Abstract
Human prion diseases are associated with a range of clinical presentations and are classified by both clinicopathological syndrome and aetiology with sub-classification according to molecular criteria. Considerable experimental evidence suggests that phenotypic diversity in human prion disease relates in significant part to the existence of distinct human prion strains encoded by abnormal PrP isoforms with differing physicochemical properties. To date, however, the conformational repertoire of pathological isoforms of wild-type human PrP and the various forms of mutant human PrP has not been fully defined. Efforts to produce a unified international classification of human prion disease are still ongoing. The ability of genetic background to influence prion strain selection together with knowledge of numerous other factors that may influence clinical and neuropathological presentation strongly emphasises the requirement to identify distinct human prion strains in appropriate transgenic models, where host genetic variability and other modifiers of phenotype are removed. Defining how many human prion strains exist allied with transgenic modelling of potentially zoonotic prion strains will inform on how many human infections may have an animal origin. Understanding these relationships will have direct translation to protecting public health.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Human prion diseases are invariably fatal conditions that include Creutzfeldt–Jakob disease (CJD), Gerstmann–Sträussler–Scheinker disease (GSS), fatal familial insomnia (FFI), kuru and variant CJD (vCJD) in humans [28]. They are associated with a range of clinical presentations and are classified by both clinico-pathological syndrome and aetiology with sub-classification according to molecular criteria [28, 45, 54, 90, 121, 139]. These diseases have attracted immense research interest for many years not only because of their unique composition and properties but also because of their impact on public health [27, 30, 100, 129]. According to the widely accepted ‘protein-only’ hypothesis [47], host-encoded cellular prion protein (PrPC) is converted to an alternative form designated PrPSc [22, 27, 30, 100, 129]. It is proposed that PrPSc is the infectious agent acting to replicate itself with high fidelity by recruiting endogenous PrPC, and that the difference between these isoforms lies purely in the monomer conformation and its state of aggregation [22, 27, 30, 99, 100, 102, 107, 129].
Central to understanding the molecular basis of prion propagation remains the conundrum of prion strains—how a protein-only infectious agent can encode information required to specify distinct disease phenotypes—and also the so-called species barrier effect which limits cross species infection. While originally considered different aspects of the prion problem, it is now clear that species barriers and prion strains are intimately related by “conformational selection” [26, 30]. Within the protein-only hypothesis of prion propagation prion strains are thought to be encoded by distinct pathogenic PrP conformations or assembly states. Conformational selection proposes that although a wide range of mammalian PrPSc conformations may be possible, only a subset will be compatible with each individual PrP primary structure. Ease of transmission of prions between species (or also within species as a result of PrP polymorphisms), therefore, relates to overlap of permissible PrPSc conformations between the structures of PrP from the source and recipient as well as heterogeneity in cellular mechanisms affecting prion propagation and clearance kinetics [26, 30]. Conformational selection has now been strongly supported by elegant studies of prions in yeast and other fungi [14, 41, 62, 66, 106, 111, 112, 132], and the wider relevance of prion-like mechanisms in other protein misfolding diseases is now becoming clear [30, 82, 85]. Understanding the molecular biology of prion disease will illuminate processes involving protein misfolding and aggregation and protein-based inheritance, which clearly have far-reaching implications in pathobiology, ageing and the evolution of cellular processes.
Aetiologies of human prion disease
Human prion diseases are biologically unique and can be divided aetiologically into inherited, sporadic and acquired forms [28]. Approximately 85% of human prion disease occurs sporadically as sporadic CJD at a rate of 1–2 cases per million population per year across the world, with an equal incidence in men and women [16, 27, 28, 38, 121]. Around 15% of human prion disease is associated with autosomal dominant pathogenic mutations in PRNP, and to date, over 30 mutations have been described [27, 28, 53, 65, 80, 121]. Acquired human prion diseases forms have, until recently, been confined to rare and unusual situations. Iatrogenic CJD has arisen as a result of transmission of CJD prions through treatment with pituitary hormones derived from human cadavers, implantation of dura mater grafts, corneal transplantation and the use of contaminated electroencephalographic electrodes [17, 18]. The most well-known incidences of acquired prion disease in humans resulting from a dietary origin have been kuru that was caused by cannibalism among the Fore linguistic group of the Eastern Highlands in Papua New Guinea [2, 3, 35, 36, 79] and more recently the occurrence of vCJD in the United Kingdom and other countries due to human exposure to BSE prions [26, 28, 120, 134]. Remarkably, kuru demonstrates that incubation periods of infection with human prions can exceed 50 years [35, 36].
Prion disease pathology and pathogenesis
The brains of patients with prion disease frequently show no recognisable abnormalities on gross examination at necropsy; however, microscopic examination typically reveals characteristic histopathologic changes, consisting of neuronal vacuolation and degeneration, which gives the cerebral grey matter a microvacuolated or ‘spongiform’ appearance, and a reactive proliferation of astroglial cells (for review see [19, 20]). Although spongiform degeneration is frequently detected, it is not an obligatory neuropathologic feature of prion disease; the presence of astro-gliosis and micro-gliosis, although not specific to the prion diseases, is more constantly seen. The lack of a lymphocytic inflammatory response is also an important characteristic. Demonstration of abnormal PrP immunoreactivity, or more specifically biochemical detection of PrPSc in brain material by immunoblotting techniques (Fig. 1) is diagnostic of prion disease, and some forms of prion disease are characterised by deposition of amyloid plaques composed of insoluble aggregates of PrP [19, 20]. Amyloid plaques are a notable feature of kuru and GSS [20, 48], but they are less frequently found in the brains of patients with sporadic CJD which typically show a diffuse pattern of abnormal PrP deposition [20, 54] (Fig. 2). The histopathological features of vCJD are remarkably consistent and distinguish it from other human prion diseases with large numbers of PrP-positive amyloid plaques that differ in morphology from the plaques seen in kuru and GSS in that the surrounding tissue takes on a micro-vacuolated appearance, giving the plaques a florid appearance [59, 134] (Fig. 2). Abundant florid plaques are established as the neuropathological hallmark of vCJD [59] and, to date, have only been found in association with BSE infection in hosts homozygous for PrP with methionine at residue 129 (in humans, primates or transgenic mice [6, 67, 119, 134]).
Immunoblot analysis of human prion disease brain. a Proteinase K digested brain homogenates analysed with monoclonal antibody 3F4 and enhanced chemiluminescence showing the PrPSc types seen in sporadic, acquired and inherited prion disease. Types 1–3 PrPSc are seen in the brain of classical forms of CJD (either sporadic or iatrogenic CJD), while type 4 PrPSc is uniquely seen in vCJD brain. Distinct PrPSc types are seen in cases with PRNP point mutations. Classification according to Hill et al. [53, 54]. b Proteinase K digestion of brain homogenate and analysis with anti-PrP monoclonal antibody 3F4 by enhanced chemifluorescence enables calculation of the proportions of di-, mono-, and non-glycosylated PrP. The key shows PrPSc type or mutation and PRNP codon 129 genotype (M methionine, V valine). Data points represent the mean relative proportions of di- and mono-glycosylated PrP as percentage ± SEM. In some cases, the error bars were smaller than the symbols used
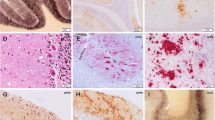
Distinct patterns of PrP deposition in brain in human prion disease. The disease aetiology, PRNP codon 129 genotype of the patient (M methionine, V valine) and the type of PrPSc detected in each sample (using the London classification of human PrPSc types [54]) is designated above each brain sample. The most common subtype of sporadic CJD (a) typically shows a diffuse, synaptic pattern of abnormal PrP deposition, while kuru (b) shows not only variably diffuse PrP deposition but also striking formation of PrP plaques in various areas of the brain. These PrP plaques are consistently distinct from those seen in vCJD (c), where PrP plaques are often surrounded by conspicuous vacuolation, designated ‘florid plaques’. Scale bars a, c 25 μm; b 50 μm
Distinct forms of prion disease show differences in lymphoreticular involvement that appear to be related to the divergent properties of distinct prion strains [1]. For example, the tissue distribution of PrPSc in vCJD differs strikingly from that in classical CJD and inherited prion disease [51–53, 55, 57, 61, 124] with uniform and prominent involvement of lymphoreticular tissues, with the highest amounts (up to 10% of brain concentrations) in tonsil [52, 124]. In contrast, in sporadic CJD, PrPSc has only been irregularly detected by immunoblotting in non-central nervous system tissues at very much lower levels [46, 94]. Tonsil biopsy is used for antemortem diagnosis of vCJD and, to date, has shown 100% sensitivity and specificity [52, 121, 124], and tonsil is the tissue of choice for prospective studies investigating the prevalence of vCJD prion infection within the UK and other populations [24, 42, 56]. The fact that lymphoreticular prion infection is not a feature of iatrogenic CJD [51, 52] or kuru [15, 36] argues that the distinct pathogenesis of vCJD relates to the effect of prion strain rather than to a peripheral route of infection [15, 36, 127]. The demonstration of extensive peripheral tissue involvement in the pathogenesis of vCJD raises concern that asymptomatically infected carriers may be transmitting the disease to others via blood transfusion, as now appears to have occurred [71, 93, 136], or other iatrogenic routes including contaminated surgical and medical instruments [4, 26, 120, 123].
Determinants of phenotypic variability in human prion disease
The clinical presentation of human prion disease varies enormously, and there is considerable overlap observed between individuals with different disease aetiologies [28, 80, 120, 121] and even in family members with the same pathogenic PRNP mutation [29, 31, 32, 65, 77, 80, 125]. Progressive dementia, cerebellar ataxia, pyramidal signs, chorea, myoclonus, extrapyramidal features, pseudobulbar signs, seizures and amyotrophic features can be seen in variable combinations. Criteria used for diagnosis of human prion disease have been defined [28, 131], and definite diagnosis of sporadic and acquired prion disease relies upon neuropathological examination and the demonstration of abnormal PrP deposition in the central nervous system by either immunoblotting or immunohistochemistry [20, 28, 60, 128, 131]. Polymorphism at residue 129 of human PrP [encoding either methionine (M) or valine (V)] powerfully affects susceptibility to human prion diseases [25, 38, 68, 79, 86, 135] with residue 129 acting to directly restrict the propagation of particular prion strains through conformational selection [26, 27, 30, 119] as well as heterozygosity conferring resistance by inhibiting homologous protein–protein interactions [25, 30, 86]. About 38% of northern Europeans are homozygous for the more frequent methionine allele, 51% are heterozygous and 11% homozygous for valine. Homozygosity at PRNP codon 129 predisposes to the development of sporadic and acquired CJD [25, 38, 68, 79, 86, 135] and is most strikingly observed in vCJD, where all neuropathologically confirmed cases studied so far have been homozygous for codon 129 methionine of PRNP [28, 81, 119, 136].
The hypothesis that alternative conformations or assembly states of PrP provide the molecular substrate for a significant part of the clinicopathological heterogeneity seen in human prion diseases and that this relates to the existence of distinct human prion strains is supported by considerable experimental evidence [30, 34, 90, 113] and also by the demonstration of protein conformation-based inheritance mechanisms of yeast prions [106, 112, 132]. Different human PrPSc isoforms, referred to as molecular strain types, have been identified in the brain of patients with phenotypically distinct forms of CJD [34, 45, 54, 88, 90, 92, 113, 122, 139] and are classified by both the fragment size and ratio of the three principal PrP bands seen after protease digestion. To date, we have characterised four types of human PrPSc that can be commonly identified in sporadic and acquired human prion diseases [27, 34, 54, 121] (Fig. 1) although much greater heterogeneity seems likely [121]. Sporadic and iatrogenic CJD and kuru are associated with PrPSc types 1-3, while type 4 PrPSc is uniquely associated with vCJD and is characterised by a fragment size and glycoform ratio that is similar to PrPSc seen in BSE and BSE when transmitted to several other species [34, 54, 122, 127]. An earlier classification of PrPSc types seen in classical CJD described only two banding patterns [88] with PrPSc types 1 and 2 that we describe corresponding with the type 1 pattern of Gambetti and colleagues and our type 3 fragment size corresponding to their type 2 pattern [87, 90]. Consensus on the nomenclature of human PrPSc types has been hindered by the fact that the N-terminal conformation of some PrPSc subtypes seen in sporadic CJD can be inter-converted in vitro via changes in metal-ion occupancy [54, 122] or solvent pH [21, 84, 138]. While it has proposed that pH alone determines the N-terminal structure of PrPSc in sporadic CJD [21, 84], this interpretation has not been supported by other studies [69, 98, 122], and the conformations of PrPSc types 1 and 2 that we describe in PRNP methionine homozygous patients show critical dependence upon the presence of copper or zinc ions under conditions, where pH 7.4 is tightly controlled [122]. While type 4 PrPSc is readily distinguished from the PrPSc types seen in classical CJD and kuru by a predominance of the di-glycosylated PrP glycoform, type 4 PrPSc also has a distinct proteolytic fragment size [54] although this is not recognised by the alternative classification which designates type 4 PrPSc as type 2b [87].
In addition to the distinct human PrPSc types associated with sporadic and acquired prion disease, molecular strain typing has also provided insights into the phenotypic heterogeneity seen inherited human prion diseases [53, 65]. Patients with inherited prion disease caused by point mutations have glycoform ratios of PrPSc fragments distinct from those seen in both classical CJD and vCJD [53]. Individuals with the same PRNP mutation can also propagate PrPSc with distinct fragment sizes [53, 96, 97]. Detection of PrPSc in the molecular mass range of ca. 21–30 kDa is, however, not a consistent feature, and some cases, in particular those in which amyloid plaques are a prominent feature, show smaller protease resistant fragments of ca. 7–15 kDa [53, 65, 89, 96, 97, 110]. The propagation of pathological isoforms of wild-type PrP may also make a significant contribution to phenotypic variability in inherited prion disease [23, 43, 108, 125].
Classification of human prion disease
Efforts to produce a unified international classification and nomenclature of human PrPSc types have been complicated. A major confounding issue in this regard has been resolving whether relatively subtle biochemical differences in PrPSc are of biological importance and accurately reflect the propagation of distinct human prion strains. This is particularly true in sporadic CJD [21, 34, 44, 54, 88, 90, 91, 138], where progress has been severely hampered by a lack of transgenic modelling data to firmly distinguish the identity of distinct prion strains and their defining molecular and neuropathological phenotypes. This fundamental problem coupled with the difficulties and variability of the biochemical methods used to distinguish PrPSc types [21, 54, 88, 91, 118, 122, 138] has so far precluded an internationally accepted classification system for human prion strains. In this regard, the increasingly recognised co-occurrence of different PrPSc types in the same brain [38, 49, 54, 90, 96, 98, 101, 105, 118, 125, 137] and the recognition that protease-sensitive pathological isoforms of PrP may have a significant role in both animal and human prion disease [9, 39, 44, 83, 103, 104, 115–117] have further confounded progress. Although agreement has yet to be reached on methodological differences, nomenclature and the biological importance of relatively subtle biochemical differences in PrPSc, there is strong agreement between laboratories that phenotypic diversity in human prion disease relates in significant part to the existence of distinct prion strains [34, 40, 45, 50, 54, 69, 88, 90, 98, 104, 113, 139]. Despite these advances, however, the ability of genetic background to influence prion strain selection [6, 72–74, 81, 109], coupled with the knowledge that route of transmission in acquired human prion disease may dramatically influence clinical and neuropathological presentation [17, 126, 127, 133], strongly emphasises the requirement to remove host genetic variability and other modifiers of phenotype (Table 1) by identifying distinct human prion strains in appropriate transgenic models [5–7, 11, 13, 33, 58, 63, 64, 76, 114, 119].
Whether the diversity of phenotypes seen in human disease can be faithfully recapitulated by transgenic modelling remains an open question. In this regard, the issue of prion strain selection or mutation will be a major factor. As recently hypothesised [30], prion strains may not exist as previously thought as molecular clones with a single PrPSc type (where strain mutation in a different host would involve generation of a distinct PrPSc type), but may consist of an ensemble of molecular species (containing a dominant PrPSc type that is preferentially propagated by its usual host) from which a less populous subspecies may be selected by an alternative host, resulting in a strain shift or mutation. In this regard, different cellular populations and tissues within a single host would provide different environments for strain selection as recently demonstrated in vitro [70]. Given the degree of molecular diversity observed in human prion disease, it may be extremely difficult to isolate the full complement of human prion strains in transgenic mice having a single genetic background.
Future perspectives
Despite major advances, the molecular basis of mammalian prion strain diversity remains unknown. To date, the conformational repertoire of pathological isoforms of wild-type human PrP and the various forms of mutant human PrP has not been fully defined. Biochemical investigation of disease-related PrP isoforms in patients allied with detailed clinical and neuropathological analysis will continue to inform on the diversity of phenotypes seen in human prion disease. As it has now become clear that prion strain type, host genetic makeup and numerous other factors may significantly influence prion disease phenotype, it is expected that the actual number of distinct human prion strains may be far less than the number of identified phenotypes. Detailed transgenic modelling will therefore be crucial to establishing how many human prion strains exist and what the defining molecular features of PrP are for each strain. This information allied with comprehensive transgenic modelling of human BSE infection and other relevant, potentially zoonotic, prion strains will inform on how many human prion strains may have an animal origin. Understanding the risks that existing and emerging animal prion diseases pose will have direct translation to protecting public health.
Development of an accurate classification for human prion disease will have major implications for epidemiological research into the causes of sporadic CJD, whose aetiology remains obscure. While spontaneous conversion of PrPC to PrPSc as a rare stochastic event, or somatic mutation of the PrP gene, resulting in expression of a pathogenic PrP mutant are plausible explanations for sporadic CJD [16, 27, 78, 125], other causes for at least some cases, include environmental exposure to human prions [37, 75, 95] or exposure to animal prions. In this regard, the number of prion strains causing sheep scrapie has yet to be established [8, 10, 12], and epidemiological data cannot exclude this as a cause of a minority of cases. Similarities between types of sporadic CJD and types of sheep scrapie have been reported [130]. As future research begins to provide a more precise understanding of the origins of human prion disease, this will facilitate re-analysis of epidemiological data to reveal important risk factors that might have been obscured by analysing sporadic CJD as a single entity.
References
Aguzzi A (2003) Prions and the immune system: a journey through gut, spleen, and nerves. Adv Immunol 81:123–171
Alpers MP (1987) Epidemiology and clinical aspects of kuru. In: Prusiner SB, McKinley MP (eds) Prions: novel infectious pathogens causing scrapie and Creutzfeldt-Jakob disease. Academic Press, San Diego, pp 451–465
Alpers MP (2008) The epidemiology of kuru: monitoring the epidemic from its peak to its end. Philos Trans R Soc Lond B Biol Sci 363:3707–3713
Armitage WJ, Tullo AB, Ironside JW (2009) Risk of Creutzfeldt-Jakob disease transmission by ocular surgery and tissue transplantation. Eye 23:1926–1930
Asano M, Mohri S, Ironside JW, Ito M, Tamaoki N, Kitamoto T (2006) vCJD prion acquires altered virulence through trans-species infection. Biochem Biophys Res Commun 342:293–299
Asante E, Linehan J, Desbruslais M et al (2002) BSE prions propagate as either variant CJD-like or sporadic CJD-like prion strains in transgenic mice expressing human prion protein. EMBO J 21:6358–6366
Asante E, Linehan J, Gowland I et al (2006) Dissociation of pathological and molecular phenotype of variant Creutzfeldt-Jakob disease in transgenic human prion protein 129 heterozygous mice. Proc Natl Acad Sci USA 103:10759–10764
Baron T, Biacabe AG, Arsac JN, Benestad S, Groschup MH (2006) Atypical transmissible spongiform encephalopathies (TSEs) in ruminants. Vaccine 25:5625–5630
Barron RM, Campbell SL, King D et al (2007) High titres of TSE infectivity associated with extremely low levels of PrPSc in vivo. J Biol Chem 282:35878–35886
Benestad SL, Arsac JN, Goldmann W, Noremark M (2008) Atypical/Nor98 scrapie: properties of the agent, genetics, and epidemiology. Vet Res 39:19
Beringue V, Le Dur A, Tixador P et al (2008) Prominent and persistent extraneural infection in human PrP transgenic mice infected with variant CJD. PLoS One 3:e1419
Beringue V, Vilotte JL, Laude H (2008) Prion agents diversity and species barrier. Vet Res 39:47
Bishop MT, Hart P, Aitchison L et al (2006) Predicting susceptibility and incubation time of human-to-human transmission of vCJD. Lancet Neurol 5:393–398
Brachmann A, Baxa U, Wickner RB (2005) Prion generation in vitro: amyloid of Ure2p is infectious. EMBO J 24:3082–3092
Brandner S, Whitfield J, Boone K et al (2008) Central and peripheral pathology of kuru: pathological analysis of a recent case and comparison with other forms of human prion disease. Philos Trans R Soc Lond B Biol Sci 363:3755–3763
Brown P, Cathala F, Raubertas RF, Gajdusek DC, Castaigne P (1987) The epidemiology of Creutzfeldt-Jakob disease: conclusion of a 15-year investigation in France and review of the world literature. Neurology 37:895–904
Brown P, Preece M, Brandel JP et al (2000) Iatrogenic Creutzfeldt-Jakob disease at the millennium. Neurology 55:1075–1081
Brown P, Preece MA, Will RG (1992) “Friendly fire” in medicine: hormones, homografts, and Creutzfeldt-Jakob disease. Lancet 340:24–27
Budka H (2003) Neuropathology of prion diseases. Br Med Bull 66:121–130
Budka H, Aguzzi A, Brown P et al (1995) Neuropathological diagnostic criteria for Creutzfeldt-Jakob disease (CJD) and other human spongiform encephalopathies (Prion diseases). Brain Pathol 5:459–466
Cali I, Castellani R, Yuan J et al (2006) Classification of sporadic Creutzfeldt-Jakob disease revisited. Brain 129:2266–2277
Caughey B, Baron GS (2006) Prions and their partners in crime. Nature 443:803–810
Chen SG, Parchi P, Brown P et al (1997) Allelic origin of the abnormal prion protein isoform in familial prion diseases. Nat Med 3:1009–1015
Clewley JP, Kelly CM, Andrews N et al (2009) Prevalence of disease related prion protein in anonymous tonsil specimens in Britain: cross sectional opportunistic survey. BMJ 338:b1442
Collinge J, Palmer MS, Dryden AJ (1991) Genetic predisposition to iatrogenic Creutzfeldt-Jakob disease. Lancet 337:1441–1442
Collinge J (1999) Variant Creutzfeldt-Jakob disease. Lancet 354:317–323
Collinge J (2001) Prion diseases of humans and animals: their causes and molecular basis. Annu Rev Neurosci 24:519–550
Collinge J (2005) Molecular neurology of prion disease. J Neurol Neurosurg Psychiatry 76:906–919
Collinge J, Brown J, Hardy J et al (1992) Inherited prion disease with 144 base pair gene insertion: II: Clinical and pathological features. Brain 115:687–710
Collinge J, Clarke A (2007) A general model of prion strains and their pathogenicity. Science 318:930–936
Collinge J, Harding AE, Owen F et al (1989) Diagnosis of Gerstmann-Straussler syndrome in familial dementia with prion protein gene analysis. Lancet 2:15–17
Collinge J, Owen F, Poulter M et al (1990) Prion dementia without characteristic pathology. Lancet 336:7–9
Collinge J, Palmer MS, Sidle KCL et al (1995) Unaltered susceptibility to BSE in transgenic mice expressing human prion protein. Nature 378:779–783
Collinge J, Sidle KC, Meads J, Ironside J, Hill AF (1996) Molecular analysis of prion strain variation and the aetiology of ‘new variant’ CJD. Nature 383:685–690
Collinge J, Whitfield J, McKintosh E et al (2006) Kuru in the 21st century—an acquired human prion disease with very long incubation periods. Lancet 367:2068–2074
Collinge J, Whitfield J, McKintosh E et al (2008) A clinical study of kuru patients with long incubation periods at the end of the epidemic in Papua New Guinea. Philos Trans R Soc Lond B Biol Sci 363:3725–3739
Collins S, Law MG, Fletcher A, Boyd A, Kaldor J, Masters CL (1999) Surgical treatment and risk of sporadic Creutzfeldt-Jakob disease: a case-control study. Lancet 353:693–697
Collins SJ, Sanchez-Juan P, Masters CL et al (2006) Determinants of diagnostic investigation sensitivities across the clinical spectrum of sporadic Creutzfeldt-Jakob disease. Brain 129:2278–2287
Cronier S, Gros N, Tattum MH et al (2008) Detection and characterization of proteinase K-sensitive disease-related prion protein with thermolysin. Biochem J 416:297–305
Demart S, Fournier JG, Creminon C et al (1999) New insight into abnormal prion protein using monoclonal antibodies. Biochem Biophys Res Commun 265:652–657
Diaz-Avalos R, King CY, Wall J, Simon M, Caspar DL (2005) Strain-specific morphologies of yeast prion amyloid fibrils. Proc Natl Acad Sci USA 102:10165–10170
Frosh A, Smith LC, Jackson CJ et al (2004) Analysis of 2000 consecutive UK tonsillectomy specimens for disease-related prion protein. Lancet 364:1260–1262
Gabizon R, Telling G, Meiner Z, Halimi M, Kahana I, Prusiner SB (1996) Insoluble wild-type and protease-resistant mutant prion protein in brains of patients with inherited prion disease. Nat Med 2:59–64
Gambetti P, Dong Z, Yuan J et al (2008) A novel human disease with abnormal prion protein sensitive to protease. Ann Neurol 63:697–708
Gambetti P, Kong Q, Zou W, Parchi P, Chen SG (2003) Sporadic and familial CJD: classification and characterisation. Br Med Bull 66:213–239
Glatzel M, Abela E, Maissen M, Aguzzi A (2003) Extraneural pathologic prion protein in sporadic Creutzfeldt-Jakob disease. N Engl J Med 349:1812–1820
Griffith JS (1967) Self replication and scrapie. Nature 215:1043–1044
Hainfellner JA, Brantner-Inthaler S, Cervenáková L et al (1995) The original Gerstmann-Straussler-Scheinker family of Austria: divergent clinicopathological phenotypes but constant PrP genotype. Brain Pathol 5:201–211
Head MW, Bunn TJ, Bishop MT et al (2004) Prion protein heterogeneity in sporadic but not variant Creutzfeldt-Jakob disease: U.K. cases 1991-2002. Ann Neurol 55:851–859
Head MW, Ironside JW (2006) Sporadic Creutzfeldt-Jakob disease: further twists and turns in a convoluted protein. Brain 129:2238–2240
Head MW, Ritchie D, Smith N et al (2004) Peripheral tissue involvement in sporadic, iatrogenic, and variant Creutzfeldt-Jakob disease: an immunohistochemical, quantitative, and biochemical study. Am J Pathol 164:143–153
Hill AF, Butterworth RJ, Joiner S et al (1999) Investigation of variant Creutzfeldt-Jakob disease and other human prion diseases with tonsil biopsy samples. Lancet 353:183–189
Hill AF, Joiner S, Beck J et al (2006) Distinct glycoform ratios of protease resistant prion protein associated with PRNP point mutations. Brain 129:676–685
Hill AF, Joiner S, Wadsworth JD et al (2003) Molecular classification of sporadic Creutzfeldt-Jakob disease. Brain 126:1333–1346
Hill AF, Zeidler M, Ironside J, Collinge J (1997) Diagnosis of new variant Creutzfeldt-Jakob disease by tonsil biopsy. Lancet 349:99–100
Hilton DA, Ghani AC, Conyers L et al (2004) Prevalence of lymphoreticular prion protein accumulation in UK tissue samples. J Pathol 203:733–739
Hilton DA, Sutak J, Smith ME et al (2004) Specificity of lymphoreticular accumulation of prion protein for variant Creutzfeldt-Jakob disease. J Clin Pathol 57:300–302
Hizume M, Kobayashi A, Teruya K et al (2009) Human prion protein (PrP) 219 K is converted to PrPSc but shows heterozygous inhibition in variant Creutzfeldt-Jakob disease infection. J Biol Chem 284:3603–3609
Ironside JW, Head MW (2004) Neuropathology and molecular biology of variant Creutzfeldt-Jakob disease. Curr Top Microbiol Immunol 284:133–159
Ironside JW, Head MW, Bell JE, McCardle L, Will RG (2000) Laboratory diagnosis of variant Creutzfeldt-Jakob disease. Histopathology 37:1–9
Joiner S, Linehan J, Brandner S, Wadsworth JD, Collinge J (2005) High levels of disease related prion protein in the ileum in variant Creutzfeldt-Jakob disease. Gut 54:1506–1508
King CY, Diaz-Avalos R (2004) Protein-only transmission of three yeast prion strains. Nature 428:319–323
Kong Q, Huang S, Zou W et al (2005) Chronic wasting disease of elk: transmissibility to humans examined by transgenic mouse models. J Neurosci 25:7944–7949
Korth C, Kaneko K, Groth D et al (2003) Abbreviated incubation times for human prions in mice expressing a chimeric mouse-human prion protein transgene. Proc Natl Acad Sci USA 100:4784–4789
Kovacs GG, Trabattoni G, Hainfellner JA, Ironside JW, Knight RS, Budka H (2002) Mutations of the prion protein gene phenotypic spectrum. J Neurol 249:1567–1582
Krishnan R, Lindquist SL (2005) Structural insights into a yeast prion illuminate nucleation and strain diversity. Nature 435:765–772
Lasmezas CI, Fournier JG, Nouvel V et al (2001) Adaptation of the bovine spongiform encephalopathy agent to primates and comparison with Creutzfeldt-Jakob disease: implications for human health. Proc Natl Acad Sci USA 98:4142–4147
Lee HS, Brown P, Cervenáková L et al (2001) Increased susceptibility to kuru of carriers of the PRNP 129 methionine/methionine genotype. J Infect Dis 183:192–196
Lewis V, Hill AF, Klug GM, Boyd A, Masters CL, Collins SJ (2005) Australian sporadic CJD analysis supports endogenous determinants of molecular-clinical profiles. Neurology 65:113–118
Li J, Browning S, Mahal SP, Oelschlegel AM, Weissmann C (2010) Darwinian evolution of prions in cell culture. Science 327:869–872
Llewelyn CA, Hewitt PE, Knight RS et al (2004) Possible transmission of variant Creutzfeldt-Jakob disease by blood transfusion. Lancet 363:417–421
Lloyd S, Collinge J (2005) Genetic susceptibility to prion diseases in humans and mice. Current Genomics 6:1–11
Lloyd S, Onwuazor ON, Beck J et al (2001) Identification of multiple quantitative trait loci linked to prion disease incubation period in mice. Proc Natl Acad Sci USA 98:6279–6283
Lloyd SE, Maytham EG, Pota H et al (2009) HECTD2 is associated with susceptibility to mouse and human prion disease. PLoS Genet 5:e1000383
Mahillo-Fernandez I, Pedro-Cuesta J, Bleda MJ et al (2008) Surgery and risk of sporadic Creutzfeldt-Jakob disease in Denmark and Sweden: registry-based case-control studies. Neuroepidemiology 31:229–240
Mallik S, Yang W, Norstrom EM, Mastrianni JA (2010) Live cell FRET predicts an altered molecular association of heterologous PrP-Sc with PrP-C. J Biol Chem 285:8967–8975
Mead S, Poulter M, Beck J et al (2006) Inherited prion disease with six octapeptide repeat insertional mutation—molecular analysis of phenotypic heterogeneity. Brain 129:2297–2317
Mead S, Webb TE, Campbell TA et al (2007) Inherited prion disease with 5-OPRI: phenotype modification by repeat length and codon 129. Neurology 69:730–738
Mead S, Stumpf MP, Whitfield J et al (2003) Balancing selection at the prion protein gene consistent with prehistoric kuru-like epidemics. Science 300:640–643
Mead S (2006) Prion disease genetics. Eur J Hum Genet 14:273–281
Mead S, Poulter M, Uphill J et al (2009) Genetic risk factors for variant Creutzfeldt-Jakob disease: a genome-wide association study. Lancet Neurol 8:57–66
Miller G (2009) Neurodegeneration. Could they all be prion diseases? Science 326:1337–1339
Nazor KE, Kuhn F, Seward T et al (2005) Immunodetection of disease-associated mutant PrP, which accelerates disease in GSS transgenic mice. EMBO J 24:2472–2480
Notari S, Capellari S, Giese A et al (2004) Effects of different experimental conditions on the PrPSc core generated by protease digestion: Implications for strain typing and molecular classification of CJD. J Biol Chem 279:16797–16804
Olanow CW, Prusiner SB (2009) Is Parkinson’s disease a prion disorder? Proc Natl Acad Sci USA 106:12571–12572
Palmer MS, Dryden AJ, Hughes JT, Collinge J (1991) Homozygous prion protein genotype predisposes to sporadic Creutzfeldt-Jakob disease. Nature 352:340–342
Parchi P, Capellari S, Chen SG et al (1997) Typing prion isoforms. Nature 386:232–233
Parchi P, Castellani R, Capellari S et al (1996) Molecular basis of phenotypic variability in sporadic Creutzfeldt-Jakob disease. Ann Neurol 39:767–778
Parchi P, Chen SG, Brown P et al (1998) Different patterns of truncated prion protein fragments correlate with distinct phenotypes in P102L Gerstmann-Sträussler-Scheinker disease. Proc Natl Acad Sci USA 95:8322–8327
Parchi P, Giese A, Capellari S et al (1999) Classification of sporadic Creutzfeldt-Jakob disease based on molecular and phenotypic analysis of 300 subjects. Ann Neurol 46:224–233
Parchi P, Notari S, Weber P et al (2008) Inter-laboratory assessment of PrP(Sc) typing in Creutzfeldt-Jakob disease: a western blot study within the NeuroPrion consortium. Brain Pathol 19:384–391
Parchi P, Zou WQ, Wang W et al (2000) Genetic influence on the structural variations of the abnormal prion protein. Proc Natl Acad Sci USA 97:10168–10172
Peden AH, Head MW, Ritchie DL, Bell JE, Ironside JW (2004) Preclinical vCJD after blood transfusion in a PRNP codon 129 heterozygous patient. Lancet 364:527–529
Peden AH, Ritchie DL, Head MW, Ironside JW (2006) Detection and localization of PrPSc in the skeletal muscle of patients with variant, iatrogenic, and sporadic forms of Creutzfeldt-Jakob disease. Am J Pathol 168:927–935
Pedro-Cuesta J, Mahillo-Fernandez I, Rabano A et al (2010) Nosocomial transmission of sporadic Creutzfeldt-Jakob disease: results from a risk-based assessment of surgical interventions. J Neurol Neurosurg Psychiatry (in press)
Piccardo P, Dlouhy SR, Lievens PMJ et al (1998) Phenotypic variability of Gerstmann-Straussler-Scheinker disease is associated with prion protein heterogeneity. J Neuropathol Exp Neurol 57:979–988
Piccardo P, Liepnieks JJ, William A et al (2001) Prion proteins with different conformations accumulate in Gerstmann-Straussler-Scheinker disease caused by A117V and F198S mutations. Am J Pathol 158:2201–2207
Polymenidou M, Stoeck K, Glatzel M, Vey M, Bellon A, Aguzzi A (2005) Coexistence of multiple PrP(Sc) types in individuals with Creutzfeldt-Jakob disease. Lancet Neurol 4:805–814
Prusiner SB (1982) Novel proteinaceous infectious particles cause scrapie. Science 216:136–144
Prusiner SB (1998) Prions. Proc Natl Acad Sci USA 95:13363–13383
Puoti G, Giaccone G, Rossi G, Canciani B, Bugiani O, Tagliavini F (1999) Sporadic Creutzfeldt-Jakob disease: co-occurrence of different types of PrPSc in the same brain. Neurology 53:2173–2176
Riesner D (2003) Biochemistry and structure of PrP(C) and PrP(Sc). Br Med Bull 66:21–33
Safar J, Wille H, Itri V et al (1998) Eight prion strains have PrPSc molecules with different conformations. Nat Med 4:1157–1165
Safar JG, Geschwind MD, Deering C et al (2005) Diagnosis of human prion disease. Proc Natl Acad Sci USA 102:3501–3506
Schoch G, Seeger H, Bogousslavsky J et al (2005) Analysis of prion strains by PrP(Sc) profiling in sporadic Creutzfeldt-Jakob disease. PLoS Med 3:e14
Shorter J, Lindquist S (2005) Prions as adaptive conduits of memory and inheritance. Nat Rev Genet 6:435–450
Silveira JR, Raymond GJ, Hughson AG et al (2005) The most infectious prion protein particles. Nature 437:257–261
Silvestrini MC, Cardone F, Maras B et al (1997) Identification of the prion protein allotypes which accumulate in the brain of sporadic and familial Creutzfeldt-Jakob disease patients. Nat Med 3:521–525
Stephenson DA, Chiotti K, Ebeling C et al (2000) Quantitative trait loci affecting prion incubation time in mice. Genomics 69:47–53
Tagliavini F, Lievens PMJ, Tranchant C et al (2001) A 7-kDa prion protein (PrP) fragment, an integral component of the PrP region required for infectivity, is the major amyloid protein in Gerstmann-Straussler-Scheinker disease A117V. J Biol Chem 276:6009–6015
Tanaka M, Chien P, Naber N, Cooke R, Weissman JS (2004) Conformational variations in an infectious protein determine prion strain differences. Nature 428:323–328
Tanaka M, Chien P, Yonekura K, Weissman JS (2005) Mechanism of cross-species prion transmission an infectious conformation compatible with two highly divergent yeast prion proteins. Cell 121:49–62
Telling GC, Parchi P, DeArmond SJ et al (1996) Evidence for the conformation of the pathologic isoform of the prion protein enciphering and propagating prion diversity. Science 274:2079–2082
Telling GC, Scott M, Mastrianni J et al (1995) Prion propagation in mice expressing human and chimeric PrP transgenes implicates the interaction of cellular PrP with another protein. Cell 83:79–90
Thackray AM, Hopkins L, Bujdoso R (2007) Proteinase K-sensitive disease-associated ovine prion protein revealed by conformation-dependent immunoassay. Biochem J 401:475–483
Tremblay P, Ball HL, Kaneko K et al (2004) Mutant PrP(Sc) conformers induced by a synthetic peptide and several prion strains. J Virol 78:2088–2099
Tzaban S, Friedlander G, Schonberger O et al (2002) Protease-sensitive scrapie prion protein in aggregates of heterogeneous sizes. Biochemistry 41:12868–12875
Uro-Coste E, Cassard H, Simon S et al (2008) Beyond PrP type 1/type 2 dichotomy in Creutzfeldt-Jakob disease. PLoS Pathog 4:e1000029
Wadsworth JD, Asante EA, Desbruslais M et al (2004) Human prion protein with valine 129 prevents expression of variant CJD phenotype. Science 306:1793–1796
Wadsworth JD, Collinge J (2007) Update on human prion disease. Biochim Biophys Acta 1772:598–609
Wadsworth JD, Hill AF, Beck JA, Collinge J (2003) Molecular and clinical classification of human prion disease. Br Med Bull 66:241–254
Wadsworth JD, Hill AF, Joiner S, Jackson GS, Clarke A, Collinge J (1999) Strain-specific prion-protein conformation determined by metal ions. Nat Cell Biol 1:55–59
Wadsworth JD, Joiner S, Fox K et al (2007) Prion infectivity in variant Creutzfeldt-Jakob disease rectum. Gut 56:90–94
Wadsworth JD, Joiner S, Hill AF et al (2001) Tissue distribution of protease resistant prion protein in variant CJD using a highly sensitive immuno-blotting assay. Lancet 358:171–180
Wadsworth JD, Joiner S, Linehan J et al (2006) Phenotypic heterogeneity in inherited prion disease (P102L) is associated with differential propagation of protease-resistant wild-type and mutant prion protein. Brain 129:1557–1569
Wadsworth JD, Joiner S, Linehan JM, Asante EA, Brandner S, Collinge J (2008) Review. The origin of the prion agent of kuru: molecular and biological strain typing. Philos Trans R Soc Lond B Biol Sci 363:3747–3753
Wadsworth JD, Joiner S, Linehan JM et al (2008) Kuru prions and sporadic Creutzfeldt-Jakob disease prions have equivalent transmission properties in transgenic and wild-type mice. Proc Natl Acad Sci USA 105:3885–3890
Wadsworth JD, Powell C, Beck JA et al (2008) Molecular diagnosis of human prion disease. Methods Mol Biol 459:197–227
Weissmann C (2004) The state of the prion. Nat Rev Microbiol 2:861–871
Wemheuer WM, Benestad SL, Wrede A et al (2009) Similarities between forms of sheep scrapie and Creutzfeldt-Jakob disease are encoded by distinct prion types. Am J Pathol 175:2566–2573
WHO (2003) WHO manual for surveillance of human transmissible spongiform encephalopathies. http://www.who.int/bloodproducts/TSE-manual2003.pdf
Wickner RB, Edskes HK, Shewmaker F, Nakayashiki T (2007) Prions of fungi: inherited structures and biological roles. Nat Rev Microbiol 5:611–618
Will RG (2003) Acquired prion disease: iatrogenic CJD, variant CJD, kuru. Br Med Bull 66:255–265
Will RG, Ironside JW, Zeidler M et al (1996) A new variant of Creutzfeldt-Jakob disease in the UK. Lancet 347:921–925
Windl O, Dempster M, Estibeiro JP et al (1996) Genetic basis of Creutzfeldt-Jakob disease in the United Kingdom: a systematic analysis of predisposing mutations and allelic variation in the PRNP gene. Hum Genet 98:259–264
Wroe SJ, Pal S, Siddique D et al (2006) Clinical presentation and pre-mortem diagnosis of variant Creutzfeldt-Jakob disease associated with blood transfusion: a case report. Lancet 368:2061–2067
Yull HM, Ritchie DL, Langeveld JP et al (2006) Detection of type 1 prion protein in variant Creutzfeldt-Jakob disease. Am J Pathol 168:151–157
Zanusso G, Farinazzo A, Fiorini M et al (2001) pH-dependent prion protein conformation in classical Creutzfeldt-Jakob disease. J Biol Chem 276:40377–40380
Zanusso G, Farinazzo A, Prelli F et al (2004) Identification of distinct N-terminal truncated forms of prion protein in different Creutzfeldt-Jakob disease subtypes. J Biol Chem 279:38936–38942
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding authors
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Wadsworth, J.D.F., Collinge, J. Molecular pathology of human prion disease. Acta Neuropathol 121, 69–77 (2011). https://doi.org/10.1007/s00401-010-0735-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-010-0735-5