Abstract
Pectinaceous agricultural residues rich in d-galacturonic acid (d-GalA), such as sugar beet pulp, are considered as promising feedstocks for waste-to-value conversions. Aspergillus niger is known for its strong pectinolytic activity. However, while specialized strains for production of citric acid or proteins are well characterized, this is not the case for the production of pectinases. We, therefore, systematically compared the pectinolytic capabilities of six A. niger strains (ATCC 1015, ATCC 11414, NRRL 3122, CBS 513.88, NRRL 3, and N402) using controlled batch cultivations in stirred-tank bioreactors. A. niger ATCC 11414 showed the highest polygalacturonase activity, specific protein secretion, and a suitable morphology. Furthermore, d-GalA release from sugar beet pulp was 75% higher compared to the standard lab strain A. niger N402. Our study, therefore, presents a robust initial strain selection to guide future process improvement of d-GalA production from agricultural residues and identifies a high-performance base strain for further genetic optimizations.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Global academic and industrial efforts to improve the sustainability of industrial processes for the modern bio-economy have sparked interest in the utilization of feedstocks that are economically viable, non-food grade, and do not compete with food resources [6, 23, 26, 44, 49, 51, 58]. As a result, agricultural waste streams have gained momentum in the recent years (reviewed by Amoah et al. [3]). Notably, downstream fermentation products derived from pectin-rich biomass are met with industrial interest by the plastics, cosmetics, and food industries, as recently reviewed by Kuivanen et al. [28], Schmitz et al. [46], and Richard and Hilditch [43].
Besides harsh thermo-chemical treatment and hydrolysis approaches [12], fermentation broths of natural pectin-degrading microorganisms containing complex enzyme mixtures can be used to liberate the constituent saccharides, such as d-galacturonic acid (d-GalA), which is the main backbone sugar of pectin. The filamentous saprophytic fungus Aspergillus niger is a well-known microorganism for pectin utilization and depolymerization as well as a well-established industrial workhorse with multiple applications in the production of enzymes, citric acid, and other organic acids [9, 31]. Pectinases from A. niger contribute to a global multi-billion dollar market for biomass-degrading enzymes [9, 31, 47] with applications ranging from fruit, vegetable, and juice processing to textile and paper treatment [22] as well as saccharification for bioethanol production [11, 50, 56]. However, with today’s perception of d-GalA shifting from an inevitable component of complex biomass feedstocks to a target product for subsequent fermentations, more versatile strains that can achieve higher d-GalA yields are needed to extend the range of commercially available saccharification enzymes, as outlined in a recent white paper on the current challenges of research on filamentous fungi in the context of a sustainable bio-economy [31].
To enable efficient industrial-scale applications, ideal base strains need to be identified for specific tasks. For A. niger, several lineages have been identified and adapted for specific purposes. The most cited lineages of A. niger encompass three main clades, namely: (i) strains adapted for easy handling and genetic manipulation in the laboratory environment, which are based on A. niger NRRL 3 (CBS 120.49, ATCC 9029); (ii) strains for improved citric acid production, based on A. niger ATCC 1015 (NRRL 328, CBS 113.46); and (iii) strains for protein production and secretion, based on A. niger NRRL 3122 (CBS 115989; ATCC 22343) (Fig. 1). To the best of our knowledge, no thorough comparison of pectinase activity between these available and highly cited A. niger strains has been conducted so far. While the commercial availability of A. niger pectinase cocktails indicates that the industry has established strains for pectinase production, almost no information on their specific origin is publically accessible. In the academic field, numerous studies on optimization of fermentation conditions for pectinase production with various strains have been published (e.g., [1, 18, 42]). However, these do not allow for direct performance comparison of individual strains due to varying study designs and fermentation conditions. Furthermore, thorough comparisons of different strains under reproducible conditions are scarce in the literature and compromised by limited morphology control (due to execution in shake flasks, for example, instead of controlled stirred-tank bioreactors) or poor description of strain origins (e.g., [17, 25]).
Highly relevant lineages of A. niger and their primary applications. Schematic overview listing selected but representative members of openly available and related A. niger strains based on phylogeny [55]. Depicted are commonly used lab strains (green, right) vs. the industrially adapted strains (blue, left) including the citric acid producer clade (light blue) and the more distantly related enzyme producers (dark blue). Yellow signs indicate instances of UV mutagenesis, grey gear wheels symbolize targeted genomic engineering steps, and red boxes indicate the strains used in this study (color figure online)
As outlined in Fig. 1, ancestral strains have been specifically isolated due to their improved citric acid production characteristics (A. niger ATCC 1015) or extracellular glucoamylase activity (A. niger NRRL 3122), respectively, and further optimized via sub-culture isolation [39] or UV mutagenesis [4, 53]. Similarly, multiple successive rounds of UV mutagenesis on the original wild-type (WT) isolate A. niger NRRL 3 [7] have resulted in a short conidiophore phenotype in strain A. niger N402 [8] and the derived strain most commonly used for laboratory-based genetic analysis, A. niger AB4.1, containing the pyrG auxotrophic marker [19, 52]. However, those mutagenic treatments likely gave rise to additional background mutations influencing gene regulation and impacting diverse phenotypic traits. Additional targeted engineering steps for improved homologous recombination (A. niger MA70.15) [32] or the introduction of auxotrophic markers [16, 35] have further expanded the genetic toolbox in the laboratory. At the same time, however, these may induce stress or alter intracellular regulation and, hence, divert these strains further from their wild-type physiological constitution [41], which might impair fungal productivity and, hence, should be considered in strain selection.
From a genomics perspective, genomic sequences are openly available for A. niger strains CBS 513.88 [38], ATCC 1015 [4], and NRRL 3 [2]—the latter with a recent advance in annotation [45]—as well as for three additional isolates [55]. Comparison of intra-species genomic data revealed the cause of some of the observed phenotypic differences between the strains, such as overproduction of glucoamylase in A. niger CBS 513.88, which was due to an additional glucoamylase gene acquired by horizontal gene transfer [4, 55]. Transcriptional profiling, however, also indicated that the regulatory networks between different strains are already highly divergent and cannot be explained purely by genomic observations [4]. Additionally, fungal productivity for different purposes depends on morphology control in submerged cultures [10, 24, 54, 57].
Accordingly, none of the current data sets provide enough information to predict superiority of any available strain for the production of pectinases.
The importance of pectin and pectin-derived sugars, as well as the predominance of A. niger in pectinase production processes, however, warrant a systematic comparison of strains under controlled and highly reproducible conditions to identify efficient host strains for larger scale pectinase production. Therefore, a total of six strains (red boxes in Fig. 1) were selected for comparison of endogenous pectinase activity based on five key prerequisites: (i) relevance to the field based on the number of publications using these strains, (ii) availability of the genomic sequence (or that of a very closely related strain) as a premise for successive genetic optimization, (iii) absence of auxotrophic markers, to avoid phenotypic differences due to mutations in central metabolism, (iv) absence of any targeted engineering of elements regulating pectinase expression, to avoid distortion of the underlying endogenous pectinase capacity, and (v) classification as biosafety level 1 to allow for universal handling.
By obtaining data on sporulation efficiency, total protein secretion, total and endo-specific polygalacturonase (PGase) activity, as well as morphology in submerged culture, this study provides essential insights into selection of suitable base strains for pectinase production. Working with controlled stirred-tank bioreactor fermentations after the initial pre-selection, we applied a robust and reproducible methodology rarely employed in phenotypic comparisons of fungal strains, resulting in the identification of a superior A. niger strain and potential chassis for additional genetic optimization to boost d-GalA release from complex pectinaceous residues.
Materials and methods
Strains, inoculum preparation, and cultivation medium
A. niger strains ATCC 1015, ATCC 11414, and NRRL 3122 were obtained from the NRRL collection, and NRRL 3, N402, and CBS 513.88 were obtained from the group of Arthur Ram at Leiden University. Fungal spores were grown on 39 g L−1 potato extract glucose agar (Carl Roth GmbH + Co. KG, Karlsruhe, Germany) supplemented with 10 g L−1 yeast extract and 1 × trace elements solution, referred to as rich complete medium. After 120 h at 30 °C, spores were harvested using sterile 0.89% NaCl solution with 0.05% Tween 80.
All experiments involving submerged fungal cultivation were carried out in 2% (w/v) pectin minimal medium containing (L−1): 20.0 g pectin C, 6.0 g NaNO3, 1.5 g KH2PO4, 0.5 g KCl, 0.5 g MgSO4 · 7 H2O, 1 mL trace-element solution, and 1 mL PPG P2000 (antifoam, only if cultivated in stirred tank fermenter). The trace-element solution was prepared as (L−1) 10 g EDTA, 4.4 g ZnSO4·7 H2O, 1.01 g MnCl2·4 H2O, 0.32 g CoCl2·6 H2O, 0.315 g CuSO4·5 H2O, 0.22 g (NH4)6Mo7O24·4 H2O, 1.47 g CaCl2·2 H2O, and 1 g FeSO4·7 H2O [5].
Cultivation conditions
Shake flask cultivation
The fungi were grown in 250 mL flasks without baffles containing 25 mL of the 20 g L−1 pectin minimal medium at 250 min−1 (25 mm shaking throw) and 30 °C for 96 h. The initial pH was set to 4.5 and the medium was inoculated to a spore density of 109 spores L−1. Strains were grown in triplicates. Data were statistically evaluated by applying an analysis of variance (one-way ANOVA) followed by a Tukey’s post hoc test using the software Origin (OriginLab). Differences among the mean activity measurements were calculated at a significance level of 0.05.
Bioreactor cultivation
A 7 L stirred-tank bioreactor equipped with three baffles and three six-blade Rushton turbines (Labfors, Infors-HT, Bottmingen, Switzerland) was used during all cultivations. All processes were performed equally under the following conditions. Three liters of the 20 g L−1 pectin mineral medium was inoculated to 109 spores L−1. Temperature was kept constant at 30 °C. The pH was controlled to a set-point of pH 4.5 by the addition of either 1 M H2SO4 or 3 M KOH. Batch processes were carried out for 86–90 h. To prevent the initial spore loss, the stirred-tank bioreactor was not aerated and only slowly mixed at 250 min−1 (~ 0.130 W L−1 [21]) during the first 6 h of batch cultivations [34]. Afterwards, the stirrer speed was set to 700 min−1 (~ 1.625 W L−1 [21]) and aeration to 0.2 vvm, which was also sufficient to keep the dissolved oxygen concentration above 30% air saturation during all cultivations conducted. Additionally, exit-gas composition (O2, CO2) was monitored (EasyLine, ABB, Zürich, Switzerland).
Biomass dry weight concentration
Biomass dry weight was determined by filtering a known volume thought pre-dried and pre-weighed filter paper (Whatman No. 1 and 5). The collected biomass was dried at 90 °C to constant weight and reweighed. The determination of the biomass dry weight was performed in triplicate and expressed as the mean with standard deviation of the measurements.
Morphological characterization
Microscopic images for morphological characterization were taken with an Axioplan microscope (Carl Zeiss AG, Jena, Germany) at 1.25 × magnification directly after sampling after 9, 12, 19, 36, and 88 h of the batch cultivation. The microscope was equipped with a 3.3-megapixel Axiocam ICc3 microscopy camera (Carl Zeiss AG, Jena, Germany).
Protein concentration of culture supernatant
Protein concentration of the culture supernatant was determined using the Coomassie (Bradford) Protein Assay Kit (Thermo Scientific) according to the manufacturer's specifications. Each sample was diluted with 0.1 M sodium citrate buffer pH 4.5, mixed with the Bradford reagent, and incubated for at least 10 min at room temperature. Afterwards, the absorbance at 595 nm was measured with a multimode microplate reader (Infinite M200, Tecan, Männedorf, Germany). Bovine serum albumin was used as the standard. The determination of the protein concentrations was performed in triplicate and expressed as the mean with standard deviation of the measurements.
Total polygalacturonase activity
The pectinase activity was determined following a miniaturized version of the Fructan Assay Kit protocol (Megazyme, Bray, Ireland) for reducing sugars. 10 µL of the culture supernatant and 10 µL of a 5 g L−1 polygalacturonic acid solution (PGA, buffered in 0.1 M sodium citrate, pH 4.5) were mixed and incubated for 40 min at 30 °C. The released reducing sugar ends were determined using a 4-hydroxybenzhydrazide solution as described in the Megazyme protocol and measured at 410 nm with a multimode microplate reader (Infinite M200, Tecan, Männedorf, Germany). Sample values were blanked against similarly prepared but non-incubated mock samples. One unit of total polygalacturonase activity was defined as the amount of enzyme that catalyzes the formation of one µmol of d-galacturonic acid per minute under the assay conditions. Assaying was performed in triplicate and results were plotted as means with standard deviations of replicate measurements.
Endo-polygalacturonase activity
Endo-polygalacturonase activity was assessed following the protocol of Ortiz [36]. 8 µL of a 5 g L−1 polygalacturonic acid solution (PGA, buffered in 0.1 M sodium acetate, pH 4.5) and 8 µL of A. niger culture supernatant were mixed and incubated for 30 min at 30 °C in a microtiter plate prior to the addition of 40 µL of freshly prepared ruthenium red working solution (Sigma-Aldrich, 1.125 mg mL−1 in ddH2O) and 100 µL of 8 mM sodium hydroxide solution. Samples were spun down at 3200 g for 10 min. 25 µL of supernatant were mixed with 175 µL ddH2O in a 96-well microtiter plate and absorbance was measured at 535 nm on a microplate reader (Infinite M200, Tecan, Männedorf, Germany). Sample values were blanked against similarly prepared but non-incubated mock samples. One enzyme unit was defined as the amount of enzyme required to hydrolyze 1 µg of polygalacturonic acid into smaller fragments unable to precipitate with the dye per minute under the assay conditions. Assaying was performed in triplicate and results were plotted as means with standard deviations of replicate measurements.
Hydrolysis of sugar beet pulp
The hydrolysis of pectin-rich residues was conducted with pre-dried (50 °C) and milled sugar beet press pulp provided by Südzucker AG (Obrigheim, Germany). 10 g of pre-sterilized pulp was mixed with 100 mL of reaction solution. The reaction solution consisted of 90 mL of sterile filtered (0.22 µm) enzyme supernatant of the strains A. niger N402 (172 ± 4 mg L−1 Protein; 20 ± 26 U L−1 total PGase activity) or A. niger ATCC 11414 (111 ± 9 mg L−1 Protein; 1476 ± 32 U L−1 total PGase activity) buffered with 10 mL of 1.0 M sodium acetate (pH 4.5). Hydrolysis was carried out at 180 rpm and 30 °C for 138 h in sterile and closed 250 mL glass bottles (DWK Life Sciences GmbH, Mainz, Germany) to prevent evaporation and contamination. Time-series samples were taken from homogenized hydrolysis mixtures and stored at − 80 °C until further use. Respective amounts of released sugars were plotted against the sampling time points and the second-order polynomial fitting curves were generated.
HPAEC-PAD analysis of the hydrolysis supernatant
Free d-GalA and neutral sugar amounts in hydrolysis samples (diluted 1:4000) were determined on a Dionex ICS 3000 HPAEC-PAD instrument setup with a Dionex AS Autosampler, a Dionex gradient mixer GM-3 (Dionex Corp., Sunnyvale, California, USA) and a CarboPac PA1 standard bore guard column (4 × 50 mm) plus a CarboPac PA1 preparative IC column (4 × 250 mm, both Thermo Fisher Scientific Inc., Waltham, Massachusetts, USA) using a 12.5 min linear gradient of 100–250 mM sodium acetate in 100 mM sodium hydroxide solution (prepared in low total organic carbion deionized water) at 1 mL min−1 flow rate and constant 30 °C elution temperature.
Results and discussion
Shake flask-based pre-selection of A. niger strains for pectinase production
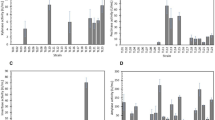
As a first step in the identification of an ideal base strain for pectinase production among the selected A. niger strains, an initial fast and cost-efficient experiment was conducted in small scale using shake flasks, as typically applied in strain screenings (e.g., [40]). Total polygalacturonase (PGase) activity was measured in supernatants of A. niger cultures after 96 h of cultivation in minimal medium supplemented with 2% (w/v) pectin C as carbon source (Fig. 2a). Strains A. niger NRRL 3122 and A. niger CBS 513.88 repeatedly had the lowest total PGase activities of all strains tested (174 ± 41 U L−1 and 345 ± 199 U L−1, respectively), while A. niger NRRL 3 (607 ± 174 U L−1), A. niger N402 (493 ± 233 U L−1), A. niger ATCC 1015 (674 ± 92 U L−1), and A. niger ATCC 11414 (543 ± 246 U L−1) had superior total PGase activities (p < 0.05). Additionally, A. niger NRRL 3122 and A. niger CBS 513.88 gave lower spore densities on rich medium, which was rated as a disadvantage for larger scale liquid culture inoculations (Fig. 2b). Based on the combination of these results, strains A. niger NRRL 3122 and A. niger CBS 513.88 were excluded from further tests.
Polygalacturonase activity and sporulation density of A. niger strains. a Total PGase activity of A. niger culture supernatants of a representative shake flask batch after 96 h of incubation in minimal medium + 2% (w/v) pectin C, plotted as means with sample standard deviations. Different capital letters indicate significant differences within the displayed data groups (p < 0.05) using a one-way ANOVA followed by a Tukey’s post hoc test. Group AB showed no significant differences either to group A and group B. b Sporulation densities of all strains on rich complete medium after 6 days
A. niger CBS 513.88 (and its ancestor A. niger NRRL 3122) differ from A. niger ATCC 1015 and its closely related A. niger NRRL 3 lineages by additional glucoamylase (glaA) genes acquired through horizontal gene transfer, as well as the upregulation of amino acid synthases (for those amino acids overrepresented in the GlaA protein) and their respective tRNAs [4]. Considering their poor performance in this study, however, A. niger CBS 513.88 and A. niger NRRL 3122 do not seem to have a universal advantage over the other tested strains in terms of protein production and secretion per se, but rather a limited one for GlaA expression.
Comparison of pre-selected A. niger strains in submerged stirred-tank bioreactor batch cultivations
A high degree of control over mechanical and physicochemical parameters influencing submerged culture morphology as well as enzyme activity is important for achieving high pectinase activity [1, 14, 54]. In the next step, pectinase production of all strains was, therefore, evaluated in batch processes on a 3 L scale in controlled stirred-tank bioreactor cultivations to perform selection under robust and reproducible conditions (see Figs. 3, 4). To this end, submerged batch cultivations in 2% (w/v) pectin minimal medium were conducted to investigate the pectinolytic properties of the four best performing A. niger strains from the pre-selection. For each batch cultivation, biomass dry weight concentration (BDW), total protein concentration (cProtein), total polygalacturonase, and endo-polygalacturonase (PGase) activity, as well as fungal morphology, were determined, all being highly relevant variables for strain productivity in submerged cultures [29, 54]. Since the level of morphology control is generally higher in stirred-tank bioreactors compared to shake flasks, we see our bioreactor approach as advantageous for in-depth strain comparison after pre-selection [14].
Biomass dry weight concentrations (BDW) and morphology of selected A. niger strains. a–d BDW of A. niger NRRL 3 (a, filled circle), A. niger N402 (b, filled diamond), A. niger ATCC 1015 (c, filled square), and A. niger ATCC 11414 (d, filled triangle) during 90 h submerged batch cultivations in a 3 L stirred-tank bioreactor with 2% pectin minimal medium, plotted as means with sample standard deviations. e Morphological changes of all four A. niger strains throughout the cultivations are shown below. White scale bars indicate 1 mm. Morphology sampling times are indicated by lines in (a–d)
Protein concentrations, total PGase, and endo-PGase activities of pre-selected A. niger strains in 3 L controlled stirred-tank batches. a–d Total secreted protein concentrations (grey) in the culture supernatants of pre-selected A. niger strains in 2% pectin minimal medium over a time course of 90 h, plotted as means with sample standard deviations. e–h Total (black) and endo-PGase activities (white) of culture supernatants throughout the cultivation, plotted as means with sample standard deviations
Biomass dry weight concentrations and morphology
In case of A. niger NRRL 3, BDW rose to a maximum of 2.28 ± 0.12 g L−1 during the first 39 h and slightly decreased about 35% afterwards until the end of the cultivation (1.47 ± 0.01 g L−1) (Fig. 3a). The BDW of the strains A. niger N402 and A. niger ATCC 1015 showed a similar behavior, increasing to 3.65 ± 0.35 g L−1 (A. niger N402) and 3.38 ± 0.07 g L−1 (A. niger ATCC 1015) within 42 h before decreasing by 30% until the end of the cultivation (Fig. 3b, c). The behavior of strain A. niger ATCC 11414 was different. Its BDW peaked only after 48 h (at 2.68 ± 0.03 g L−1), with a drastic decrease of 70% to 0.81 ± 0.02 g L−1 at the end of the observation period (Fig. 3d). The loss of BDW in the cultivations correlates with the dissolved oxygen concentration (DO) and the carbon dioxide fraction in the exit gas measured online. When the BDW loss started, the DO concentration increased and the carbon dioxide fraction in the exit gas decreased, even though there was no change in the aeration of the process, indicating depletion of pectin C as the sole energy source provided (data not shown).
Since the productivity of filamentous fungi in submerged cultivations depends on their morphology [13, 24, 37, 57], this parameter was monitored throughout each submerged cultivation. Figure 3e depicts the morphology of all four strains after 9, 12, 19, 36, and 88 h of the cultivation. After 36 h, all strains had a stable morphology. A. niger NRRL 3 and A. niger ATCC 11414 showed disperse, filamentous growth, A. niger N402 strongly pellet-like growth, and A. niger ATCC 1015 a less dense but still pelleted form of growth. Towards the end of the cultivation, the morphology of A. niger NRRL 3 shifted to a more pelleted structure. Notably, strong yellow pigmentation occurred after 34 h in all culture supernatants with the exception of A. niger ATCC 11414, where only mild pigmentation was observed after 58 h. The characteristic color of the pigment matches the description of Aurasperones, which are well-known side products in many A. niger fermentations [48]. As all four batch cultivations were run under identical conditions, these results indicate significant intra-species differences in physiological regulation among the selected strains that cannot be explained by comparison of their genomic sequences alone [4].
Secreted protein concentrations and PGase activities
Next, the total secreted protein concentration in the supernatants (Fig. 4a–d) as well as the total and endo-specific PGase activities were assessed (Fig. 4e–h). The protein concentration of the cultivations with A. niger NRRL 3, A. niger N402, and A. niger ATCC 1015 increased throughout the cultivation, peaking at 157 ± 10 mg L−1 (90 h), 162 ± 6 mg L−1 (81 h), and at 167 ± 5 mg L−1 (71 h), respectively. Following the observed BDW decrease towards the end of cultivation, A. niger N402 and A. niger ATCC 1015 also displayed a mild decrease in protein concentration. In accordance with its delayed biomass generation (Fig. 3d), A. niger ATCC 11414 showed its maximal protein titer (111 ± 10 mg L−1) only after 87 h (Fig. 4d). This protein concentration was 32% lower than the maximal concentration observed for the other three investigated strains. However, considering specific secretion rates (normalized to fungal biomass), A. niger ATCC 11414 performed on par with the other strains—particularly towards the end of the incubation time. Disperse mycelial growth and pellet-like growth did not correlate with the overall amount of secreted protein between different strains of A. niger. However, dispersed mycelial growth, as observed for strains NRRL3 and ATCC 11414, is beneficial for the secretion of specific proteins in submerged cultures of A. niger and other filamentous fungi compared to pellet-like growth [29, 54].
Intriguingly, in terms of PGase activities, the dispersedly growing A. niger NRRL 3 and A. niger ATCC 11414 strains showed superior performance. Total PGase activity of A. niger NRRL 3 increased sharply between 36 and 59 h, with a maximum activity of 1379 ± 98 U L−1 (90 h). Endo-PGase activity of A. niger NRRL 3, however, showed a moderate and continuous increase to a maximum of 185 ± 39 U L−1 throughout the cultivation. The increase of endo-PGase activity appeared to be decoupled from the increase of the total PGase activity, indicating differences in expressional regulation of individual pectinase classes. The highest total PGase activity was observed for A. niger ATCC 11414, continuously increasing throughout the cultivation to a maximum of 1524 ± 35 U L−1 (82 h). Endo-PGase activity peaked at 389 ± 22 U L−1 at 82 h. A. niger ATCC 11414 thus exceeded the maximal total PGase activity of NRRL 3—as the second strongest polygalacturonase producer—by 13% and the maximal endo-PGase activity by 111%. Total secreted protein as well as biomass generation, therefore, did not correlate with detected polygalacturonase activity in this study. Moreover, the different maximum activities for endo- and total PGase as well as the different time profiles of PGase activity for A. niger NRRL3 and A. niger ATCC 11414 throughout the cultivation indicate significant differences in the regulation and expression of pectinases between these two strains. Considering the lower biomass accumulations and total protein production during the cultivation of A. niger ATCC 11414, this strain also generated the highest specific total and endo-PGase activities.
Assuming similar total capacities of the secretory machineries in all tested A. niger strains, A. niger ATCC 11414 is, therefore, the most promising candidate for genetic improvements towards pectinase overexpression. Less off-target secondary metabolism activity (as judged by pigment formation; see Fig. S1) and highest specific polygalacturonase production hold promise for additional metabolic capacities which might be exploitable for enhanced pectinase expression.
Comparison of hydrolytic performance of culture supernatants for d-GalA release from complex pectinaceous substrates
A. niger ATCC 11414 was determined as the most promising base strain for pectinase production out of six strains under the tested cultivation conditions. A. niger ATCC 11414 not only had the highest total and specific PGase activity, but also a disperse morphology desirable for protein secretion.
To test whether the selection of A. niger ATCC 11414 based on defined substrate assay conditions would translate into improved activity also on complex pectinaceous biomass, d-GalA release from milled dry sugar beet press pulp (SBPP) using A. niger ATCC 11414 culture supernatant was compared against that of the A. niger N402 culture supernatant as the ancestor of today’s standard laboratory strains. Using a 96 h A. niger ATCC 11414 buffered culture, an average of 8.8 g L−1 of free d-GalA was released from 9% (w/v) SBPP within 138 h, as compared to an average of 4.9 g L−1 for A. niger N402 culture supernatant (Fig. 5). Taking into account the water molecules incorporated during hydrolytic cleavage of d-GalA from polymers, this corresponded to degraded amounts of 8.0 g L−1 and 4.5 g L−1 of the provided biomass, respectively. Considering a total d-GalA content in SBPP of approximately 22% (w/w) [27], ~ 36.4% of the expected d-GalA was released using A. niger ATCC 11414 culture supernatant (vs. ~ 20% with A. niger N402 culture supernatant). In other words, the same release level was obtained in less than 45% of the process time. In summary, thorough screening and activity-driven selection of A. niger strains from a set of readily available and highly referenced strains resulted in a 75% higher d-GalA release compared to that obtained with the standard lab strain. Additionally, higher quantities of the main sugars of SBPP (hemi-)cellulosic fractions, such as l-arabinose, d-glucose, d-fructose, and d-galactose [27], were released using A. niger ATCC 11414 supernatant compared to A. niger N402 supernatant (see Table 1, Fig. S4–7).
d-GalA release from 9% (w/v) sugar beet press pulp (SBPP) using A. niger ATCC 11414 vs. A. niger N402 culture supernatant. Black and white triangles (open triangle, filled triangle) represent replicates of hydrolysis using 96 h culture supernatants of A. niger ATCC 11414 stirred-tank batch cultivations in 2% pectin minimal medium, with respective dashed lines in grey (y = − 0.1545x2 + 2.411x + 0.2364; R2 = 0.993) and black (y = − 0.1772x2 + 2.4832x + 0.069; R2 = 0.988). Black and white diamonds (filled diamond, open diamond) represent two replicates using supernatants of N402 with respective dotted lines in black (y = − 0.0901x2 + 1.3532x + 0.2218; R2 = 0.990) and grey (y = − 0.122x2 + 1.4493x + 0.5205; R2 = 0.960)
Monomeric d-GalA release from SBPP of up to 79% within 48 h has been reported in highly optimized saccharification conditions using combinations of commercial pectinase mixes of the Aspergillus genus at enzyme concentrations comparable to those used in this study [30], while no comparable efficiency data exist for direct application of crude Aspergillus culture supernatants on SBPP. Furthermore, the hydrolysis setup presented in this study was used as a pectinase production benchmark in strain selection only and has not yet undergone optimization for ideal d-GalA release conditions. Additional optimization of culture conditions for A. niger ATCC 11414 may further improve its performance, e.g., in terms of protein secretion, for which titers of up to 20 g L−1 have been reported [33]. Pectins, furthermore, comprise a variety of sugars and sugar acids with a highly diverse set of linkages between them [20]. Hence, a highly complex enzyme cocktail is required to facilitate efficient sequential and synergistic removal of decorating side chains from the polygalacturonic acid backbone for subsequent d-GalA release. Media supplementation with (hemi-)cellulosic substrates and thereby induced expression of (hemi-)cellulases during pectinase production could further contribute to d-GalA release from complex biomasses [11, 30]. Research in this field is still actively ongoing, but mostly focusing on process engineering, as recently demonstrated for continuous generation of d-GalA from SBPP pectin extracts in membrane enzyme reactors [15]. Genetic engineering may provide additional means to induce (hemi-)cellulase expression for improved d-GalA release from complex pectinaceous biomasses. Via systematic screening and strain performance evaluation under controlled conditions, we complemented this research with an important comparison of highly cited, openly available and readily applied A. niger strains. Endeavors to use SBPP and other complex pectinaceous biomasses of interest for industrial d-GalA supply in the context of the bio-economy, hence, could highly benefit from this work.
Conclusion
Considering the lack of systematic screening for A. niger strains with high pectinase production, we have implemented a robust protocol for the discrimination of competing strains in controlled stirred-tank bioreactors. Superior performance of A. niger ATCC 11414 was verified in a realistic setting using complex sugar beet press pulp. This strain shows potentially untapped metabolic and secretory reservoirs that could be exploited for improved pectinase production via targeted genetic engineering. However, to foster transfer of research results to industrial applications, it will be necessary to establish genetic tools, such as non-homologous end-joining suppressors or genetic markers, in this non-standard host strain.
E-supplementary data of this work can be found in the online version of the paper.
References
Aguilar G, Huitrón C (1990) Constitutive exo-pectinase produced by Aspergillus sp. CH-Y-1043 on different carbon source. Biotechnol Lett 12(9):655–660
Aguilar-Pontes MV, Brandl J, McDonnell E, Strasser K, Nguyen TTM, Riley R, Mondo S, Salamov A, Nybo JL, Vesth TC, Grigoriev IV, Andersen MR, Tsang A, de Vries RP (2018) The gold-standard genome of Aspergillus niger NRRL 3 enables a detailed view of the diversity of sugar catabolism in fungi. Stud Mycol 91:61–78
Amoah J, Kahar P, Ogino C, Kondo A (2019) Bioenergy and biorefinery: feedstock, biotechnological conversion, and products. Biotechnol J 14(6):1800494
Andersen MR, Salazar MP, Schaap PJ, van de Vondervoort PJI, Culley D, Thykaer J, Frisvad JC, Nielsen KF, Albang R, Albermann K, Berka RM, Braus GH, Braus-Stromeyer SA, Corrochano LM, Dai Z, van Dijck PWM, Hofmann G, Lasure LL, Magnuson JK, Menke H, Meijer M, Meijer SL, Nielsen JB, Nielsen ML, van Ooyen AJJ, Pel HJ, Poulsen L, Samson RA, Stam H, Tsang A, van den Brink JM, Atkins A, Aerts A, Shapiro H, Pangilinan J, Salamov A, Lou Y, Lindquist E, Lucas S, Grimwood J, Grigoriev IV, Kubicek CP, Martinez D, van Peij NNME, Roubos JA, Nielsen J, Baker SE (2011) Comparative genomics of citric-acid-producing Aspergillus niger ATCC 1015 versus enzyme-producing CBS 513.88. Genome Res 21(6):885–897
Arentshorst M, Ram AFJ, Meyer V (2012) Using non-homologous end-joining-deficient strains for functional gene analyses in filamentous fungi. In: Bolton M, Thomma B (eds) Plant fungal pathogens. Methods in molecular biology (Methods and Protocols), vol 835. Humana Press, Totowa, pp 133–150
Aristidou A, Penttilä M (2000) Metabolic engineering applications to renewable resource utilization. Curr Opin Biotechnol 11(2):187–198
Bos CJ (1986) Induced mutation and somatic recombination as tools for genetic analysis and breeding of imperfect fungi. PhD, Wageningen University
Bos CJ, Debets AJ, Swart K, Huybers A, Kobus G, Slakhorst SM (1988) Genetic analysis and the construction of master strains for assignment of genes to six linkage groups in Aspergillus niger. Curr Genet 14(5):437–443
Cairns TC, Nai C, Meyer V (2018) How a fungus shapes biotechnology: 100 years of Aspergillus niger research. Fungal Biol Biotechnol 5:13
Cairns TC, Zheng X, Zheng P, Sun J, Meyer V (2019) Moulding the mould: understanding and reprogramming filamentous fungal growth and morphogenesis for next generation cell factories. Biotechnol Biofuels 12:77
Chen Q, Jin Y, Zhang G, Fang Y, Xiao Y, Zhao H (2012) Improving production of bioethanol from duckweed (Landoltia punctata) by pectinase pretreatment. Energies 5(8):3019–3032
Diaz JV, Anthon GE, Barrett DM (2007) Nonenzymatic degradation of citrus pectin and pectate during prolonged heating: effects of pH, temperature, and degree of methyl esterification. J Agric Food Chem 55(13):5131–5136
El-Enshasy H, Kleine J, Rinas U (2006) Agitation effects on morphology and protein productive fractions of filamentous and pelleted growth forms of recombinant Aspergillus niger. Process Biochem 41(10):2103–2112
El-Enshasy HA (2007) Filamentous fungal cultures—process characteristics, products, and applications. In: Yang S-T (ed) Bioprocessing for value-added products from renewable resources. Elsevier, Amsterdam, pp 225–261
Elst K, Babbar N, Van Roy S, Baldassarre S, Dejonghe W, Maesen M, Sforza S (2018) Continuous production of pectic oligosaccharides from sugar beet pulp in a cross flow continuous enzyme membrane reactor. Bioprocess Biosyst Eng 41(11):1717–1729
Fiedler MRM, Gensheimer T, Kubisch C, Meyer V (2017) HisB as novel selection marker for gene targeting approaches in Aspergillus niger. BMC Microbiol 17(1):57
Friedrich J, Cimerman A, Perdih A (1986) Comparison of different cellulolytic fungi for bioconversion of apple distillery waste. Appl Microbiol Biotechnol 24(5):432–434
Friedrich J, Cimerman A, Steiner W (1989) Submerged production of pectolytic enzymes by Aspergillus niger: effect of different aeration/agitation regimes. Appl Microbiol Biotechnol 31(5–6):490–494
Goosen T, Bloemheuvel G, Gysler C, de Bie DA, van den Broek HWJ, Swart K (1987) Transformation of Aspergillus niger using the homologous orotidine-5′-phosphate-decarboxylase gene. Curr Genet 1(6–7):499–503
Harholt J, Suttangkakul A, Vibe Scheller H (2010) Biosynthesis of Pectin. Plant Physiol 153(2):384–395
Hortsch R, Weuster-Botz D (2010) Power consumption and maximum energy dissipation in a milliliter-scale bioreactor. Biotechnol Progress 26(2):595–599
Kashyap DR, Vohra PK, Chopra S, Tewari R (2001) Applications of pectinases in the commercial sector: a review. Bioresour Technol 77(3):215–227
Keasling JD (2008) Synthetic biology for synthetic chemistry. ACS Chem Biol 3(1):64–76
Kelly S, Grimm LH, Hengstler J, Schultheis E, Krull R, Hempel DC (2004) Agitation effects on submerged growth and product formation of Aspergillus niger. Bioprocess Biosys Eng 26(5):315–323
Khairnar Y, Krishna VK, Boraste A, Gupta N, Trivedi S, Patil P, Gupta G, Gupta M, Jhadav A, Mujapara A (2009) Study of pectinase production in submerged fermentation using different strains of Aspergillus niger. Int J Microbiol Res 1(2):13–17
Klein-Marcuschamer D, Blanch HW (2015) Renewable fuels from biomass: technical hurdles and economic assessment of biological routes. AIChE J 61(9):2689–2701
Kühnel S (2011) Characterization of cell wall degrading enzymes from Chrysosporium lucknowense C1 and their use to degrade sugar beet pulp. PhD, Wageningen University
Kuivanen J, Biz A, Richard P (2019) Microbial hexuronate catabolism in biotechnology. AMB Express 9(1):16
Kurt T, Marbà-Ardébol A-M, Turan Z, Neubauer P, Junne S, Meyer V (2018) Rocking Aspergillus: morphology-controlled cultivation of Aspergillus niger in a wave-mixed bioreactor for the production of secondary metabolites. Microb Cell Factories 17(1):128
Leijdekkers AGM, Bink JPM, Geutjes S, Schols HA, Gruppen H (2013) Enzymatic saccharification of sugar beet pulp for the production of galacturonic acid and arabinose; a study on the impact of the formation of recalcitrant oligosaccharides. Bioresour Technol 128:518–525
Meyer V, Andersen MR, Brakhage AA, Braus GH, Caddick MX, Cairns TC, de Vries RP, Haarmann T, Hansen K, Hertz-Fowler C, Krappmann S, Mortensen UH, Peñalva MA, Ram AFJ, Head RM (2016) Current challenges of research on filamentous fungi in relation to human welfare and a sustainable bio-economy: a white paper. Fungal Biol Biotechnol 3(1):6
Meyer V, Arentshorst M, El-Ghezal A, Drews A-C, Kooistra R, van den Hondel CAMJJ, Ram AFJ (2007) Highly efficient gene targeting in the Aspergillus niger kusA mutant. J Biotechnol 128(4):770–775
Meyer V, Fiedler M, Nitsche B, King R (2015) The cell factory Aspergillus enters the big data era: opportunities and challenges for optimising product formation. In: Krull R, Bley T (eds) Filaments in bioprocesses. Advances in biochemical engineering/biotechnology, vol 149. Springer, Berlin, pp 91–132
Nitsche BM, Jørgensen TR, Akeroyd M, Meyer V, Ram AFJ (2012) The carbon starvation response of Aspergillus niger during submerged cultivation: insights from the transcriptome and secretome. BMC Genomics 13:380
Niu J, Arentshorst M, Seelinger F, Ram AFJ, Ouedraogo JP (2016) A set of isogenic auxotrophic strains for constructing multiple gene deletion mutants and parasexual crossings in Aspergillus niger. Arch Microbiol 198(9):861–868
Ortiz GE, Guitart ME, Alberto E, Fernandez Lahore HM, Blasco M (2014) Microplate assay for endo-polygalacturonase activity determination based on ruthenium red method. Anal Biochem 454:33–35
Papagianni M, Mattey M, Kristiansen B (1998) Citric acid production and morphology of Aspergillus niger as functions of the mixing intensity in a stirred tank and a tubular loop bioreactor. Biochemical Eng J 2(3):197–205
Pel HJ, de Winde JH, Archer DB, Dyer PS, Hofmann G, Schaap PJ, Turner G, de Vries RP, Albang R, Albermann K, Andersen MR, Bendtsen JD, Benen JAE, van den Berg M, Breestraat S, Caddick MX, Contreras R, Cornell M, Coutinho PM, Danchin EGJ, Debets AJM, Dekker P, van Dijck PWM, van Dijk A, Dijkhuizen L, Driessen AJM, d'Enfert C, Geysens S, Goosen C, Groot GSP, de Groot PWJ, Guillemette T, Henrissat B, Herweijer M, van den Hombergh JPTW, van den Hondel CAMJJ, van der Heijden RTJM, van der Kaaij RM, Klis FM, Kools HJ, Kubicek CP, van Kuyk PA, Lauber J, Lu X, van der Maarel MJEC, Meulenberg R, Menke H, Mortimer MA, Nielsen J, Oliver SG, Olsthoorn M, Pal K, van Peij NNME, Ram AFJ, Rinas U, Roubos JA, Sagt CMJ, Schmoll M, Sun J, Ussery D, Varga J, Vervecken W, van de Vondervoort PJJ, Wedler H, Wösten HAB, Zeng A-P, van Ooyen AJJ, Visser J, Stam H (2007) Genome sequencing and analysis of the versatile cell factory Aspergillus niger CBS 513.88. Nat Biotechnol 25(2):221–231
Perlman D, Kita DA, Peterson WH (1946) Production of citric acid from cane molasses. Arch Biochem 11:123–129
Posch AE, Herwig C, Spadiut O (2013) Science-based bioprocess design for filamentous fungi. Trends Biotechnol 31(1):37–44
Pronk JT (2002) Auxotrophic yeast strains in fundamental and applied research. Appl Environ Microbiol 68(5):2095–2100
Reginatto C, Rossi C, Miglioranza BG, dos Santos M, Meneghel L, da Silveira MM, Malvessi E (2017) Pectinase production by Aspergillus niger LB-02-SF is influenced by the culture medium composition and the addition of the enzyme inducer after biomass growth. Process Biochem 58:1–8
Richard P, Hilditch S (2009) d-galacturonic acid catabolism in microorganisms and its biotechnological relevance. Appl Microbiol Biotechnol 82(4):597–604
Rude MA, Schirmer A (2009) New microbial fuels: a biotech perspective. Curr Opin Microbiol 12(3):274–281
Schäpe P, Kwon MJ, Baumann B, Gutschmann B, Jung S, Lenz S, Nitsche B, Paege N, Schütze T, Cairns TC, Meyer V (2019) Updating genome annotation for the microbial cell factory Aspergillus niger using gene co-expression networks. Nucleic Acids Res 47(2):559–569
Schmitz K, Protzko R, Zhang L, Benz JP (2019) Spotlight on fungal pectin utilization-from phytopathogenicity to molecular recognition and industrial applications. Appl Microbiol Biotechnol 103(6):2507–2524
Srivastava A, Srivastava M (2018) Enzymes market type (protease, carbohydrase, lipase, polymerase and nuclease, and other types), source (Microorganisms, plants, and animals), reaction type (hydrolase, oxidoreductase, transferase, lyase, and other teaction types), and application (food and beverages, household care, bioenergy, pharmaceutical and biotechnology, feed, and other applications)—global opportunity analysis and industry forecast, 2017–2024
Tanaka H, Wang P-L, Yamada Late O, Tamura T (1966) Yellow pigments of Aspergillus niger and Asp. awamori. Agric Biol Chem 30(2):107–113
Tao L, Aden A (2009) The economics of current and future biofuels. Vitro Cell Dev Biol Plant 45(3):199–217
Thite VS, Nerurkar AS (2018) Physicochemical characterization of pectinase activity from Bacillus spp. and their accessory role in synergism with crude xylanase and commercial cellulase in enzyme cocktail mediated saccharification of agrowaste biomass. J Appl Microbiol 124(5):1147–1163
Tilman D, Socolow R, Foley JA, Hill J, Larson E, Lynd L, Pacala S, Reilly J, Searchinger T, Somerville C, Williams R (2009) Energy. Beneficial biofuels—the food, energy, and environment trilemma. Science 325(5938):270–271
van Hartingsveldt W, Mattern IE, van Zeijl CM, Pouwels PH, van den Hondel CA (1987) Development of a homologous transformation system for Aspergillus niger based on the pyrG gene. Mol Gen Genet 206(1):71–75
Van Lanen JM, Smith MB (1965) Process of producing glucamylase and an alcohol product USA patent application US3418211A
Veiter L, Rajamanickam V, Herwig C (2018) The filamentous fungal pellet-relationship between morphology and productivity. Appl Microbiol Biotechnol 102(7):2997–3006
Vesth TC, Nybo JL, Theobald S, Frisvad JC, Larsen TO, Nielsen KF, Hoof JB, Brandl J, Salamov A, Riley R, Gladden JM, Phatale P, Nielsen MT, Lyhne EK, Kogle ME, Strasser K, McDonnell E, Barry K, Clum A, Chen C, LaButti K, Haridas S, Nolan M, Sandor L, Kuo A, Lipzen A, Hainaut M, Drula E, Tsang A, Magnuson JK, Henrissat B, Wiebenga A, Simmons BA, Mäkelä MR, de Vries RP, Grigoriev IV, Mortensen UH, Baker SE, Andersen MR (2018) Investigation of inter- and intraspecies variation through genome sequencing of Aspergillus section Nigri. Nat Genet 50(12):1688–1695
Wang J, Chio C, Chen X, Su E, Cao F, Jin Y, Qin W (2019) Efficient saccharification of agave biomass using Aspergillus niger produced low-cost enzyme cocktail with hyperactive pectinase activity. Bioresour Technol 272:26–33
Wang L, Ridgway D, Gu T, Moo-Young M (2003) Effects of process parameters on heterologous protein production in Aspergillus niger fermentation. J Chem Technol Biotechnol 78(12):1259–1266
Willke T, Vorlop K (2008) Biotransformation of glycerol into 1,3-propanediol. Eur J Lipid Sci Technol 110(9):831–840
Acknowledgements
Open Access funding provided by Projekt DEAL. This work was supported by the German Federal Ministry of Education and Research [Grant numbers 031B0342A and 031B0342C]. We furthermore want to thank Petra Arnold, Sabrina Paulus, and Nadine Griesbacher (Wood Bioprocesses, TUM, Germany), as well as Markus Amann und Norbert Werth (Institute of Biochemical Engineering, TUM, Germany) for excellent technical assistance. Also, we would like to acknowledge Nathalie Hafner (Institute of Biochemical Engineering, TUM, Germany) for assisting with the assays and fermentations. Additionally, we want to acknowledge the Südzucker AG, Obrigheim, Germany, for kindly providing the sugar beet press pulp and Arthur Ram (Leiden University, The Netherlands), Vera Meyer and Tabea Schütze (both TU Berlin, Germany) for providing strains and lab trainings for this study. The support of Dominik Schäfer and Kevin Schmitz by the TUM Graduate School is acknowledged as well.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they do not have any competing interests.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
449_2020_2347_MOESM1_ESM.tif
Supplementary file1 Fig. S1: Yellow pigment formation in A. niger culture supernatants. Accumulation of yellow pigmentation in culture supernatants of A. niger NRRL3, N402, ATCC1015 and ATCC11414 submerged stirred tank bioreactor batch cultivations in 2% pectin minimal medium. (TIF 1877 kb)
449_2020_2347_MOESM2_ESM.tif
Supplementary file2 Fig. S2: Reproduced submerged stirred tank batch cultivations of A. niger N402 and A. niger ATCC 11414. Biomass dry weight concentrations (a, b), total protein concentration (c) and total PGase activity (d) of the reproduced cultivations of A. niger N402 (filled diamond) and A. niger ATCC 11414 (filled triangle). Grey symbols display the values of the cultivations mentioned in the results and discussion. (TIF 308 kb)
449_2020_2347_MOESM3_ESM.tif
Supplementary file3 Fig. S3: Specific protein concentrations of selected A. niger strains. A. niger NRRL 3 (a, filled circle), A. niger N402 (b, filled diamond), A. niger ATCC 1015 (c, filled square) and A. niger ATCC 11414 (d, filled triangle) specific protein concentrations normalized by BDW during 90 h submerged batch cultivations in a 3 L stirred tank bioreactor in 2% pectin minimal medium. (TIF 295 kb)
449_2020_2347_MOESM4_ESM.tif
Supplementary file4 Fig. S4: L-arabinose release from 9% sugar beet press pulp (SBPP) using A. niger ATCC 11414 vs. A. niger N402 culture supernatant. Black and white triangles (open triangle, filled triangle) represent replicates of hydrolysis using 96 h culture supernatants of A. niger ATCC 11414 stirred-tank batch cultivations in 2% pectin minimal medium, with respective dashed lines in grey (y = − 0.1378x2 + 2.0842x − 0.3886; R² = 0.964) and black (y = − 0.1745x2 + 2.3955x − 0.4715; R² = 0.977). Black and white diamonds (filled diamond, open diamond) represent two replicates using supernatants of N402 with respective dotted lines in black (y = − 0.0048x2 + 1.3076x − 0.215; R² = 0.997) and grey (y = − 0.0481x2 + 1.3925x + 0.0902; R² = 0.994). (TIF 2185 kb)
449_2020_2347_MOESM5_ESM.tif
Supplementary file5 Fig. S5: D-glucose release from 9% sugar beet press pulp (SBPP) using A. niger ATCC 11414 vs. A. niger N402 culture supernatant. Black and white triangles (open triangle, filled triangle) represent replicates of hydrolysis using 96 h culture supernatants of A. niger ATCC 11414 stirred tank batch cultivations in 2% pectin minimal medium, with respective dashed lines in grey (y = 0.0453x2 + 0.5357x − 0.0448; R² = 0.954) and black (y = − 0.0679x2 + 0.9492x − 0.2538; R² = 0.945). Black and white diamonds (filled diamond, open diamond) represent two replicates using supernatants of N402 with respective dotted lines in black (y = − 0.0241x2 + 0.4811x − 0.0788; R² = 0.996) and grey (y = − 0.0146x2 + 0.3509x + 0.2036; R² = 0.929). (TIF 2185 kb)
449_2020_2347_MOESM6_ESM.tif
Supplementary file6 Fig. S6: D-galactose release from 9% sugar beet press pulp (SBPP) using A. niger ATCC 11414 vs. A. niger N402 culture supernatant. Black and white triangles (open triangle, filled triangle) represent replicates of hydrolysis using 96 h culture supernatants of A. niger ATCC 11414 stirred tank batch cultivations in 2% pectin minimal medium, with respective dashed lines in grey (y = − 0.0435x2 + 0.4411x − 0.0222; R² = 0.948) and black (y = − 0.046x2 + 0.4297x − 0.0008; R² = 0.981). Black and white diamonds (filled diamond, open diamond) represent two replicates using supernatants of N402 with respective dotted lines in black (y = − 0.018x2 + 0.2303x - 0.0102; R² = 0.998) and grey (y = − 0.0322x2 + 0.3022x + 0.035; R² = 0.990). (TIF 2190 kb)
449_2020_2347_MOESM7_ESM.tif
Supplementary file7 Fig. S7: D-fructose release from 9% sugar beet press pulp (SBPP) using A. niger ATCC 11414 vs. A. niger N402 culture supernatant. Black and white triangles (open triangle, filled triangle) represent replicates of hydrolysis using 96 h culture supernatants of A. niger ATCC 11414 stirred tank batch cultivations in 2% pectin minimal medium, with respective dashed lines in grey (y = − 0.0423x2 + 0.4679x − 0.1405; R² = 0.954) and black (y = − 0.0233x2 + 0.3521x − 0.1109; R² = 0.892). Black and white diamonds (filled diamond, open diamond) represent two replicates using supernatants of N402 with respective dotted lines in black (y = − 0.0204x2 + 0.2856x − 0.1176; R² = 0.954) and grey (y = − 0.0342x2 + 0.3475x + 0.0389; R² = 0.973). (TIF 2188 kb)
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Schäfer, D., Schmitz, K., Weuster-Botz, D. et al. Comparative evaluation of Aspergillus niger strains for endogenous pectin-depolymerization capacity and suitability for d-galacturonic acid production. Bioprocess Biosyst Eng 43, 1549–1560 (2020). https://doi.org/10.1007/s00449-020-02347-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00449-020-02347-z