveng PL, Costa DP, Crocker DE, Haulena M, Hindell M, Hindle AG, Holser RR, Hooker SK, Hückstädt LA, Johnson S, Lea MA, McDonald BI, McMahon CR, Robinson PW, Sattler RL, Shuert CR, Steingass SM, Thompson D, Tuomi PA, Williams CL, Womble JN (2019) Best practice recommendations for the use of external telemetry devices on pinnipeds. Anim Biotelem 7:20
Hurlbert SH (1971) The non-concept of species diversity: a critique and alternative parameters. Ecology 52:577–586
Jenouvrier S, Che-Castaldo J, Wolf S, Holland M, Labrousse S, LaRue M, Wienecke B, Fretwell P, Barbraud C, Greenwald N, Stroeve J, Trathan PN (2021) The call of the emperor penguin: Legal responses to species threatened by climate change. Glob Change Biol 27:5008–5029
Jonsen ID, Patterson TA (2020) foieGras: fit latent variable movement models to animal tracking data for location quality control and behavioural inference. R package version 0.7–6. The Comprehensive R Archive Network. https://CRAN.R-project.org/package=foieGras. Accessed 07 Jul 2021
Jonsen ID, McMahon CR, Patterson TA, Auger-Méthé M, Harcourt R, Hindell MA, Bestley S (2019) Movement responses to environment: fast inference of variation among southern elephant seals with a mixed effects model. Ecology 100:e02566
Jonsen ID, Patterson TA, Costa DP, Doherty PD, Godley BJ et al (2020) A continuous-time state-space model for rapid quality control of Argos locations from animal-borne tags. Move Ecol 8:31
Kearse M, Moir R, Wilson A, Stones-Havas S, Cheung M, Sturrock S, Buxton S, Cooper A, Markowitz S, Duran C, Thierer T, Ashton B, Meintjes P, Drummond A (2012) Geneious basic: an integrated and extendable desktop software platform for the organisation and analysis of sequence data. Bioinformatics 28:1647–1649
Kumar S, Stecher G, Li M, Knyaz C, Tamura K (2018) MEGA X: Molecular Evolutionary Genetics Analysis across computing platforms. Mol Biol Evol 35:1547–1549
Labrousse S, Sallee JB, Fraser AD, Massom RA, Reid P, Sumner M, Guinet C, Harcourt R, McMahon CR, Bailleul F, Hindell MA, Charrassin JB (2017) Under the sea ice: Exploring the relationship between sea ice and the foraging behaviour of southern elephant seals in East Antarctica. Progr Oceanogr 156:17–40
Laidre KL, Born EW, Atkinson SN, Wiig Ø, Andersen LW, Lunn NJ, Dyck M, Regehr EV, McGovern R, Heagerty P (2018) Range contraction and increasing isolation of polar bear subpopulation in an era of sea-ice loss. Ecol Evol 8:2062–2075
Le Boeuf BJ, Law RM (1994) Elephant seals: population ecology, behaviour, and physiology. University of California Press, Berkley
Leigh JW, Bryant D (2015) POPART: full-feature software for haplotype network construction. Methods Ecol Evol 6:1110–1116
McIntyre T, Bornemann H, de Bruyn PJN, Reisinger RR, Steinhage D, Márquez MEI, Bester MN, Plötz J (2014) Environmental influences on the at-sea behaviour of a major consumer, Mirounga leonina, in a rapidly changing environment. Polar Res 33:23808
McMahon CR, Burton H (2005) Climate change and seal survival: evidence for environmentally mediated changes in elephant seal, Mirounga leonina, pup survival. Proc R Soc B 272:923–928
McMahon CR, Burton HR, McLean S, Slip D, Bester MN (2000) Field immobilisation of southern elephant seals with intravenous tiletamine and zolazepam. Vet Rec 146:251–254
McMahon CR, Burton H, Bester MN (2003) A demographic comparison of two southern elephant seal populations. J Anim Ecol 72:61–74
McMahon CR, Bester MN, Burton HR, Hindell MA, Bradshaw CJA (2005a) Population status, trends and a re-examination of the hypotheses explaining the recent declines of the southern elephant seal Mirounga leonina. Mammal Rev 35:82–100
McMahon CR, Hindell MA, Burton HR, Bester MN (2005b) Comparison of southern elephant seal populations, and observations of a population on a demographic knife-edge. Mar Ecol Progr Ser 288:273–283
McMahon CR, Field IC, Bradshaw CJA, White GC, Hindell MA (2008) Tracking and data-logging devices attached to elephant seals do not affect individual mass gain or survival. J Exp Mar Biol Ecol 360:71–77
McMahon CR, Thums M, Bradshaw M, Busby S, Chapple V, Evans M, Goodlich S, Holland C, Raudino H, Rebuck P, Hindell MA (2017) It’s a girl! A female southern elephant seal born in Western Australia. Aust J Zool 65:179–182
McMahon CR, Roquet F, Baudel S, Belbeoch M, Bestley S, Blight C, Boehme L, Carse F, Costa DP, Fedak MA, Guinet C, Harcourt R, Heslop E, Hindell MA, Hoenner X, Holland K, Holland M, Jaine FRA, Jeanniard Du Dot T, Jonsen ID, Keates TR, Kovacs KM, Labrousse S, Lovell P, Lydersen C, March D, Mazloff M, McKinzie MK, Muelbert MMC, O’Brien KM, Phillips LR, Portela E, Pye J, Rintoul S, Sato K, Sequeira AMM, Simmons SE, Tsontos VM, Turpin V, van Wijk E, Vo D, Wege M, Whoriskey FG, Wilson K, Woodward BE (2021) Animal Borne Ocean Sensors – AniBOS – an essential component of the Global Ocean Observing System (GOOS) Front Mar Sci 8:751840
Mestre J, Authier M, Cherel Y, Harcourt R, McMahon CR, Hindell MA, Charrassin JB, Guinet C (2020) Decadal changes in blood δ13C values, at-sea distribution, and weaning mass of southern elephant seals from Kerguelen Islands. Proc R Soc B 287:20201544
Myers AE, Lovell P, Hays GC (2006) Tools for studying animal behaviour: validation of dive profiles relayed via the Argos satellite system. Anim Behav 71:989–993
Oksanen J, Blanchet FG, Friendly M, Kindt R, Legendre P, McGlinn D, Minchin PR, O'Hara RB, Simpson GL, Solymos P, Stevens MHH, Szoecs E, Wagner H (2020) vegan: Community ecology package. R package version 2.5–6. 2019. The Comprehensive R Archive Network. https://cran.r-project.org/package=vegan. Accessed 07 Jul 2021
Oosthuizen WC, Bester MN, Tosh CA, Guinet C, Besson D, de Bruyn PJN (2011) Dispersal and dispersion of southern elephant seals in the Kerguelen province, Southern Ocean. Antarct Sci 23:567–577
O’Toole M, Hindell MA, Charrassin JB, Guinet C (2014) Foraging behaviour of the southern elephant seals over the Kerguelen Plateau. Mar Ecol Progr Ser 502:281–294
Parkinson CL (2019) A 40-y record reveals gradual Antarctic sea ice increases followed by decreases at rates far exceeding the rates seen in the Arctic. Proc Natl Acad Sci USA 116:14414–14423
Reisinger RR, Bester MN (2010) Long distance breeding dispersal of a southern elephant seal. Polar Biol 33:1289–1291
Rodríguez JP, Fernández-Garcia J, Thums M, Hindell MA, Sequeira AMM, Meekan MG, Costa DP, Guinet C, Harcourt RG, McMahon CR, Muelbert M, Duarte CM, Eguíluz VM (2017) Big data analyses reveal patterns and drivers of the movements of southern elephant seals. Sci Rep 7:112
Rogers AD, Frinault BAV, Barnes DKA, Bindoff NL, Downie R, Ducklow HW, Friedlaender AS, Hart T, Hill SL, Hofmann EE, Linse K, McMahon CR, Murphy EJ, Pakhomov EA, Reygondeau G, Staniland IJ, Wolf-Gladrow DA, Wright R (2020) Antarctic futures: an assessment of climate-driven changes in ecosystem structure, function, and service provisioning in the Southern Ocean. Ann Rev Mar Sci 12:87–120
Rozas J, Ferrer-Mata A, Sanchez-Delbarrio JC, Guirao-Rico S, Librado P, Ramos-Onsins SE, Sanchez-Gracia A (2017) DnaSP v6: DNA sequence polymorphism analysis of large datasets. Mol Biol Evol 34:3299–3302
Siegert M, Atkinson A, Banwell A, Brandon M, Convey P, Davies B, Downie R, Edwards T, Hubbard B, Marshall G, Rogelj J, Rumble J, Stroeve J, Vaughan D (2019) The Antarctic Peninsula under a 1.5°C global warming scenario. Front Environ Sci 7:102
Slade RW, Moritz C, Heideman A (1994) Multiple nuclear-gene phylogenies: application to pinnipeds and comparison with a mitochondrial DNA gene phylogeny. Mol Biol Evol 11:341–356
Slade RW, Moritz C, Hoelzel AR, Burton HR (1998) Molecular population genetics of the southern elephant seal Mirounga leonina. Genetics 149:1945–1957
Stowell SML, Bentley EG, Gagne RB, Gustafson KD, Rutledge LY, Ernest HB (2018) Optimal DNA extractions from blood on preservation paper limits conversation genomic but not conservation genetic applications. J Nat Conserv 46:89–96
Tierney TJ (1977) The southern elephant seal, Mirounga leonina (L.), in the Vestfold Hills, Antarctica. Aust Wildl Res 4:13–24
van den Hoff J, Burton H (2007) The population trend of southern elephant seals (Mirounga leonina L.) at Macquarie Island (1952–2004). Polar Biol 30:1275–1283
Volzke S, McMahon CR, Hindell MA, Burton HR, Wotherspoon SJ (2021) Climate influences on female survival in a declining population of southern elephant seals (Mirounga leonina). Ecol Evol 11:11333–11344
Wilkinson IS, Bester MN (1997) Tag-loss in southern elephant seals, Mirounga leonina, at Marion Island. Antarct Sci 9:162–167
Zappes IA, Fabiani A, Sbordoni V, Rakaj A, Palozzi R, Allegrucci G (2017) New data on Weddell seal (Leptonychotes weddellii) colonies: a genetic analysis of a top predator from the Ross Sea. Antarctica PLOS ONE 12:e0182922
Acknowledgements
We thank the Integrated Marine Observing System (IMOS) team for collecting and providing the elephant seal samples and tracking data. IMOS is a national collaborative research infrastructure, supported by the Australian Government and operated by a consortium of institutions as an unincorporated joint venture, with the University of Tasmania as Lead Agent. S.Y.W.H. was funded by the Australian Research Council. Colleagues in the Molecular Ecology, Evolution, and Phylogenetics Lab provided advice and support throughout this project. We also thank Dieter Piepenburg, Jilda Alicia Caccavo, and an anonymous reviewer for their constructive comments on our manuscript.
Funding
Open Access funding enabled and organized by CAUL and its Member Institutions.
Author information
Authors and Affiliations
Contributions
CRM conducted Antarctic fieldwork and collected data. MC did the molecular lab work, conducted molecular and tracking data analyses, and wrote the first draft of the manuscript. IJ provided guidance on and contributed to tracking data analysis. SYWH, CRM, and MdB supervised the research project. All authors revised and contributed to the writing of the final manuscript.
Corresponding authors
Ethics declarations
Conflict of interest
The authors declare that they have no conflicts of interest.
Ethical approval
The animal capture, handing and tag deployment procedures were reviewed and approved by Macquarie University under permit number: ARA 2014_057.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Chua, M., Ho, S.Y.W., McMahon, C.R. et al. Movements of southern elephant seals (Mirounga leonina) from Davis Base, Antarctica: combining population genetics and tracking data. Polar Biol 45, 1163–1174 (2022). https://doi.org/10.1007/s00300-022-03058-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00300-022-03058-9