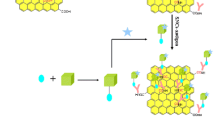
Abstract
Cathepsin G (CatG) is an endopeptidase that is associated with the early immune response. The synthetic compound cathepsin G inhibitor I (CGI-I) was tested for its ability to inhibit the activity of CatG via a new surface plasmon resonance imaging assay. CGI-I was immobilized on the gold surface of an SPR sensor that was first modified with 1-octadecanethiol. A concentration of CGI-I equal to 4.0 μg·mL-1 and a pH of 8.0 were found to give the best results. The dynamic response of the sensor ranges from 0.25 to 1.5 ng·mL-1, and the detection limit is 0.12 ng·mL-1. The sensor was applied to detect CatG in human saliva and white blood cells.

The synthetic compound cathepsin G inhibitor I (CGI-I) was tested for its ability to inhibit the activity of cathepsin G via a newly developed surface plasmon resonance imaging assay. The sensor was applied to detect cathepsin G in human saliva and white blood cells.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cathepsin G (EC 3.4.21.20) is one of the serine endopeptidases. It is stored in the azurophilic granules of neutrophilic polymorphonuclear leukocytes, but is also present in monocytes and mast cells [1].
The physiological role of cathepsin G (CatG) in the living organism is mainly associated with early immune response, but its biological activity is not limited to this. CatG is involved in several physiological processes such as tissue degradation, platelet activation, monocyte and neutrophil chemotaxis, proteolysis of blood coagulation factors, and generation of angiotensin II. It also displays a broad spectrum of antiviral, antibacterial, and antifungal activities. The potent antibacterial activity of CatG is independent of its proteolytic activity. The antibacterial action of CatG is thought to contribute significantly to the nonoxidative antibacterial capacity of neutrophils [2–5].
Together with similarly localized human neutrophil elastase and proteinase 3, CatG represents a major class of enzymes utilized in the phagocythic pathways to degrade foreign organisms and dead tissues during inflammatory reactions. Neutrophils play a key role in host defense by migrating to sites of infection, where they phagocytose invading microorganisms. After a neutrophil encloses a microbe, the resulting phagosome fuses with granules containing microbicidal and digestive enzymes to form a phagolysosome. The azurophilic granules of neutrophils are rich in serine proteases, including CatG and neutrophil elastase, which play critical roles in killing bacteria. Neutrophils are also known to invade many tumor tissues and influence tumor development, whereas CatG in neutrophils has a role in the dissemination of tumor aggregates from orimary tumor sites later to be secreted out by infiltrated neutrophils [5–7].
Activity of CatG, as well as other serine endopeptidases, is controlled by one or more members of the serine proteinase inhibitor (serpin) protein superfamily. However, an imbalance between proteases and serpins is implicated in tissue damage during inflammation [6]. In ischaemic diseases the impaired balance between CatG and its physiological inhibitors leads to tissue destruction and platelet aggregation. Inhibitors of CatG are thus suitable for the treatment of inflammatory diseases and procoagulant conditions [8]. Serpins can fail quantitatively or qualitatively in down-regulation due to genetic or acquired deficiencies. This results in the disturbed digestion of extracellular proteins that could contribute to the development of connective tissue diseases, such as emphysema, rheumatoid arthritis and perionditis [6].
Information concerning the determination of CatG is very scanty. Activities of CatG were determined using highly sensitive Abz-peptidyl-EDDnp fluorescence resonance energy transfer [9]. Elastase, proteinase 3 and CatG activities at the surface of human neutrophils were determined with fluorescence resonance energy transfer, and micellar electrokinetic chromatography (MEKC). MEKC was also use for simultaneous determination of Pseudomonas aeruginosa elastase, human leukocyte elastase and CatG activities [10]. In most cases activities of CatG are determined jointly by modification of the Barrett’s method [11], or using indirect immunofluorescence and fixed granulocyte ELISA [12].
The aim of this work was to develop an SPRI biosensor for the highly selective determination of CatG and to demonstrate the sensor applicability for the determination of the enzyme in different human body fluids. Highly selective interaction of CatG with immobilized cathepsin G inhibitor I (CGI-I) was utilised for sensor development. CGI-I, a potent, selective, reversible, competitive, non-peptide inhibitor of CatG (IC50 = 53 nmol·L-1 and Ki = 63 nmol·L-1) is a 620.6 Da-molecular compound. CGI-I weakly inhibits chymotrypsin (Ki = 1.5 μmol·L-1) and poorly inhibits thrombin, factor Xa, factor IXa, plasmin, rypsin, tryptase, proteinase 3, and human leukocyte elastase (IC50 > 100 μmol·L-1) [13]. The strategy for designing the sensor involved the immobilization of thiol (1-octadecanethiol) on a gold surface, and then the immobilization of CGI-I using hydrophobic interactions.
The Surface Plasmon Resonance Imaging (SPRI) technique is a promising tool for the development of sensitive sensors for biologically active species. It was applied for the determination of β-2-microglobulin, mucin, novel protein immunostimulating factor, microRNAs, DNA, cystatin C, cysteine cathepsins, cathepsin D and E, papain [14–22].
Experimental
Reagents
CatG from human sputum, 1-octadecanethiol (ODM), Tween 20, (ALDRICH, Steinheim, Germany, www.sigmaaldrich.com), cathepsin B (CatB) from human liver, CGI-I (CALBIOCHEM, Merck group, Merck Sp. z.o.o., Warsaw, Poland, www.merck-chemicals.pl) were used, as well as absolute ethanol, acetic acid, sodium hydroxide, sodium chloride, sodium carbonate, sodium phosphate, potassium phosphate, sodium acetate, potassium chloride, magnesium chloride (all POCh, Gliwice, Poland, www.poch.com.pl), HBS-ES solution pH = 7.4 (0.01 M HEPES, 0.15 mol·L-1 sodium chloride, 0.005% Tween 20, 3 mmol·L-1 EDTA), acetic buffer pH = 3.79–5.57, phosphate buffer pH = 7.17–8.04, carbonate buffer pH = 8.50–9.86 (all BIOMED, Lublin, Poland, www.biomed.lublin.pl), photopolymer ELPEMER SD 2054, hydrophobic protective paint SD 2368 UV SG-DG (PETERS, Kempen, Germany, www.peters.de) were used as received. Aqueous solutions were prepared with milliQ water (Simplicity®MILLIPORE). Whole citrated blood was provided by the Haematology Clinic of the Medical University in Bialystok, Poland and saliva samples by the volunteers. All samples were provided after obtaining the requisite consent of the Bioethical Commission.
Chip preparation
Gold chips were manufactured as described in a previous paper [24]. The gold surface of the chip was covered with photopolymer and hydrophobic paint as described in a previous paper [23]. 9 × 12 free gold surfaces were obtained. Using this chip, nine different solutions can be simultaneously measured without mixing the tested solutions. Twelve single SPRI measurements can be performed from one solution.
SPRI measurements
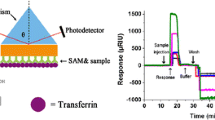
SPRI measurements for protein array were performed as described in the previous papers [21–23]. A schematic diagram of the apparatus is given in Fig. 1. The SPRI set-up consists of HeNe laser (JDS Uniphase, Edmund Industrial Optics, USA), two glass lenses L1 (f = 3 mm) and L2 (f = 300 mm), two polarizers (P1 and P2), mirror, prism and a CCD camera.
The chip consists of 9 detection areas, each containing an array of 12 spots. Spots are separated with photopolymer while detection areas are separated by hydrophobic paint. The picture of the chip is shown in Fig. 2a, while the image of the chip obtained by the CCD camera is shown in Fig. 2b.
Briefly, the measurements were performed at a fixed angle of incident light and reflectivity was simultaneously measured across an entire chip surface. The contrast values obtained for all pixels across a particular sample single spot were integrated. Thus the SPRI signal was integrated over the single spot area. Background correction was applied. NIH Image J version 1.42 software was used to evaluate the SPRI images in 2D form. The signal was measured twice on the basis of registered images, after immobilisation of CGI-I and then after interaction with CatG. The SPRI signal, which is proportional to coupled biomolecules, was obtained from subtraction between the signal before and after interaction with a biomolecule for each spot separately.
CGI-I immobilization
Chips were rinsed with ethanol and water and dried under a stream of nitrogen. They were then immersed in 10 mL of 2.7 mmol·L-1 ODM ethanolic solutions for at least 24 h. The chips were further rinsed with ethanol and water and dried under a stream of nitrogen. CGI-I solution (3 μL of CGI-I solution for each array of 12 spots) was placed on the thiol-modified surface and incubated at room temperature for 24 h [23].
Preparation of the standard curve
The response of the analytical SPRI signal for CatG concentration was measured within a concentration range between 0.25 and 2.5 ng·mL-1. The chip surface was covered by a monolayer of ODM and a layer of immobilized CGI-I (4.0 μg·mL-1). Experiments were performed at pH = 8.0. The interaction time was 10 min.
Preparation of biological samples
White blood cells
2 mL of whole blood was centrifuged over 15 min at 1,000 × g. As a result, plasma was separated and white blood cells were collected. The solution of white blood cells was diluted two-fold with a buffer of pH = 8.0 and transferred onto the sensor surface for 10 min. The volume of the sample applied on each detection area was 3 μL. The surface of the biosensor was then washed six times with the HBS-ES buffer. The SPRI measurement was performed. Concentration was evaluated using a calibration curve.
Saliva
3 mL of saliva (in fasting state) was centrifuged over 15 min at 100 × g and a supernatant was separated. Finally, the sample was filtered three times through a paper filter of medium density. The prepared saliva was diluted two-fold with a buffer of pH = 8.0 and transferred onto the sensor surface for 10 min. The volume of sample applied on each detection area was 3 μL. The surface of the biosensor was then washed six times with the HBS-ES buffer. The SPRI measurement was performed. Concentration was evaluated using a calibration curve.
Results and discussion
Influence of CGI-I concentration on the analytical signal of CatG
The purpose of this investigation was to find the best conditions for CatG determination. The investigation was performed within a CGI-I concentration range of 0.25 to 6.0 μg·mL-1 at constant CatG concentration (1.0 ng·mL-1). A series of 9 CGI-I solutions of increasing concentration were measured (0.25, 0.75, 1.50, 2.25, 3.00, 3.75, 4.50, 5,25 and 6.00 μg·mL-1). These solutions were transferred onto different points of the surface of a chip, which had been previously modified with ODM. CGI-I was immobilised during this treatment. The chip was then treated with the CatG solution (1.0 ng·mL-1) for 10 min, rinsed with water and dried, and the SPRI measurement was performed. The results are given in Fig. 3.
The obtained curve is a Langmuir isotherm type, with the plateau of the signal for CGI-I concentration above 3.00 μg·mL-1. A concentration of CGI-I equal to 4.00 μg·mL-1 was selected as optimal for further investigation.
Influence of solution pH on the interaction process
The influence of pH of CatG solution on the SPRI signal was studied for 13 different pH values (within the range 3.0–10.0) under conditions of constant concentrations of CatG (1 ng·mL-1) and CGI-I (4.0 μg·mL-1). Solutions of CGI-I were transferred onto different points of a chip which had been previously modified with ODM. In this manner CGI-I was immobilised onto the chip surface. The chip was then treated with CatG solutions (1 ng·mL-1) with different pH values, for 10 min. The results are shown in Fig. 4.
As is apparent from the diagram, the maximum of the SPRI signal for CGI-I—CatG complex is for pH = 8.0. The value of pH = 8.0 was selected as optimal for further investigation.
Analytical response of the sensor to CatG concentration. Calibration curves
The response of the analytical SPRI signal for CatG concentration was measured within a range of concentration of between 0.25 and 2.5 ng·mL-1. The chip surface was covered by a monolayer of ODM and a layer of immobilized CGI-I (4.0 μg·mL-1). The experiments were performed at pH = 8.0. The time of interaction was 10 min. The obtained calibration curve is shown in Fig. 5.
The calibration curves represent a Langmuir isotherm type with the dynamic response range between 0.25 and 1.5 ng·mL-1 (0.01–0.06 nmol·L-1). This section is useful for analytical purposes. The plateau of the curve corresponds to saturation of active points of the sensor. The detection limit calculated on the 3 S.D. basis is equal to 0.12 ng·mL-1.
Selectivity of the CGI-I—CatG interaction
In order to be sure that the interaction between CGI-I and CatG is selective and the sensor does not react with the other enzymes of the cathepsin family, the chip containing immobilised CGI-I was treated with mixtures of CatG and CatB. Various excesses of CatB within the range from 1:1 to 1:1000 were examined. Results are shown in Table 1.
The results show no influence of CatB on the results of determination of CatG even at a 1000-fold excess. Thus sensor high selectivity was confirmed.
The problem of the development of a specific sensor for CatG requires a specific inhibitor for this cathepsin. There is only one commercially available inhibitor of CatG, which was used in this research. However, there are reports about developments of new potent inhibitors of CatG [2]. Construction of a biosensor with the use of one of them is the subsequent goal of the authors.
Precision of the method for CatG determination
Precision of the method was tested under optimal conditions, i.e. pH = 8.0 and CGI-I concentration at the chip preparation stage equal to 4.0 μg·mL-1. The precision of CatG determination was tested for the concentration of this enzyme equal to 1.00 ng·mL-1. The results are shown in Table 2.
Standard deviation and the confidence limit, assuming that the confidence level equals 95%, are relatively high. This indicates that the biosensor has a relatively low precision. However, the effect of relatively poor precision is compensated by large number of repetitions, because the sensor is an array of 12 analyzed spots. Therefore, recoveries of spikes are good.
Recovery of the method was checked by addition of spikes of CatG into a real samples of white blood cells. The results are given in Table 3.
Atomic force microscopy measurements
Atomic Force Microscopy (AMF) enables observation of the surface of the biosensor. AFM was used to confirm the creation of each layer: thiol (ODM), inhibitor (CGI-I) and the enzyme (CatG). Pictures were taken of bare gold (Fig. 6a), ODM (Fig. 6b), CGI-I (Fig. 6c) and CatG (Fig. 6d). The presented AFM pictures confirm that the described stages of the creation of each layer on the biosensor surface have really taken place. This may be concluded on the basis of the creation of different structures after each stage.
CatG determination in real samples
White blood cells samples
Samples of white blood cells from 9 patients suffering from leukemia were analysed for CatG determination using the sensor. The results for patients are given in Table 4. Concentration of CatG in white blood cells of donors was below the detection limit, so it could not be estimated.
Obtained results for CatG in white blood cells of patients suffering from leukemia are within the range of 2.37 to 234.14 10−5 ng·103 white blood cells. Due to different type and stage of leukemia, these concentrations varied within a wide range.
Saliva samples
CatG was determined in 6 samples of saliva in order to demonstrate the sensor’s applicability to this purpose. The results are shown in Table 5.
The range of CatG concentration in the saliva of donors aged 19–24 is between 1.25 and 2.17 ng·mL-1, while donors over 75 present concentration between 3.18 and 3.51 ng·mL-1. The changes of CatG activity in saliva can play an important role in the pathobiochemistry and diagnostics of salivary gland, gingiva, and oral mucosa diseases. It is worth stressing that an elevated CatG concentration in saliva can be a symptom of oral cavity diseases [25].
Conclusions
A selective SPRI array biosensor for the determination of CatG has been developed. The sensor works on the basis of a highly selective reaction between immobilised CGI-I and CatG in solution. Under optimised conditions, the sensor exhibits selectivity to cathepsin G, and the precision and accuracy is well suited to enzyme determination. The sensor can be used for the determination of CatG in white blood cells and saliva.
References
Polanowska J, Krokoszyńska I, Czapińska H, Watorek W, Dadlez M, Otlewski J (1998) Specificity of human cathepsin G. Biochim Biophys Acta 1386:189–198
Sieńczyk M, Lesner A, Wysocka M, Łęgowska A, Pierusiewicz E, Rolka K, Oleksyszyn J (2008) New potent cathepsin G phosphonate inhibitors. Bioorg Med Chem 16:8863–8867
Lesner A, Wysocka M, Guzow K, Wiczk W, Łęgowska A, Rolka K (2008) Development of sensitive cathepsin G fluorogenic substrate using combinatorial chemistry methods. Anal Biochem 375:306–312
Gütschow M, Kuerschner L, Pietsch M, Ambrożak A, Neumann U, Günther R, Hofmann HJ (2002) Inhibition of cathepsin G by 2-amino-3, 1-benzoxazin-4-ones: kinetic investigations and docking studies. Arch Biochem Biophys 402:180–191
Kudo T, Kigoshi H, Hagiwara T, Takino T, Yamazaki M, Yui S (2009) Cathepsin G, a neutrophil protease, induces compact cell-cell adhesion in MCF-7 human breast cancer cells. Mediat Inflamm 2009:1–11
Martins FT, Assis DM, Santos MH, Camps I, Veloso MP, Juliano MA, Alves LC, Doriguetto AC (2009) Natural polyprenylated benzophenones inhibiting cysteine and serine proteases. Eur J Med Chem 44:1230–1239
Shao B, Belaaouaj A, Verlinde CLMJ, Fu X, Heinecke JW (2005) Methionine sulfoxide and proteolytic cleavage contribute to the inactivation of cathepsin G by hypochlorous acid: an oxidative mechanism for regulation of serine proteinases by myeloperoxidase. J Biol Chem 280:29311–29321
Gatto B, Vianini E, Lucatello L, Sissi C, Moltrasio D, Pescador R, Porta R, Palumbo M (2008) Effective DNA inhibitors of cathepsin G by in vitro selection. Int J Mol Sci 9:1008–1023
Korkmaz B, Attucci S, Juliano MA, Kalupov T, Jourdan ML, Juliano L, Gauthier F (2008) Measuring elastase, proteinase 3 and cathepsin G activities at the surface of human neutrophils with fluorescence resonance energy transfer substrates. Nat Protoc 3:991–1000
Viglio S, Luisetti M, Zanaboni G, Döring G, Worlitzsch D, Cetta G, Iadarola P (1999) Simultaneous determination of pseudomonas aeruginosa elastase, human leukocyte elastase and cathepsin G activities by micellar electrokinetic chromatography. J Chromatogr A 846:125–134
Barrett AJ (1981) Cathepsin G. In: Colowick SP, Kaplan NO (eds) Methods in enzymology, volume 80. Academic, New York, pp 561–565
Halbwachs-Mecarelli L, Nusbaum P, Noël H, Reumaux D, Erlinger S, Grünfeld JP, Lesavre P (1992) Antineutrophil cytoplasmic antibodies (ANCA) directed against cathepsin G in ulcerative colitis, Crohn’s disease and primary sclerosing cholangitis. Clin Exp Immunol 90:79–84
Greco MN, Hawkins MJ, Powell ET, Almond HR, Corcoran TW Jr, de Garavilla L, Kauffman JA, Recacha R, Chattopadhyay D, Andrade-Gordon P, Maryanoff BE (2002) Nonpeptide inhibitors of cathepsin G: optimization of a novel β-ketophosphonic acid lead by structure-based drug design. J Am Chem Soc 124:3810–3811
Fernández-González A, Rychlowska J, Badía R, Salzer R (2007) SPR imaging as a tool for detecting mucin—anti-mucin interaction. Outline of the development of a sensor for near-patient testing for mucin. Microchim Acta 158:219–225
Chen H, Lee J, Jo WS, Jeong MH, Koh K (2010) Development of surface plasmon resonance immunosensor for the novel protein immunostimulating factor. Microchim Acta. doi:10.1007/s00604-010-0476-0
Lee HJ, Yan Y, Marriott G, Corn RM (2005) Quantitative functional analysis of protein complexes on surfaces. J Physiol 563:61–71
Lee HJ, Nedelkov D, Corn RM (2006) Surface plasmon resonance imaging measurements of antibody arrays for the multiplexed detection of low molecular weight protein biomarkers. Anal Chem 78:6504–6510
Fang S, Lee J, Wark AW, Corn RM (2006) Attomole microarray detection of MicroRNAs by nanoparticle-amplified SPR imaging measurements of surface polyadenylation reactions. J Am Chem Soc 128:14044–14046
Gorodkiewicz E, Regulska E, Roszkowska-Jakimiec W (2010) Determination of the active form concentration of cathepsins D and B by SPRI biosensors. J Lab Diagn 46:107–109
Gorodkiewicz E, Regulska E (2010) SPR imaging biosensor for aspartyl cathepsins: sensor development and application for biological material. Protein Pept Lett 17:1148–1154
Gorodkiewicz E (2007) The surface plasmon resonance imaging sensor for papain based on immobilized cystatin. Protein Pept Lett 14:443–445
Gorodkiewicz E (2009) Surface plasmon resonance imaging sensor for cathepsin determination based on immobilized cystatin. Protein Pept Lett 16:1379–1385
Gorodkiewicz E, Fernández-González A, Akkoyun A, Salzer R (2005) Systematic evaluation of a surface plasmon resonance imaging biochip reader: study of gold surface modifications. Chem Anal (Warsaw) 50:103–116
Ozmeric N (2004) Advances in periodontal disease markers. Clin Chim Acta 343:1–16
Acknowledgements
Doctor J. Svieklo is greatly thanked for offering invaluable help with Atomic Force Microscope. Prof. J. Kloczko and dr J. Piszcz from Hematology Clinic of Medical University of Bialystok are greatly thanked for making available to us the blood samples of leukaemia patients. Very fruitful consultations with Professor Zenon Lukaszewski from Poznan University of Technology are greatly appreciated. Professor Zbigniew Artur Figaszewski from the University in Bialystok is thanked for creating a stimulating climate for our research.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Gorodkiewicz, E., Regulska, E. & Wojtulewski, K. Development of an SPR imaging biosensor for determination of cathepsin G in saliva and white blood cells. Microchim Acta 173, 407–413 (2011). https://doi.org/10.1007/s00604-011-0569-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s00604-011-0569-4