Abstract
Main conclusion
The biochemical characterization of glycolate oxidase in Ricinus communis hints to different physiological functions of the enzyme depending on the organ in which it is active.
Abstract
Enzymatic activities of the photorespiratory pathway are not restricted to green tissues but are present also in heterotrophic organs. High glycolate oxidase (GOX) activity was detected in the endosperm of Ricinus communis. Phylogenetic analysis of the Ricinus l-2-hydroxy acid oxidase (Rc(l)-2-HAOX) family indicated that Rc(l)-2-HAOX1 to Rc(l)-2-HAOX3 cluster with the group containing streptophyte long-chain 2-hydroxy acid oxidases, whereas Rc(l)-2-HAOX4 clusters with the group containing streptophyte GOX. Rc(l)-2-HAOX4 is the closest relative to the photorespiratory GOX genes of Arabidopsis. We obtained Rc(l)-2-HAOX4 as a recombinant protein and analyze its kinetic properties in comparison to the Arabidopsis photorespiratory GOX. We also analyzed the expression of all Rc(l)-2-HAOXs and conducted metabolite profiling of different Ricinus organs. Phylogenetic analysis indicates that Rc(l)-2-HAOX4 is the only GOX encoded in the Ricinus genome (RcGOX). RcGOX has properties resembling those of the photorespiratory GOX of Arabidopsis. We found that glycolate, the substrate of GOX, is highly abundant in non-green tissues, such as roots, embryo of germinating seeds and dry seeds. We propose that RcGOX fulfills different physiological functions depending on the organ in which it is active. In autotrophic organs it oxidizes glycolate into glyoxylate as part of the photorespiratory pathway. In fast growing heterotrophic organs, it is most probably involved in the production of serine to feed the folate pathway for special demands of those tissues.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Glycolate is a metabolic intermediate produced in high amounts during photorespiration in all oxygenic photosynthetic organisms (Zelitch et al. 2009; Maurino and Peterhansel 2010). As part of the photorespiratory pathway, land plants and Charophyta convert glycolate to glyoxylate through glycolate oxidase (GOX) (Leliaert et al. 2012; Esser et al. 2014).
GOX belongs to the l-2-hydroxy acid oxidase ((l)-2-HAOX) family (Esser et al. 2014). The Arabidopsis thaliana genome encodes three GOXs that oxidize glycolate and l-lactate using oxygen as the electron acceptor (Engqvist et al. 2015). GOX1 and GOX2 are mostly involved in photorespiration, while GOX3 supports l-lactate metabolism in roots (Engqvist et al. 2015; Dellero et al. 2016). Animals also possess GOX (Kohler et al. 1999; Ryan et al. 2001), which has a common eukaryotic ancestor with plant GOX (Esser et al. 2014). Animal GOX produces glyoxylate, which is then used for the peroxisomal synthesis of glycine (Williams et al. 2000). Animal and plant GOX convergently duplicated to evolve long-chain 2-hydroxy acid oxidases (lHAOX), which have a broad substrate specificity (Jones et al. 2000; Esser et al. 2014). In mammals, lHAOX play a role in the degradation of food components or xenobiotic compounds (Belmouden and Lederer 1996; Verhoeven et al. 1997, 1998). Plant lHAOX are likely involved in the conversion or degradation of 2-hydroxy acids produced during the metabolism of fatty acids or amino acids (Esser et al. 2014). Arabidopsis possesses two lHAOX, which prefer long-chain fatty acids and short-chain hydroxy acids such as l-lactate, leucic acid, valic acid, and isoleucic acid over glycolate as substrates (Esser et al. 2014).
In plants several enzymes associated with the photorespiratory pathway are present in roots and other heterotrophic tissues (Lernmark et al. 1991; Igamberdiev et al. 1997). For example, in the scutellum of maize, glycine decarboxylase is involved in the oxidation of glycine formed from glyoxylate (Igamberdiev et al. 1997). Glyoxylate, the product of the GOX reaction, was shown to be metabolized to glycine and serine by different plant heterotrophic organs, such as germinating cotyledons, roots, coleoptiles, and storage tissue (Sinha and Cossins 1965). Previous work indicated the existence of GOX activity in germinating castor bean (Ricinus communis) endosperm (Tanner and Beevers 1965), which was also related to the synthesis of glycine (Cossins and Sinha 1967). Thus, as in the case of animals, in plant heterotrophic organs GOX may be involved in metabolic processes other than photorespiration (Maurino and Engqvist 2015).
Here, we aimed the identification and biochemical characterisation of the protein responsible for the GOX activity of heterotrophic organs of Ricinus. We found that Rc(l)-2-HAOX4 is the protein with GOX characteristics and is encoded in the Ricinus genome by a single copy gene. We cloned Rc(l)-2-HAOX4 from Ricinus endosperm and found that it has properties similar to Arabidopsis photorespiratory GOX. We propose that in heterotrophic organs, RcGOX is likely involved in the metabolism of glycolate to produce serine to feed the folate pathway for special demands of those tissues.
Materials and methods
Sequence alignment and phylogenetic analysis
Rc(l)-2-HAOX1 (EEF33207.1), Rc(l)-2-HAOX2 (EEF33202.1), Rc(l)-2-HAOX3 (EEF33208.1) and Rc(l)-2-HAOX4 (EEF42631.1) sequences were retrieved from NCBI using Arabidopsis thaliana and Homo sapiens GOX as queries in blastp searches. Sequences were aligned with previously characterized GOX sequences from higher plants and Animalia according to Esser et al. (2014). 21 sequences were aligned with MAFFT using Guidance2, performing 400 bootstrap iterations and an alignment score threshold for non-reliable columns of 0.8 (Katoh et al. 2002; Sela et al. 2015). Evolutionary analyses were conducted in MEGA7 (Kumar et al. 2016). The evolutionary gene tree was inferred using the Maximum Likelihood method with the Le Gascuel 2008 model (Le and Gascuel 2008). A discrete Gamma distribution was used to model evolutionary rate differences among sites [5 categories (+ G, parameter = 1.3901)].
Plant growth and sample collection
Seeds of Ricinus communis var. zanzibariensis (Samen Aders GmbH & Co.KG, Düsseldorf, Germany) were surface sterilized with a 1% (v/v) sodium hypochlorite solution for 5 min and subsequently imbibed for 48 h in running water. Imbibed seeds were transferred to boxes of watered Vermiculite and incubated 4 days at 30 °C in darkness. Germinated seeds were either harvested and separated into germinated embryo and endosperm or transferred to vermiculite pots for further growth under a 12/12 h light/dark period and 23/20 °C cycle. The light intensity was around 90–100 µmol photons·m−1 s−2 from Spectralux Plus NL 36 W/840 (Radium) light bulbs. Plants were fertilized on a weekly basis with N/P/K fertilizer (Bayer). Leaf and root material was harvested after 5 weeks in the light. Three to four individual samples were pooled for one biological replicate, and all plant material was immediately shock frozen in liquid nitrogen.
RNA extraction, reverse transcription and quantitative real-time PCR
Total RNA was isolated from grinded plant material according to Logemann et al. (1987). Quality and quantity of total RNA was verified in gel and photospectrometrically, respectively. Genomic DNA contaminations were removed using Ambion DNA-free DNA Removal Kit. 1.5 µg of total RNA were used for reverse transcription into complementary DNA (cDNA) using RevertAid Reverse Transcriptase (Thermo Scientific) and OligodT primer according to the manufacturer’s instruction. KAPA SYBR FAST qPCR Master Mix (Kapabiosystems) was used for real-time PCR in an Applied Biosystems Step one Plus real time PCR system with 2 µl of 1:10 diluted cDNA according to the manufacturer’s instructions. The reference gene elongation factor 1b (EF1b), with the most stable expression pattern in seed development, and primer pair (EF1b_F + EF1b_R; Table S1) were taken from Cagliari et al. (2010) and compared to a second reference gene, ubiquitin conjugating enzyme E2 (UBC, XM_002509871.3, UBC_F + UBC_R; Table S1), for which the Arabidopsis homolog (UBC9, At4g27960) has been shown to be the most stably expressed gene in developmental series. Primers for Rc(L)-2-HAOX genes (Table S1) were designed using Primer-Blast (Ye et al. 2012), and relative transcript abundance in particular tissue types was analysed from three biological samples and expressed as fold change in delta CT.
Heterologous expression and purification of RcHAOX4
Complementary DNA (cDNA) of Ricinus endosperm was used as a template for PCR with the primer pair Rc4-pET_F + Rc4-pET_R (Table S1), which introduces a BamHI restriction site at the 5′ and 3′ termini of the full-length coding sequence. The PCR products were sub-cloned into pCR-Blunt-II-TOPO (Life Technologies, Invitrogen) resulting in pTOPO-Rc(l)-2-HAOX4 and confirmed via sequencing. The obtained Rc(l)-2-HAOX4 sequence revealed the presence of a valine instead of the isoleucine at position 216 expected from the sequence retrieved from NCBI. As valine at position 216 is conserved in Arabidopsis, maize, and spinach GOX proteins, the substitution is not expected to affect the protein function and most likely represents the actual GOX sequence in Ricinus communis var. zanzibariensis. The plasmid pTOPO-Rc(l)-2-HAOX4 was digested with BamHI and the purified DNA fragment was ligated into the BamHI linearized pET16b.
The expression vector pET16b-HIS:Rc(l)-2-HAOX4 was transformed into E. coli BL21 Rosetta pLysS strain (Novagen). Protein expression and Histidine-tag (HIS) based purification was performed as described in Schmitz et al. (2017a). Protein purity was verified via SDS-PAGE according to Laemmli (1970). Protein concentration was determined using the amidoblack precipitation protocol of (Schaffner and Weissmann 1973).
Determination of enzymatic parameters of Rc(l)-2-HAOX4
Enzymatic activity was determined as described in Schmitz et al. (2017b) by measuring the change in absorption at 324 nm by the formation of phenylhydrazone using a Synergy HT Plate reader (Biotek). The standard assay mixture contained 0.5 mM EDTA, 5 mM MgSO4, 3 mM phenylhydrazin, 2 mM FMN, and 5 mM of either glycolate, l-lactate, or d-lactate. The assay mixture was buffered with 100 mM MES-NaOH to work at pH 5.5–6.5 or with TRIS–HCl to work at pH 7.0–9.0. Substrate affinity (Km) was deduced from saturation kinetics using nonlinear regression, fitting the Michaelis–Menten equation [f(x) = a·x/(b + x)] to the combined data from three independent protein purifications. Arabidopsis GOX1 (AtGOX1) (Engqvist et al. 2015) was assayed in parallel as a positive control.
Cloning, transient expression, and subcellular localization of RcHAOX4
Rc(l)-2-HAOX4 was cloned as a C-terminal and N-terminal fusion protein into the vector peqFP611 (Forner and Binder 2007) containing the red fluorescent protein FP611. To generate the N-terminal fusion, two inserts were generated: the primers Rc4_N-term_fusion_F and Rc4_N-term_fusion_R (Table S1) were used to amplify Rc(l)-2-HAOX4 coding sequence from pTOPO-Rc(l)-2-HAOX4 and the primers FP611_F and FP611_R (Table S1) were used to generate a stop codon free FP611 coding sequence from peqFP611. The vector peqFP611 was digested with the restriction enzymes XbaI and SmaI. These two inserts and the backbone were combined via Gibson assembly (Gibson et al. 2009) and the resulting pFP611:RcHAOX4 was verified by sequencing. The primers Rc4_C-term_fusion_F and Rc4_C-term_fusion_R were used to amplify Rc(l)-2-HAOX4 coding sequence from pTOPO-Rc(l)-2-HAOX4. The vector peqFP611 was digested using XhoI and SacI and both fragments combined as above. The resulting pRc(l)-2-HAOX4:FP611 was verified by sequencing. Both pFP611:Rc(l)-2-HAOX4 and pRc(l)-2-HAOX4:FP611 were transformed into the Agrobacterium tumefaciens strain GV3101 (pMP90) and cultivated for transient transformation of Nicotiana benthamiana plants. Tobacco plants were grown in green house conditions during 4 weeks. Infiltration and protoplastation of tobacco leaves was conducted after 2 and 3 days of transient protein expression according to Waardt and Kudla (2008). As Rc(l)-2-HAOX4 is strongly predicted to be localized to the peroxisomes, N-bodipy ((8-(4-Nitrophenyl) bodipy, Biozol) was used as an organelle marker for peroxisomes (Landrum et al. 2010). Staining was achieved by adding 5 µM N-bodipy to the isolated protoplast 10 min before imaging.
Imaging was performed at the LSM 780 confocal microscope (Zeiss) with acquisition settings as described in Schmitz et al. (2017a) and N-bodipy (excitation: 488 nm, emission: 503–530 nm) fluorescence detected in a separate frame. False color images were processed using the Fiji software with ImageJ version 1.53c (Schindelin et al. 2012).
Metabolite analysis
Total metabolites were extracted from ground material of seeds, embryo of germinated seeds, endosperm, leaf, and root according to Fiehn (2007) and Weckwerth et al. (2004) in a methanol/chloroform/water mix (5:2:2, by vol.). Ribitol was used for internal standardization. In brief a dried fraction of the extraction mixture was derivatisized with methoxyamine hydrochloride at 37 °C for 90 min and subsequently with N-methyl-N-(trimethylsilyl)trifluoroacetamide (MSTFA) at 37 °C for 30 min. Extracts were analysed by GC/MS using an accurate mass Q-TOF GC/MS system with a HP-5MS column with 5% phenyl methyl siloxane film (Agilent 7890A GC system, Agilent Technologies). A metabolite standard mixture including all metabolite targets was measured every 30 samples as retention and response reference. Raw data was processed with MassLynx and QuanLynx software (Waters GmbH) to identify metabolites in comparison to the NIST14 Mass Spectral Library (https://www.nist.gov/srd/nist-standard-reference-database-1a-v14). Response peak areas were integrated and were normalized to the internal ribitol standard and to gram of dry weight as a measure of relative metabolite content. Relative metabolite levels of three biological replicates were averaged. For comparison across different tissue types, the highest abundance of each particular metabolite was set to 1.
Results
The genome of Ricinus encodes four l-2-hydroxy acid oxidases
GOX and lHAOX in plants and animals most likely arose from an ancestral eukaryotic peroxisomal (l)-2-HAOX (Esser et al. 2014). A phylogenetic analysis using sequences of Ricinus (l)-2-HAOX and experimentally confirmed lHAOX and GOX proteins separate Animalia and Plantae sequences into two clades (Fig. 1a). Animalia and Streptophyta sequences each split further into GOX and lHAOX groups (Fig. 1a). This points to the convergent diversifications of the (l)-2-HAOX gene family into GOX and lHAOX subfamilies in these two eukaryotic kingdoms (Esser et al. 2014). From the four Ricinus (l)-2-HAOX proteins, Rc(l)-2-HAOX1 to Rc(l)-2-HAOX3 cluster with the group containing streptophyte lHAOX, whereas Rc(l)-2-HAOX4 clusters with the group containing streptophyte GOX.
Phylogenetic and sequence analysis of (l)-2-HAOXs proteins. a Selected (L)-2-HAOXs sequences (Esser et al. 2014) were aligned with MAFFT using Guidance2 and the alignment filtered for unreliably aligned positions was used for subsequent analysis (Katoh et al. 2002; Sela et al. 2015). The evolutionary history was inferred using the Maximum Likelihood method based on the Le_Gascuel_2008 model (Le and Gascuel 2008). A discrete Gamma distribution was used to model evolutionary rate differences among sites (5 categories (+ G, parameter = 1.3901)). The tree with the highest log likelihood (-6484.48) is shown. The percentage of trees in which the associated taxa clustered together is shown above the branches (1000 Bootstraps). The tree is drawn to scale, with branch lengths proportional to the number of substitutions per site (scale bar). The analysis involved a reduced set of 21 l-(2)-HAOX amino acid sequences (gene identifier in brackets; Esser et al. 2014). There were in total 355 positions in the final dataset. b Conserved amino acid positions that allow a discrimination of (l)-2-HAOX into lHAOX and GOX and their different predicted substrate specificities, and sequence of the terminal tripeptide and the probability of peroxisomal targeting via a PTS1 signal, of (l)-2-HAOXs of Ricinus and Arabidopsis. The targeting prediction was performed with PredPlantPTS1 (https://ppp.gobics.de/, Threshold score 0.412, max score Arabidopsis 1.188) (Reumann et al. 2012). Exper. verified means that the PTS1 signal [SML (Lingner et al. 2011), ARL (Lingner et al. 2011), AKL (Lingner et al. 2011), PRL (Reumann et al. 2007)] has already been verified experimentally as a functional plant PTS1 tripeptide for peroxisomal targeting. c False color visualization of fluorescence in isolated N. benthamiana protoplasts. C-terminal fusion of FP611 to Rc(l)-2-HAOX4 (FP611:Rc(l)-2-HAOX4, magenta) results in cytosolic localization (top row). N-terminal fusion (Rc(l)-2-HAOX4:FP611, magenta, bottom row) colocalizes with N-bodipy fluorescence (green) in peroxisomes. Chlorophyll fluorescence depicted as blue. Scale bar = 10 µm
Apart from the phylogenetic relationships, our previous work showed that four amino acid positions allow the discrimination of (l)-2-HAOX with different predicted substrate specificities (Esser et al. 2014). The analysis of these positions further confirmed that Rc(l)-2-HAOX1 to -3 possess the conserved amino acids of lHAOX sequences, while Rc(l)-2-HAOX4 possesses the conserved amino acids of GOX sequences (Fig. 1b). Moreover, computational prediction of the subcellular localization indicate that all examined sequences contain a peroxisomal targeting signal type 1 PTS1) sequence at the C-terminus (Fig. 1b; Reumann et al. 2012). All PTS1 signals [SML (Lingner et al. 2011), ARL (Lingner et al. 2011), AKL (Lingner et al. 2011), PRL (Reumann et al. 2007)] have been verified experimentally as a functional plant PTS1 tripeptide before (Fig. 1b).
To support the predicted subcellular localization, we cloned and transiently expressed fluorescent fusion proteins of Rc(l)-2-HAOX4 in tobacco leaves (Fig. 1c). We found that a C-terminal fusion protein (Rc(l)-2-HAOX4:FP611, Fig. 1c top row) localizes to the cytosol and an N-terminal fusion protein (FP611:Rc(l)-2-HAOX4, Fig. 1c, bottom row), leaving the C-terminal PTS1 of Rc(l)-2-HAOX4 free, colocalizes with an established peroxisomal marker in tobacco protoplasts. Taken together, these findings support the localization and physiological role of Rc(l)-2-HAOXs in peroxisomes.
Rc(l)-2-HAOX4, the ortholog of Arabidopsis photorespiratory GOX, is highly expressed in heterotrophic organs
The Arabidopsis genome encodes three GOX isoforms; photorespiratory AtGOX1 and AtGOX2 are highly expressed in photosynthetic organs, while AtGOX3 is expressed in roots, where it supports l-lactate oxidation (Engqvist et al. 2015). Analysis of the unambiguous phylogenetic clustering and the conservation of amino acids determining substrate specificity point to the existence of a single GOX and three (l)-2-HAOX proteins in Ricinus (Fig. 1a).
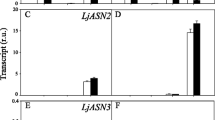
To gain knowledge on the abundance of (l)-2-HAOX mRNA in Ricinus, we performed transcriptional analysis via qPCR using heterotrophic and autotrophic organs. The expression strength of all l-(2)-HAOX was compared to Ubiquitin (UBC) (Fig. 2a). Rc(l)-2-HAOX1 and Rc(l)-2-HAOX3 are expressed at very low levels in roots and leaves, and expression is negligible in embryos and the endosperm of germinated seeds. The expression levels of Rc(l)-2-HAOX2 is low in all organs analysed and is comparable to that of Rc(l)-2-HAOX1 and Rc(l)-2-HAOX3 in roots and leaves. Rc(l)-2-HAOX4 is highly expressed in leaves. Intriguingly, we also found expression of Rc(l)-2-HAOX4 in roots, embryos, and the endosperm of germinated seeds, with levels similar to UBC.
Transcript analyses of Ricinus (l)-2-HAOXs in heterotrophic and autotrophic tissues. a Quantitative real-time PCR data expressed in fold change of ΔCT, visualising expression strength across different organs. RcUBC expression is shown as a reference. n = 3, ± SE; x not determined, E embryo of germinated seeds, ES endosperm of germinated seeds, L leaf, R root. All genes of interest were normalized to the housekeeping gene EF1b and tested for significant differences with one-way Anova with subsequent Tukey post-hoc test. * Indicate significant difference to all others: a = significant difference to E, b = significant difference to ES, c = significant difference to L, d = significant difference to R. b Rc(l)-2-HAOX1 to Rc(l)-2-HAOX4 transcript abundances in fragments per kilobase of transcript per million mapped reads (FPKM) compared to the housekeeping gene RcUBC (Brown et al. 2012). ES II/III = endosperm of developing seeds at stage II/III, ES V/VI = endosperm of developing beans at stage V/VI. S whole germinating seed, L leaf, F flower. Stages of developing beans were taken from (Greenwood and Bewley 1982)
We broadened the study by analyzing available deep-sequencing RNAseq data derived from different Ricinus autotrophic and heterotrophic organs (Fig. 2b) (Brown et al. 2012). In line with the qPCR results, the expression of Rc(l)-2-HAOX1 to Rc(l)-2-HAOX3 is very low (FPKM < 20, i.e., less than 20 fragments per kilobase million) in all organs analysed. Rc(l)-2-HAOX1 transcripts are only detected in leaves, while Rc(l)-2-HAOX2 and Rc(l)-2-HAOX3 transcripts are found at comparable low levels in all organs analysed. Rc(l)-2-HAOX4 expression is very high in leaves (FPKM = 1966), followed by whole germinating seeds (FPKM = 424).
Rc(l)-2-HAOX4 is a glycolate oxidase
Rc(l)-2-HAOX4 is highly expressed in leaves (Fig. 2), and phylogenetic analysis (Fig. 1a) indicates that it clusters with the group containing streptophyte GOX; as AtGOX1 and Rc(l)-2-HAOX4 share 90 % aa identity we postulate that Rc(l)-2-HAOX4 represents the (l)-2-HAOX responsible for the detoxification of glycolate formed in the photorespiratory pathway. As this putative GOX is also expressed at high levels in heterotrophic organs, we analyzed the enzymatic properties of the recombinant enzyme. Rc(l)-2-HAOX4 was expressed in E. coli and the recombinant protein was isolated by affinity chromatography. The purified protein presented the expected molecular weight of 44 kDa after separation by SDS-PAGE (Fig. 3a).
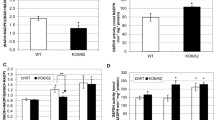
Biochemical properties of Rc(l)-2-HAOX4. a Coomassie-stained SDS–polyacrylamide gel of different steps during the isolation of recombinant Rc(L)-2-HAOX4. Crude protein extract of E. coli before (1) and after (2) induction of expression with 1 mM IPTG, non-soluble fraction (3) and soluble fraction (4) after cell disruption, flow-through of washing steps (5–7), and eluted recombinant Rc(l)-2-HAOX4 (8 and 9; 44 kDa); M molecular mass standards. b Enzymatic activity of recombinant Rc(l)-2-HAOX4 with the possible substrates glycolate, l-lactate, and d-lactate, tested for significant differences with one-way Anova with subsequent Tukey post-hoc test. *Indicate significant difference to all others: a significant difference to glycolate. c Dependence of Rc(l)-2-HAOX4 activity on the pH in the presence of glycolate as substrate. d Saturation kinetic of Rc(l)-2-HAOX4 using glycolate as substrate. e Kinetic parameters of Rc(l)-2-HAOX4 and AtGOX1 measured in parallel using glycolate as substrate. n = 3, ± SE; R2 coefficient of determination
Activity measurements using different 2-hydroxy acids as substrates at pH 7.5, the pH optimum of all three AtGOX (Engqvist et al. 2015), demonstrate that Rc(l)-2-HAOX4 has the highest activity with glycolate as substrate (100%), followed by l-lactate (25%). In line with the strict stereospecificity of (l)-2-HAOX (Maurino and Engqvist 2015), Rc(l)-2-HAOX4 does not use d-lactate (Fig. 3b). In summary, the substrate specificities of Rc(l)-2-HAOX4 are characteristic of an (l)-2-HAOX with GOX activity (Esser et al. 2014; Engqvist et al. 2015; Maurino and Engqvist 2015). The pH-dependent activity profile of Rc(l)-2-HAOX4 determined using glycolate follows a bell-shaped curve with a pH optimum of 7.5 (Fig. 3c). Further kinetic characterization of recombinant Rc(l)-2-HAOX4 conducted with the preferred substrate glycolate at the optimum pH indicated a Michaelis–Menten response (Fig. 3d). Rc(l)-2-HAOX4 exhibits a high affinity for glycolate (Km = 0.24 mM) and a high catalytic rate (kcat = 60.1 min−1), and it thus possesses a high catalytic efficiency (kcat/Km = 255 min−1 mM−1). These parameters are highly similar to those of the photorespiratory AtGOX1, which we analyzed in parallel (Fig. 3e and Fig. S1).
Our phylogenetic, expressional, and biochemical analyses strongly indicate that the genome of Ricinus possesses a single copy gene encoding a GOX that is found in heterotrophic and autotrophic organs, and has comparable biochemical features to the photorespiratory AtGOX. Thus, in the following we rename Rc(l)-2-HAOX4 to RcGOX.
The photorespiratory metabolite glycolate is abundant in Ricinus heterotrophic organs
To understand the need for GOX activity in heterotrophic organs of Ricinus, we performed a comparative metabolite profile analysis in leaf, roots, dry seeds, and endosperm and embryo of germinated seeds (Fig. 4 and Table S2). A general profile of amino acids shows that their abundance is highest in the embryonal tissue of germinating seeds, followed by the endosperm of germinating seeds (Fig. 4a); this pattern reflects the high metabolic activity of these fast-growing tissues. The highly abundant amino acids are likely derived from the metabolism of seed storage proteins or through de novo synthesis, as their contents in dry seeds are particularly low (Fig. 4a). Only shikimate, the precursor of the aromatic amino acids, shows a tenfold higher relative amount in leaf than in the heterotrophic organs (Fig. 4a), fitting the fact that the plant shikimate pathway is largely active in chloroplasts (Richards et al. 2006).
Relative metabolite levels in different organs of Ricinus communis. Relative content normalized per dry weight (DW) of amino acids (a) and photorespiratory and TCA metabolites and free sugars (b) determined by GC/MS. For ease of comparison, the highest relative content measured for each metabolite was set to 1.00 (see Suppl. Table S1 for normalized peak areas). n = 3; ± SE; S dry seed; gE embryo of germinated seed; ES endosperm of germinated seed; L leaf; R root. Values were tested for significant differences with two-way Anova with subsequent Tukey post-hoc test. *Indicate significant difference to all others: a = significant difference to S, b = significant difference to gE, c = significant difference to ES, d = significant difference to L and e = significant difference to R
We found that relative amounts of TCA cycle intermediates are highest in the root (Fig. 4b), indicating high mitochondrial respiratory activity in this organ. The TCA cycle intermediates show a comparable abundance in leaves and the embryo of germinating seeds, whereas the abundances in dry seeds and the endosperm of germinating seeds are relatively low. The soluble sugars, such as fructose, glucose, sucrose, and xylose show their highest abundances in roots and the embryo of germinating seeds (Fig. 4b).
The photorespiratory metabolites show different profiles in the examined organs (Fig. 4b). Glycerate is predominantly found in the leaf and might derive mainly from photorespiration. As in the case of most amino acids, glycine and serine relative contents are highest in the embryo and endosperm of germinating seeds. Glycolate is predominantly found in roots followed by the embryo of germinating seeds and leaves. Only small amounts of glycolate are present in dry seeds. The presence of glycolate in heterotrophic tissues is not exclusive of Ricinus, as high relative levels of glycolate were also measured in roots of Arabidopsis thaliana (Engqvist et al. 2015).
Discussion
Different physiological functions of RcGOX in autotrophic and heterotorphic tissues
Our phylogenetic, expressional, and biochemical analyses indicate that the genome of Ricinus possesses a single copy gene, Rc(l)-2-HAOX4, encoding a GOX. RcGOX is highly active in autotrophic and heterotrophic organs, where it most likely serves different functions depending on the physiology of the organs.
In plant metabolism, the known origin and fate of glycolate is found so far within the photorespiratory pathway. In photosynthetic tissues, the sequential action of Rubisco and 2-PG phosphatase (PGLP) is the major pathway for the formation of glycolate in chloroplasts (Fig. 5) and GOX is involved in its detoxification in the peroxisomes (Zelitch et al. 2009). Thus, the function of RcGOX in leaves is the same as that of Arabidopsis GOX1/GOX2: it oxidizes glycolate into glyoxylate as part of the photorespiratory pathway.
Putative route for respiration of glycolate metabolism to feed mitochondrial demands for NADPH and folates for purine synthesis in fast-growing heterotrophic tissue. 1: Ribulose-1,5-bisphosphat-carboxylase/-oxygenase (RubisCO); 2: phosphoglycolate phosphatase (PGLP); 3: glyoxylate reductase (GLYR); 4: glycolate oxidase (GOX); 5: glutamate:glyoxylate aminotransferase (GGAT)/serine:glyoxylate aminotransferase (SGAT); 6: serine hydroxymethyl transferase (SHMT); 7: glycine decarboxylase complex (GDC); 8: methyltetrahydrofolate dehydrogenase/5,10-methenyltetrahydrofolate cyclohydrolase (MTHF); 9: citrate synthase (CYS); 10: aconitase (ACO); 11: isocitrate lyase (ICL); 12: malate synthase (MSY); 13: malate dehydrogenase (MDH)
In heterotrophic organs RcGOX is most probably involved in the respiration of glycolate for special demands of those tissues. Our results indicate the presence of a substantial amount of glycolate in the embryo during germination, which hints to the production of this 2-hydroxy acid in this non-photosynthetic tissue. Other heterotrophic organs, such as root, may produce or import glycolate from autotrophic sources. Glycolate could be produced through the action of Rubisco and PGLP in heterotrophic tissues, as the small and large subunits of Rubisco are present in those tissues in Arabidopsis (Baerenfaller et al. 2008). Alternatively, glycolate can be produced through the action of cytosolic glyoxylate reductase (Hoover 2007; Simpson et al. 2008; Allan et al. 2009). In this case, glyoxylate would arise through the glyoxylate cycle, which is highly active in germinating oilseeds (Fig. 5) (Eastmond and Graham 2001; Kunze et al. 2006). In this scenario, a fraction of glyoxylate produced in the glyoxylate cycle likely escape to the cytosol, where glyoxylate reductase converts it into glycolate (Fig. 5). Glycolate is most probably transported to the peroxisome and converted back to glyoxylate by the action of GOX (Fig. 5). With this shunt, glyoxylate produced by GOX most probably enter a different peroxisomal metabolon separated from the glyoxylate cycle, which would subsequently enable channeling of glyoxylate to a different downstream part of the metabolic pathway (Fig. 5) (Kunze and Hartig 2013). This theory is strengthened by the fact that the expression of photorespiratory enzymes in heterotrophic tissues as well as the transfer of metabolites across the peroxisomal membrane during the glyoxylate cycle has been already acknowledged (Courtois-Verniquet and Douce 1993; Kunze et al. 2006; Pracharoenwattana et al. 2007; Nunes-Nesi et al. 2014).
Glycolate respiration most probably fill the demands for mitochondrial NADPH and folate in fast growing heterotrophic organs: a hypothesis
The abundance of glycolate and the presence of a highly active GOX in the embryo and endosperm of germinating seeds and the root of Ricinus indicates an important role of glycolate metabolism in those heterotrophic organs. Feeding experiments using endosperm of Ricinus and radiolabeled glycolate indicated that glycolate is rapidly converted into glyoxylate, glycine, serine, and carbon dioxide (Cossins and Sinha 1967). Glyoxylate directly fed to diverse heterotrophic organs is also rapidly metabolized to serine and glycine (Sinha and Cossins 1965). In agreement with these findings, our metabolite profile indicated large amounts of glycine and serine specifically in the embryo and endosperm of germinating seeds. These amino acids can be produced from glyoxylate through the action of glutamate:glyoxylate aminotransferase/serine:glyoxylate aminotransferase, glycine decarboxylase, and serine hydroxymethyltransferase (Fig. 5).
In addition to amino acid synthesis, the action of the glycine cleavage system is an essential source of one-carbon units (Fig. 5) (Hanson and Roje 2001). In plants, folate metabolism contributes to NADPH production through the conversion of 5,10-methylene-THF to 5,10-methenyl-THF by 5,10-methylene-THF dehydrogenase (Gorelova et al. 2017). In animal cells, serine was also shown to be the major carbon sources for the formation of 10-formyl-THF, the further metabolization of which makes an important contribution to the production of mitochondrial NADPH (Fan et al. 2014). In animal and plant cells, NADPH can be further used in the reduction of glutathione, one of the most important reactive oxygen species (ROS) scavengers (Fan et al. 2014; Gorelova et al. 2017). Controlled levels of ROS are necessary to regulate specific processes during plant cell growth, such as germination (Yazdanpanah et al. 2018), root hair growth (Foreman et al. 2003) and leaf cell expansion (Rodriguez et al. 2002). Thus, folate metabolism can contribute to NADPH production needed for glutathione reduction to maintain ROS levels especially in fast-growing cells. Furthermore, 10-formyl-THF can be used for de novo purine synthesis feeding the demands of nucleotide metabolism in fast-growing embryonic tissue (Hanson and Roje 2001).
In Arabidopsis the phosphorylated pathway of serine biosynthesis is essential to sustain root growth and embryo development (Munoz-Bertomeu et al. 2013; Cascales-Minana et al. 2013). This pathway likely provides the serine involved in folate metabolism as has been demonstrated in mammals (Fan et al. 2014). Nevertheless, it cannot be ruled out that in other plant species there exist a special necessity of strengthening folate metabolism for specific metabolic demands of heterotrophic organs. The results obtained in our work together with existing literature let us propose a hypothesis for a likely function of RcGOX in Ricinus heterotrophic organs (Fig. 5). In this model, glycolate is channeled through RcGOX to fill the NADPH and purine pools. Glycolate is converted by RcGOX into glyoxylate, which is used for the synthesis of serine and glycine. The metabolism of these amino acids likely produces 5,10-methylene-THF, which further transformations produce NADPH and purines (Fig. 5). In summary, we suggest that in heterotrophic organs of Ricinus RcGOX is likely involved in the production of serine to feed the folate pathway for special demands of those tissues; a proposal that deserves further investigations.
Author contributions statement
VGM conceived and led the project. VGM and JS designed the work. JS, MH and DM produced data. JS, MH, NL, and VGM contributed to writing the manuscript and generation of the figures.
Abbreviations
- GOX:
-
Glycolate oxidase
- (l)-2-HAOX:
-
L-2-hydroxy acid oxidase
- lHAOX:
-
Long-chain 2-hydroxy acid oxidases
References
Allan WL, Clark SM, Hoover GJ, Shelp BJ (2009) Role of plant glyoxylate reductases during stress: a hypothesis. Biochem J 423(1):15–22. https://doi.org/10.1042/BJ20090826
Baerenfaller K, Grossmann J, Grobei MA, Hull R, Hirsch-Hoffmann M, Yalovsky S, Zimmermann P, Grossniklaus U, Gruissem W, Baginsky S (2008) Genome-scale proteomics reveals Arabidopsis thaliana gene models and proteome dynamics. Science 320(5878):938–941. https://doi.org/10.1126/science.1157956
Belmouden A, Lederer F (1996) The role of a beta barrel loop 4 extension in modulating the physical and functional properties of long-chain 2-hydroxy-acid oxidase isozymes. Eur J Biochem 238(3):790–798
Brown AP, Kroon JT, Swarbreck D, Febrer M, Larson TR, Graham IA, Caccamo M, Slabas AR (2012) Tissue-specific whole transcriptome sequencing in castor, directed at understanding triacylglycerol lipid biosynthetic pathways. PLoS ONE 7(2):e30100. https://doi.org/10.1371/journal.pone.0030100
Cagliari A, Margis-Pinheiro M, Loss G, Mastroberti AA, de Araujo Mariath JE, Margis R (2010) Identification and expression analysis of castor bean (Ricinus communis) genes encoding enzymes from the triacylglycerol biosynthesis pathway. Plant Sci 179(5):499–509. https://doi.org/10.1016/j.plantsci.2010.07.015
Cascales-Minana B, Munoz-Bertomeu J, Flores-Tornero M, Anoman AD, Pertusa J, Alaiz M, Osorio S, Fernie AR, Segura J, Ros R (2013) The phosphorylated pathway of serine biosynthesis is essential both for male gametophyte and embryo development and for root growth in Arabidopsis. Plant Cell 25(6):2084–2101. https://doi.org/10.1105/tpc.113.112359
Cossins EA, Sinha SK (1967) Studies of glycollate utilization and some associated enzymes of C1 metabolism in the endosperm of Ricinus communis L. J Exp Bot 18:215–228. https://doi.org/10.1093/jxb/18.2.215
Courtois-Verniquet F, Douce R (1993) Lack of aconitase in glyoxysomes and peroxisomes. Biochem J 294(Pt 1):103–107. https://doi.org/10.1042/bj2940103
Dellero Y, Jossier M, Schmitz J, Maurino VG, Hodges M (2016) Photorespiratory glycolate-glyoxylate metabolism. J Exp Bot 67(10):3041–3052. https://doi.org/10.1093/jxb/erw090
Eastmond PJ, Graham IA (2001) Re-examining the role of the glyoxylate cycle in oilseeds. Trends Plant Sci 6(2):72–78
Engqvist MK, Schmitz J, Gertzmann A, Florian A, Jaspert N, Arif M, Balazadeh S, Mueller-Roeber B, Fernie AR, Maurino VG (2015) GLYCOLATE OXIDASE3, a glycolate oxidase homolog of yeast l-lactate cytochrome c oxidoreductase, supports l-lactate oxidation in roots of Arabidopsis. Plant Physiol 169(2):1042–1061. https://doi.org/10.1104/pp.15.01003
Esser C, Kuhn A, Groth G, Lercher MJ, Maurino VG (2014) Plant and animal glycolate oxidases have a common eukaryotic ancestor and convergently duplicated to evolve long-chain 2-hydroxy acid oxidases. Mol Biol Evol 31(5):1089–1101. https://doi.org/10.1093/molbev/msu041
Fan J, Ye J, Kamphorst JJ, Shlomi T, Thompson CB, Rabinowitz JD (2014) Quantitative flux analysis reveals folate-dependent NADPH production. Nature 510(7504):298–302. https://doi.org/10.1038/nature13236
Fiehn O (2007) Validated high quality automated metabolome analysis of Arabidopsis thaliana leaf disks. In: Nikolau BJ, Wurtele ES (eds) Concepts in plant metabolomics. Springer, Dordrecht, pp 1–18
Foreman J, Demidchik V, Bothwell JH, Mylona P, Miedema H, Torres MA, Linstead P, Costa S, Brownlee C, Jones JD, Davies JM, Dolan L (2003) Reactive oxygen species produced by NADPH oxidase regulate plant cell growth. Nature 422(6930):442–446. https://doi.org/10.1038/nature01485
Forner J, Binder S (2007) The red fluorescent protein eqFP611: application in subcellular localization studies in higher plants. BMC Plant Biol 7:28. https://doi.org/10.1186/1471-2229-7-28
Gibson DG, Young L, Chuang RY, Venter JC, Hutchison CA, Smith HO (2009) Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat Methods 6(5):343–345
Gorelova V, De Lepeleire J, Van Daele J, Pluim D, Mei C, Cuypers A, Leroux O, Rebeille F, Schellens JHM, Blancquaert D, Stove CP, Van Der Straeten D (2017) Dihydrofolate reductase/thymidylate synthase fine-tunes the folate status and controls redox homeostasis in plants. Plant Cell 29(11):2831–2853. https://doi.org/10.1105/tpc.17.00433
Greenwood JS, Bewley JD (1982) Seed development in Ricinus communis (castor bean). 1. Descriptive morphology. Can J Bot 60(9):1751–1760. https://doi.org/10.1139/b82-222
Hanson AD, Roje S (2001) One-carbon metabolism in higher plants. Annu Rev Plant Physiol Plant Mol Biol 52:119–137. https://doi.org/10.1146/annurev.arplant.52.1.119
Hoover GJ, Van Cauwenberghe OR, Breitkreuz KE, Clark SM, Merrill AR, Shelp BJ (2007) Characteristics of an Arabidopsis glyoxylate reductase: general biochemical properties and substrate specificity for the recombinant protein, and developmental expression and implications for glyoxylate and succinic semialdehyde metabolism in planta. Can J Bot 85:883–895
Igamberdiev AU, Bykova NV, Gardestrom P (1997) Involvement of cyanide-resistant and rotenone-insensitive pathways of mitochondrial electron transport during oxidation of glycine in higher plants. FEBS Lett 412(2):265–269
Jones JM, Morrell JC, Gould SJ (2000) Identification and characterization of HAOX1, HAOX2, and HAOX3, three human peroxisomal 2-hydroxy acid oxidases. J Biol Chem 275(17):12590–12597
Katoh K, Misawa K, Kuma K, Miyata T (2002) MAFFT: a novel method for rapid multiple sequence alignment based on fast Fourier transform. Nucleic Acids Res 30(14):3059–3066
Kohler SA, Menotti E, Kuhn LC (1999) Molecular cloning of mouse glycolate oxidase. High evolutionary conservation and presence of an iron-responsive element-like sequence in the mRNA. J Biol Chem 274(4):2401–2407
Kumar S, Stecher G, Tamura K (2016) MEGA7: molecular evolutionary genetics analysis version 7.0 for bigger datasets. Mol Biol Evol 33(7):1870–1874. https://doi.org/10.1093/molbev/msw054
Kunze M, Hartig A (2013) Permeability of the peroxisomal membrane: lessons from the glyoxylate cycle. Front Physiol 4:204. https://doi.org/10.3389/fphys.2013.00204
Kunze M, Pracharoenwattana I, Smith SM, Hartig A (2006) A central role for the peroxisomal membrane in glyoxylate cycle function. Biochim Biophys Acta 1763(12):1441–1452. https://doi.org/10.1016/j.bbamcr.2006.09.009
Laemmli UK (1970) Cleavage of structural proteins during the assembly of the head of bacteriophage T4. Nature 227(5259):680–685
Le SQ, Gascuel O (2008) An improved general amino acid replacement matrix. Mol Biol Evol 25(7):1307–1320. https://doi.org/10.1093/molbev/msn067
Leliaert F, Smith DR, Moreau H, Herron MD, Verbruggen H, Delwiche CF, De Clerck O (2012) Phylogeny and molecular evolution of the green algae. Crit Rev Plant Sci 31(1):1–46. https://doi.org/10.1080/07352689.2011.615705
Lernmark U, Henricson D, Wigge B, Gardestrom P (1991) Glycine oxidation in mitochondria isolated from light grown and etiolated plant-tissue. Physiol Plant 82(3):339–344
Lingner T, Kataya AR, Antonicelli GE, Benichou A, Nilssen K, Chen XY, Siemsen T, Morgenstern B, Meinicke P, Reumann S (2011) Identification of novel plant peroxisomal targeting signals by a combination of machine learning methods and in vivo subcellular targeting analyses. Plant Cell 23(4):1556–1572. https://doi.org/10.1105/tpc.111.084095
Logemann J, Schell J, Willmitzer L (1987) Improved method for the isolation of RNA from plant tissues. Anal Biochem 163(1):16–20
Landrum M, Smertenko A, Edwards R, Hussey PJ, Steel PG (2010) BODIPY probes to study peroxisome dynamics in vivo. Plant J 62(3):529–538
Maurino VG, Engqvist MK (2015) 2-Hydroxy acids in plant metabolism. The Arabidopsis book/Am Soc Plant Biol 13:e0182. https://doi.org/10.1199/tab.0182
Maurino VG, Peterhansel C (2010) Photorespiration: current status and approaches for metabolic engineering. Curr Opin Plant Biol 13(3):249–256. https://doi.org/10.1016/j.pbi.2010.01.006
Munoz-Bertomeu J, Anoman A, Flores-Tornero M, Toujani W, Rosa-Tellez S, Fernie AR, Roje S, Segura J, Ros R (2013) The essential role of the phosphorylated pathway of serine biosynthesis in Arabidopsis. Plant Signal Behav 8(11):e27104. https://doi.org/10.4161/psb.27104
Nunes-Nesi A, Florian A, Howden A, Jahnke K, Timm S, Bauwe H, Sweetlove L, Fernie AR (2014) Is there a metabolic requirement for photorespiratory enzyme activities in heterotrophic tissues? Mol Plant 7:248–251. https://doi.org/10.1093/mp/sst111
Pracharoenwattana I, Cornah JE, Smith SM (2007) Arabidopsis peroxisomal malate dehydrogenase functions in beta-oxidation but not in the glyoxylate cycle. Plant J 50(3):381–390. https://doi.org/10.1111/j.1365-313X.2007.03055.x
Reumann S, Babujee L, Ma C, Wienkoop S, Siemsen T, Antonicelli GE, Rasche N, Luder F, Weckwerth W, Jahn O (2007) Proteome analysis of Arabidopsis leaf peroxisomes reveals novel targeting peptides, metabolic pathways, and defense mechanisms. Plant Cell 19(10):3170–3193. https://doi.org/10.1105/tpc.107.050989
Reumann S, Buchwald D, Lingner T (2012) PredPlantPTS1: a Web Server for the prediction of plant peroxisomal proteins. Front Plant Sci 3:194. https://doi.org/10.3389/fpls.2012.00194
Richards TA, Dacks JB, Campbell SA, Blanchard JL, Foster PG, McLeod R, Roberts CW (2006) Evolutionary origins of the eukaryotic shikimate pathway: gene fusions, horizontal gene transfer, and endosymbiotic replacements. Eukaryot Cell 5(9):1517–1531. https://doi.org/10.1128/EC.00106-06
Rodriguez AA, Grunberg KA, Taleisnik EL (2002) Reactive oxygen species in the elongation zone of maize leaves are necessary for leaf extension. Plant Physiol 129(4):1627–1632. https://doi.org/10.1104/pp.001222
Ryan P, Delhaize E, Jones D (2001) Function and mechanism of organic anion exudation from plant roots. Annu Rev Plant Physiol Plant Mol Biol 52:527–560. https://doi.org/10.1146/annurev.arplant.52.1.527
Schaffner W, Weissmann C (1973) A rapid, sensitive, and specific method for the determination of protein in dilute solution. Anal Biochem 56(2):502–514
Schindelin J, Arganda-Carreras I, Frise E, Kaynig V, Longair M, Pietzsch T, Preibisch S, Rueden C, Saalfeld S, Schmid B, Tinevez JY, White DJ, Hartenstein V, Eliceiri K, Tomancak P, Cardona A (2012) Fiji: an open-source platform for biological-image analysis. Nat Methods 9(7):676–682. https://doi.org/10.1038/nmeth.2019
Schmitz J, Dittmar IC, Brockmann JD, Schmidt M, Hudig M, Rossoni AW, Maurino VG (2017) Defense against reactive carbonyl species involves at least three subcellular compartments where individual components of the system respond to cellular sugar status. Plant Cell 29(12):3234–3254. https://doi.org/10.1105/tpc.17.00258
Schmitz J, Srikanth NV, Hudig M, Poschmann G, Lercher MJ, Maurino VG (2017) The ancestors of diatoms evolved a unique mitochondrial dehydrogenase to oxidize photorespiratory glycolate. Photosynth Res 132(2):183–196. https://doi.org/10.1007/s11120-017-0355-1
Sela I, Ashkenazy H, Katoh K, Pupko T (2015) GUIDANCE2: accurate detection of unreliable alignment regions accounting for the uncertainty of multiple parameters. Nucleic Acids Res 43(W1):W7-14. https://doi.org/10.1093/nar/gkv318
Simpson JP, Di Leo R, Dhanoa PK, Allan WL, Makhmoudova A, Clark SM, Hoover GJ, Mullen RT, Shelp BJ (2008) Identification and characterization of a plastid-localized Arabidopsis glyoxylate reductase isoform: comparison with a cytosolic isoform and implications for cellular redox homeostasis and aldehyde detoxification. J Exp Bot 59(9):2545–2554. https://doi.org/10.1093/jxb/ern123
Sinha SK, Cossins EA (1965) The importance of glyoxylate in amino acid biosynthesis in plants. Biochem J 96:254–261
Tanner WH, Beevers H (1965) Glycolic acid oxidase in ccastor bean endosperm. Plant Physiol 40(6):971–976
Verhoeven NM, Schor DS, Roe CR, Wanders RJ, Jakobs C (1997) Phytanic acid alpha-oxidation in peroxisomal disorders: studies in cultured human fibroblasts. Biochim Biophys Acta 1361(3):281–286
Verhoeven NM, Jakobs C, ten Brink HJ, Wanders RJ, Roe CR (1998) Studies on the oxidation of phytanic acid and pristanic acid in human fibroblasts by acylcarnitine analysis. J Inherit Metab Dis 21(7):753–760
Waadt R, Kudla J (2008) In planta visualization of protein interactions using bimolecular fluorescence complementation (BiFC). CSH Protoc. https://doi.org/10.1101/pdb.prot4995
Weckwerth W, Wenzel K, Fiehn O (2004) Process for the integrated extraction, identification and quantification of metabolites, proteins and RNA to reveal their co-regulation in biochemical networks. Proteomics 4(1):78–83. https://doi.org/10.1002/pmic.200200500
Williams E, Cregeen D, Rumsby G (2000) Identification and expression of a cDNA for human glycolate oxidase. Biochim Biophys Acta 1493(1–2):246–248
Yazdanpanah F, Maurino VG, Mettler-Altmann T, Buijs G, Bailly M, Jashni MK, Willems L, Sergeeva LI, Rajjou L, Hilhorst HWM, Bentsink L (2018) NADP-MALIC ENZYME 1 affects germination after seed storage in Arabidopsis thaliana. Plant Cell Physiol 60(2):318–328. https://doi.org/10.1093/pcp/pcy213
Ye J, Coulouris G, Zaretskaya I, Cutcutache I, Rozen S, Madden TL (2012) Primer-BLAST: a tool to design target-specific primers for polymerase chain reaction. BMC Bioinform 13:134. https://doi.org/10.1186/1471-2105-13-134
Zelitch I, Schultes NP, Peterson RB, Brown P, Brutnell TP (2009) High glycolate oxidase activity is required for survival of maize in normal air. Plant Physiol 149:195–204. https://doi.org/10.1104/pp.108.128439
Acknowledgements
This work was funded by grants of the Deutsche Forschungsgemeinschaft (DFG, German Research Foundation) under Germany´s Excellence Strategy – EXC 2048/1 – Project ID: 390686111 and EXC 1028 to VGM. The authors thank the Center for Advanced Imaging (Düsseldorf, Germany) for technical assistance with fluorescence microscopy.
Funding
Open Access funding enabled and organized by Projekt DEAL.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Additional information
Communicated by Dorothea Bartels.
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
425_2020_3504_MOESM1_ESM.docx
Supplementary Fig. S1 Purification and kinetic measurements of AtGOX1. a Coomassie-stained SDS-polyacrylamide gel of different steps during the isolation of recombinant AtGOX1. Crude protein extract of E. coli before (lane 1) and after (lane 2) induction of expression with 1 mM IPTG, non-soluble fraction (lane 3) and soluble fraction (lane 4) after cell disruption, flow-through of washing steps (lanes 5-7), and eluted recombinant AtGOX1 (lanes 8 and 9; 43 kDa); M=Molecular mass standards. b Dependence of AtGOX1 activity on the pH in the presence of glycolate as substrate. c Saturation kinetic of AtGOX1 using glycolate as substrate. R2 = coefficient of determination (DOCX 3151 KB)
425_2020_3504_MOESM3_ESM.xlsx
Supplementary Table S2 Normalized peak areas of metabolites normalized per DW and FW measured by GC/MS including results of two-way Anova with subsequent Tukey post-hoc test (XLSX 35 KB)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schmitz, J., Hüdig, M., Meier, D. et al. The genome of Ricinus communis encodes a single glycolate oxidase with different functions in photosynthetic and heterotrophic organs. Planta 252, 100 (2020). https://doi.org/10.1007/s00425-020-03504-0
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s00425-020-03504-0