Abstract
Main conclusion
During re-establishment of desiccation tolerance (DT), early events promote initial protection and growth arrest, while late events promote stress adaptation and contribute to survival in the dry state.
Mature seeds of Arabidopsis thaliana are desiccation tolerant, but they lose desiccation tolerance (DT) while progressing to germination. Yet, there is a small developmental window during which DT can be rescued by treatment with abscisic acid (ABA). To gain temporal resolution and identify relevant genes in this process, data from a time series of microarrays were used to build a gene co-expression network. The network has two regions, namely early response (ER) and late response (LR). Genes in the ER region are related to biological processes, such as dormancy, acquisition of DT and drought, amplification of signals, growth arrest and induction of protection mechanisms (such as LEA proteins). Genes in the LR region lead to inhibition of photosynthesis and primary metabolism, promote adaptation to stress conditions and contribute to seed longevity. Phenotyping of 12 hubs in relation to re-establishment of DT with T-DNA insertion lines indicated a significant increase in the ability to re-establish DT compared with the wild-type in the lines cbsx4, at3g53040 and at4g25580, suggesting the operation of redundant and compensatory mechanisms. Moreover, we show that re-establishment of DT by polyethylene glycol and ABA occurs through partially overlapping mechanisms. Our data confirm that co-expression network analysis is a valid approach to examine data from time series of transcriptome analysis, as it provides promising insights into biologically relevant relations that help to generate new information about the roles of certain genes for DT.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
During seed development, orthodox seeds acquire the remarkable ability to tolerate desiccation. This means that during development, as a part of the maturation phase, these seeds experience slow reduction of their water content to less than 5 % of their dry weight with little or no loss of viability (Ooms et al. 1993). In this dry state, orthodox seeds can survive for years or centuries, which permit their storage and ensure better dispersal (Ramanjulu and Bartels 2002). During seed imbibition and germination, increasing water availability allows metabolic processes to resume, eventually leading to the emergence of the radicle and the associated progressive loss of desiccation tolerance (DT) (Buitink et al. 2003, 2006; Maia et al. 2011). Yet, after radicle protrusion, there is a small developmental window during which DT can be rescued by treatment with an osmoticum (polyethylene glycol, PEG) and/or the plant hormone abscisic acid (ABA) (Buitink et al. 2003; Maia et al. 2011, 2014).
Incubation in PEG induces membrane changes, inhibits radicle growth, down-regulates genes related to energy metabolism and cell wall modification, up-regulates genes related to antioxidant activity, response to stress and seed storage, and induces synthesis of protective molecules, such as non-reducing sugars and certain proteins, such as late embryogenesis abundant (LEA) and heat shock proteins (HSPs) (Buitink et al. 2003, 2006; Maia et al. 2011). Furthermore, genes coding for ABA signal transduction elements and drought/stress-induced transcription factors (TFs) are also up-regulated, leading to the hypothesis of a partial overlap of ABA-dependent and ABA-independent regulatory pathways involved in both drought and DT (Buitink et al. 2003, 2006; Maia et al. 2011). Mutants compromised in ABA sensitivity or synthesis were shown to be still able to produce desiccation-tolerant seeds that, however, are impaired in re-establishment of DT during germination (Maia et al. 2014). Moreover, ABA perception and signaling, more than ABA content, are influencing the ability to re-establish DT (Maia et al. 2014). In previous studies, the re-establishment of DT was brought about by the simultaneous application of cold and osmotic stress (Buitink et al. 2003), or osmotic stress and/or ABA (Maia et al. 2011, 2014). Although treatment of germinated seeds with ABA alone is able to re-establish DT (Maia et al. 2014), the regulatory and mechanistic pathways activated via ABA remain to be elucidated.
Here, we used microarrays to characterize global gene expression during the re-establishment of DT in germinated A. thaliana seeds by ABA, and used the data for network analysis of gene co-expression, to gain more resolution on and identify relevant genes in this process. Gene-expression data derived from microarray analysis have been commonly used to provide insight into biological processes at a system-wide level (Wang et al. 2006; Freeman et al. 2007) and is well suited to reconstruct gene regulatory networks to explore potential biological relations (Wang et al. 2006). These networks consist of nodes that represent genes connected by edges that infer co-expression based on a correlation threshold (Freeman et al. 2007). The analysis of this kind of networks is based on the assumption that genes with similar expression patterns have similar functions (Freeman et al. 2007).
In this study, the construction and analysis of a co-expression network proved to be a valid approach to examine data from microarrays of a time series of DT re-establishment by ABA. Using a combination of physiology and transcriptomic approaches, we generated and partially validated a co-expression network, which revealed two major patterns of gene expression during the re-establishment of DT by ABA: an early and a late response. Genes could be divided into those that respond earlier to the incubation in ABA and provide initial protection and signal transduction, and those that respond later and provide adaptation to the stress condition. Validation of the network by functional characterization of genes identified based on the network analysis revealed important roles of some of these genes in the re-establishment of DT in germinated seeds.
Materials and methods
Plant growth conditions and germination assays
Arabidopsis thaliana plants, accession Columbia (Col-0, N60000), were grown on Rockwool plugs (MM40/40; Grodan), in a climate cell (20 °C day, 18 °C night), under 16 h of light, and watered with Hyponex nutrient solution (1 g l−1, http://www.hyponex.co.jp). Seeds were bulk harvested in three replicates of at least two plants. Seeds used in germination assays were cold stratified for 72 h at 4 °C in 9-cm Petri dishes on two layers of blue filter paper (Blue Blotter Paper, Anchor Paper Company, http://www.seedpaper.com) and 10 ml of distilled water. After stratification, seeds were transferred to germination cabinets with constant white light at 22 °C.
Re-establishment of DT using ABA
To assess the re-establishment of DT using ABA, germinated seeds at the stage of radicle protrusion (stage II, Maia et al. 2011) were incubated for a maximum of 3 days in 6-cm Petri dishes containing 1.3 ml of an ABA solution (10 µM) on two sheets of white filter paper (grade 3hw, Biolab Products, Sartorius Stedim Biotec) in the dark at 20 °C. The incubation in ABA was done in the dark to reduce oxidative damage. After incubation, seeds were rinsed in distilled water, transferred to new Petri dishes with one dry sheet of white filter paper and dried for 3 days at 40 % relative humidity (RH) at 20 °C, resulting in water content levels as low as 0.08 g H2O g−1 dry weight. After drying, seeds were pre-humidified in air of 100 % RH for 24 h at 22 °C in the dark and subsequently rehydrated in water on a Copenhagen Table under a 12/12 h dark/light regime at 20 °C. Germinated seeds were evaluated according to the survival of their primary root, presence of green and fully expanded cotyledons (cotyledon survival) and growth resumption with both green and fully expanded cotyledons and development of a root system (seedling survival).
Longevity of germinated seeds
Longevity of germinated seeds was evaluated by an accelerated aging test after incubation in ABA for 24 h and 72 h, drying for 3 days at 40 % RH, storage for 24 h at 80 % RH and 40 °C in the dark and rehydration. The parameters evaluated were survival of the primary root, cotyledons survival and seedling survival.
RNA extraction and microarray hybridization
Seeds at the stage of radicle protrusion (control) and these seeds after four periods (2, 12, 24 and 72 h) of incubation in ABA were used for RNA extraction. Total RNA was extracted from three replicates of approximately 1000 germinated seeds for each time point following a modified hot borate protocol (Wan and Wilkins 1994; Maia et al. 2011). The seeds were ground and mixed with 800 µl of extraction buffer (0.2 N Na borate decahydrate (Borax), 30 mM EGTA, 1 % SDS, 1 % Na deoxycolate) containing 1.76 mg DTT and 52.8 mg PVP40, and heated to 80 °C. In the next step, 4 mg proteinase K was added to this solution before incubation for 15 min at 42 °C. After the addition of 64 µl of 2 M KCL, the samples were incubated on ice for 30 min and subsequently centrifuged for 20 min at 12,000g. The supernatant was transferred to a new tube, 260 µl of ice-cold 8 M LiCl was added, and the tubes were incubated overnight on ice. After centrifugation at 4 °C for 20 min at 12,000g, the pellets were washed with 750 ml of ice-cold 2 M LiCl and re-suspended in 100 µl milliQ water. The samples were phenol–chloroform extracted, DNAse treated (RQ1 DNase, Promega) and further purified with RNeasy spin columns (Qiagen) according to the manufacturer’s instructions. RNA quality and concentration were assessed by agarose gel electrophoresis and a NanoDrop ND-1000 spectophotometer (Nanodrop® Technologies, Wilmington, DE, USA). RNA was processed for the use on Affymetrix ARAGene 1.1ST Arrays as described by the manufacturer. In brief, reverse transcription was employed to generate double-stranded cDNA that was in vitro transcribed to biotinylated cRNA. The biotinylated cRNA was used for hybridization. The Affymetrix HWS Kit for GeneTitan (part nr. 901622) was used for hybridization, washing and staining of the array plates. The array plates were scanned using the Affymetrix Command Console v3.2 software.
All data are MIAME compliant as detailed on the MGED Society website http://www.mged.org/Workgroups/MIAME/miame.html. The microarray data have been deposited on the NCBI’s Gene Expression Omnibus (Edgar et al. 2002) and is accessible through the GEO Series accession number GSE62876.
Microarray analysis
Signal intensities were extracted and analyzed using the Bioconductor packages of R (Gentleman et al. 2004). The data were normalized using the RMA algorithm (Irizarry et al. 2003) with the TAIRG v17 cdf file (http://www.brainarray.mbni.med.umich.edu).
To equalize background noise, gene expression values less than four were replaced with four (Dekkers et al. 2013). After this transformation, fold changes were calculated comparing each time-point with the control (germinated seeds at the stage of radicle protrusion non-treated with ABA). A gene was considered differentially expressed (the differentially expressed genes: DEGs) if the difference between its mean expression in at least one time-point and the control was statistically significant at P ≤ 0.01 after application of linear modeling with thresholds for absolute fold change of 2.0 (on a log2 scale).
The resulting gene set was used for an over-representation analysis (ORA) to recover over-represented biological processes (using Bonferroni-corrected P value at 0.05) based on gene ontologies using Gene Trail (Keller et al. 2008). The term’s semantic distance with respect to other semantically close terms (“Dispensability”) was calculated using the online tool ReviGO (Supek et al. 2011) and redundant gene ontology (GO) terms were removed applying a cut-off of <0.1 for this “Dispensability” value.
Network construction and analysis
Pearson correlation coefficients between all pairs of DEGs were calculated. A table with correlation coefficient values was exported to Cytoscape v.2.8.2 (Smoot et al. 2011) and correlation coefficients above a threshold of 0.97 (determined according to Freeman et al. 2007) were used to filter the connections that were used to determine the edges between nodes in the network. The resulting network was displayed with a yGraph Organic layout. The Cytoscape plug-in NetworkAnalyzer (Assenov et al. 2008) was used to compute node degree. The 100 nodes with the highest degrees were considered “hubs”.
Self-organizing maps (SOMs) were calculated using GeneMaths software (version 2.1, Applied Maths BVBA, Sint-Martens-Latem, Belgium) by importing gene expression data and mapping them into six groups (2 × 3 node format) that provided optimal representation of gene expression patterns in a reasonably small number of independent bins.
Mutant analysis
T-DNA insertion lines (Suppl. Table S1; Supp. Fig. S1) for selected genes were obtained from the Nottingham Arabidopsis Stock Centre (Scholl et al. 2000). Plants with homozygous T-DNA insertions were selected and confirmed using PCR and grown as described above for wild-type plants. Seeds were bulk-harvested in three replicates of at least two plants. Germinated seeds at the stage of appearance of first root hairs (stage IV, Maia et al. 2011) were phenotyped to evaluate the ability to re-establish DT after treatment with PEG as described by Maia et al. (2011). The evaluated parameters were survival of their primary root, cotyledon survival and seedling survival.
Seeds were also phenotyped for seed dormancy, longevity, and vigor. Seed dormancy was evaluated as the number of days of Seed Dry Storage required to reach 50 % germination (DSDS50) (Alonso-Blanco et al. 2003). Seed longevity was estimated based on an accelerated aging assay (germination percentage after storage for 6 days at 80 % RH and 40 °C in the dark) (Bentsink et al. 2000). Seed vigor was measured as the ability of seeds to germinate at high temperature (at 33 °C) or on NaCl (130 mM). Germination experiments and scoring of germination were performed with the GERMINATOR as described by Joosen et al. (2010) in a fully randomized setup. For each measurement, three replicates of 40–60 seeds per seed batch were used.
Results
Re-establishment of DT in germinated A. thaliana seeds
Mature A. thaliana seeds are desiccation tolerant and imbibition and progression into germination change their status to desiccation sensitive already at the phase of testa rupture (Maia et al. 2011). However, a treatment with ABA fully rescues DT in germinated seeds till the phase of radicle protrusion (Maia et al. 2014). In subsequent developmental stages, DT can no longer be rescued in all organs (Maia et al. 2014).
After 12 h, the treatment with ABA had already led to the re-establishment of DT in 80 % of germinated A. thaliana seeds at the stage of radicle protrusion (Fig. 1). After 24 h, all the seeds had re-established DT. To further assess the capacity of survival in the dry state, storability was determined by an accelerated aging test. Survival rates after the accelerated aging test were lower for seeds incubated in ABA for 24 h than for seeds incubated in ABA for 72 h (Fig. 2).
Longevity of germinated A. thaliana seeds at the stage of radicle protrusion incubated in ABA for 24 or 72 h. Longevity was estimated as germination percentage after aging (24 h of storage at 80 % RH and 40 °C). Control germinated seeds were incubated in ABA, but not aged. Asterisks indicate significant differences at P ≤ 0.05 for one, P ≤ 0.01 for two asterisks comparing 24 h and 72 of incubation in ABA for each parameter in control and aged seeds separated
To gain temporal resolution of the changes in gene expression induced by incubation in ABA, gene expression analysis was performed on a time series. Based on the results shown in Fig. 1, five time-points were chosen for microarray analysis: 0 (control), 2, 12, 24 and 72 h of incubation in ABA.
Principal component analysis was used to compare global changes between the transcriptomes of the different time-points and to evaluate group clustering. One replicate of time-point 2 h was a statistically significant outlier (data not shown) and was removed from further analysis. Without this replicate, the first principal component described 61.2 % of the variation (Fig. 3).
In total, 1177 genes were considered differentially expressed in response to the ABA treatment for at least one time-point as compared to the control. The number of DEGs was the lowest after 2 h of incubation and increased until 24 h, after which no substantial changes occurred until 72 h of incubation (Table 1). To verify the accuracy of the microarray data, the expression of 20 genes with different expression patterns were also analyzed by qPCR. Both microarray data and qPCR analysis showed comparable trends (Suppl. Fig. S2).
Over-representation analysis (ORA) of functional GO categories
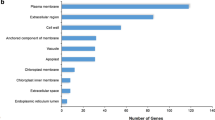
ORA was used to get an overview of the enriched functional GO categories comparing the expression data of genes from each time-point relative to the control. Among the genes with accumulating transcript levels, the category lipid storage was highly overrepresented after 2 h of incubation in ABA but, over time, this category became less overrepresented (Fig. 4). The category dormancy process is over-represented in the genes with accumulating transcript levels only after 24 h of incubation, when DT was fully re-established. Within the genes represented by declining transcript levels, categories related to cell wall (wax biosynthetic process, cell wall organization or biogenesis, plant-type cell wall organization, and syncytium formation) were over-represented. Genes in these categories are mainly involved in depolymerisation of cell wall components during the beginning of storage mobilization (Sreenivasulu et al. 2008). Categories related to photosynthesis and metabolism (chlorophyll metabolic process, generation of precursor metabolites and energy, and photosynthesis) were also over-represented within the DEGs represented by declining transcript levels after 12 and 24 h, when 80 and 100 % of the seeds had re-established DT, respectively. Overall, this analysis revealed significant accumulation of transcripts of genes involved in protection, response to stimulus, seed development, and seed dormancy and decline of transcripts of genes related to cell growth, photosynthesis and response to stimulus (Fig. 4).
Heat-map of ORA of biological processes on differences between different time-points during incubation of germinated A. thaliana seeds in ABA relative to the control (non-treated with ABA). Functional classes were determined using Gene Trail and ReviGO. P values were adjusted for multiple testing by the Bonferroni method applying a cut-off of <0.05. a Results for genes with accumulating transcript levels in each time-point compared to the control. b Results for genes with declining transcript levels in each time-point compared to the control. Shades of gray indicate the extent (ratio of expected to observed number of genes) to which the category was overrepresented in each comparison
Gene co-expression network
Transcriptional studies generate vast amounts of gene expression data. This type of data can be used to build co-expression networks which help to study coordinated gene expression and to identify key genes, functional modules or relations between the network structure, and additional information (Mutwil et al. 2010; Villa-Vialaneix et al. 2013). Using Cytoscape, we constructed a gene co-expression network from the correlation coefficients calculated between all pairs of DEGs. The DEGs are represented as nodes connected by edges that model significant correlation coefficients (Villa-Vialaneix et al. 2013). The network was built from 1083 genes/nodes (Suppl. Table S2) completely connected by 35,296 edges, meaning that any node could be reached from any other node by a path passing along the edges (Villa-Vialaneix et al. 2013) and visualized using an organic layout in Cytoscape (Fig. 5a).
Gene co-expression network (a) and SOM (b) of transcriptomes of germinated A. thaliana seeds incubated in ABA for different intervals. The network is visualized using an yGraph Organic layout in Cytoscape and the temporal analysis of modules was obtained by coloring each node by the profile specific for the SOM groups. The dashed line and the arrows in the network indicate the two regions, namely early response and late response. For the SOM, the probe sets were grouped into six clustered patterns. Horizontal bars at the top of each graph represent the number of probe sets belonging to a certain SOM bin. The SOM bins having the highest number of genes have the largest bar. Vertical bars represent standard deviation in average expression of the genes in each time point per group. The three color shades in each graph ranging from dark to light correspond to the 98th, 90th and 75th percentiles, respectively
SOMs can be used to categorize gene expression data into groups that show similar gene expression profiles and thereby may also contain functionally related genes (Törönen et al. 1999). We colored each node of the co-expression network by the profile group according to the SOM (Fig. 5b). In this way, two main regions of highly interconnected transcripts were identified: early response (ER) and late response (LR). The ER region is the most tightly co-regulated one with nodes representing genes with a sharp increase or decrease in expression after 2 h of incubation in ABA. This region concentrates the hubs of the network, meaning that it contains all 100 nodes with the highest degrees (number of nodes directly connected to a given node, ranging from 165 to 209). The LR region contains genes with slow increase or decrease in expression during incubation in ABA.
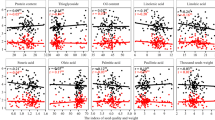
To assess the cross-link between re-establishment of DT with ABA and seed and stress-related processes, we projected sets of DEGs on the network comparing: dormancy (dormant seeds vs. after-ripened seeds; Cadman et al. 2006, Fig. 6a); DT acquisition (the cotyledon stage vs. the post mature-green stage of seed development; Le et al. 2010, Fig. 6b); re-establishment of DT with PEG [germinated seeds at the stage of radicle protrusion before (desiccation sensitive) vs. after (desiccation tolerant) treatment with PEG; Maia et al. 2011, Fig. 6c]; and drought (control plants vs. plants subjected to drought for 10 h; Matsui et al. 2008, Fig. 6d). Genes with increased transcript levels in these comparisons locate mainly in the ER region of the network, while genes with decreased transcript levels locate mainly in the LR region. Moreover, we found an overlap of 16 genes differentially expressed in relation to all these physiological processes (Table 2).
Gene co-expression network of incubation in ABA of germinated A. thaliana seeds. Nodes are colored according to genes with accumulating (purple) or declining (green) transcript levels in different datasets comparing: a Dormancy (dormant seeds vs. after-ripened seeds; Cadman et al. 2006). b DT acquisition (cotyledon stage vs. post mature-green stage of seed development; Le et al. 2010). c Re-establishment of DT with PEG (germinated seeds in the stage of radicle protrusion before (desiccation sensitive) vs. after (desiccation tolerant) treatment with PEG; Maia et al. 2011). d Drought (control plants vs. plants dried for 10 h; Matsui et al. 2008). The dashed line separates the early response (left) and the late response (right) region
After the projection on the network of the DEGs in the re-establishment of DT with PEG, 84 % of the nodes were highlighted, with a clear overlap between genes with increasing and decreasing transcript levels in both processes (re-establishment of DT by ABA and PEG) (Fig. 6c). A comparison between over-represented GO categories in DEGs with accumulating transcript levels in response to incubation in ABA and in PEG revealed a large overlap in categories related to response to stimulus and seed development (Maia et al. 2011). However, the categories circadian rhythm, dormancy process and macromolecule localization were over-represented only in response to incubation in ABA, while categories related to respiration were over-represented only in response to incubation in PEG. ABA regulates the expression of genes related to circadian clock through TFs such as ABI5, AP2/ERF and NAC (Fujita et al. 2011), and the induction and maintenance of dormancy (Finch-Savage and Leubner-Metzger 2006). The osmotic stress and a possible hypoxia caused by the incubation in PEG may have induced the expression of genes related to respiration, such as AT1G19530, AT1G33055, AT3G10020, ADH1, leading to the over-representation of the categories anaerobic respiration and cellular respiration (Maia et al. 2011). Considering DEGs with declining transcripts, GO categories related to cell wall, photosynthesis and response to stimulus were over-represented in response to both incubation in ABA and in PEG (Maia et al. 2011). The categories heterocycle biosynthetic process and syncytium were over-represented only in response to incubation in ABA, while the categories fatty acid metabolic process and sulfur metabolic process were over-represented only in response to incubation in PEG. The over-representation of these categories is an indication that incubation in either ABA or PEG leads to growth arrest by affecting different genes. Incubation in ABA reduces abundance of transcripts related to expansins (ATEXPA1, ATEXPA8 and ATEXPA 10), extensins (AT2G43150, AT3G28550 and AT3G54580) and certain photosynthesis-related genes (APT3, APT5, GUN4, GUN5, HEMA1 and PORB) that are not significantly affected by PEG treatment, while PEG reduced the abundance of gene transcripts related to fatty-acid metabolism (AT5G10160, ACP4, HCD1, OPR1 and OPR2) and related to the synthesis of compounds that contain sulfur (ATGSTU27, CYP79B2, CYP83B1, MAT3 and MTO3), such as the amino acid cysteine.
Considering that in plant genomes many TFs are master regulators of signaling and regulatory pathways of stress adaptation and act in an ABA-dependent manner (Lindemose et al. 2013), we searched for the presence of known and predicted TFs as listed in the database of Arabidopsis transcription factors (DATF, http://www.datf.cbi.pku.edu.cn, Guo et al. 2005) in our network. From the 2290 TFs gene model identifiers available in the DATF, we found 69 in our network (Fig. 7a). Of these, 46 are located in the ER region and are mainly related to response and tolerance to abiotic stress, such as Abscisic acid insensitive 5 (ABI5), members of the APETALA 2/ethylene-responsive element-binding factor (AP2/ERF) family (AT5G18450, AT5G51190 and ERF5), members of the NAM/ATF1/CUC2 (NAC) class (NAC032, NAC048, NAC053, NAC060, NAC083 and NAC089) and WRKY TFs (WRKY 18, WRKY29 and WRKY36). TFs located in the LR region are mainly related to abiotic stress tolerance, growth regulation and light signaling pathways, such as ABRE-binding factor 1 (ABF1), enhanced Em levels (EEL), LATERAL ORGAN BOUNDARY DOMAIN 41 (LBD41), LATE ELONGATED HYPOCOTYL (LHY), PHYTOCHROME-INTERACTING FACTOR-LIKE 2 (PIL2), PHYTOCHROME INTERACTING FACTOR 4 (PIF4), PSEUDO-RESPONSE REGULATORS (PRR5), TIMING OF CAB EXPRESSION 1 (TOC1) and WUSCHEL RELATED HOMEOBOX 12 (WOX12).
Gene co-expression network of germinated A. thaliana seeds incubated in ABA. Nodes are colored according to genes encoding transcription factors (a), members of the ABI3 regulon (b), and LEA proteins (c). The dashed line separates the early response (left) and the late response (right) region of the network
The TF ABI3 is a key regulator of seed development and maturation that, together with ABA, plays an essential role in the protection of embryonic structures from desiccation (Mönke et al. 2012). Targets of ABI3 include genes with a protective role in DT, such as LEA genes. We projected the set of target genes from the ABI3 regulon (Mönke et al. 2012) (Fig. 7b) and the set of genes from the inventory of all LEA proteins identified in the A. thaliana genome on our network (Bies-Ethève et al. 2008) (Fig. 7c). Most of the genes representing the ABI3 regulon and LEA proteins are concentrated in the ER region of the network. Within the hubs identified in the network, genes of the ABI3 regulon and LEA genes are significantly (P value <0.01) enriched with 44 and 19 genes respectively.
We selected 12 hub genes that are members of the ABI3 regulon, six of which are LEA genes, and investigated their function in the re-establishment of DT and seed-related traits such as dormancy, longevity and vigor (Suppl. Table S3). These genes were chosen based on the indications of their involvement in DT from ongoing studies in our laboratory. The evaluation of seed-related traits is important since the acquisition of DT by seeds requires a series of mechanisms that are also necessary for seed survival in the dry state (Kalemba and Pukacka 2011; Delahaie et al. 2013) and the disruption of one or more of these mechanisms can impact other aspects important for the completion of germination. We characterized T-DNA insertion lines for each of the aforementioned 12 genes. ECP63, AT2G42560, AT3G15670, AT3G53040, AT4G36600, and AT5G44310 are the ones that are both members of the ABI3 regulon and the LEA gene family. The remaining six (AT1G27990, CBSX4, RTNLB13, AT2G25890, AT3G54940, and AT4G25580) are members of the ABI3 regulon that are not LEA genes: AT1G27990 encodes an unknown protein; CBSX4 encodes a cystathionine-β-synthase (CBS) domain-containing protein involved in reproduction and is highly expressed in dry seeds (Fang et al. 2011); RTNLB13 codes for a plant reticulon localized in the tubular endoplasmic reticulum and is predicted to be expressed in seeds (Sparkes et al. 2010); AT2G25890 codes for a sugar-regulated oleosin (Huang et al. 2010); AT3G54940 encodes a papain-like proteinase up-regulated in senescing siliques (Trobacher et al. 2006) and AT4G25580 encodes a protein related with response to stress and is highly similar to a CAP160 protein linked with cold acclimation (Kaye et al. 1998).
For the phenotypic characterization in relation to re-establishment of DT, germinated seeds in the stage of appearance of first root hairs were used instead of germinated seeds in the stage of radicle protrusion. At the stage of radicle protrusion, germinated seeds of all the lines, including the wild-type, were able to re-establish DT in more than 90 % of germinated seeds (data not shown). Therefore, no differences were detected between them. On the other hand, at the stage of appearance of first root hairs, germinated seeds were not able to fully re-establish DT, allowing a clearer separation between wild-type seeds and seeds from T-DNA insertion lines. During evaluation of the survival, frequently, when the primary root fails to survive drying, the hypocotyl remains able to generate lateral roots, allowing normal seedling formation. Therefore, the combined evaluation of root survival, cotyledon survival and seedling formation provides a complete representation of the phenotypic influence of the T-DNA insertion on the re-establishment of DT. Three lines had a significant increase in the ability to re-establish DT compared to the wild-type: cbsx4, at3g53040 and at4g25580 (Fig. 8; Table 3). Three different lines had a decrease in seed vigor, with at1g27990 being affected in high temperature seed vigor, and at3g54940 and at5g44310 showing severely reduced germination under salt stress (Table 3). The genes disrupted in these six lines had a fast increase in transcript level (expression level increase of more than fourfold) already after 2 h of incubation in ABA and remained highly expressed but without significant changes until the last time-point. None of the lines showed phenotypes for seed dormancy and longevity (Suppl. Table S3).
Re-establishment of desiccation tolerance in germinated seeds of wild type (WT) and T-DNA insertion lines scored as survival of primary root (root survival) and cotyledons and seedling formation. Bars represent standard error of three replicates. Asterisks indicate significant differences at P ≤ 0.05 for one and P ≤ 0.01 for two asterisks
Discussion
Studies focusing on re-establishment of DT often use PEG alone or in combination with low temperature or ABA (Buitink et al. 2003; Vieira et al. 2010; Maia et al. 2011, 2014). In each case, seeds are exposed to a mild osmotic stress that often triggers a series of ABA-related responses (Huang et al. 2008; Matsui et al. 2008; Dalal et al. 2009). To evaluate the responses to ABA alone, without additional stressors, we studied germinated A. thaliana seeds at the stage of radicle protrusion during incubation in ABA. In this stage, DT can be re-established in all the seeds incubated in ABA already after 24 h of incubation (Fig. 1), while approximately 85 % of seeds incubated in PEG for 24 h had re-established DT (Maia et al. 2011). DT re-established in all the seeds by incubation in PEG is only obtained after 72 h of incubation (Maia et al. 2011).
In general, osmotic stresses trigger a series of physiological responses in ABA-dependent and ABA-independent manners with the existence of a cross-talk between them (Buitink et al. 2003; Yamaguchi-Shinozaki and Shinozaki 2005). A comparison between over-represented GO categories in DEGs with accumulating transcripts in response to incubation in ABA and in PEG reinforced the idea that these treatments lead to the re-establishment of DT through partially overlapping mechanisms.
A closer look at the main processes that occur during incubation in ABA revealed enrichment of different GO categories of biological processes (Fig. 4), attesting the growth arrest and the partial return to a quiescent stage similar to the dry seed (Buitink et al. 2006; Maia et al. 2011).
Lipid storage was the most over-represented GO category within the genes with accumulating transcript levels and the genes inside this category are oleosins and oleosin family proteins. Oleosins are typical for the later stages of seed development and, in desiccation tolerant seeds, they prevent lipid bodies from coalescing on dehydration and prevent the disruption of cellular structures during rehydration (Leprince et al. 1998; Pammenter and Berjak 1999).
Despite the fact that all the germinated seeds incubated in ABA had re-established DT after 24 h, the incubation was maintained for a total of 72 h, which improves longevity compared with incubation for 24 h (Fig. 2). Also during development, acquisition of DT precedes acquisition of longevity, likely to ensure timely response of stress-related genes (such as LEAs, HSPs and oxidative stress-related genes) that also improve seed storability (Verdier et al. 2013).
Transcriptomic studies generate large datasets. Visualization and analysis of this kind of data as networks is an important approach to explore a wide variety of biological relations (Freeman et al. 2007; Dekkers et al. 2013). To construct a network, the similarity between individual expression profiles may be determined and used as edges that connect nodes, or genes (Freeman et al. 2007). Once the network is given, an analysis of its structure can indicate key genes, functional modules or relations between the network structure and additional information (Villa-Vialaneix et al. 2013). Several centrality measures or topological indexes have been used to analyze networks. The degree centrality or just degree is thought to be simplest (Scardoni et al. 2009; Piraveenan et al. 2013; Bass et al. 2013). It refers to the number of links a given node has with other nodes and allows an immediate evaluation of the relevance of the node to the network (Scardoni et al. 2009; Piraveenan et al. 2013). Nodes with high degrees are called hubs and are thought to play important roles in organizing the behavior of the network (Dong and Horvath 2007; Bass et al. 2013; Villa-Vialaneix et al. 2013).
The ER region of the network contains genes related mainly to wax biosynthetic processes, lipid storage, seed development and response to abscisic acid stimulus. Most of the genes with accumulating transcript levels in relation with dormancy, DT acquisition, re-establishment of DT with PEG, and drought also locate in this region (Fig. 6). Also, all the hubs and most of the members of the ABI3 regulon, LEA proteins and TFs found in the network are in this region (Fig. 7). ABA is a key component of responses to abiotic stresses and regulation of seed dormancy and germination. The responses to abiotic stresses elicited by ABA include minimization of water transpiration (for example, by the accumulation of cuticular wax), and synthesis of protectants (such as LEA proteins) and antioxidants (Dalal et al. 2009; Seo et al. 2011). During seed development and germination, several genes under the control of ABA are related to the acquisition and loss of dormancy and DT (Toorop et al. 2005; Buitink et al. 2006; Maia et al. 2011; Terrasson et al. 2013). Moreover, ABA-responsive gene expression is directly regulated by TFs (Fujita et al. 2011). The TFs in the ER region of the network, such as ABI5, AP2/ERF TFs and NAC TFs, are known to be related with response and tolerance to abiotic stress possibly by a rapid amplification and broadening of signal responses (Fujita et al. 2011). Taken together, these observations reinforce the relation between early stress response and seed development and the view that early responsive genes may provide initial protection and amplification of signals (Buitink et al. 2006).
The LR region contains genes related mainly with response to abiotic stimulus (especially light stimulus), and aromatic amino acid family metabolic processes. The GO category aromatic amino acid family metabolic processes refer to chemical reactions and pathways involving amino acids with an aromatic ring, such as phenylalanine, tyrosine, and tryptophan, which could be involved in protein stability (Khuri et al. 2001; Carbon et al. 2009). TFs located in this region are related to abiotic stress tolerance, growth regulation and light signaling pathways and might influence the plant’s ability to adapt to daily changes in water status in a coordinated action with the circadian clock (Alabadí et al. 2001; Fujita et al. 2011). These results are in agreement with the hypothesis that genes which are responding later may be involved in adaptation to stress conditions (Buitink et al. 2006).
We considered the 100 nodes with the highest degrees as hubs. They are all concentrated in the ER region and most of them follow the expression pattern corresponding to the 5th clustered pattern of the SOM analysis (fast increase in expression in the first 2 h of incubation in ABA followed by certain stabilization in the following hours).
Three T-DNA insertion lines showed a higher ability to re-establish DT compared to wild-type, suggesting the existence of de-repression mechanisms and redundancy between highly similar genes. These lines are cbsx4, at3g53040 and at4g25580. CBSX4 is one of the two A. thaliana homologous genes (the other is AT1G15330) that encode CBS domain-containing proteins that belongs to the PV42 class of gamma subunits of SnRK1 (Fang et al. 2011). CBS domain-containing proteins have been found to act in a variety of biological processes, such as metabolic enzymes, transcriptional regulators, ion channels, and transporters (Rosnoblet et al. 2007; Fang et al. 2011). A. thaliana mutants in HISTONE ACETYLTRANSFERASE1 (HAC1) had reduced expression of both AT1G15330 and CBSX4, suggesting a role of these genes in sugar sensing and fertility (Heisel et al. 2013). Phenotyping a T-DNA insertion line for AT1G15330 did not show any phenotypes (data not shown). The observed phenotype of higher ability to re-establish DT compared to wild-type seeds could be due to compensatory effects of one homologous gene over the other. Interestingly, cbsx4 had an increased expression of the mutated gene compared to the wild-type in a RT-PCR experiment. We speculate that the amplification of this gene generates a product with a non-functional sub-unit, as the presence and location of the insertion in the last exon of the gene was confirmed.
AT3G53040 codes for a cytosolic LEA domain-containing protein that shares typical features of the LEAM protein class A α-helix motifs, being classified as LEA_4 family (Candat et al. 2014). Proteins of this family are likely to interact and protect various cellular membranes during dehydration (Candat et al. 2014). In A. thaliana seedlings, the expression of AT3G53040 was shown to respond to ABA, but without inducible expression to stress conditions (Huang and Wu 2006; Bies-Ethève et al. 2008). AT3G53040 and AT2G36640 are very similar to each other in sequence and expression pattern and are considered to form a pair derived from a whole genome duplication (Hundertmark and Hincha 2008; Bies-Ethève et al. 2008). It is expected that duplicated genes have redundant functions and the higher ability to re-establish DT observed in at3g53040 compared to wild-type could be caused by this redundancy and compensatory effects. It is likely that the phenotyping of a T-DNA insertion line of AT2G36640 did not result in differences compared to the wild-type (data not shown) due to this redundancy.
AT4G25580 codes for a stress-responsive protein-related with weak similarity (less than 20 % identity) with LEA group 2 (Bies-Ethève et al. 2008) and high similarity to a CAP160 protein (Mönke et al. 2012) and to a CDeT11-24 from Craterostigma plantagineum (Röhrig et al. 2006). In spinach, CAP160 proteins are predominantly cytosolic, induced by drought stress and related to stabilization of membranes, ribosomes and cytoskeletal elements (Kaye et al. 1998). CDeT11-24 is thought to contribute to the plant’s DT possibly by interacting with other proteins such as dehydrins (Röhrig et al. 2006). However, at4g25580 seeds had an increased ability to re-establish DT compared to wild-type seeds, suggesting the operation of derepression mechanisms.
None of the T-DNA lines analyzed showed phenotypes for seed dormancy or longevity. Only one of the analyzed genes (AT3G53040) was reported to be differentially expressed comparing dormant and after-ripened states (Cadman et al. 2006; Bassel et al. 2011). Considering the increase in longevity between 24 and 72 h of incubation in ABA and the location in the network (ER region) of the genes chosen for phenotypic characterization, it is conceivable that the disruption of single genes would not influence longevity. Besides, as only three of the T-DNA lines had the vigor affected, we believe that seed development was not significantly affected by the disruption of the single genes.
Taken together, these results confirm that the analysis of co-expression network structures can bring insights to biological processes. We showed that the re-establishment of DT in germinated A. thaliana seeds can be divided into two phases. In the first phase, a series of stress-responsive genes that are also related to seed development as well as other biological processes (such as dormancy, acquisition of DT, drought and the circadian clock) is induced, promoting amplification of signals, growth arrest and protection mechanisms (such as LEA proteins). In the second phase, photosynthesis and primary metabolism are strongly inhibited and another set of stress-responsive genes promotes adaptation to stress conditions that also contribute to seed survival in the dry state, improving longevity. Moreover, we suggest that redundancy and compensatory mechanisms may be operating when genes important for DT are disrupted.
Author contribution statement
MCDC designed research, conducted experiments, analyzed data and wrote the manuscript. KR analyzed data and wrote the manuscript. HN designed scripts for microarray analysis. FY conducted experiments. WL, JB and HWMH conceived and designed research, analyzed data and wrote the manuscript. All authors read and approved the manuscript.
Abbreviations
- ABA:
-
Abscisic acid
- DT:
-
Desiccation tolerance
- DEG:
-
Differentially expressed gene
- ER:
-
Early response
- GO:
-
Gene ontology
- HSP:
-
Heat shock protein
- LR:
-
Late response
- LEA:
-
Late embryogenesis abundant
- ORA:
-
Over-representation analysis
- PEG:
-
Polyethylene glycol
- SOM:
-
Self-organizing map
- TF:
-
Transcription factors
References
Alabadí D, Oyama T, Yanovsky MJ et al (2001) Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293:880–883
Alonso-Blanco C, Bentsink L, Hanhart CJ et al (2003) Analysis of natural allelic variation at seed dormancy loci of Arabidopsis thaliana. Genetics 164:711–729
Assenov Y, Ramírez F, Schelhorn S-E, Lengauer Thomas, Albrecht M (2008) Computing topological parameters of biological networks. Bioinformatics 24:282–284
Bass JIF, Diallo A, Nelson J et al (2013) Using networks to measure similarity between genes: association index selection. Nat Methods 10:1169–1176
Bassel GW, Lan H, Glaab E et al (2011) Genome-wide network model capturing seed germination reveals coordinated regulation of plant cellular phase transitions. Proc Natl Acad Sci USA 108:9709–9714
Bentsink L, Alonso-Blanco C, Vreugdenhil D et al (2000) Genetic analysis of seed-soluble oligosaccharides in relation to seed storability of Arabidopsis. Plant Physiol 124:1595–1604
Bies-Ethève N, Gaubier-Comella P, Debures A et al (2008) Inventory, evolution and expression profiling diversity of the LEA (late embryogenesis abundant) protein gene family in Arabidopsis thaliana. Plant Mol Biol 67:107–124
Buitink J, Vu BL, Satour P, Leprince O (2003) The re-establishment of desiccation tolerance in germinated radicles of Medicago truncatula Gaertn. seeds. Seed Sci Res 13:273–286
Buitink J, Leger JJ, Guisle I et al (2006) Transcriptome profiling uncovers metabolic and regulatory processes occurring during the transition from desiccation-sensitive to desiccation-tolerant stages in Medicago truncatula seeds. Plant J 47:735–750
Cadman CSC, Toorop PE, Hilhorst HWM, Finch-Savage WE (2006) Gene expression profiles of Arabidopsis Cvi seeds during dormancy cycling indicate a common underlying dormancy control mechanism. Plant J 46:805–822
Candat A, Paszkiewicz G, Neveu M et al (2014) The ubiquitous distribution of late embryogenesis abundant proteins across cell compartments in Arabidopsis offers tailored protection against abiotic stress. Plant Cell 26:3148–3166
Carbon S, Ireland A, Mungall CJ et al (2009) AmiGO: online access to ontology and annotation data. Bioinformatics 25:288–289
Dalal M, Tayal D, Chinnusamy V, Bansal KC (2009) Abiotic stress and ABA-inducible Group 4 LEA from Brassica napus plays a key role in salt and drought tolerance. J Biotechnol 139:137–145
Dekkers BJW, Pearce S, van Bolderen-Veldkamp RP et al (2013) Transcriptional dynamics of two seed compartments with opposing roles in Arabidopsis seed germination. Plant Physiol 163:205–215
Delahaie J, Hundertmark M, Bove J et al (2013) LEA polypeptide profiling of recalcitrant and orthodox legume seeds reveals ABI3-regulated LEA protein abundance linked to desiccation tolerance. J Exp Bot 64:4559–4573
Dong J, Horvath S (2007) Understanding network concepts in modules. BMC Syst Biol 1:24
Edgar R, Domrachev M, Lash AE (2002) Gene expression omnibus: NCBI gene expression and hybridization array data repository. Nucleic Acids Res 30:207–210
Fang L, Hou X, Lee LYC et al (2011) AtPV42a and AtPV42b redundantly regulate reproductive development in Arabidopsis thaliana. PLoS One 6:e19033
Finch-Savage WE, Leubner-Metzger G (2006) Seed dormancy and the control of germination. New Phytol 171:501–523
Freeman TC, Goldovsky L, Brosch M et al (2007) Construction, visualisation, and clustering of transcription networks from microarray expression data. PLoS Comput Biol 3:2032–2042
Fujita Y, Fujita M, Shinozaki K, Yamaguchi-Shinozaki K (2011) ABA-mediated transcriptional regulation in response to osmotic stress in plants. J Plant Res 124:509–525
Gentleman RC, Carey VJ, Bates DM et al (2004) Bioconductor: open software development for computational biology and bioinformatics. Genome Biol 5:R80
Guo A, He K, Liu D et al (2005) DATF: a database of Arabidopsis transcription factors. Bioinformatics 21:2568–2569
Heisel TJ, Li CY, Grey KM, Gibson SI (2013) Mutations in HISTONE ACETYLTRANSFERASE1 affect sugar response and gene expression in Arabidopsis. Front Plant Sci 4:245
Hellemans J, Mortier G, De Paepe A et al (2007) qBase relative quantification framework and software for management and automated analysis of real-time quantitative PCR data. Genome Biol 8:R19
Huang M-D, Wu W-L (2006) Genome-wide in silico identification and experimental confirmation of abscisic acid-regulated genes in Arabidopsis. Plant Sci 170:986–993
Huang D, Wu W, Abrams SR, Cutler AJ (2008) The relationship of drought-related gene expression in Arabidopsis thaliana to hormonal and environmental factors. J Exp Bot 59:2991–3007
Huang Y, Li CY, Pattison DL et al (2010) SUGAR-INSENSITIVE3, a RING E3 ligase, is a new player in plant sugar response. Plant Physiol 152:1889–1900
Hundertmark M, Hincha DK (2008) LEA (late embryogenesis abundant) proteins and their encoding genes in Arabidopsis thaliana. BMC Genom 9:1–18
Irizarry RA, Hobbs B, Collin F et al (2003) Exploration, normalization, and summaries of high density oligonucleotide array probe level data. Biostatistics 4:249–264
Joosen RVL, Kodde J, Willems LAJ et al (2010) GERMINATOR: a software package for high-throughput scoring and curve fitting of Arabidopsis seed germination. Plant J 62:148–159
Kalemba EM, Pukacka S (2011) Association of protective proteins with dehydration and desiccation of orthodox and recalcitrant category seeds of three Acer genus species. J Plant Growth Regul 31:351–362
Kaye C, Neven L, Hofig A et al (1998) Characterization of a gene for spinach CAP160 and expression of two spinach cold-acclimation proteins in tobacco. Plant Physiol 116:1367–1377
Keller A, Backes C, Al-Awadhi M et al (2008) GeneTrailExpress: a web-based pipeline for the statistical evaluation of microarray experiments. BMC Bioinformatics 9:552
Khuri S, Bakker FT, Dunwell JM (2001) Phylogeny, function, and evolution of the cupins, a structurally conserved, functionally diverse superfamily of proteins. Mol Biol Evol 18:593–605
Le BH, Cheng C, Bui AQ et al (2010) Global analysis of gene activity during Arabidopsis seed development and identification of seed-specific transcription factors. Proc Natl Acad Sci USA 107:8063–8070
Leprince O, Van Aelst AC, Pritchard HW, Murphy DJ (1998) Oleosins prevent oil-body coalescence during seed imbibition as suggested by a low-temperature scanning electron microscope study of desiccation-tolerant and -sensitive oilseeds. Planta 204:109–119
Lindemose S, O’Shea C, Jensen MK, Skriver K (2013) Structure, function and networks of transcription factors involved in abiotic stress responses. Int J Mol Sci 14:5842–5878
Maia J, Dekkers BJW, Provart NJ et al (2011) The re-establishment of desiccation tolerance in germinated Arabidopsis thaliana seeds and its associated transcriptome. PLoS One 6:e29123
Maia J, Dekkers BJW, Dolle M et al (2014) Abscisic acid (ABA) sensitivity regulates desiccation tolerance in germinated Arabidopsis seeds. New Phytol 203:81–93
Matsui A, Ishida J, Morosawa T et al (2008) Arabidopsis transcriptome analysis under drought, cold, high-salinity and ABA treatment conditions using a tiling array. Plant Cell Physiol 49:1135–1149
Mönke G, Seifert M, Keilwagen J et al (2012) Toward the identification and regulation of the Arabidopsis thaliana ABI3 regulon. Nucleic Acids Res 40:8240–8254
Mutwil M, Usadel B, Schütte M et al (2010) Assembly of an interactive correlation network for the Arabidopsis genome using a novel heuristic clustering algorithm. Plant Physiol 152:29–43
Ooms JJJ, Léon-Kloosterziel KM, Bartels D et al (1993) Acquisition of desiccation tolerance and longevity in seeds of Arabidopsis thaliana. Plant Physiol 102:1185–1191
Pammenter NW, Berjak P (1999) A review of recalcitrant seed physiology in relation to desiccation-tolerance mechanisms. Seed Sci Res 9:13–37
Piraveenan M, Thedchanamoorthy G, Uddin S, Chung KSK (2013) Quantifying topological robustness of networks under sustained targeted attacks. Soc Netw Anal Min 3:939–952
Ramanjulu S, Bartels D (2002) Drought- and desiccation-induced modulation of gene. Plant, Cell Environ 25:141–151
Röhrig H, Schmidt J, Colby T et al (2006) Desiccation of the resurrection plant Craterostigma plantagineum induces dynamic changes in protein phosphorylation. Plant, Cell Environ 29:1606–1617
Rosnoblet C, Aubry C, Leprince O et al (2007) The regulatory gamma subunit SNF4b of the sucrose non-fermenting-related kinase complex is involved in longevity and stachyose accumulation during maturation of Medicago truncatula seeds. Plant J 51:47–59
Scardoni G, Petterlini M, Laudanna C (2009) Analyzing biological network parameters with CentiScaPe. Bioinformatics 25:2857–2859
Scholl RL, May ST, Ware DH (2000) Seed and molecular resources for Arabidopsis. Plant Physiol 124:1477–1480
Seo PJ, Lee SB, Suh MC et al (2011) The MYB96 transcription factor regulates cuticular wax biosynthesis under drought conditions in Arabidopsis. Plant Cell 23:1138–1152
Smoot ME, Ono K, Ruscheinski J et al (2011) Cytoscape 2.8: new features for data integration and network visualization. Bioinformatics 27:431–432
Sparkes I, Tolley N, Aller I et al (2010) Five Arabidopsis reticulon isoforms share endoplasmic reticulum location, topology, and membrane-shaping properties. Plant Cell 22:1333–1343
Sreenivasulu N, Usadel B, Winter A et al (2008) Barley grain maturation and germination: metabolic pathway and regulatory network commonalities and differences highlighted by new MapMan/PageMan profiling tools. Plant Physiol 146:1738–1758
Supek F, Bošnjak M, Škunca N, Šmuc T (2011) REVIGO summarizes and visualizes long lists of gene ontology terms. PLoS One 6:e21800
Terrasson E, Buitink J, Righetti K et al (2013) An emerging picture of the seed desiccome: confirmed regulators and newcomers identified using transcriptome comparison. Front Plant Sci 4:1–16
Toorop PE, Barroco RM, Engler G et al (2005) Differentially expressed genes associated with dormancy or germination of Arabidopsis thaliana seeds. Planta 221:637–647
Törönen P, Kolehmainen M, Wong G, Castre E (1999) Analysis of gene expression data using self-organizing maps. FEBS Lett 451:142–146
Trobacher CP, Senatore A, Greenwood JS (2006) Masterminds or minions? Cysteine proteinases in plant programmed cell death. Can J Bot 84:651–667
Verdier J, Lalanne D, Pelletier S et al (2013) A regulatory network-based approach dissects late maturation processes related to the acquisition of desiccation tolerance and longevity of Medicago truncatula seeds. Plant Physiol 163:757–774
Vieira CV, da Silva EAA, de Alvarenga AA et al (2010) Stress-associated factors increase after desiccation of germinated seeds of Tabebuia impetiginosa Mart. Plant Growth Regul 62:257–263
Villa-Vialaneix N, Liaubet L, Laurent T et al (2013) The structure of a gene co-expression network reveals biological functions underlying eQTLs. PLoS One 8:e60045
Wan C-Y, Wilkins TA (1994) A modified hot borate method significantly enhances the yield of high quality RNA from cotton (Gossypium hirsutum L.). Anal Biochem 223:7–12
Wang Y, Joshi T, Zhang X-S et al (2006) Inferring gene regulatory networks from multiple microarray datasets. Bioinformatics 22:2413–2420
Yamaguchi-Shinozaki K, Shinozaki K (2005) Organization of cis-acting regulatory elements in osmotic- and cold-stress-responsive promoters. Trends Plant Sci 10:88–94
Acknowledgments
This work was supported by the ‘Conselho Nacional de Desenvolvimento Científico e Tecnológico’ (CNPq, Brazil).
Author information
Authors and Affiliations
Corresponding author
Additional information
Special topic: Desiccation Biology. Guest editors: Olivier Leprince and Julia Buitink
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2015_2283_MOESM1_ESM.tif
Supplemental Fig. S1 Schematic illustration and relative abundance of transcripts of mutated genes in cbsx4, at3g53040 and at4g25580. a Mapped T-DNA insertion sites (triangles) are indicated on the top of the genomic structure. Black boxes on the black solid line indicate the exons, gray boxes indicate 3′and 5′ untranslated regions and arrows indicate location of qPCR primers (fwd: forward; rev: reverse). b Normalized relative expression levels of transcripts of mutated genes in cbsx4, at3g53040 and at4g25580 compared to wild-type (WT) calculated with the qBase software (Hellemans et al. 2007). Asterisks indicate significant differences at P ≤ 0.05 for one, P ≤ 0.01 for two asterisks (TIFF 220 kb)
425_2015_2283_MOESM2_ESM.tif
Supplemental Fig. S2 Temporal expression profiles of 20 genes measured by qPCR and microarray after incubation of germinated seeds in the stage of radicle protrusion for 0 h, 2 h and 24 h. Blue lines indicate expression levels measured by qPCR and red lines indicate expression levels measured by microarray (TIFF 551 kb)
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Costa, M.C.D., Righetti, K., Nijveen, H. et al. A gene co-expression network predicts functional genes controlling the re-establishment of desiccation tolerance in germinated Arabidopsis thaliana seeds. Planta 242, 435–449 (2015). https://doi.org/10.1007/s00425-015-2283-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00425-015-2283-7