Abstract
Pediatric central nervous system (CNS) tumors represent the most common cause of cancer-related death in children aged 0–14 years. They differ from their adult counterparts, showing extensive clinical and molecular heterogeneity as well as a challenging histopathological spectrum that often impairs accurate diagnosis. Here, we use DNA methylation-based CNS tumor classification in combination with copy number, RNA-seq, and ChIP-seq analysis to characterize a newly identified CNS tumor type. In addition, we report histology, patient characteristics, and survival data in this tumor type. We describe a biologically distinct pediatric CNS tumor type (n = 31 cases) that is characterized by focal high-level amplification and resultant overexpression of either PLAGL1 or PLAGL2, and an absence of recurrent genetic alterations characteristic of other pediatric CNS tumor types. Both genes act as transcription factors for a regulatory subset of imprinted genes (IGs), components of the Wnt/β-Catenin pathway, and the potential drug targets RET and CYP2W1, which are also specifically overexpressed in this tumor type. A derived PLAGL-specific gene expression signature indicates dysregulation of imprinting control and differentiation/development. These tumors occurred throughout the neuroaxis including the cerebral hemispheres, cerebellum, and brainstem, and were predominantly composed of primitive embryonal-like cells lacking robust expression of markers of glial or neuronal differentiation (e.g., GFAP, OLIG2, and synaptophysin). Tumors with PLAGL1 amplification were typically diagnosed during adolescence (median age 10.5 years), whereas those with PLAGL2 amplification were diagnosed during early childhood (median age 2 years). The 10-year overall survival was 66% for PLAGL1-amplified tumors, 25% for PLAGL2-amplified tumors, 18% for male patients, and 82% for female patients. In summary, we describe a new type of biologically distinct CNS tumor characterized by PLAGL1/2 amplification that occurs predominantly in infants and toddlers (PLAGL2) or adolescents (PLAGL1) which we consider best classified as a CNS embryonal tumor and which is associated with intermediate survival. The cell of origin and optimal treatment strategies remain to be defined.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
The current 2021 edition of the World Health Organization (WHO) Classification of Central Nervous System (CNS) Tumors comprises more than 100 distinct pediatric and adult tumor types based on combined phenotypic and genotypic classification [1]. This updated classification is reflective of the notion that adult-type and pediatric-type tumors are markedly different and can be distinguished based on their molecular features, as is the case for diffuse gliomas, such as the H3 G34-mutant diffuse hemispheric glioma or the H3 K27-altered diffuse midline glioma. The latter is predominant in pediatric patients and restricted to certain anatomic locations [25, 26, 48, 68]. While gliomas constitute the majority of malignant CNS tumors, further CNS tumor categories such as glioneuronal and neuronal tumors, embryonal tumors, pineal tumors, and mesenchymal tumors also contribute substantially to mortality [35, 40]. According to the latest report from the CBTRUS, the CNS is the most common cancer site in children aged 0–14 years, and CNS tumors are the most common cause of cancer-related death in this age group [40]. Considerable heterogeneity can be found between the different tumor types in terms of both molecular alterations and clinical outcomes. Molecular analyses classify ependymomas, for example, into at least ten subgroups despite shared histological features, and revealed that medulloblastomas comprise multiple distinct molecular groups and subgroups [11, 33, 52]. In addition to the above-mentioned H3-altered gliomas, extensive heterogeneity in IDH- and histone H3-wild-type pediatric high-grade gliomas (HGG) has been observed, with multiple subgroups displaying differential enrichment of affected oncogenes and clinical features [14, 60]. Such molecular heterogeneity, together with a broad histological spectrum of many CNS tumors, can make histopathological diagnosis highly challenging and observer-dependent. To address this, a DNA methylation-based CNS tumor classification system was developed, seeking to improve diagnostic accuracy and objectivity across all CNS tumor types and age groups [12]—a principle which was widely adopted by the latest WHO classifications [35, 41].
Here, we utilized a broad genome-wide DNA methylation cohort, combined with copy number profiling, targeted next-generation DNA sequencing, and RNA sequencing, to identify a rare CNS tumor type characterized by amplification and overexpression of either PLAGL1 (located at chromosome 6q24.2) or PLAGL2 (located at chromosome 20q11.21). The pleomorphic adenoma gene (PLAG) family of transcription factors (TFs) comprises three genes, namely PLAG1, PLAGL1, and PLAGL2, whose roles are multifaceted and dependent on their different DNA-binding capacities as well as on the cellular context [20]. While PLAGL1 has been suggested as a putative tumor suppressor gene, PLAG1 and PLAGL2 are presumed proto-oncogenes [4]. In the brain tumor context, our evidence supports an oncogenic role also for PLAGL1, which is further substantiated by the recent discovery of recurrent PLAGL1 fusions in a subset of pediatric-type supratentorial neuroepithelial tumors [54].
Using ChIP-seq data, we show that both PLAGL1 and PLAGL2 act as TFs for the kinase RET that is specifically overexpressed in our cohort of PLAGL1/2-amplified tumors. RET functions as an oncogenic driver in a variety of cancers and serves as a therapeutic target, with selective RET inhibitors showing promising results in patients [51, 57,58,59]. We also demonstrate that PLAGL1 and PLAGL2 potentially act as TFs for components of the Wnt/β-Catenin pathway and a set of imprinted genes (IG) that was shown to be controlled by Plagl1 and to regulate the imprinted gene network (IGN) in mouse models [36, 63, 64]; this set of genes, including IGF2, is also deregulated in the PLAGL-amplified tumors. In addition, we derive a gene-signature (n = 250) characteristic for PLAGL-amplified tumors that indicates dysregulation of imprinting control and differentiation/development as a prominent feature, and shed light on the molecular, histopathologic, and clinical parameters of this novel CNS tumor type.
Materials and methods
Patients and samples
Patient samples and retrospective clinical information were either obtained through the databases of the Heidelberg University Department of Neuropathology and the German Cancer Research Center (DKFZ), or directly collected from the respective collaborating national or international institutions in compliance with local regulations. The sample set included CNS tumors enrolled in the INFORM, PTT2.0, and MNP2.0 molecular profiling studies [50, 67].
Histology and immunohistochemistry
Detailed histopathological review was retrospectively performed on a subset of the tumors (n = 15) to investigate their histological and immunohistochemical features. Representative hematoxylin and eosin (H&E)-stained sections and immunohistochemical stains from the 15 tumors were digitally scanned on an Aperio slide scanner to assemble a digital pathology library from which histological and immunohistochemical features were reviewed and annotated using ImageScope software (Leica Biosystems). H&E and immunohistochemical staining was performed at the UCSF Histology Laboratory and the Heidelberg University Department of Neuropathology or received from other participating medical centers. Immunohistochemistry was performed on formalin-fixed, paraffin-embedded tissue sections using the following antibodies: glial fibrillary acidic protein (GFAP, DAKO, polyclonal, 1:3000 dilution, no antigen retrieval), oligodendrocyte transcription factor 2 (OLIG2, Immuno Bio Labs, polyclonal, 1:200 dilution, ER1 antigen retrieval), synaptophysin (Cell Marque, polyclonal, 1:100 dilution, ER2 antigen retrieval), neurofilament (Cell Marque, clone 2F11, undiluted, ER1 antigen retrieval), epithelial membrane antigen (EMA, Leica, clone GP1.4, undiluted, ER1 antigen retrieval), S100 (DAKO, polyclonal, 1:2,000 dilution, no antigen retrieval), CD99 (Signet, clone CD99, 1:400 dilution, ER1 antigen retrieval), BCOR (Santa Cruz Biotechnology, sc-514576, clone C-10, 1:200 dilution, ER2 antigen retrieval), desmin (Cell Marque, clone D33, undiluted, ER1 antigen retrieval or DAKO, M0760, clone D33, 1:50 dilution, CC1 antigen retrieval), GAB1 (Abcam, ab27439, polyclonal, 1:50 dilution, CC2 antigen retrieval), YAP1 (Cell Signaling, #14074, clone D8H1X, 1:100 dilution), beta-catenin (BD Bioscience, 610153, clone 14, 1:100 dilution, CC1 antigen retrieval), and Ki-67 (Dako, clone Mib1, 1:50 dilution, ER2 antigen retrieval). Immunostaining was performed in Leica BOND-III or Ventana BenchMark Ultra automated stainers. Diaminobenzidine was used as the chromogen, followed by hematoxylin counterstain. Scanned image files of H&E and select immunostained sections from a subset of the tumors in this cohort are available for downloading and viewing at the following link: https://figshare.com/projects/CNS_embryonal_tumor_with_PLAGL_amplification/151806.
Extraction of DNA/RNA
DNA and RNA of samples processed in Heidelberg were extracted using the automated Maxwell nucleic acid purification platform (Promega, Madison, WI, USA). RNA was extracted from fresh–frozen tissue samples with the Maxwell RSC simply RNA Tissue kit and DNA was extracted from fresh–frozen or FFPE tissue samples with the Maxwell RSC Tissue DNA kit or the Maxwell RSC DNA FFPE kit, respectively, according to the manufacturer’s instructions. Other external samples were extracted according to standard local procedures with corresponding QC measures.
Genome-wide DNA methylation profiling
Fresh–frozen or formalin-fixed paraffin-embedded (FFPE) tissue samples were subjected to genome-wide DNA methylation profiling and were either processed at the DKFZ Genomics and Proteomics Core Facility using the Infinium Methylation EPIC (EPIC) BeadChip or Infinium Human Methylation 450 k Bead Chip arrays (Illumina) according to the manufacturer’s instructions, or at the University of California, San Francisco (UCSF) using the same arrays and methodology. A subset of cases were obtained through uploads to the online http://www.molecularneuropathology.org platform. Methylation array processing was performed as previously described [28]. t-Distributed Stochastic Neighbor Embedding (t-SNE) dimensionality reduction as well as copy number variation (CNV) analysis based on the raw intensities of the methylation array probes were performed as described before [12]. The raw methylation array data of the ET, PLAGL samples have been deposited in NCBI’s Gene Expression Omnibus [16] and are accessible through GEO Series accession number GSE212621 (https://www.ncbi.nlm.nih.gov/geo/query/acc.cgi?acc=GSE212621).
Copy number analysis
The Integrative Genomics Viewer (IGV) was used to visualize copy number variants (CNVs) of the respective amplified PLAG-family genes [46]. Amplifications and deletions were visualized across the entire genome by summary plots. Copy number profiles generated by the conumee R-package were segmented with the circular binary segmentation (CBS) algorithm using the default settings of the conumee package for the 450 k and EPIC array [23]. Resulting segments for each sample were combined to a cohort file using R version 3.6.2 [42], which was then analyzed with the GISTIC_2.0 method available within the GenePattern cloud tool (https://cloud.genepattern.org/) [37, 43]. Human_Hg19.mat was used as reference gene file. Maxspace was set to 10,000. Amplification/deletion thresholds were set to 0.1 and focal length cutoff to distinguish broad from focal events was set to 0.5 (fraction of chromosome arm). Gene GISTIC algorithm was used to calculate the significant regions of deletion. A confidence level of 0.99 as well as a false discovery rate (FDR) q value of < 0.25 were used for a region to be considered as significant. Join segment size was set to 4, cap values were set to 1.5, and the maximum number of segments allowed per sample was 2000. Arm level peel off was performed to reduce noise. Amplifications/deletions were rated (0, 1, 2) and divided into three different classes: no amplification/deletion (0; log2 ratio < 0.1/log2 ratio > − 0.1), low-level amplification (1; 0.1 < log2 ratio < 0.9) or low-level deletion (1; − 0.1 > log2 ratio > − 1.3), high-level amplification (2; log2 ratio > 0.9) or high-level deletion (2; log2 ratio < − 1.3). All focal regions and genes identified by GISTIC2.0 are summarized in Supplementary Tables S2, S3, and S4.
Targeted next-generation DNA sequencing
Genomic DNA extracted from formalin-fixed, paraffin-embedded tumor tissue or frozen tissue was used for targeted next-generation DNA sequencing (NGS) at the UCSF, DKFZ (NPHD gene panel), and PMC for a subset of the patients. For 6 patients (A108, A110, A112, A113, A387, A388), capture-based NGS was performed using the UCSF500 NGS Panel that targets all coding exons of 479 cancer-related genes, select introns and upstream regulatory regions of 47 genes to enable detection of structural variants including gene fusions, and DNA segments at regular intervals along each chromosome to enable genome-wide copy number and zygosity analysis, with a total sequencing footprint of 2.8 Mb [27, 38]. For 5 patients (A93, A94, A96, A379, A380), targeted NGS was performed using the NPHD gene panel developed at the Neuropathology department of the University Hospital Heidelberg that targets the coding exons of 201 cancer-related genes, 9 gene fusions, and 1 upstream regulatory region. For three additional patients (A100, A105, A390), complete exome sequencing was performed. NGS data may be made available upon request.
RNA sequencing and analysis
All RNAs used for quantitative gene expression analysis were extracted from fresh–frozen tissues. Quality of input RNA was assessed using the Agilent Bioanalyzer System and transcriptome analysis was performed using Illumina TruSeq strand-specific PolyA + libraries on an Illumina HiSeq4000 or NovaSeq device. Differential gene expression analysis was performed with the R2 Genomics Analysis and Visualization Platform (http://r2.amc.nl) using a reference cohort of embryonal tumors (n = 117), glial tumors (n = 126), and normal fetal and adult brain tissues (n = 36) that had been processed the same way. TPM values of the ET, PLAGL samples from our cohort are provided in the supplementary materials (Supplementary Table S8).
ChIP-seq analysis
Chromatin immunoprecipitation (ChIP) was performed at Active Motif (Carlsbad, CA, USA) using antibodies against H3K27ac (AM#39133, Active Motif), PLAGL1 (HPA055706, Sigma), and PLAGL2 (SAB3500815, Sigma) according to Active Motif’s established ChIP protocol, which includes validation of ChIP reactions via quantitative PCR (qPCR). Twenty pooled CNS tumor samples were used as an Input control. Illumina sequencing libraries were prepared from ChIPs and Input at Active Motif using their standard protocol. Libraries were sequenced via paired-end sequencing with a read length of 100 bp or 75 bp at the DKFZ Genomics and Proteomics Core Facility on the Illumina HiSeq 4000 or NextSeq550, respectively.
Paired end reads were aligned to hg38 using bowtie2. Alignment SAM files were converted to BAM files using samtools. Sambamba was used to sort and remove multimapped, unmapped and duplicated reads from the resulting BAM files. MACS2 was used to call narrow peaks on sorted and processed BAMS with input as the reference. Processed BAM files were normalized with bamCompare (—normalizeUsing BPM—scaleFactorsMethod None—smoothLength 60—extendReads—centerReads) using input as reference for visual comparison of ChIP-seq signal around gene loci.
Survival analysis
Survival analysis was performed using R version 3.6.2 [42]. The Kaplan–Meier method was used to determine overall survival (OS) and progression-free survival (PFS) for the PLAGL1- and PLAGL2-amplified tumors separately, as well as stratified by sex. The log-rank test (p-value) was used to identify differences between the Kaplan–Meier curves. Overall survival was defined as the time between first diagnosis and last follow-up date or death, and PFS was defined as the time between first diagnosis and time point of first relapse. A swimmer plot was used to display survival times, treatment, and outcome for each patient.
Results
Methylation analysis
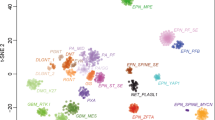
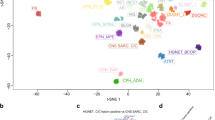
Unsupervised visualization of genome-wide DNA methylation data using t-distributed stochastic neighbor embedding (t-SNE) of > 90,000 pediatric and adult tumor samples of numerous types revealed a subset of 46 tumor samples clustering closely together, but away from established DNA methylation reference classes. Investigation of copy number alterations in each sample indicated amplification of the genetic loci corresponding to one of two different PLAG-family genes (PLAGL1 at 6q24.2 or PLAGL2 at 20q11.21) in the majority of tumors—a genetic aberration not known to be a characteristic feature in any of the currently defined CNS tumor types. We, therefore, provisionally named this novel DNA methylation class “CNS embryonal tumor with PLAG-family gene amplification”—ET, PLAGL (Fig. 1a).
DNA methylation clustering identifies a novel epigenetically distinct subtype of CNS embryonal tumor characterized by focal PLAG-family gene amplification. a Left: DNA methylation-based t-SNE analysis of > 90,000 pediatric and adult tumor samples. Circled are different medulloblastoma (MB) and embryonal tumor with multilayered rosettes (ETMR) subtypes, the ET, PLAGL type, and various low grade and high grade glioma subtypes—pilocytic astrocytoma (PA), pleomorphic Xanthoastrocytoma (PXA), H3 G34-mutant diffuse hemispheric glioma (G34), H3 K27-altered diffuse midline glioma (K27), diffuse pediatric-type high grade glioma, RTK subtype (pedRTK). Right: enlarged depiction of samples belonging to the ET, PLAGL type. The arrows mark two slightly outlying samples based on t-SNE. Methylation classes are color-coded as described in [12], grey color means the sample could not be matched to any of the existing methylation classes. b DNA methylation-based analysis using t-SNE dimensionality reduction on 33 ET, PLAGL tumors and a reference cohort of 910 different CNS tumors including 780 gliomas/glioneuronal tumors and 130 medulloblastomas. Methylation classes are color-coded and labeled using the respective group abbreviations. ET, PLAGL tumors are differentially colored according to their amplified PLAG-family gene. Two outlying ET, PLAGL samples are circled and marked with an arrow. Samples belonging to the ET, PLAGL type are depicted enlarged on the right. Full group names are: adult-type diffuse high grade glioma, IDH-wild type, subtype E (HGG_E), diffuse pediatric-type high grade glioma, RTK1 and 2 subtype (pedRTK1, pedRTK2), HGG-IDH wild type-subclass midline (HGG_MID), diffuse hemispheric glioma, H3 G34-mutant (G34), diffuse midline glioma, H3 K27-altered, subtype EGFR-altered (EGFR), diffuse midline glioma, H3 K27-altered (K27), glioblastoma, IDH-wild type, subtype posterior fossa (CBM), Glioblastoma, IDH-wild type, RTK1 and 2 subtype (RTK1, RTK2), Glioblastoma, IDH-wild type, mesenchymal subtype (MES), diffuse pediatric-type high grade glioma, MYCN subtype (pedMYCN), embryonal tumor, not otherwise specified (EMB), high-grade astrocytoma with piloid features (HGAP), Pleomorphic Xanthoastrocytoma (PXA), diffuse leptomeningeal glioneuronal tumor, subtype 1 and 2 (DLGNT_1, DLGNT_2), Medulloblastoma, SHH-activated (MB_SHH), Medulloblastoma, WNT-activated (MB_WNT), Medulloblastoma, non-WNT/non-SHH, Group 3 and 4 subtype (MBg34), Inflammatory microenvironment (LYMPH_HI), neuroepithelial tumor with PATZ1 fusion (PATZ), embryonal tumor with PLAG-family gene amplification (ET, PLAGL)
Out of the 46 samples initially belonging to the ET, PLAGL cluster, 11 samples were found to be duplicate or relapse samples based on genotype matches. One additional sample was excluded based on quality control indicating array hybridization issues. Another tumor with primary extracranial location was also excluded. This resulted in a set of 33 individual tumors classified as ET, PLAGL that were subjected to further analysis. Including information about the PLAGL1/PLAGL2 amplification status of each sample, we repeated t-SNE analysis using a select subset of 910 reference tumors of various types—including HGGs, medulloblastomas, and a set of the recently published neuroepithelial tumors with PATZ1 fusions [6]—together with the 33 ET, PLAGL tumors (Fig. 1b). All ET, PLAGL tumors formed one distinct cluster regardless of their PLAG gene amplification status, which confirmed their group affiliation and epigenetic similarity. The ET, PLAGL cluster was not located in proximity to any of the HGG, medulloblastoma, or other embryonal tumor clusters (Fig. 1a, b) underlining its epigenetic divergence from those tumors—an important point to stress since apart from HGG, medulloblastoma or other embryonal tumors were frequently among the initial histopathological diagnoses for the PLAGL-amplified cases, especially when occurring in the cerebellum. Two samples were found to be outliers that clustered close to ET, PLAGL, but slightly apart from the core group (Fig. 1a) as well as further apart in the refined t-SNE analysis (Fig. 1b). Both outlying samples were PLAGL1-amplified tumors, one of which was from an adult patient (age 59 years) and one with unknown age. These two samples were subsequently excluded and the remaining analyses were focused on the core cluster of 31 samples (Fig. 1b). When investigating possible further substructure within this cluster, there was some evidence that the ET, PLAGL cluster could potentially be subdivided into two different sub-clusters based on their location on the t-SNE plot, separating the PLAGL1-amplified from the PLAGL2-amplified samples. Three samples without apparent PLAG-family gene amplification were also part of the core group based on their DNA methylation pattern, with two seemingly PLAGL1-like and one PLAGL2-like. In a further t-SNE analysis, which also included a set of the recently published supratentorial ependymoma-like tumors with PLAGL1 fusions [54], as well as 1,124 sarcomas in addition to the previous reference cohort of 910 tumors, the PLAGL-amplified samples maintained its own unique cluster (Supplementary Fig. 1).
Copy number analysis
We derived copy number (CN) plots and assessed CN status for all 31 samples based on the raw intensities of the DNA methylation array probes, which revealed focal amplification of PLAGL1 or PLAGL2 in 28 of the 31 core samples (90.3%) with 11 samples being PLAGL1-amplified (35.5%) and 17 samples being PLAGL2-amplified (54.8%). Three samples showed no amplification of any PLAG-family gene (9.7%). CN summary plots were derived for PLAGL1- and PLAGL2-amplified samples separately to visualize broad chromosomal copy number changes in each subtype (Fig. 2a). As the segmentation algorithm used to produce the summary plots does not always recognize amplicons of very small size as a segment, only a subset of the PLAGL2 amplifications were detected automatically, but manual screening of the PLAGL-regions confirmed focal amplification of PLAGL1 or PLAGL2 as described above (Fig. 2b, c). Differential comparative analysis was performed using GISTIC2.0 to compare PLAGL1-amplified versus PLAGL2-amplified samples and detect significantly altered regions across all samples and per subtype (Fig. 2d, Supplementary Fig. 3).
Copy number analysis of CNS embryonal tumors with PLAGL gene amplification. a Copy number summary plots were derived per subgroup for PLAGL1-amplified and PLAGL2-amplified tumors. b, c Chromosome 6 and chromosome 20 amplifications in ET, PLAGL tumors were verified using IGV. Samples are grouped according to their amplification status. b Shown are focal amplifications on chromosome 6 and chromosome 20 for PLAGL1 and PLAGL2, respectively. c Zooming in on the amplified regions around PLAGL1 and PLAGL2 (with co-amplification). d GISTIC amplification plot of all 31 samples belonging to the ET, PLAGL type. The genome is displayed vertically on the y-axis and genomic positions of amplified regions are indicated. Normalized amplification signals (G-score) and q values (log scale) are indicated on the X-axis on the top and bottom, respectively. The green line represents the significance cutoff (q value = 0.25)
Ten of the 17 PLAGL2-amplified samples (58.8%) showed co-amplification of a region immediately downstream of PLAGL2 on chromosome 20, which mainly affected the gene CBFA2T2 (Fig. 2b, c). GISTIC2.0 analysis confirmed the region containing PLAGL1 (6q24.2; q value 4.83*10–23), PLAGL2 (20q11.21; q value 2.20*10–22), and the downstream region of co-amplification on 20q11.21 (q value 2.98*10–16) (Fig. 2d; Supplementary Tables S2, S3) as significantly amplified segments in the PLAGL1- and PLAGL2-amplified tumors, respectively. Multiple ET, PLAGL group-wide and subgroup-specific deletions were identified, including a common deletion of region 11p15.4 (q value 2.07*10–11) (Supplementary Fig. 3, Supplementary Tables S2, S4), but this region encompassing various olfactory receptors as well as further affected genomic regions are likely to represent copy number polymorphisms and/or technical artifacts rather than functional somatic alterations.
Patient and sample characteristics
Patient characteristics were summarized (n = 31, Table 1) and visualized (Supplementary Fig. 2). The median age as well as the age range differed between PLAGL1- and PLAGL2-amplified tumors (p = 0.005497, Wilcoxon rank sum test). PLAGL1-amplified cases occurred mainly in school age children and teenagers, with only a few younger patients (1–19 years, median age of 10.5 years), while PLAGL2-amplified cases were mostly prevalent in infants and toddlers, with an age range from 1 to 5 years (with the exception of one adult case of 36 years; median age of 2 years). The three ET, PLAGL tumors without PLAGL1/2 amplification occurred in very young patients (0–2 years, median age of 1). The incidence of PLAGL1 tumors was higher in girls than in boys (M:F 3:8), while the opposite trend was seen for the incidence of PLAGL2 tumors (M:F 10:7), but this difference in sex distribution was not statistically significant (p = 0.1021, Chi-square test). Tumors occurred at several anatomic sites, mainly the cerebral hemispheres (35.5%) and the cerebellum (25.8%), but were also found in the brainstem (6.5%), other midline structures (9.7%), or growing into the ventricles (6.5%). Nine out of ten hemispheric PLAGL-amplified tumors occurred in female patients, compared to only three out of seven cerebellar PLAGL-amplified tumors and one tumor that was growing into the ventricles (Supplementary Fig. 2). Initially rendered histopathological diagnoses based on morphological features comprised various tumor types—medulloblastoma (16.1%), other embryonal tumors (22.6%), HGGs (19.4%), indeterminate neuroepithelial tumors (9.7%), and sarcoma (3.2%) (Table 1).
Targeted next-generation DNA sequencing analysis
A subset of the tumors (n = 14) were further analyzed by targeted next-generation DNA sequencing to interrogate the genomic landscape beyond PLAGL amplification (Supplementary Table S1). One tumor with PLAGL2 amplification (#A113—excluded from the core DNA methylation cohort due to QC issues, but with a clear signal for ET, PLAGL), harbored additional focal high-level amplifications of the MDM4 oncogene on chromosome 1q32.1 and the MYCN oncogene on chromosome 2p24.3. In one tumor with PLAGL1 amplification (#A388), we found separate focal high-level amplification of the GLI2 oncogene on chromosome 2q14.2, and in another tumor with PLAGL1 amplification (#A93), we found focal amplification of MYB on chromosome 6q23.3. One PLAGL2-amplified tumor (#A105) and one PLAGL1-amplified tumor (#A93) harbored deleterious missense mutations in the TP53 tumor suppressor gene. The remaining seven tumors with PLAGL2 amplification and three tumors with PLAGL1 amplification harbored no additional likely oncogenic amplifications, homozygous deletions, mutations, insertions/deletions, or gene fusions among any of the evaluated genes. Specifically, all 14 evaluated tumors were wild type for the IDH1 and IDH2 genes, as well as the histone H3 genes (H3F3A, H3F3B, HIST1H3B, and HIST1H3C). None harbored amplifications, fusions, or mutations of receptor tyrosine kinase genes including EGFR, PDGFRA, FGFR1, MET, ALK, ROS1, or NTRK2 that are common in pediatric HGG. None harbored alterations within genes of the MAP kinase signaling pathway (e.g., BRAF, KRAS, NF1, and PTPN11) that are also common in pediatric gliomas. None harbored mutation or deletion of the SMARCB1 or SMARCA4 genes, thereby distinguishing these tumors from atypical teratoid/rhabdoid tumors. None harbored DICER1 mutation or amplification of the chromosome 19 microRNA cluster (C19MC), thereby distinguishing these tumors from embryonal tumor with multilayered rosettes. None harbored BCOR fusions or internal tandem duplication, and none of the evaluated tumors (n = 6) harbored MN1 or BEND2 fusions. Genetic alterations known to contribute to telomere maintenance (TERT promoter mutation or ATRX mutation/deletion) were also not identified in any of the tumors.
Histopathological characterization
Histopathological review was performed on a subset of the tumors, including 6 with PLAGL2 amplification, 8 with PLAGL1 amplification, and 1 with no PLAG gene family amplification. The predominant morphological pattern was a densely cellular neoplasm with solid growth composed of primitive, embryonal-like cells with brisk mitotic activity (Fig. 3, Supplementary Fig. 4, and Supplementary Table S5). Less common patterns included spindled and more uniform/monotonous round cells, but tumors with these patterns always had other areas with more primitive, embryonal-like cells. While most tumors demonstrated a solid growth pattern with a paucity of entrapped neuropil and a sharply circumscribed border with adjacent brain parenchyma, a couple of tumors displayed focal infiltrative growth. Many tumors had regions of necrosis, usually without palisading of tumor cells at the periphery. No well-developed microvascular proliferation was observed in any of the reviewed tumors. Ependymal canals or perivascular pseudorosettes (characteristic histological features of ependymoma) were not observed.
Imaging and histologic features of CNS embryonal tumors with PLAGL gene amplification. Shown are pre-operative T2-weighted MR images and low/high resolution H&E-stained histology images of a a PLAGL2-amplified tumor in a 2-year-old female patient (#A110) and b a PLAGL1-amplified tumor in a 13-year-old female patient (#A387)
Immunostaining for markers of glial differentiation (GFAP and OLIG2) was mostly negative, with only a few PLAGL2-amplified tumors showing labeling of rare scattered tumor cells (Fig. 4, Supplementary Table S5). Several tumors demonstrated patchy weak staining for synaptophysin, while others were negative. Neurofilament expression was often seen in scattered tumor cells, but otherwise revealed an absence of entrapped neuropil, confirming the solid growth pattern of these tumors. Two PLAGL2-amplified tumors demonstrated focal collections of tumor cells with paranuclear dot-like positivity for EMA staining, while the majority of tumors lacked EMA expression. All evaluated tumors had intact/retained expression of INI1/SMARCB1 and BRG1/SMARCA4. All evaluated tumors had minimal to absent immunostaining for LIN28A, BCOR, and CD99. A subset of tumors demonstrated positivity for YAP1 and GAB1, while no tumors had nuclear beta-catenin staining. Desmin expression was present in the majority of evaluated tumors (9/12, 75%), which ranged from rare scattered cells to diffuse strong labeling of all tumor cells in a small number of the PLAGL2-amplified cases. Other markers of myogenic differentiation (myogenin, smooth muscle actin, and MyoD1) were negative in all evaluated tumors. Ki-67 labeling indices ranged from 30 to 70%.
Immunohistochemical features of CNS embryonal tumors with PLAGL gene amplification. Shown are representative immunostains of a a PLAGL2-amplified tumor in a 1-year-old female patient and b a PLAGL1-amplified tumor in a 13-year-old female patient. c Summary of IHC results in PLAGL1/2-amplified tumor samples
Gene expression analysis
Differential gene expression between tumors with PLAG-family gene amplification and a selection of other CNS tumor types was examined using the R2 Genomics Analysis and Visualization Platform (http://r2.amc.nl). In concordance with the observed gene amplification, ET, PLAGL tumors showed overexpression of the respective amplified PLAG-family gene as assessed by RNA-seq (Fig. 5a, c), while both PLAGL1 and PLAGL2 are downregulated postnatally in normal brain and cerebellar tissues (https://apps.kaessmannlab.org/evodevoapp/) [13] (Supplementary Fig. 5). Leveraging our expression data set of 11 PLAGL1- and PLAGL2-amplified tumor samples and 279 samples from other CNS tumor and normal tissue types (HGGs with H3 G34R/V or K27M mutation and GBM_pedRTK1 or 2 (n = 76), PA with BRAF fusion (n = 25), PXA (n = 25), normal brain tissue (n = 36), embryonal tumors such as ATRT, ETMR, or medulloblastomas (n = 117)), we first compared gene expression of the PLAGL-amplified tumors to our subset of embryonal tumors. We derived a gene set specific to the ET, PLAGL type (Fig. 5a) as well as a PLAGL-specific gene-signature consisting of the top 250 differentially expressed genes (Supplementary Table S6). In addition to PLAGL1/2 overexpression, we found differential expression of several genes involved in developmental and differentiation processes such as CDX1, NR5A1, TLX1, TBX1, FGF19, and DLK1 (Fig. 5a, Supplementary Table S6); known direct PLAGL target genes such as IGF2, H19, CDKN1C and DLK1 [64] (Fig. 5a, Supplementary Fig. 6), as well as CYP2W1 and the kinase RET, both putative treatment targets (Fig. 5a, c). We screened expression of 86 human IGs in the PLAGL1/2-amplified samples (Fig. 5a). A subset of 13 IGs (Meg3, Ndn, Grb10, Dlk1, Igf2, Cdkn1c, Plagl1, Peg3, Mest, Nnat, Asb4, H19, and Ppp1r9a) described as having high connectivity with other IGs [5] were differentially expressed in the PLAGL1/2-amplified tumors (Fig. 5a, Supplementary Figs. 6, 7). We also ran the same differential expression analyses comparing ET, PLAGL versus glial tumors as well as versus normal fetal and adult brain tissues. This analysis yielded similar results regarding the overrepresentation of imprinted genes as well as developmental and differentiation-related genes (Supplementary Fig. 8). Expression of classical pan-neuronal, glial, sarcoma/mesenchymal, neural stem cell, and proliferation marker genes was also examined in the ET, PLAGL tumors versus our subset of CNS embryonal tumors, gliomas, and normal tissues, but was inconclusive in terms of possible cell/lineage of origin, as there was no set of marker genes that was clearly differentially expressed in the ET, PLAGL tumor type—with the exception of high Desmin expression in the PLAGL-amplified tumors in all three comparisons (Supplementary Fig. 9). Overexpression of the myogenic marker Desmin was more pronounced in the PLAGL2-amplified samples (Supplementary Fig. 9d). Furthermore, this analysis showed a lack of glial marker expression in the PLAGL tumors. We compared bulk RNA-seq data of the ET, PLAGL tumors to a single-nucleus sequencing atlas containing transcriptomes from different cell types, differentiation states, and subtypes of the developing human cerebellum to map the cellular origins of the ET, PLAGL tumors (Supplementary Fig. 10) as described in Okonechnikov et al. [39]. None of the lineages that were used as reference could be identified as the origins of ET, PLAGL tumors. Consequently, we analyzed the expression of genes representing different developmental states and locations as markers for pluripotency, germ layers (ectoderm, mesoderm, endoderm), neuroectoderm, forebrain and pallium, subpallium (including the ganglionic eminence), midbrain, hindbrain, spinal cord, as well as various additional pan-neuronal and glial markers [56] (Fig. 5b, Supplementary Fig. 11). The early neural genes OTX2, TLX1, SIX3, MSI1, and DACH1 were overexpressed in the PLAGL-amplified tumors, as were some subpallial neural markers such as DLX5, DLX6, the lateral ganglionic eminence (LGE) marker ISL1, and the germ layer markers KRT18 and GATA4 pointing to a cell of origin at an early and rather undifferentiated developmental stage.
Gene expression profiles of CNS embryonal tumors with PLAGL gene amplification. a, b Volcano plots showing fold-change and p-value for the comparison of differential gene expression of 11 PLAGL1/2-amplified tumors versus 117 embryonal tumors from different types and subtypes. Highlighted are a 86 human IGs (ocher) and 13 IGs with high connectivity (lilac) as described in reference [5]. Shown in black: selection of genes with large magnitude fold-changes (x axis) and high statistical significance (− log10 of p-value, y-axis). b Genes with differential expression in different brain regions and during different developmental states as described in reference [56] c Boxplots comparing gene expression between CNS tumor types for a select set of genes. The subset of 117 embryonal tumor samples (atrt, etmr, med) is identical to a and b. plagl, ET,PLAGL; pa, pilocytic astrocytoma; pxa, pleomorphic xanthoastrocytoma; hgg, high-grade gliomas (G34R/V, K27M, pedRTK1/2); norm, normal brain tissues; atrt, atypical teratoid rhabdoid tumor; etmr, embryonal tumor with multilayered rosettes; med, medulloblastomas (WNT, SHH, group 3, group 4); red: samples with PLAGL1 amplification, blue: samples with PLAGL2 amplification. Significance bars indicate groups whose differences in gene expression are statistically significant when compared to samples with PLAGL1/2 amplification (t-test, Bonferroni-corrected p-value = 0.00714286). PLAGL1/2 upregulation is statistically significant compared to all other groups when looking at PLAGL1 or PLAGL2 tumors separately
ChIP-seq analysis
Chromatin immunoprecipitation studies using antibodies against PLAGL1 and PLAGL2 proteins (n = 5) was performed to identify gene loci bound by these two transcription factors in PLAGL-amplified tumors. This ChIP-seq data confirmed the known Plagl1 targets IGF2, CDKN1C, and DLK1, as well as most of the other IGs with high connectivity, as being direct targets of PLAGL1 and PLAGL2 TF binding in the PLAGL-amplified tumors (Supplementary Fig. 6, Supplementary Fig. 12). The receptor tyrosine kinase RET and the cytochrome P450 family member CYP2W1—both potential drug targets—were also revealed as further direct PLAGL1/2 targets (Supplementary Fig. 13). In addition, we identified components of the Wnt/β-Catenin signaling pathway, FZD2 and FZD9, to be targets of PLAGL1/2 TF binding that are also differentially expressed in the PLAGL-amplified tumors (Supplementary Fig. 14).
Survival analysis
Clinical outcome data were available for 21 patients with PLAGL1/2-amplified tumors. Five-year and 10-year OS for patients with PLAGL1- and PLAGL2-amplified tumors as well as for male and female patients was determined. Survival rates across the cohort remained constant after 5 years, hence both 5- and 10-year OS was 66% for patients with PLAGL1-amplified tumors, 25% for patients with PLAGL2-amplified tumors, 18% for male patients, and 82% for female patients, respectively. Although a trend towards a worse prognosis for patients with PLAGL2-amplified tumors was noticeable—with 5 out of 12 patients with a PLAGL2-amplified tumor being deceased compared to 2 out of 9 patients with a PLAGL1-amplified tumor (Fig. 6b)—PFS and OS did not differ significantly between the two different groups (Fig. 6a, p value = 0.096 and 0.44, respectively). Patient sex was also not a significant predictor for PFS or OS (Fig. 6a, p value = 0.12 and 0.2), but more deaths in male patients were recorded irrespective of the subgroup. With respect to different treatment regimens, the inclusion of chemotherapy agents beyond temozolomide (TMZ) early on in treatment showed a potential benefit for patient survival (Fig. 6b, Supplementary Table S7) while the inclusion of radiotherapy as part of the initial treatment seemed to have limited effect (Supplementary Fig. 15), but this should be judged with caution given the overall low numbers.
Clinical outcomes of patients with CNS embryonal tumor with PLAGL gene amplification. a Kaplan–Meier plots showing OS and PFS stratified by subgroup and sex. The log-rank test was used to show differences between the curves, p-values of the log-rank test are shown in each graph. b Swimmer plot showing available OS and PFS times per patient, including treatment information and clinical response/relapse. Samples are stratified by sex, PLAGL1/2 amplification status is indicated. Information about surgical resection (SUR) and presence of metastasis (MET) at the time point of primary diagnosis is displayed in the squares on the left where available (resections or metastases at later time points are not displayed), GTR, gross total resection; STR, subtotal resection; RES, resection (unknown, if GTR or STR). Information about chemotherapy (CT) and radiotherapy (RT) treatment regarding the entire follow-up time is displayed in the squares on the left where available
Discussion
We describe a rare, novel type of pediatric CNS tumor with a distinct methylation pattern that we name “CNS embryonal tumor with PLAGL amplification” (abbreviated as ET, PLAGL). The described new type is epigenetically divergent from all other described CNS tumor types and in the vast majority of cases marked by amplification of one of the PLAG-family genes, PLAGL1 or PLAGL2. Despite the initial diagnosis as medulloblastoma, other embryonal tumor, or HGG in more than half of cases, PLAGL-amplified tumors differ both epigenetically and in terms of outcome from these other tumor types. Initial histopathologic diagnoses were variable, and many tumors had been considered not classifiable despite multiple differential diagnoses being discussed. t-SNE dimensionality reduction of DNA methylation profiles unraveled further substructure within the PLAGL group in accordance with the respective gene amplification—PLAGL1 or PLAGL2. A subset of the PLAGL2-amplified cases also harbored co-amplification of a downstream region on chromosome 20 as possible evidence for more complex structural rearrangements, which will be of interest to explore further in larger tumor cohorts.
PLAGL1, PLAGL2, and also PLAG1 belong to the PLAG gene family [20]. PLAG1 and PLAGL2 have been suggested to be proto-oncogenes with comparable DNA-binding capacities and partly overlapping functions [4, 20]. PLAG1 promoter swapping and subsequent activation is reported as playing a key role in the development of pleomorphic adenomas of the salivary gland, lipoblastomas, hepatoblastomas, and some leukemias, and PLAG1 overexpression has been found in uterine leiomyomas, leiomyosarcomas, and other smooth muscle tumors [4, 65, 66]. PLAGL2 has been found to be amplified in a small subset of cancers, and has been reported to promote tumorigenesis together with POFUT1 in colorectal cancer [4, 32]. Overexpression of PLAGL2 has been reported to play a role in lung adenocarcinoma development, and may represent a poor prognostic marker in prostate cancer [18, 69]. The role of PLAGL1 is less clear, and it may potentially act as both tumor suppressor and oncogene depending on the context. PLAGL1 was first discovered as a zinc finger protein that concurrently induces apoptosis and cell cycle arrest [55]. The maternally imprinted PLAGL1 is expressed in normal tissues and downregulated in various tumors, such as breast cancer, ovarian cancer, nonfunctioning pituitary tumors, basal cell carcinomas, head and neck squamous cell carcinoma, and diffuse large B-cell lymphoma [3, 8, 30, 64]. It was shown to downregulate proliferation through induction of PPARɣ in human colon carcinoma cells [8]. In contrast to these findings, overexpression of PLAGL1 was found to contribute to the tumorigenesis of glioma-initiating cells [21] and recurrent PLAGL1 fusions (most commonly with EWSR1) were found to characterize a novel subtype of supratentorial ependymoma-like tumor in pediatric patients [54]. Our findings further support that in a brain tumor context, both PLAGL1 and PLAGL2 likely act as oncogenes, whose amplification and resultant overexpression drive tumor development.
In addition to the apparent epigenetic differences between PLAGL1- and PLAGL2-amplified tumors discovered through t-SNE analysis, we noted further distinctions between the two subtypes. PLAGL1-amplified tumors rarely occurred in young children and were more prevalent in school-aged children and teenagers, while PLAGL2-amplified cases were mostly prevalent in infants and toddlers. While PLAGL1 is ubiquitously expressed in many normal fetal and adult tissues, PLAGL2 is only expressed in fetal tissues [2]. Accordingly, the median age at diagnosis was significantly lower for the PLAGL2-amplified tumors than for the PLAGL1-amplified tumors. Although not significant, a trend towards a more favorable clinical outcome in PLAGL1-amplified tumors was noted. One possibility is that amplification of PLAGL2 leading to tumor formation can only occur during a small spatiotemporal window during development when PLAGL2 is expressed, while amplification of PLAGL1 might be less temporally limited in terms of subsequent tumor formation. Alternatively, considering the less aggressive phenotype and more favorable outcome, PLAGL1 activation might lead to slower growing tumors that only become symptomatic in older children, even if they arise around the same time during development and target the same cell type of origin.
We further report clinical differences between male and female patients within this tumor type. The incidence of PLAGL1 tumors was higher in female patients, while the incidence of PLAGL2 tumors was higher in male patients. In terms of location, cerebral hemispheric tumors were more prevalent in females, while cerebellar, brainstem, and other midline tumors were more prevalent in males. Sex-specific differences were also noted in terms of outcome, with female patients showing a trend towards more favorable survival. Since the patients who succumbed to their disease more often had tumors in the cerebellum or midline structures, one possible explanation apart from molecular differences between the tumors might be the more surgically accessible tumor location in the cerebral hemispheres, which is predominantly found in female patients and might better enable total surgical excision. Outcomes do not seem to be related to differences in the specific PLAGL gene that is amplified or treatment between the two sexes based on the patient cohort to date, although it should be noted that there was some evidence of improved outcomes for early chemotherapeutic intervention for incompletely resected tumors.
Gene expression profiles of 11 tumors with amplification of PLAGL1 (n = 5) or PLAGL2 (n = 6) were compared to gene expression profiles of a total of 279 tumor samples from various other CNS tumor types and normal tissues. Overexpression of a subset of 13 IGs—MEG3, NDN, GRB10, DLK1, IGF2, CDKN1C, PLAGL1, PEG3, MEST, NNAT, ASB4, H19, PPP1R9A—was specific for the PLAGL-amplified tumors. These 13 genes are imprinted in humans and are reported to have high connectivity with other genes belonging to the IGN in mouse [5]. Plagl1 was previously shown to regulate expression of Cdkn1c, Igf2, H19, and Dlk1 and to belong to a subset of IGs that control embryonic growth and differentiation, and loss of Plagl1 function resulted in intrauterine growth restriction [64]. We show through analysis of ChIP-seq data that PLAGL1 and PLAGL2 bind directly upstream of the majority of the 13 IGs in human PLAGL1/2-amplified tumors, which underlines similarities in the mouse and human networks. Furthermore, 9 of the 13 genes—Igf2, H19, Plagl1, Mest, Peg3, Dlk1, Grb10, Ndn, Cdkn1c—were reported to belong to a subset of 11 IGs which were downregulated postnatally in an age-dependent fashion accompanying a decline in growth rate [36]. The same set of nine genes was found differentially expressed in different types of mouse and human somatic stem cells compared to their differentiated counterparts, and the expression of those IGs correlated with stem cell properties [9]. A subset (MEST, PLAGL1, PEG3, DLK1, IGF2) showed elevated expression in various embryonal cancers, especially rhabdomyosarcoma, as compared to non-embryonal cancers and normal tissues. The whole set of nine genes was found overexpressed in mouse embryoid bodies—aggregates of embryonic stem cells (ESCs) undergoing differentiation and comprising differentiated cell phenotypes of all three germ lineages [10, 29]—but not in ESCs [9, 44]. Further literature confirms the role of PLAGL1 as a “master switch”, a TF that regulates a substantial fraction of the IGN genes and extracellular matrix (ECM) genes, and may regulate a subset of neuroblastoma signature genes [63].
IGs are considered to be key regulators of embryonic development and global loss of imprinting (LOI) as well as LOI of IGF2, which is regulated by PLAGL1, can lead to tumor formation [5, 22, 64]. It was also shown that overexpression of Plagl1 abolished the neuronal commitment of non-glioma-initiating cells and caused them to become malignant [21]. Therefore, we conclude that PLAGL1 amplification and subsequent overexpression may contribute to tumor formation depending on the cell of origin and developmental state. The above-named subset of 13 IGs was also overexpressed to the same extent in PLAGL2-amplified samples, despite the fact that PLAGL2 is not an IG in humans and has not been associated with the mouse IGN of 409 (85 imprinted and 324 non-imprinted) genes published by Al Adhami et al. [5]. However, significant overlap was found between Plagl1 target genes and PLAG1 and Plagl2 targets [63]. PLAGL2 was shown to upregulate IGF2 expression levels in hematopoietic progenitors of acute myeloid leukemia and IGF2 harbors eight PLAG1/PLAGL2 consensus binding sites [31]. PLAGL2 was also reported to activate the IGF2 signaling pathway in colorectal cancer [34]. It is, therefore, possible that amplification and overexpression of PLAGL2 as well as subsequent upregulation of IGF2 in turn lead to both upregulation of genes of the IGN, specifically of those 13 genes with high connectivity, and activation of the IGF mitotic signaling pathway. Amplification and subsequent overexpression of PLAGL2 was also shown to suppress differentiation in neural stem cells and glioma-initiating cells, in part through aberrant Wnt/β-Catenin signaling in malignant glioma [71]. In keeping herewith, we show through ChIP-seq data that components of the Wnt-pathway such as FZD2 and FZD9 are direct targets of PLAGL1 and PLAGL2 and are overexpressed in the PLAGL1/2-amplified samples compared to the other CNS tumor types. Further genes that were found to be overexpressed as well as direct PLAGL1/2 targets are the receptor tyrosine kinase rearranged during transfection (RET) and the cytochrome P450 family gene CYP2W1. RET plays a role in the development of the nervous system, where it is expressed in neural crest cells. It was reported to be oncogenic through gene rearrangements, activating mutations, or overexpression of the wild-type gene in multiple cancers, and various small molecule inhibitors targeting RET are available or being tested in clinical trials [15, 51, 57]. CYP2W1 belongs to the cytochrome P450 superfamily of monooxygenases that are involved in xenobiotic metabolism [53]. CYP2W1 was found to be selectively expressed in colon cancer tissues, but not in healthy tissues making it a putative tumor-specific drug target [61]. The fact that it was also found highly overexpressed in our cohort of PLAGL-amplified tumors indicates its potential for targeted treatment, for example, with seco-duocarmycin based prodrugs that are converted into cytotoxic metabolites upon bioactivation by CYP2W1, or with antibody drug conjugates (ADC) whose payload gets released upon antibody cleavage [7, 49, 70].
Since our intent to map the cellular origins using a single-nucleus atlas of the developing human cerebellum did not yield informative results, we screened the expression of classical marker genes as well as multiple genes with differential expression in various regions and developmental states of the brain to investigate the potential underlying cell of origin of PLAGL-amplified CNS tumors. Glial, astroglial, and oligodendrocytic markers were clearly under-expressed compared to glial tumors and normal brain tissues, making it unlikely that PLAGL-amplified tumors are of purely glial origin. The neural stem cell markers SOX2 and Nestin were also either not differentially expressed or under-expressed in the PLAGL-amplified type compared to the other CNS tumor types. Neuronal marker expression did not unambiguously point to a neuronal tumor, as classical neuronal markers were not over-represented among the differentially expressed genes specific to the PLAGL-amplified tumors. The germ layer markers GATA4 and KRT18 were over-represented in the ET, PLAGL tumors and concurrently, some neuronal genes were also expressed to some extent—TUBB3, which is expressed in neuronal restricted progenitor cells [47], and several early neural genes [56] such as TLX1, which is a key TF in the determination of neuronal cell fates [19, 45], were overexpressed in the PLAGL type, potentially suggesting a neuronal lineage at an early developmental state. During mouse brain development (E12-E14—corresponds to 36–48 days post conception in humans) the mouse orthologue Plagl1 shows strong expression in the neural tube and several neuroepithelia such as the telencephalic vesicles (forebrain region) as well as the third (midbrain) and fourth (hindbrain) ventricles. Plagl1-expressing cells are further found in the spinal cord, co-localized with Tubb3 in the outer cell layer of the subventricular zone of the neuroepithelia, in the brainstem and many more proliferative areas [62], which is consistent with our finding of high expression of early neural genes as well as marker genes localized in the forebrain, midbrain, hindbrain, or spinal cord and could also be an explanation for the diverse tumor sites we report. We found overexpression and differential expression of several genes that are highly expressed in the human ganglionic eminence (GE) at around 7–9 p.c.w.—DLX5, DLX6, and the aforementioned SIX3—as well as DLX2, which is expressed before 7 weeks p.c. and ISL1, a neuroblast marker expressed in the lateral ganglionic eminence (LGE) around 8 weeks p.c. [24, 56]. In general, all three PLAG-family genes are found at higher levels in neural progenitor cells than in post-mitotic neurons [4]. Data derived from scRNA-seq from the first trimester of human development showed a gradual transition from neuroepithelial to radial glia populations rather than specific gene programs that distinguish progenitor groups—Eze et al. describe a DLK1 overexpressing cortical progenitor cluster, where DLK1 in combination with low SOX2 levels, a combination we also see in the PLAGL-amplified tumors, was highly enriched in early samples and disappeared after CS16 [17]. In addition, we find concurrent expression of BCL11B and to some extent DCX in the PLAGL-amplified tumors—which were described as markers of maturing and newborn neurons, respectively. Overall, these data support that PLAGL1/2-amplified tumors may arise from early neural progenitor cells with possible early commitment to a neuronal lineage. However, the morphology and other histological features confirm only an undifferentiated, embryonal-like pattern. We, therefore, propose the name ET, PLAGL, for CNS embryonal tumor with PLAGL gene amplification.
Conclusion
We describe a novel, biologically distinct type of CNS embryonal tumor—ET, PLAGL—that is presumably driven by amplification and subsequent overexpression of PLAGL1 or PLAGL2, the only recurrent molecular event detected in these tumors. Dysregulation of imprinted genes is a common feature in the PLAGL1/2-amplified tumors that might contribute to tumor formation. Histopathological appearance is that of a primitive, embryonal-like neoplasm without specific morphologic features or routine immunohistochemical markers that enable the accurate distinguishment of ET, PLAGL from other CNS tumor types. Further studies will be needed to identify ET, PLAGL-specific biomarkers that can be used in routine neuropathology diagnostic practice. Finally, the oncogenic RET kinase as well as CYP2W1 are direct PLAGL1/2 targets and may be potential drug targets in this tumor type, which should be top priorities for future functional validation.
Change history
14 February 2023
A Correction to this paper has been published: https://doi.org/10.1007/s00401-023-02538-4
References
Central Nervous System Tumours (2021) WHO classification of tumours, 5th edition, volume 6. Edited by the WHO Classification of Tumours Editorial Board. 2021: International Agency for Research on Cancer
Abdollahi A (2007) LOT1 (ZAC1/PLAGL1) and its family members: mechanisms and functions. J Cell Physiol 210(1):16–25. https://doi.org/10.1002/jcp.20835
Abdollahi A, Pisarcik D, Roberts D, Weinstein J, Cairns P, Hamilton TC (2003) LOT1 (PLAGL1/ZAC1), the candidate tumor suppressor gene at chromosome 6q24-25, is epigenetically regulated in cancer. J Biol Chem 278(8):6041–6049. https://doi.org/10.1074/jbc.M210361200
Adnani L, Dixit R, Chen X, Balakrishnan A, Modi H, Touahri Y et al (2018) Plag1 and Plagl2 have overlapping and distinct functions in telencephalic development. Biol Open. https://doi.org/10.1242/bio.038661
Al Adhami H, Evano B, Le Digarcher A, Gueydan C, Dubois E, Parrinello H et al (2015) A systems-level approach to parental genomic imprinting: the imprinted gene network includes extracellular matrix genes and regulates cell cycle exit and differentiation. Genome Res 25(3):353–367. https://doi.org/10.1101/gr.175919.114
Alhalabi KT, Stichel D, Sievers P, Peterziel H, Sommerkamp AC, Sturm D et al (2021) PATZ1 fusions define a novel molecularly distinct neuroepithelial tumor entity with a broad histological spectrum. Acta Neuropathol 142(5):841–857. https://doi.org/10.1007/s00401-021-02354-8
Bart AG, Morais G, Vangala VR, Loadman PM, Pors K, Scott EE (2022) Cytochrome P450 binding and bioactivation of tumor-targeted duocarmycin agents. Drug Metab Dispos 50(1):49–57. https://doi.org/10.1124/dmd.121.000642
Barz T, Hoffmann A, Panhuysen M, Spengler D (2006) Peroxisome proliferator-activated receptor gamma is a Zac target gene mediating Zac antiproliferation. Cancer Res 66(24):11975–11982. https://doi.org/10.1158/0008-5472.CAN-06-1529
Berg JS, Lin KK, Sonnet C, Boles NC, Weksberg DC, Nguyen H et al (2011) Imprinted genes that regulate early mammalian growth are coexpressed in somatic stem cells. PLoS ONE 6(10):e26410. https://doi.org/10.1371/journal.pone.0026410
Bratt-Leal AM, Carpenedo RL, McDevitt TC (2009) Engineering the embryoid body microenvironment to direct embryonic stem cell differentiation. Biotechnol Prog 25(1):43–51. https://doi.org/10.1002/btpr.139
Cacciotti C, Fleming A, Ramaswamy V (2020) Advances in the molecular classification of pediatric brain tumors: a guide to the galaxy. J Pathol 251(3):249–261. https://doi.org/10.1002/path.5457
Capper D, Jones DTW, Sill M, Hovestadt V, Schrimpf D, Sturm D et al (2018) DNA methylation-based classification of central nervous system tumours. Nature 555(7697):469–474. https://doi.org/10.1038/nature26000
Cardoso-Moreira M, Halbert J, Valloton D, Velten B, Chen C, Shao Y et al (2019) Gene expression across mammalian organ development. Nature 571(7766):505–509. https://doi.org/10.1038/s41586-019-1338-5
Castel D, Kergrohen T, Tauziede-Espariat A, Mackay A, Ghermaoui S, Lechapt E et al (2020) Histone H3 wild-type DIPG/DMG overexpressing EZHIP extend the spectrum diffuse midline gliomas with PRC2 inhibition beyond H3–K27M mutation. Acta Neuropathol 139(6):1109–1113. https://doi.org/10.1007/s00401-020-02142-w
Drilon A, Hu ZI, Lai GGY, Tan DSW (2018) Targeting RET-driven cancers: lessons from evolving preclinical and clinical landscapes. Nat Rev Clin Oncol 15(3):150. https://doi.org/10.1038/nrclinonc.2017.188
Edgar R, Domrachev M, Lash AE (2002) Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30(1):207–210. https://doi.org/10.1093/nar/30.1.207
Eze UC, Bhaduri A, Haeussler M, Nowakowski TJ, Kriegstein AR (2021) Single-cell atlas of early human brain development highlights heterogeneity of human neuroepithelial cells and early radial glia. Nat Neurosci 24(4):584–594. https://doi.org/10.1038/s41593-020-00794-1
Guo J, Wang M, Wang Z, Liu X (2016) Overexpression of pleomorphic adenoma gene-like 2 is a novel poor prognostic marker of prostate cancer. PLoS ONE 11(8):e0158667. https://doi.org/10.1371/journal.pone.0158667
Guo Z, Zhao C, Huang M, Huang T, Fan M, Xie Z et al (2012) Tlx1/3 and Ptf1a control the expression of distinct sets of transmitter and peptide receptor genes in the developing dorsal spinal cord. J Neurosci 32(25):8509–8520. https://doi.org/10.1523/JNEUROSCI.6301-11.2012
Hensen K, Van Valckenborgh IC, Kas K, Van de Ven WJ, Voz ML (2002) The tumorigenic diversity of the three PLAG family members is associated with different DNA binding capacities. Cancer Res 62(5):1510–1517
Hide T, Takezaki T, Nakatani Y, Nakamura H, Kuratsu J, Kondo T (2009) Sox11 prevents tumorigenesis of glioma-initiating cells by inducing neuronal differentiation. Cancer Res 69(20):7953–7959. https://doi.org/10.1158/0008-5472.CAN-09-2006
Holm TM, Jackson-Grusby L, Brambrink T, Yamada Y, Rideout WM 3rd, Jaenisch R (2005) Global loss of imprinting leads to widespread tumorigenesis in adult mice. Cancer Cell 8(4):275–285. https://doi.org/10.1016/j.ccr.2005.09.007
Hovestadt VZM, conumee: enhanced copy-number variation analysis using Illumina DNA methylation arrays. R package version 1.9.0. http://bioconductor.org/packages/conumee/. Accessed 6 Jan 2021
Jansky S, Sharma AK, Korber V, Quintero A, Toprak UH, Wecht EM et al (2021) Single-cell transcriptomic analyses provide insights into the developmental origins of neuroblastoma. Nat Genet 53(5):683–693. https://doi.org/10.1038/s41588-021-00806-1
Jones C, Baker SJ (2014) Unique genetic and epigenetic mechanisms driving paediatric diffuse high-grade glioma. Nat Rev Cancer. https://doi.org/10.1038/nrc3811
Jones C, Karajannis MA, Jones DTW, Kieran MW, Monje M, Baker SJ et al (2017) Pediatric high-grade glioma: biologically and clinically in need of new thinking. Neuro Oncol 19(2):153–161. https://doi.org/10.1093/neuonc/now101
Kline CN, Joseph NM, Grenert JP, van Ziffle J, Talevich E, Onodera C et al (2017) Targeted next-generation sequencing of pediatric neuro-oncology patients improves diagnosis, identifies pathogenic germline mutations, and directs targeted therapy. Neuro Oncol 19(5):699–709. https://doi.org/10.1093/neuonc/now254
Koelsche C, Schrimpf D, Stichel D, Sill M, Sahm F, Reuss DE et al (2021) Sarcoma classification by DNA methylation profiling. Nat Commun 12(1):498. https://doi.org/10.1038/s41467-020-20603-4
Kurosawa H (2007) Methods for inducing embryoid body formation: in vitro differentiation system of embryonic stem cells. J Biosci Bioeng 103(5):389–398. https://doi.org/10.1263/jbb.103.389
Landrette SF, Madera D, He F, Castilla LH (2011) The transcription factor PlagL2 activates Mpl transcription and signaling in hematopoietic progenitor and leukemia cells. Leukemia 25(4):655–662. https://doi.org/10.1038/leu.2010.301
Landrette SF, Kuo YH, Hensen K, Barjesteh van Waalwijk van Doorn-Khosrovani S, Perrat PN, Van de Ven WJ et al (2005) Plag1 and Plagl2 are oncogenes that induce acute myeloid leukemia in cooperation with Cbfb-MYH11. Blood 105(7):2900–2907. https://doi.org/10.1182/blood-2004-09-3630
Li D, Lin C, Li N, Du Y, Yang C, Bai Y et al (2019) PLAGL2 and POFUT1 are regulated by an evolutionarily conserved bidirectional promoter and are collaboratively involved in colorectal cancer by maintaining stemness. EBioMedicine 45:124–138. https://doi.org/10.1016/j.ebiom.2019.06.051
Liu KW, Pajtler KW, Worst BC, Pfister SM, Wechsler-Reya RJ (2017) Molecular mechanisms and therapeutic targets in pediatric brain tumors. Sci Signal. https://doi.org/10.1126/scisignal.aaf7593
Liu X, Chen X, Zeng K, Xu M, He B, Pan Y et al (2018) DNA-methylation-mediated silencing of miR-486-5p promotes colorectal cancer proliferation and migration through activation of PLAGL2/IGF2/beta-catenin signal pathways. Cell Death Dis 9(10):1037. https://doi.org/10.1038/s41419-018-1105-9
Louis DN, Perry A, Wesseling P, Brat DJ, Cree IA, Figarella-Branger D et al (2021) The 2021 WHO classification of tumors of the central nervous system: a summary. Neuro Oncol 23(8):1231–1251. https://doi.org/10.1093/neuonc/noab106
Lui JC, Finkielstain GP, Barnes KM, Baron J (2008) An imprinted gene network that controls mammalian somatic growth is down-regulated during postnatal growth deceleration in multiple organs. Am J Physiol Regul Integr Comp Physiol 295(1):R189-196. https://doi.org/10.1152/ajpregu.00182.2008
Mermel CH, Schumacher SE, Hill B, Meyerson ML, Beroukhim R, Getz G (2011) GISTIC2.0 facilitates sensitive and confident localization of the targets of focal somatic copy-number alteration in human cancers. Genome Biol 12(4):R41. https://doi.org/10.1186/gb-2011-12-4-r41
Mondal G, Lee JC, Ravindranathan A, Villanueva-Meyer JE, Tran QT, Allen SJ et al (2020) Pediatric bithalamic gliomas have a distinct epigenetic signature and frequent EGFR exon 20 insertions resulting in potential sensitivity to targeted kinase inhibition. Acta Neuropathol 139(6):1071–1088. https://doi.org/10.1007/s00401-020-02155-5
Okonechnikov K, Joshi P, Sepp M, Leiss K, Sarropoulos I, Murat F et al (2021) Mapping pediatric brain tumors to their origins in the developing cerebellum. bioRxiv 2021.2012.2019.473154. https://doi.org/10.1101/2021.12.19.473154
Ostrom QT, Cioffi G, Waite K, Kruchko C, Barnholtz-Sloan JS (2021) CBTRUS statistical report: primary brain and other central nervous system tumors diagnosed in the United States in 2014–2018. Neuro Oncol 23(12 Suppl 2):iii1–iii105. https://doi.org/10.1093/neuonc/noab200
Pfister SM, Reyes-Mugica M, Chan JKC, Hasle H, Lazar AJ, Rossi S et al (2021) A summary of the inaugural WHO classification of pediatric tumors: transitioning from the optical into the molecular era. Cancer Discov. https://doi.org/10.1158/2159-8290.CD-21-1094
R Core Team, R (2019) A language and environment for statistical computing. 2019, R Foundation for Statistical Computing: Vienna, Austria
Reich M, Liefeld T, Gould J, Lerner J, Tamayo P, Mesirov JP (2006) GenePattern 2.0. Nat Genet 38(5):500–501. https://doi.org/10.1038/ng0506-500
Rezvani G, Lui JC, Barnes KM, Baron J (2012) A set of imprinted genes required for normal body growth also promotes growth of rhabdomyosarcoma cells. Pediatr Res 71(1):32–38. https://doi.org/10.1038/pr.2011.6
Riz I, Lee HJ, Baxter KK, Behnam R, Hawley TS, Hawley RG (2009) Transcriptional activation by TLX1/HOX11 involves Gro/TLE corepressors. Biochem Biophys Res Commun 380(2):361–365. https://doi.org/10.1016/j.bbrc.2009.01.099
Robinson JT, Thorvaldsdottir H, Winckler W, Guttman M, Lander ES, Getz G et al (2011) Integrative genomics viewer. Nat Biotechnol 29(1):24–26. https://doi.org/10.1038/nbt.1754
Schiffer D, Mellai M, Boldorini R, Bisogno I, Grifoni S, Corona C et al (2018) The significance of chondroitin sulfate proteoglycan 4 (CSPG4) in human gliomas. Int J Mol Sci. https://doi.org/10.3390/ijms19092724
Schwartzentruber J, Korshunov A, Liu XY, Jones DT, Pfaff E, Jacob K et al (2012) Driver mutations in histone H3.3 and chromatin remodelling genes in paediatric glioblastoma. Nature 482(7384):226–231. https://doi.org/10.1038/nature10833
Scribner JA, Brown JG, Son T, Chiechi M, Li P, Sharma S et al (2020) Preclinical development of MGC018, a duocarmycin-based antibody-drug conjugate targeting B7–H3 for solid cancer. Mol Cancer Ther 19(11):2235–2244. https://doi.org/10.1158/1535-7163.MCT-20-0116
Selt F, Deiss A, Korshunov A, Capper D, Witt H, van Tilburg CM et al (2016) Pediatric Targeted Therapy: Clinical Feasibility of Personalized Diagnostics in Children with Relapsed and Progressive Tumors. Brain Pathol 26(4):506–516. https://doi.org/10.1111/bpa.12326
Seoane J, Capdevila J (2018) The right compound for the right target: tackling RET. Ann Oncol 29(8):1623–1625. https://doi.org/10.1093/annonc/mdy188
Sharma T, Schwalbe EC, Williamson D, Sill M, Hovestadt V, Mynarek M et al (2019) Second-generation molecular subgrouping of medulloblastoma: an international meta-analysis of Group 3 and Group 4 subtypes. Acta Neuropathol 138(2):309–326. https://doi.org/10.1007/s00401-019-02020-0
Sheldrake HM, Travica S, Johansson I, Loadman PM, Sutherland M, Elsalem L et al (2013) Re-engineering of the duocarmycin structural architecture enables bioprecursor development targeting CYP1A1 and CYP2W1 for biological activity. J Med Chem 56(15):6273–6277. https://doi.org/10.1021/jm4000209
Sievers P, Henneken SC, Blume C, Sill M, Schrimpf D, Stichel D et al (2021) Recurrent fusions in PLAGL1 define a distinct subset of pediatric-type supratentorial neuroepithelial tumors. Acta Neuropathol 142(5):827–839. https://doi.org/10.1007/s00401-021-02356-6
Spengler D, Villalba M, Hoffmann A, Pantaloni C, Houssami S, Bockaert J et al (1997) Regulation of apoptosis and cell cycle arrest by Zac1, a novel zinc finger protein expressed in the pituitary gland and the brain. EMBO J 16(10):2814–2825. https://doi.org/10.1093/emboj/16.10.2814
Straccia M, Garcia-Diaz Barriga G, Sanders P, Bombau G, Carrere J, Mairal PB et al (2015) Quantitative high-throughput gene expression profiling of human striatal development to screen stem cell-derived medium spiny neurons. Mol Ther Methods Clin Dev 2:15030. https://doi.org/10.1038/mtm.2015.30
Subbiah V, Yang D, Velcheti V, Drilon A, Meric-Bernstam F (2020) State-of-the-art strategies for targeting RET-dependent cancers. J Clin Oncol 38(11):1209–1221. https://doi.org/10.1200/JCO.19.02551
Subbiah V, Velcheti V, Tuch BB, Ebata K, Busaidy NL, Cabanillas ME et al (2018) Selective RET kinase inhibition for patients with RET-altered cancers. Ann Oncol 29(8):1869–1876. https://doi.org/10.1093/annonc/mdy137
Subbiah V, Gainor JF, Rahal R, Brubaker JD, Kim JL, Maynard M et al (2018) Precision targeted therapy with BLU-667 for RET-driven cancers. Cancer Discov 8(7):836–849. https://doi.org/10.1158/2159-8290.CD-18-0338
Tauziede-Espariat A, Debily MA, Castel D, Grill J, Puget S, Sabel M et al (2019) An integrative radiological, histopathological and molecular analysis of pediatric pontine histone-wildtype glioma with MYCN amplification (HGG-MYCN). Acta Neuropathol Commun 7(1):87. https://doi.org/10.1186/s40478-019-0738-y
Travica S, Pors K, Loadman PM, Shnyder SD, Johansson I, Alandas MN et al (2013) Colon cancer-specific cytochrome P450 2W1 converts duocarmycin analogues into potent tumor cytotoxins. Clin Cancer Res 19(11):2952–2961. https://doi.org/10.1158/1078-0432.CCR-13-0238
Valente T, Junyent F, Auladell C (2005) Zac1 is expressed in progenitor/stem cells of the neuroectoderm and mesoderm during embryogenesis: differential phenotype of the Zac1-expressing cells during development. Dev Dyn 233(2):667–679. https://doi.org/10.1002/dvdy.20373
Varrault A, Dantec C, Le Digarcher A, Chotard L, Bilanges B, Parrinello H et al (2017) Identification of Plagl1/Zac1 binding sites and target genes establishes its role in the regulation of extracellular matrix genes and the imprinted gene network. Nucleic Acids Res 45(18):10466–10480. https://doi.org/10.1093/nar/gkx672
Varrault A, Gueydan C, Delalbre A, Bellmann A, Houssami S, Aknin C et al (2006) Zac1 regulates an imprinted gene network critically involved in the control of embryonic growth. Dev Cell 11(5):711–722. https://doi.org/10.1016/j.devcel.2006.09.003
Voz ML, Agten NS, Van de Ven WJ, Kas K (2000) PLAG1, the main translocation target in pleomorphic adenoma of the salivary glands, is a positive regulator of IGF-II. Cancer Res 60(1):106–113
Voz ML, Mathys J, Hensen K, Pendeville H, Van Valckenborgh I, Van Huffel C et al (2004) Microarray screening for target genes of the proto-oncogene PLAG1. Oncogene 23(1):179–191. https://doi.org/10.1038/sj.onc.1207013
Worst BC, van Tilburg CM, Balasubramanian GP, Fiesel P, Witt R, Freitag A et al (2016) Next-generation personalised medicine for high-risk paediatric cancer patients—the INFORM pilot study. Eur J Cancer 65:91–101. https://doi.org/10.1016/j.ejca.2016.06.009
Wu G, Broniscer A, McEachron TA, Lu C, Paugh BS, Becksfort J et al (2012) Somatic histone H3 alterations in pediatric diffuse intrinsic pontine gliomas and non-brainstem glioblastomas. Nat Genet 44(3):251–253. https://doi.org/10.1038/ng.1102
Yang YS, Yang MC, Weissler JC (2011) Pleiomorphic adenoma gene-like 2 expression is associated with the development of lung adenocarcinoma and emphysema. Lung Cancer 74(1):12–24. https://doi.org/10.1016/j.lungcan.2011.02.006
Yao HP, Zhao H, Hudson R, Tong XM, Wang MH (2021) Duocarmycin-based antibody-drug conjugates as an emerging biotherapeutic entity for targeted cancer therapy: Pharmaceutical strategy and clinical progress. Drug Discov Today 26(8):1857–1874. https://doi.org/10.1016/j.drudis.2021.06.012
Zheng H, Ying H, Wiedemeyer R, Yan H, Quayle SN, Ivanova EV et al (2010) PLAGL2 regulates Wnt signaling to impede differentiation in neural stem cells and gliomas. Cancer Cell 17(5):497–509. https://doi.org/10.1016/j.ccr.2010.03.020
Acknowledgements
We thank the German Cancer Research Centre (DKFZ) Genomics and Proteomics Core Facility and Laura Dörner, Katharina Müller, Lea Hofmann (Department of Neuropathology, Institute of Pathology at the University Hospital Heidelberg). This work was supported by the German Childhood Cancer Foundation (“Neuropath 2.0—Increasing diagnostic accuracy in pediatric neurooncology”; DKS 2015.01), the Everest Centre for Low-Grade Paediatric Brain Tumour Research (The Brain Tumour Charity, UK; GN-000382), the Brain Tumor Charity (UK), grant number QfC_2019/1_10507, the German Federal Ministry of Education and Research (BMBF), and Cancéropôle Lyon Auvergne Rhône-Alpes (CLARA). A subset of the human tissue was obtained from the CRB-Cancer, CHU de Bordeaux. A subset of the human tissue was obtained from University College London NHS Foundation Trust as part of the UK Brain Archive Information Network (BRAIN UK, Ref: 19/001) which is funded by the Medical Research Council and Brain Tumour Research UK. S. Brandner was also supported by the Department of Health’s NIHR Biomedical Research Centre's funding scheme to University College London Hospitals. Ulrich Schüller was supported by the Fördergemeinschaft Kinderkrebszentrum Hamburg, Germany. Pediatric brain tumor research in the Solomon Lab is supported by the Morgan Adams Foundation, the Yuvaan Tiwari Foundation, and a Developmental Research Program award from the UCSF Brain Tumor SPORE grant from the National Cancer Institute, National Institutes of Health (P50 CA097257). DNA methylation profiling at NYU was in part supported by grants from the Friedberg Charitable Foundation, the Sohn Conference Foundation and the Making Headway Foundation (to M. Snuderl.). Pediatric brain tumor research at Karolinska Institutet as well as The Swedish Childhood Tumor Biobank (Karolinska Institutet, Stockholm, Sweden) were supported by the Swedish Childhood Cancer Fund. The molecular biology lab at the Olgahospital/Klinikum Stuttgart (Stuttgart, Germany) was supported by the Förderkreis Krebskranke Kinder e.V Stuttgart.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Conflict of interest
Stephen Yip is a member of advisory boards and has received honorarium from Amgen, AstraZeneca, Bayer, Incyte, and Roche. P. Wesseling, U. Schüller, A. Perry, F. Sahm, and D.A. Solomon are members of the Editorial Board of Acta Neuropathologica. They were not involved in the assessment or decision making for this manuscript.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
The original online version of this article was revised: Figure 6 in the original version of this article has been replaced.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Keck, MK., Sill, M., Wittmann, A. et al. Amplification of the PLAG-family genes—PLAGL1 and PLAGL2—is a key feature of the novel tumor type CNS embryonal tumor with PLAGL amplification. Acta Neuropathol 145, 49–69 (2023). https://doi.org/10.1007/s00401-022-02516-2
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00401-022-02516-2