Abstract
It has been suggested that the phytohormone abscisic acid (ABA) plays an important role in the ripening of climatic fruit, although relevant genetic/molecular evidence is lacking. In the present study, a peach gene homologous to the putative Arabidopsis ABA receptor gene ABAR/CHLH, named PpCHLH, was isolated and characterized. PpCHLH is expressed ubiquitously as a single-copy gene in peach. Using tobacco rattle virus-induced gene silencing (VIGS), the PpCHLH gene was silenced in both peach leaves and fruit. The silenced PpCHLH gene affected leaf stomatal movement and delayed fruit ripening significantly. Although exogenously applied ABA promoted the ripening of the wild-type fruits, it could not rescue the RNAi chimeric fruit ripening. Collectively, these results demonstrate that PpCHLH plays a critical role in peach fruit ripening, and suggest that ABA might function as an important signal in the regulation of climacteric fruit development.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
As a plant phytohormone, abscisic acid (ABA) not only responds to adverse environmental conditions, but it also regulates various aspects of plant growth and development (Leung and Giraudat 1998; Finkelstein and others 2002; Himmelbach and others 2003). In fleshy fruits, ABA promotes photosynthetic assimilation uptake in sinks and can trigger fruit ripening (Yamaki and Asakura 1991; Coombe 1992; Davies and others 1997; Opaskornkul and others 1999; Peng and others 2003; Pan and others 2005; Zhang and others 2009). Nevertheless, the genetic/molecular evidence for ABA regulation of fruit ripening has not yet been established.
The most obvious visual changes in developing fruit are the degradation of chlorophyll and accumulation of newly synthesized pigments, including carotenes, lycopenes, and anthocyanins. Chlorophyll molecules are central players in harvesting light energy for photosynthesis in leaves; however, the role of photosynthesis in fruit appears to be limited, accounting for only a few percent of the sugar accumulation in fruit (Piechulla and others 1987). The regulation of chlorophyll breakdown coupled with both the onset of fruit ripening and color break is poorly understood (Azoulay-Shemer and others 2008).
Photosynthetic organisms exhibit green color due to the accumulation of chlorophyll pigments in chloroplasts. The first committed step in chlorophyll biosynthesis is the insertion of a magnesium ion into protoporphyrin IX (Proto), which is catalyzed by magnesium-protoporphyrin IX (ProtoIX) chelatase (Mg-chelatase). Notably, this chelatase is not only at the branch point of multiple metal ion chelations (Fig. 1), but it is also a three-component enzyme with various homologous forms in anoxygenic phototrophs (40-kDa BchI, 70-kDa BchD, and 150-kDa BchH) and oxygenic phototrophs [40-kDa CHLI, 70-kDa CHLD, and 150-kDa CHLH; 40-kDa XanH, 70-kDa XanG, and 150-kDa XanF (Walker and Willows 1997)]. Both the Mg-chelatase I and D subunits belong to the AAA protein superfamily that may participate in various cellular processes, including DNA replication, membrane fusion, microtubule processing, and proteolysis (Reid and Hunter 2002; Axelsson and others 2006). BchI and BchD form an ATP-dependent complex that transiently interacts with the Proto-binding BchH subunit, at which point Mg2+ is chelated (Sirijovski and others 2008). The binding of the H subunit to the ID complex may trigger ATP hydrolysis and lead to a conformational change in the whole complex (ID + H), thereby resulting in the release of the H subunit from the three-subunit complex (Lundqvist and others 2010). Moreover, in recent years, much progress has been made in uncovering the multiple functions of the Mg-chelatase H subunit in cells. Specifically, it may play an important role in mediating plastid-to-nucleus retrograde signaling in Arabidopsis (Mochizuki and others 2001), although recently, this notion has been challenged by Mochizuki himself and by another group (Mochizuki and others 2008; Moulin and others 2008). It may also serve as an ABA receptor (ABAR/CHLH) to mediate ABA responses in Arabidopsis seed germination, post-germination growth, and stomatal movement, which is distinct from the tetrapyrrole/chlorophyll biosynthesis pathways (Shen and others 2006; Wu and others 2009; Shang and others 2010). In addition, the Mg-chelatase H subunit functions as an antisigma factor to relay light signals to SigE, which is involved in sugar catabolism in Synechocystis sp. PCC 6803 (Osanai and others 2009). Finally, it has been reported to act as a molecular switch to interact with TOC1, which plays an important role in connecting the circadian clock with Arabidopsis responses to drought (Legnaioli and others 2009).
Position of Mg-chelatase at the branch point of multiple metal ion chelations. Metal ion chelatases can be separated into two classes of enzymes sharing a common intermediate uroporphyrinogen-III for the biosynthesis of heme, chlorophyll, and cobalamin (Debussche and others 1992; Walker and Willows 1997): a single-subunit enzyme, namely, ferrochelatase (HemH), and the heterotrimeric chelatases, namely, the cobalt chelatase (Cob N, S, T) and magnesium chelatase (Chl H, I, D)
In the current study, we cloned and characterized a homolog of the ABA receptor ABAR/CHLH, namely, the H subunit of the magnesium chelatase gene (PpCHLH) in peach. We demonstrated that PpCHLH is involved in a classic response to ABA in stomatal movement, and found that PpCHLH acts as a positive regulator in peach fruit ripening. This new function of the H subunit of magnesium chelatase could be of great importance to agriculture by allowing the control of fruit ripening.
Material and Methods
Plant Materials
Ten-year-old peach trees were grown in a greenhouse under standard culture conditions [28/15°C, relative humidity (RH) 65%]. About 40 flowers were tagged on five peach trees, and 13 fruits of uniform size were selected at each experimental time point (one replicate). Roots, stems, leaves, and fruits sampled at approximately 2 cm of fruit diameter were cut into small sections and then frozen quickly in liquid nitrogen for RNA extraction.
RNA Extraction, RT-PCR, and Sequencing
Total RNA was isolated using an RNA Extraction Kit (SV Total RNA Isolation System; Promega, Madison, WI) from 10 g of fresh fruit, or from 1.0 g of root, stem, or leaf. Genomic DNA was removed through 15-min incubation at 37°C with RNase-free DNase (TaKaRa, Kyoto, Japan). Next, an RNA-Clean Purification Kit was used to purify sample RNA (BioTeke Corporation, Beijing, China). The RNA purity and integrity were analyzed by agarose gel electrophoresis and by the A260:A230 and A260:A280 ratios. To generate first-strand cDNA, 3 μg of total RNA was reverse-transcribed using a SMARTTM RACE cDNA Synthesis Kit (TaKaRa) according to the manufacturer’s protocols. The cDNA was used as a template for amplification of a partial PpCHLH gene using degenerate primers [forward, 5′-GTC(C/A)GTGA/GATA(T/C)CAAAGT(C/G)TAACTCC-3′; reverse, 5′-AA(C/G)TCC(T/C)TCAACTT(C/A)TC(A/G)ATGTTCTC-3′] designed from the conserved sequences of plant CHLH (XM002284042, AJ001091, and AF014052). PCR was performed under the following conditions: 94°C for 5 min, followed by 35 cycles at 94°C for 30 s, 56°C for 40 s, and 72°C for 3 min, followed by a final incubation at 72°C for 10 min. The PCR products were ligated into a pMD19-T vector and subsequently transformed into Escherichia coli DH5α. The full-length PpCHLH gene was cloned using universal primers (UPM; TaKaRa) with 3′-RACE primer (forward, 5′-ATGCAAGGACCAAGTTGTTGAACC-3′) or with 5′-RACE primer (reverse, 5′-CAAGCCAAAACTGCAAACTGAGAA-3′), respectively.
Probe Preparation and Northern and Southern Hybridizations
Digoxigenin (DIG)-labeled probes were synthesized using a PCR-DIG Probe Synthesis Kit (Roche Diagnostics, Indianapolis, IN) with primers specific to the cloned PpCHLH gene (forward, 5′-TGGAGAACTCCTCGTGGAATGATG-3′ and reverse, 5′-ATCGATTCCCTCAATTTTGTCTTC-3′). For Northern hybridization analysis, total RNA (15 μg) was separated by electrophoresis on 1% (w/v) agarose gels containing 2.2 M formaldehyde and blotted onto nylon membranes (Hybond N+; Amersham Biosciences, Piscataway, NJ). For Southern hybridization analysis, the digested DNA was separated by electrophoresis on 1% (w/v) agarose gels containing 1% ethidium bromide and blotted onto nylon membranes (Amersham Biosciences). The filters were hybridized overnight with the DIG-labeled DNA probes (0.3–1 μg/ml) in high hybridization solution (50% formamide, 2× SSPE buffer, 10 mM DTT, 1 mg/ml herring sperm DNA, 500 μg/ml yeast tRNA, and 1 mg/ml BSA) in a shaking water bath at 50°C. Following hybridization, the filters were washed twice at 50°C for 15 min in each of 2× SSC, 1× SSC, and 0.1× SSC. The membranes were then subjected to immunological detection according to the manufacturer’s instructions using NBT/BCIP stock solution as a chemiluminescent substrate for alkaline phosphatase (Roche Diagnostics).
RT-PCR and SYBR® Real-Time RT-PCR
Total RNA was isolated respectively from peach fruits, roots, stems, or leaves, as described above. For RT-PCR analysis of PpCHLH gene expression, the first-strand cDNA was used as a template for PCR amplification through 26 cycles. For real-time RT-PCR, the reactions (20 μl) contained 10 μl SYBR® Premix Ex Taq (TaKaRa), 0.4 μl forward-specific primer, 0.4 μl reverse-specific primer, and 2 μl cDNA template. The mixture was placed in an iQ Sequence Detector (Bio-Rad, Hercules, CA), and DNA amplification was conducted using the following thermocycling program: 95°C for 2 min, followed by 40 cycles at 94°C for 20 s, 54°C for 20 s, and 72°C for 30 s, followed by 71 cycles increasing from 60 to 95°C at increments of 0.5°C per cycle for 30 s. The primers used for RT-PCR and real-time RT-PCR were as follows: CHLH (271-bp DNA) forward, 5′-TCCTCGTGGAATGATGAGAAGC-3′ and reverse, 5′-GTGGTGGTGTCAGCAATGTAAG-3′; actin (internal control, 120-bp DNA) forward, 5′-ATGCTCTGATGAAGATTCTAAC-3′ and reverse, 5′-TTGTAGGTAGTCTCATGAATAC-3’.
Stomatal Movement
Three-week-old RNAi and control seedling leaves were prepared as described by Jia and others (2010). The sampled leaves were placed in epidermal buffer for use. For stomatal aperture assays, leaves were floated in the buffer containing 50 mM KCl and 10 mM MES-Tris (pH 6.15) under a halogen cold-light source (Cole-Parmer, Vernon Hills, IL) at 200 μmol m−2 s−1 for 2 h, followed by the addition of various concentrations of (±)-ABA. Apertures were recorded on epidermal strips after 2 h of further incubation to estimate ABA-induced closure. To study the inhibition of opening, leaves were floated in the same buffer in the dark for 2 h before being transferred to the cold-light for 2 h in the presence of ABA, and then apertures were measured.
Virus Vector Construction
For the TRV vector, a 684-bp cDNA fragment of the PpCHLH gene (from 2861 to 3531) was amplified using forward (5′-CTGACAATGAGATAGGGAGTTT-3′) and reverse (5′-CAAATGACTTCCTGCTTAGATACA-3′) primers. The amplified fragment was then cloned into T-vector pMD19-T (TaKaRa). Subsequently, this construct was digested with SacI and XbaI, and the fragment was ligated into the virus vector pTRV2 cut with SacI and XbaI, as described previously (Liu and others 2002).
VIGS
VIGS assays were conducted according to the protocols reported by Liu and others (2002), with minor modifications. pTRV1, pTRV2, or the recombinant plasmid pTRV2–PpCHLH were transformed into Agrobacterium strain GV3101 using the freeze–thaw method (Fire and others 1998). A 5-ml culture of each strain was grown overnight at 28°C in LB medium containing 50 mg/ml kanamycin, 50 mg/ml rifampicin, 10 mM MES, and 20 mM acetosyringone. Each overnight culture was inoculated into 50 ml of LB medium and incubated at 28°C overnight. Cells were harvested by centrifugation at 5000×g for 5 min at 20°C, resuspended in infiltration buffer containing 10 mM MgCl2, 10 mM MES (pH 5.6), and 200 μM acetosyringone, adjusted to an A600 of 1.0–2.0, and then left at room temperature for 3 h. Next, 0.1–0.2 ml of a 1:1 (v/v) mixture of induced Agrobacterium mixture containing pTRV1 plus pTRV2 or pTRV2-PpCHLH was infiltrated into leaves or fruits using a needleless 5-ml syringe.
Results
Cloning and Bioinformatic Analysis of the PpCHLH Gene
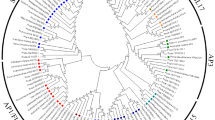
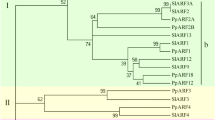
A 4445-bp cDNA encoding the H subunit of magnesium chelatase, designated PpCHLH (GenBank accession No. FJ842115), was cloned from young fruits through a combination of RT-PCR and 3′/5′-RACE. The PpCHLH gene includes an open reading frame that encodes a deduced protein of 1382 amino acids. The deduced amino acid sequence homologous analysis using protein Blast at the NCBI web site (http://blast.ncbi.nlm.nih.gov/Blast.cgi) showed that the PpCHLH protein shares high similarity with not only higher plants such as Vitis vinifera (ADE05291, 90%), Glycine max (CAA04526, 89%), Antirrhinum majus (CAA51664, 88%), Nicotiana tabacum (AAB97152, 88%), Ricinus communis(XP_002532078, 88%), Arabidopsis thaliana (CAA92802, 86%), Hordeum vulgare (AAK72401, 83%), and Oryza sativa (ABF95686, 83%), but also with lower organisms such as Synechococcus (YP_001734277, 67%) and Cyanobacteria (YP_002485640, 66%). This suggested that this protein was evolutionarily conserved (Fig. 2). An analysis of phylogenetic relationships among these plants based on amino acid sequence alignments showed that PpCHLH and its orthologs were classified largely into two groups, namely, dicots and monocots, in which A. thaliana was separated from the other dicots, and Prunus persica and R. communis were classified into the same cluster (Fig. 3).
Conserved regions in CHLH proteins. Alignment of the derived amino acid sequences of CHLH proteins using Clustal 2.0 tools. Asterisks indicate the conserved regions; Ant. (Antirrhinum majus, CAA51664), Nic. (Nicotiana tabacum, AAB97152), Gly. (Glycine max, CAA04526), Hor. (Hordeum vulgare, AAK72401), Vit. (Vitis vinifera, ADE05291), Ric. (Ricinus communis, XP_002532078), Pru. (Prunus persica, FJ842115), Ara. (Arabidopsis thaliana, CAA92802), Ory. (Oryza sativa, ABF95686), Syn. (Synechococcus, YP_001734277), and Cya. (Cyanobacteria, YP_002485640)
Phylogenetic tree of PpCHLH proteins from Prunus persica and other plant species. A phylogenetic tree with distances was analyzed using the neighbor-joining method (Saitou and Nei 1987). The numbers at the nodes represent bootstrap values, and the bar represents 0.02 substitution per amino acid position. The confidence values represent bootstrap values from 1000 replicates
Bioinformatic analysis of the PpCHLH protein indicated that its theoretical isoelectric point (pI) and molecular weight (mw) are 5.82 and 153.75 kDa, respectively (http://www.expasy.ch/tools/pi_tool.html). A putative signal peptide cleavage site was found in the PpCHLH protein between residues 16 and 17 (http://www.cbs.dtu.dk/services/SignalP/). Transmembrane region/helix prediction analysis indicated that PpCHLH is a soluble protein (http://bp.nuap.nagoya-u.ac.jp/sosui/sosui_submit.html), possibly having several transmembrane segments (http://www.sbc.su.se/~miklos/DAS/; Fig. 4). This predictive result is consistent with a recent report in which a PpCHLH homolog ABAR in Arabidopsis was demonstrated to be a transmembrane protein that spans the chloroplast envelope, exposing its N- and C-termini to the cytosolic side of cells (Shang and others 2010).
Prediction of the potential transmembrane domains in PpCHLH. The prediction was carried out using the “DAS” Transmembrane Prediction server (http://www.sbc.su.se/≃miklos/DAS). PpCHLH has several potential transmembrane sites with higher probability (profile score) at amino acid residues 142–145, 390–392, 419–423, 524–528, and 1051–1055, and lower probability at residues 80–85, 241–246, 326–324, 826–832, and 941–947
Expression Pattern of PpCHLH in Peach
To explore the mRNA expression pattern of the PpCHLH gene, total RNA was extracted from roots, stems, leaves, and fruits (Fig. 5a). Based on the PpCHLH gene sequence (GenBank FJ842115), the expression of a PCR-amplified 684-bp fragment was detected in both green and in nongreen tissues (Fig. 5b). Real-time RT-PCR using a 120-bp fragment specific to the PpCHLH gene revealed that PpCHLH expression levels were the highest in leaves, followed by stems and green fruits, and was lowest in roots (in the proportion of 100:30:15:1; Fig. 5c), suggesting that the PpCHLH protein might function at the whole-plant level.
mRNA gene expression pattern of the PpCHLH gene. a Total RNA was extracted from roots, stems, leaves, and fruits of peach trees. b A 684-bp fragment of PpCHLH from roots, stems, leaves, and fruits was each amplified by RT-PCR. c Real-time RT-PCR analysis of the expression level of PpCHLH by amplification of a 120-bp fragment specific to PpCHLH in roots, stems, leaves, and fruits. The error bars represent the SE (n = 3)
Genomic Southern Analysis of PpCHLH
To investigate the copy number of the PpCHLH gene in the peach genome, Southern blot analysis was performed using a probe specific to the PpCHLH gene. Genomic DNA was digested with NotI and KpnI restriction enzymes. The probe generated only one hybridizing band (Fig. 6), suggesting that the peach genome contains one copy of CHLH-related genes.
Southern blot analysis of the copy number of the PpCHLH gene. Peach genomic DNA (15 μg) was digested with NotI and KpnI, electrophoresed in a 0.8% agarose gel, then transferred onto a nylon membrane. The membrane was hybridized with a digoxigenin (DIG)-labeled cDNA specific to PpCHLH. The DNA marker (M, kb) is shown at the right of the section
The PpCHLH Gene Is Involved in Stomatal Movement
To explore whether the PpCHLH gene is involved in stomatal movement, we generated peach transgenic degreening RNAi leaves by VIGS, which led to a 70% reduction in the expression levels of the PpCHLH gene (Fig. 7A2, C2) compared with dark-green control leaves (Fig. 7A1, C1). The lower epidermis of leaves was separated and placed in buffer for ABA treatment. In the control leaves, ABA treatment induced stomatal closure significantly (Fig. 7E1) and inhibited stomatal opening (Fig. 7E3) in a concentration-dependent manner. In RNAi leaves, a much weaker effect of ABA-induced stomatal closure was observed (Fig. 7E2) and ABA inhibited stomatal opening (Fig. 7E4). That is, variations in the transcription levels of the PpCHLH gene could alter stomatal sensitivity to ABA, suggesting that the PpCHLH gene is involved in stomatal movement.
Changes in PpCHLH expression alter plant sensitivity to ABA. Peach leaves attached to plants were infiltrated with Agrobacterium containing TRV alone (Control) or TRV carrying a fragment of PpCHLH (RNAi). Total RNA was extracted from natural control leaves (A1) or VIGS-RNAi leaves (A2). rRNA indicates the loading control of the RNA samples stained with ethidium bromide (B1, B2). Total RNA (20 mg) was transferred onto a nylon membrane after agarose electrophoresis and hybridized with the digoxigenin (DIG)-labeled cDNA specific to PpCHLH, which detected the mRNA expression levels of PpCHLH in control leaves (C1) and the RNAi leaves (C2). Guard cells were treated with ABA concentrations of 0, 0.4, 0.8, 2.5, 5, and 10 μmol/L, respectively, for 2 h. The ABA-induced stomatal closure (E1, E2) and ABA-inhibited stomatal opening (E3, E4) were evaluated. SD values were calculated from three individual experiments; each experiment consisted of ten guard cells from one leaf of each plant (n = 10)
The PpCHLH Gene Is Involved in the Regulation of Fruit Ripening
To explore whether the PpCHLH gene is involved in fruit development, we generated peach transgenic RNAi fruit infiltrated with TRV-PpCHLH, and control fruit infiltrated with an empty TRV vector. As shown in Fig. 8, degreening fruits were selected for infiltration 2 weeks before the onset of ripening (Fig. 8a). About 3 weeks after infiltration, the infiltrated sector turned red on the surface of control fruits (Fig. 8b). In contrast, the infiltrated sector remained light green on the surface of RNAi fruits (Fig. 8c).
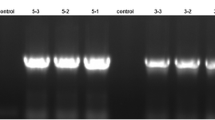
Silencing the PpCHLH gene in peach fruits attached to plants by VIGS. Peach fruits attached to plants were infiltrated with Agrobacterium containing TRV alone (Control) or TRV carrying a fragment of PpCHLH (RNAi) at 7–8 weeks after anthesis (a). Photographs of the infiltrated peach fruits were taken 3 weeks after infiltration (b, control; c, RNAi). Semiquantitative qRT-PCR, Northern blot RT-PCR, and real-time qRT-PCR analyses of the reduced PpCHLH transcript levels in RNAi fruits compared with control fruits (d, f, and h). rRNA indicates a loading control of the RNA samples stained with ethidium bromide (g). Actin was used as an internal control (e). The error bars represent the SE (n = 3). Arrows indicate the infiltrated site. Scale bar = 1 cm in a, b, and c
To confirm the PpCHLH gene suppression at the molecular level, semiquantitative RT-PCR, real-time qRT-PCR, and Northern blot analyses were carried out. As shown in Fig. 8, the mRNA expression of the PpCHLH gene in RNAi fruits was downregulated by 80% as compared to that in control fruits (Fig. 8d, f, h). The combination of transgenic phenotype observation and molecular detection indicated that compared with the control fruits (Fig. 8b), the drastically reduced PpCHLH gene transcription could lead to the loss of red coloring in infiltrated regions of RNAi fruits (Fig. 8c), which significantly affected fruit ripening. These results suggested that the PpCHLH gene might be an important positive regulator for peach fruit ripening.
To explore the relationship of ABA with PpCHLH during peach fruit ripening, we detected the effects of exogenous ABA on both PpCHLH-silenced RNAi fruits and control fruits attached to plants. As shown in Table 1, of the 12 control fruits treated with exogenous 0.5 mM ABA at the light green stage, all turned full red 15 days after treatment (advancing 3 days compared with naturally ripening fruits). In contrast, among 12 chimeric fruits treated with exogenous 0.5 mM ABA, only one fruit turned full red 15 days after treatment. Thereby, these results not only demonstrated that ABA could significantly promote peach fruit ripening, but also confirmed that the disruption of the PpCHLH gene could markedly alter the response of peach fruit to ABA. In conclusion, we have provided both physiological and forceful molecular evidence to verify a key role of ABA in the regulation of peach fruit ripening, suggesting that ABA might be an important regulator for controlling ripening of climatic fruit.
Discussion
Characteristics of the PpCHLH Gene and Protein
Based on bioinformatic, PCR, and Southern blot analyses, we identified the following distinct features of the PpCHLH gene and its deduced amino acid sequence: the PpCHLH protein is an evolutionarily conserved protein (Fig. 2), the ubiquitous expression pattern of the PpCHLH gene suggests that its coding protein might function at the whole-plant level (Fig. 5), a single copy of the PpCHLH gene exists in the peach genome (Fig. 6), and the PpCHLH protein is a soluble protein containing several potential transmembrane regions (Fig. 4).
One interesting question is the copy number of CHLH gene in plants. The current and previous studies show that the CHLH gene is a single-copy gene in peach (P. persica), Arabidopsis (Shen and others 2006), soybean (G. max; Nakayama and others 1998), and barley (H. vulgare; Henningsen and others 1993), but two copies were found only in rice (O. sativa; Wu and others 2009). Whether multiple copies of the CHLH gene also occur in other species requires verification. Another interesting question is the localization of CHLH protein in plant cells. As noted above, the PpCHLH protein was predicted to be a soluble protein with several potential transmembrane regions, suggesting to some extent that its localization in plant cells is flexible. Consistent with this notion, previous studies have shown that in vitro, the soybean CHLH localizes in the envelope and stroma fractions in a Mg2+-dependent manner; in 1 mM Mg2+, CHLH is largely present in the stroma fraction, whereas in 5 mM Mg2+, a significant proportion of the CHLH protein is membrane-associated (Gibson and others 1996). A more recent study has reported that Arabidopsis CHLH/ABAR is a transmembrane protein that spans the chloroplast envelope, exposing its N- and C-termini to the cytosol (Shang and others 2010). It is notable that some proteins similar to CHLH, such as Src kinase, protein kinase C, phospholipase C, vinculin, and DnaA, also have two apparent localizations: an aqueous compartment or the cell membrane. These proteins are reportedly involved in important cellular functions, including catalytic function, the assembly of signaling complexes, and regulation of access to substrates. A transmembrane helix may serve as a membrane anchor (Johnson and Cornell 1999), and thus the mechanism underlying the movement of CHLH protein from envelope membranes to stroma should to be elucidated.
ABA Might Be a Vital Regulator in Climacteric Fruit Ripening
Fleshy fruits have been classically defined as either climacteric or nonclimacteric fruits based on the physiological processes of ripening. Climacteric fruits show a sudden increase in respiration at the onset of ripening, with a concomitant increase in production of the gaseous hormone ethylene, which is typically regarded as necessary for climacteric ripening (Giovannoni 2001; Alexander and Grierson 2002). Peach fruit ripening shows an increase in ethylene production, and it is a typical climacteric fruit (Miller and others 1988; Tonutti and others 1991). Furthermore, peach fruit softening is completely ethylene-dependent (Hayama and others 2006).
A recent study has reported a role for ABA in peach fruit. Specifically, a 9-cis-epoxycarotenoid dioxygenase gene, PpNCED1, encoding a key enzyme for ABA biosynthesis is highly expressed at the beginning of ripening, which leads to the rapid accumulation of ABA. Such accumulated ABA can accelerate ethylene production in peach fruit, suggesting that ABA might play a key role in the regulation of ripeness and senescence of peach fruits (Zhang and others 2009). ABA action is known to be initiated by ABA perception, which triggers downstream signaling cascades to induce physiological effects. In the current study, RNAi peach leaves with a 70% reduction in PpCHLH transcript levels affected ABA-mediated stomatal movement, demonstrating that the PpCHLH protein is involved in the ABA signaling pathway. Based on this fact, direct evidence for the role of ABA in the regulation of peach fruit ripening has been established by silencing of the PpCHLH gene, not only because the reduced transcription of the PpCHLH gene significantly disrupts peach fruit red-coloring, but also in that the loss of color in RNAi fruits is not rescued by exogenously applied ABA that can promote the natural control of red-coloring in fruit (Table 1). Indeed, these results provide convincing evidence for a key role of ABA in the regulation of peach fruit ripening, and suggest that ABA might play an important role in the control of climacteric fruit ripening.
In conclusion, based on the results of our previous and current studies, we have demonstrated that the H subunit of magnesium chelatase has at least three functions in peach: chlorophyll biosynthesis, stomatal movement, and fruit ripening. The detection of ABA binding to PpCHLH protein may provide more information about the regulation of ABA in fruit development.
References
Alexander L, Grierson D (2002) Ethylene biosynthesis and action in tomato: a model for climacteric fruit ripening. J Exp Bot 53:2039–2055
Axelsson E, Lundqvist J, Sawicki A, Nilsson S, Schröder I et al (2006) Recessiveness and dominance in barley mutants deficient in Mg-chelatase subunit D, an AAA protein involved in chlorophyll biosynthesis. Plant Cell 18:3606–3616
Azoulay-Shemer T, Harpaz-Saad S, Belausov E, Lovat N, Krokhin O et al (2008) Citrus chlorophyllase dynamics at ethylene-induced fruit color-break: a study of chlorophyllase expression, posttranslational processing kinetics, and in situ intracellular localization. Plant Physiol 148:108–118
Coombe BG (1992) Research on development and ripening of the grape berry. Am J Enol Vitic 43:101–110
Davies C, Boss PK, Robinson SP (1997) Treatment of grape berries, anon-climacteric fruit with asynthetic auxin, retards ripening and alters the expression of developmentally regulated genes. Plant Physiol 115:1155–1161
Debussche L, Couder M, Thibaut D, Cameron B, Crouzet J et al (1992) Assay, purification, and characterization of cobaltochelatase, a unique complex enzyme catalyzing cobalt insertion in hydrogenobyrinic acid a, c-diamide during coenzyme B12 biosynthesis in Pseudomonas denitrificans. J Bacteriol 174:7445–7451
Finkelstein RR, Gampala SS, Rock CD (2002) Abscisic acid signaling in seeds and seedlings. Plant Cell 14:S15–S45
Fire A, Xu S, Montgomery MK, Kostas SA, Driver SE et al (1998) Potent and specific genetic interference by double-stranded RNA in Caenorhabditis elegans. Nature 391:806–811
Gibson LC, Marrison J, Leech RM, Jensen PE, Bassham DC et al (1996) A putative Mg-chelatase subunit from Arabidopsis thaliana cv C24. Plant Physiol 111:61–71
Giovannoni J (2001) Molecular regulation of fruit ripening. Annu Rev Plant Physiol Plant Mol Biol 52:725–749
Hayama H, Shimada T, Fujii H, Ito A, Kashimura Y (2006) Ethylene regulation of fruit softening and softening-related genes in peach. J Exp Bot 57:4071–4077
Henningsen KW, Boynton JE, von Wettstein D (1993) Mutants at xantha and albina loci in relation to chloroplast biogenesis in barley (Hordeum vulgare L.). R Danish Acad Sci Lett Copenhagen 42:1–349
Himmelbach A, Yang Y, Grill E (2003) Relay and control of abscisic acid signaling. Curr Opin Plant Biol 6:470–479
Jia HF, Guo JX, Qin L, Shen YY (2010) Virus-induced PpCHLH gene silencing in peach leaves (Prunus persica). J Hortic Sci Biotechnol 85:528–532
Johnson JE, Cornell RB (1999) Amphitropic proteins: regulation by reversible membrane interactions (review). Mol Membr Biol 16:217–235
Legnaioli T, Cuevas J, Mas P (2009) TOC1 functions as a molecular switch connecting the circadian clock with plant responses to drought. EMBO J 28:3745–3757
Leung J, Giraudat J (1998) Abscisic acid signal transduction. Ann Rev Plant Physiol Plant Mol Biol 49:199–222
Liu YL, Schiff M, Dinesh-Kumar SP (2002) Virus-induced gene silencing in tomato. Plant J 31:777–786
Lundqvist J, Elmlund H, Wulff RP, Berglund L, Elmlund D et al (2010) ATP-induced conformational dynamics in the AAA + Motor Unit of magnesium chelatase. Structure 18:354–365
Miller AN, Krizek BA, Walsh CS (1988) Whole-fruit ethylene evolution and ACC content of peach pericarp and seeds during development. J Am Soc Hortic Sci 113:119–124
Mochizuki N, Brusslan JA, Larkin R, Nagatani A, Chory J (2001) Arabidopsis genomes uncoupled 5 (GUN5) mutant reveals the involvement of Mg-chelatase H subunit in plastid-to-nucleus signal transduction. Proc Natl Acad Sci USA 98:2053–2058
Mochizuki N, Tanaka R, Tanaka A, Masuda T, Nagatani A (2008) The steady-state level of Mg-protoporphyrin IX is not a determinant of plastid-to-nucleus signaling in Arabidopsis. Proc Natl Acad Sci USA 105:15184–15189
Moulin M, McCormac AC, Terry MJ, Smith AG (2008) Tetrapyrrole profiling in Arabidopsis seedlings reveals that retrograde plastid nuclear signaling is not due to Mg-protoporphyrin IX accumulation. Proc Natl Acad Sci USA 105:15178–15183
Nakayama M, Masuda T, Bando T, Yamagata H, Ohta H et al (1998) Cloning and expression of the soybean chlH gene encoding a subunit of Mg-chelatase and localization of the Mg2+ concentration-dependent ChlH protein within the chloroplast. Plant Cell Physiol 39:275–284
Opaskornkul C, Linberg S, Tiliberg JE (1999) Effect of ABA on the distribution of sucrose and protons across the plasmalemma of pea mesophyll protoplasts, suggesting a sucrose / proton symport. J Plant Physiol 154:447–453
Osanai T, Imashimizu M, Seki A, Sato S, Tabata S et al (2009) CHLH, the H subunit of the Mg-chelatase, is an anti-sigma factor for SigE in Synechocystis sp. PCC 6803. Proc Natl Acad Sci USA 106:6860–6865
Pan QH, Li MJ, Peng CC, Zhang N, Zou X et al (2005) Abscisic acid activates acid invertases in developing grape berry. Physiol Plant 125:157–170
Peng YB, Lu YF, Zhang DP (2003) Abscisic acid activities ATPase in developing apple fruit especially in fruit phloem cells. Plant Cell Environ 26:1329–1342
Piechulla B, Glick RE, Bahl H, Melis A, Gruissem W (1987) Changes in photosynthetic capacity and photosynthetic protein pattern during tomato fruit ripening. Plant Physiol 84:911–917
Reid JD, Hunter CN (2002) Current understanding of the function of magnesium chelatase. Biochem Soc Trans 30:643–645
Saitou N, Nei M (1987) The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol Biol Evol 4:406–425
Shang Y, Yan L, Liu ZQ, Cao Z, Mei C et al (2010) The Mg-chelatase H subunit of Arabidopsis antagonizes a group of WRKY transcription repressors to relieve ABA-responsive genes of inhibition. Plant Cell 22:1909–1935
Shen YY, Wang XF, Wu FQ, Du SY, Cao Z et al (2006) The Mg-chelatase H subunit is an abscisic acid receptor. Nature 443:823–826
Sirijovski N, Lundqvist J, Rosenbäck M, Elmlund H, Al-Karadaghi S et al (2008) Substrate-binding model of the chlorophyll biosynthetic magnesium chelatase BchH subunit. J Biol Chem 283:11652–11660
Tonutti P, Casson P, Ramina A (1991) Ethylene biosynthesis during peach fruit development. J Am Soc Hortic Sci 116:274–279
Walker CJ, Willows RD (1997) Mechanism and regulation of Mg-chelatase. Biochem J 327:321–333
Wu FQ, Xin Q, Cao Z, Liu ZQ, Du SY et al (2009) The Mg-chelatase H subunit binds abscisic acid and functions in abscisic acid signaling: New evidence in Arabidopsis. Plant Physiol 150:1940–1954
Yamaki S, Asakura T (1991) Stimulation of the uptake of sorbitol into vacuoles from apple fruit flesh by abscisic acid and into protoplasts by indoleacetic acid. Plant Cell Physiol 32:315–318
Zhang M, Leng P, Zhang G, Li X (2009) Cloning and functional analysis of 9-cis-epoxycarotenoid dioxygenase (NCED) genes encoding a key enzyme during abscisic acid biosynthesis from peach and grape fruits. J Plant Physiol 166:1241–1252
Acknowledgments
We thank Dr. Yu-Le Liu (Qinghua University, China) for pTRV vectors. This work was supported financially by the China National Natural Science Foundation (grant No. 30971977), the Beijing Natural Science Foundation and Scientific Research Key Program of the Beijing Commission of Education (grant No. KZ200910020001), and the Beijing Natural Science Foundation (grant No. 6082005).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
H.-F. Jia and Y.-M. Chai have contributed equally to this work.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Jia, HF., Chai, YM., Li, CL. et al. Cloning and Characterization of the H Subunit of a Magnesium Chelatase Gene (PpCHLH) in Peach. J Plant Growth Regul 30, 445–455 (2011). https://doi.org/10.1007/s00344-011-9207-y
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00344-011-9207-y