Abstract
Surface protein display C (SpdC) protein was described as a novel virulence factor of Staphylococcus aureus that affects biofilm formation and pathogenesis and favors resistance to antimicrobials targeting cell wall. We evaluated the possible correlation between spdC gene expression level and virulence as well as antibiotic resistance phenotypes in S. aureus clinical isolates. The antimicrobial susceptibility of S. aureus clinical isolates (n = 100) was determined by the disk diffusion method. Vancomycin susceptibility was determined by the broth microdilution method. The level of the extracellular proteases and delta-hemolysin was evaluated by measuring the proteolysis and hemolysis zone diameters in skim milk and blood agar plates, respectively. Biofilm formation was assayed using the 96-well microtiter plate method. Most of the isolates (81%) were multidrug-resistant and about half of the isolates (49%) were methicillin-resistant S. aureus. Hemolysin, protease, and biofilm production were detectable in 79%, 71%, and 96% of the isolates. No significant correlation was detectable between the level of spdC gene expression and the activity of tested virulence factors or the antimicrobial resistance phenotype. Therefore, the role of SpdC protein as a virulence regulator in S. aureus needs further evaluation together with the determination of the predominant regulators for each virulence factor.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Staphylococcus aureus is a highly virulent human pathogen implicated in many diseases; it causes both nosocomial and community-acquired infections (Uhlemann et al. 2014) and can result in bacteremia with significant rates of morbidity and mortality (van Hal et al. 2012). S. aureus infections range from localized skin infections such as folliculitis, furuncles, carbuncles, and impetigo to deep infections that spread systemically. Systemic infections involve bacteremia (with and without endocarditis); bones, joints, deep organs, and tissue infections; scalded skin syndrome in neonates; toxic shock syndrome; and food poisoning (Tong et al. 2015). According to Centers for Disease Control and Prevention (CDC), people with chronic conditions and hospitalized patients are at high risk of S. aureus infections (CDC 2011).
S. aureus is increasingly becoming resistant to several antimicrobial agents which is a challenging problem in clinical practice (Naimi et al. 2017). Methicillin-resistant S. aureus (MRSA) has been listed by the World Health Organization as one of the high priority pathogens for which new antibiotics are urgently needed (World Health Organization 2017). People infected with MRSA are 64% more likely to die than those infected with methicillin susceptible S. aureus (MSSA)(World Health Organization 2020). MRSA has also been listed by CDC as a serious threat (CDC 2019).
Vancomycin is the drug of choice for treatment of MRSA infection. The emergence of vancomycin-resistant S. aureus has posed additional challenge in the treatment of S. aureus infections, where other antimicrobial agents have been approved for the treatment of MRSA infections including daptomycin, linezolid, tedizolid, oritavancin, dalbavancin, ceftaroline, and ceftobiprole (Boswihi and Udo 2018).
Many virulence determinants were identified in S. aureus. Surface proteins such as adhesins, clumping factors, iron-regulated surface determinant (IsdA), fibrinogen-binding proteins, and fibronectin-binding proteins are responsible for attachment to host cells and tissue colonization. Invasins like leukocidin kinase and hyaluronidase promote bacterial spread in tissues. Other virulence determinants include surface factors (capsule and protein A) that inhibit phagocytic engulfment; and molecules that enhance their survival in phagocytes (carotenoids and catalase production). In addition, S. aureus produces several immunological disguises (protein A and coagulase), membrane-damaging toxins (hemolysins, leukotoxin, and leukocidin) that lyse eukaryotic cell membranes, and exotoxins that damage host tissues and provoke symptoms of disease (Reddy et al. 2017). Different types of sortase enzyme, that are considered virulence determinants through anchoring different proteins into the bacterial cell wall, were reported in S. aureus; these proteins have many functions in evasion of host defense and adhesion to target tissues and may function in biofilm formation (Nitulescu et al. 2021).
S. aureus also encodes about 16 two-component systems (TCS), representing an important regulatory mechanism that affects survival and virulence under infection conditions. The TCS encodes a histidine kinase that is usually membrane bound and senses different signals causing its autophosphorylation and a response regulator that is subsequently accepting the phosphoryl group from the histidine kinase and affects the activity of many chromosomal operons (Bleul et al. 2022).
Recently, Poupel and colleagues have described spdC gene (also known as LyrA) as a global S. aureus virulence factor (Poupel et al. 2018). It encodes a transmembrane protein with ABI domain, an element first described in lactococci for its role in phage exclusion function (Frankel et al. 2010). Expression of spdC gene affects biofilm formation and pathogenesis and is positively regulated by the WalKR system, one of the S. aureus TCS (Dubrac et al. 2008). SpdC protein interacts with the WalK histidine kinase to inhibit the activity of the WalKR TCS and regulates nine other histidine kinases of S. aureus, suggesting its role as a global regulator (Poupel et al. 2018). spdC gene was found to be encoded in all sequenced staphylococci strains (Gründling et al. 2006); however, deletion of spdC gene was reported in a vancomycin intermediate–resistant strain (Yamaguchi et al. 2019). It was hypothesized that deletion of spdC gene may impact vancomycin resistance through its effect on WalKR regulon.
Poupel and colleagues reported one hundred genes whose expression is regulated by SpdC protein. The spdC mutant, prepared by deletion of spdC gene in the parent S. aureus strain HG001, displayed altered resistance towards compounds targeting the cell wall; it was highly sensitive to oxacillin and tunicamycin, but not to fosfomycin, which inhibits the first step of cell wall biosynthesis. Additionally, the spdC mutant had a diminished biofilm formation and reduced virulence (Poupel et al. 2018). Attenuation of S. aureus virulence can be utilized as a strategy for treatment of resistant infections (Mahdally et al. 2021).
In this study, the possible correlation between spdC gene expression level and the virulence as well as the resistance to different antimicrobials, in S. aureus clinical isolates, was evaluated.
Materials and methods
Microbial strains and culture conditions
S. aureus standard strain ATCC 25923 was used as a reference strain. S. aureus clinical isolates (n = 100) were obtained from Ain Shams Specialized Hospital (H1) and El-Demerdash Specialized Hospital (H2), between November 2018 and August 2019. The source of each isolate is indicated in Supplementary Table 1. The identification of isolates as S. aureus was confirmed phenotypically by Gram staining, DNase test, positive coagulase test, and mannitol fermentation (yellow colonies on mannitol salt agar media) (Procop et al. 2017) and genotypically by detection of thermonuclease (nuc) gene using polymerase chain reaction (PCR). The isolates were stored in tryptone soya broth (TSB) containing 25% glycerol at − 80 °C. Unless otherwise described, they were isolated on brain–heart infusion agar plates and incubated at 37 °C prior to use.
Antimicrobial susceptibility testing
Antimicrobial susceptibility testing was carried out using the Kirby-Bauer disk diffusion method, according to Clinical and Laboratory Standards Institute’s (CLSI) guidelines (CLSI 2012a). S. aureus standard strain ATCC 25923 was used as a control strain. Fresh bacterial colonies were suspended in saline to reach an optical density (OD) equivalent to that of 0.5 McFarland turbidity standard (about 1 to 2 × 108 CFU/mL) and then spread onto the surface of agar plates using sterile swabs. Antibiotic disks were placed onto the surface of the agar plates; the tested antibiotics were penicillin (10 µg), gentamicin (10 μg), erythromycin (15 µg), ciprofloxacin (5 μg), tetracycline (30 µg), cefoxitin (30 μg), sulfamethoxazole/trimethoprim (1.25/23.75 μg), clindamycin (2 µg), linezolid (30 µg), and chloramphenicol (30 µg). The selection of antimicrobial agents was based on CLSI-suggested antimicrobials for routine testing of S. aureus (CLSI 2017) and the suggested agents for defining multidrug resistance (MDR) in S. aureus according to the definition of Magiorakos et al. (2012). All disks were obtained from Bioanalyse, Turkey. The plates were incubated at 37 °C for 24 h. The diameters of the inhibition zones were measured and the results were interpreted according to CLSI guidelines (CLSI 2017). The breakpoints used for defining the susceptibility pattern of tested isolates are given in Supplementary Table 2.
The antimicrobial susceptibility of the isolates to vancomycin was determined by the broth microdilution method according to CLSI guidelines (CLSI 2012b). S. aureus ATCC 29213 was used as a quality control strain (CLSI 2017). Vancomycin (Mylan, Ireland) was tested in concentration range from 0.5 to 64 μg/mL. The 96-well microtiter plates were incubated at 37 °C for 24 h. The minimum inhibitory concentration (MIC) was defined as the least concentration that completely inhibited the growth of the microorganisms. S. aureus isolates with vancomycin MICs ≤ 2 μg/mL were considered susceptible. Vancomycin intermediate S. aureus had MICs of 4–8 μg/mL, while vancomycin-resistant S. aureus had MIC of ≥ 16 μg/mL (CLSI 2017).
MRSA strains were defined as S. aureus strains that were resistant to methicillin; cefoxitin was used as methicillin surrogate in MRSA testing (CLSI 2017). S. aureus isolates were categorized as MDR if they were resistant to at least one agent in three or more antimicrobial categories or were MRSA, according to the definition of Magiorakos et al. (2012).
Determination of virulence in tested S. aureus isolates
Delta-hemolysin activity
Overnight cultures of the tested isolates in Muller Hinton broth (MHB; HiMedia, India) were diluted to reach an OD equivalent to one at 600 nm. The diluted culture (50 µL) was added into holes (5 mm in diameter) made in 7% sheep-blood agar plates. The plates were incubated overnight at 37 °C followed by storage overnight at 4 °C. S. aureus standard strain ATCC 25923 was used as a positive control (Zhang et al. 2016), while uninoculated MHB was used as negative control. Diameters of the hemolysis zones, in blood agar, were measured as indicative of hemolysin activity (Quiblier et al. 2011). Isolates were given arbitrary scores according to zone diameters as follows: score = 0 if hemolysis zone diameter ≤ 5 mm, score = 1 (weak hemolytic activity) if hemolysis zone diameter > 5 mm and ≤ 10 mm, score = 2 (moderate hemolytic activity) if hemolysis zone diameter > 10 mm and ≤ 20 mm, score = 3 (high hemolytic activity) if hemolysis zone diameter > 20 mm.
Protease activity
The assay of protease activity was performed by applying the same method described under hemolysin activity except that skim milk agar plates (Conda, Spain) were used instead of blood agar plates. Plates were incubated at 37 °C for 24 h. S. aureus ATCC 25923 was used as a positive control (Kaur et al. 2017), while uninoculated MHB was used as negative control. Diameters of the proteolysis zones, in skim milk agar, were measured as indicative of protease activity (Quiblier et al. 2011). Isolates were given arbitrary scores according to zone diameters as follows: score = 0 if proteolysis zone diameter ≤ 5 mm, score = 1 (weak proteolytic activity) if proteolysis zone diameter ˃ 5 mm and ≤ 10 mm, score = 2 (moderate proteolytic activity) if proteolysis zone diameter ˃ 10 mm and ≤ 20 mm, score = 3 (high proteolytic activity) if proteolysis zone diameter ˃ 20 mm.
Assay of biofilm production
Biofilm production was determined according to the method of Christensen et al. (1985). Briefly, overnight cultures of the tested isolates in TSB (HiMedia, India) were diluted 1:100 with the same media. Diluted samples were added in the wells of a sterile 96-polystyrene microtiter plate. Each strain was tested in triplicate. Wells with uninoculated TSB were used as a negative control. S. aureus ATCC 25923 was used as a positive control (Kashef et al. 2020). The plates were incubated at 37 °C for 24 h and the OD of the cultures was measured at 600 nm. The culture was then discarded, and the wells were washed with phosphate buffered saline three times. The plates were left to dry followed by addition of 200 µL of 1% crystal violet for 30 min. Excess dye was removed and the plates were washed with distilled water three times and left to dry. To measure the OD of the stained biofilms, 200 µL of 95% ethyl alcohol was added to each well, and the plate was left for 15 min; then, 125 µL of each solution was transferred to a well in a new plate. The absorbance was measured at 570 nm using enzyme-linked immunosorbent assay plate reader (Torlak et al. 2017). The cutoff OD value (Odc) was determined as the mean plus three times the standard deviations of the negative control. The OD of stained biofilm formed by each sample was normalized to the OD of the culture measured at 600 nm. If normalized sample optical density (Ods) ≤ Odc, then it is non-biofilm producer (score = 0). If Odc < Ods ≤ 2 × Odc, the sample is a weak biofilm producer (score = 1). If 2 × Odc < Ods ≤ 4 × Odc, the sample is a moderate biofilm producer (score = 2) and the sample is considered strong biofilm producer if 4 × Odc < Ods (score = 3) (Mesrati et al. 2018).
Detection of spdC gene and its expression level
Detection of spdC gene
The detectability of spdC gene in the collected isolates was determined using PCR. To ensure conservation of spdC gene in S. aureus isolates, 78 sequences of spdC gene were randomly selected from the spdC gene sequences available at the GenBank and aligned using Sequence Alignment Tool of Clustal Omega software (https://www.ebi.ac.uk/Tools/msa/clustalo/).
Oligonucleotides were designed using the Primer Quest Tool of Integrated DNA Technologies (IDTDNA, Coralville, USA). Primer-Blast (National Center for Biotechnology Information, https://www.ncbi.nlm.nih.gov/) was used to test the specificity of the designed primers. The sequences of the used primers were as follows: SpdC_Forward: GCTTCAATGACATTTGGCCTTA, and SpdC_Reverse: CTGCAACGATTGCTGTTGAAATG. Genomic DNA was extracted by the boiling method (Sambrook and Russell 2001). The PCR reaction mixture (25 µL) contained 5 µL of bacterial lysate, 5 µL of 5 × Green GoTaq Reaction Buffer, 1 mM MgCl2, 0.2 mM dNTP mix, 10 pmol of each primer, and 0.625 U GoTaq DNA Polymerase. All PCR reagents were from Promega (USA). The reaction was carried out in a thermal cycler using initial denaturation at 94 °C for 3 min, 30 cycles of denaturation at 94 °C for 30 s, annealing at 60 °C for 30 s and extension at 72 °C for 1 min, and a final extension step at 72 °C for 10 min. Amplicons with the expected size (≈100 bp) were visualized by electrophoresis on 1.5% agarose gel stained with ethidium bromide.
To further confirm the specificity of the PCR product, the product from the standard S. aureus ATCC 25923 was purified using QIAquick PCR Purification Kit (Qiagen, Germany) and sequenced by 3730 × l DNA Analyzer (Applied Biosystems, USA). Similarity searches for the nucleotide sequences were performed with BLASTN program (http://www.ncbi.nlm.nih.gov/blast) using default settings.
Determination of spdC gene expression
The level of spdC gene expression was determined by quantitative real-time PCR (qRT-PCR). The previously described primers for spdC gene detection, in the “Detection of spdC gene” section, were used in qRT-PCR. 16S rRNA was used as a housekeeping gene; the sequences of the primers used for 16S rRNA amplification were as follows: 16S_Forward: GTGGAGGGTCATTGGAAACT, and 16S_Reverse: CACTGGTGTTCCTCCATATCTC.
Tested strains were allowed to grow in TSB with shaking at 180 rpm until the OD of the culture at 600 nm reached 1 (Poupel et al. 2018). RNA extraction was carried out using RNeasy mini kit (Qiagen, Germany) as per manufacturer’s protocol. The extracted RNA was used (400 ng) for reverse-transcription and amplification of the produced cDNA using iTaq™ Universal SYBR® Green One-Step Kit (Bio-Rad, USA) following manufacturer’s protocol. The amplification of cDNA was conducted with Rotor-Gene 6000 real-time thermal cycler (Corbett Life Science, Mortlake, Australia) according to the manufacturer’s instructions. The level of spdC gene expression in each tested isolate was normalized to the 16S rRNA gene expression level as a housekeeping gene and expressed as fold change relative to that in the reference S. aureus ATCC 25923 strain as a calibrator.
Statistical analysis
Kendall’s Tau correlation coefficient was used to test the correlation between the fold change of spdC gene expression level and the activity of different virulence factors, as well as the resistance to different antibiotics. Kendall’s Tau correlation coefficient measures the strength of the relationship between two variables by assessing the statistical associations based on the ranks of data and is preferred for small sample size (Puka 2011). The Wilcoxon rank-sum test was used for the determination of the significance of the difference between the median of fold change in spdC gene expression level in isolates with different antimicrobial susceptibility phenotypes. The Wilcoxon rank-sum test is used for comparing the median of non-parametric data (Neuhäuser 2011).
Results
A total of 100 S. aureus clinical isolates were used in the study. Identification of isolates as S. aureus was confirmed by detection of nuc gene in all the collected isolates. Most of the isolates were MDR (81%) and nearly half of which were MRSA. Vancomycin resistance was detectable in 31% of the isolates. Most of S. aureus isolates were sensitive to linezolid (91%). Penicillin resistance was detectable in 81% of the tested isolates. Sensitivity to sulfamethoxazole/trimethoprim and ciprofloxacin was detected in 70% of the isolates (Fig. 1). Supplementary Table 1 describes the antimicrobial susceptibility pattern of each tested isolate.
Virulence of S. aureus isolates
The activity of different virulence factors produced by the tested isolates was determined and the isolates were given arbitrary scores based on the activity of each virulence factor. The scores of each virulence factor activity in each isolate are given in Supplementary Table 3.
Delta-hemolysin activity
Delta-hemolysin activity was determined by measuring the diameter of hemolysis zone in sheep-blood agar plates. About 42% of the isolates had moderate hemolysin activity while 26% and 11% of the isolates had high and low hemolysin activity, respectively. The remaining isolates (21%) lacked any hemolysin activity under the same conditions. Representative results with different scores are given in Supplementary Fig. 1.
Extracellular protease activity
The extracellular protease activity was tested in skim milk agar plates. The protease activity was high in 17% of the isolates, moderate in 43% of the isolates, and weak in 11% of the isolates, while 29% of the isolates lacked any protease activity. Representative results with different scores are given in Supplementary Fig. 2.
Biofilm production
Out of 100 tested isolates, only 4% were non-biofilm producers including 6% of MRSA isolates, while 62% were weak biofilm producers including 57% of MRSA isolates. Moderate biofilm formation was detectable in 31% of the tested isolates (n = 31) including nearly 35% of MRSA isolates, while strong biofilm production was detectable in three isolates including 2% of the isolates that were MRSA. Representative example of biofilms formed by isolates with different scores is given in Supplementary Fig. 3.
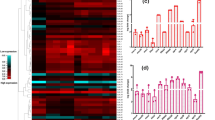
The level for production of each virulence factor by tested S. aureus clinical isolates is given in Fig. 2. The total virulence score of each isolate was determined by sum up of the scores recorded for the three measured virulence determinants (protease, hemolysin, and biofilm) and ranged from 0 to 9 (Supplementary Table 3).
Detection and expression level of spdC gene
Detection of spdC gene
spdC gene was detected in all the tested isolates by conventional PCR. Supplementary Fig. 4 shows representative examples of PCR products of spdC gene amplification visualized on agarose gel electrophoresis. Sequencing of the resulting PCR product from S. aureus ATCC 25923 confirmed 100% identity of the amplified product to the corresponding sequence of spdC gene of S. aureus. The sequenced product was deposited in GenBank under accession number ON367942.
Level of spdC gene expression
Due to financial limitations, the level of spdC gene expression was determined in eight selected isolates only. These isolates represented different virulence levels and antimicrobial resistance (AMR) patterns as follows: MRSA and high virulence score (isolates 13 and 78, total score = 8), MRSA and low virulence score (isolates 33 and 83, total score = 1 and 2, respectively), MSSA and high virulence score (isolates 65 and 25, total score = 8 and 7, respectively), and MSSA and low virulence score (isolates 93 and 30, total score = 0 and 1, respectively).
The specificity of the used primers was confirmed by the melting curves of the amplification products that showed only one single peak (Supplementary Fig. 5). spdC expression level varied between 0.08 and 1.18 that recorded in the standard S. aureus ATCC 25923. Five isolates had spdC expression level < 0.6 that of the standard strain. Three of these isolates were MRSA including two with a high virulence score (isolates 78 and 13) and one with a low virulence score (isolate 33). The other two isolates were MSSA and had a high virulence score (isolates 25 and 65). Three isolates had spdC expression level > 0.6 that of the standard S. aureus ATCC 25923; all had a low virulence score: one was MRSA (isolate 83) and the other two were MSSA (isolates 30 and 93; Table 1).
Correlation between spdC gene expression level and virulence
Discordant moderate correlation was recorded between the level of expression of spdC gene and the activity of the tested virulence factors, using Kendall’s Tau correlation coefficient. However, this correlation was not significant (p = 0.16, 0.23, 0.39, or 0.24 for protease, hemolysin, biofilm, or the total virulence scores, respectively; Table 1).
Correlation between spdC gene expression level and AMR
Moderate concordant correlation was recorded between spdC gene expression level and the AMR to vancomycin, linezolid, tetracycline, gentamicin, clindamycin, and chloramphenicol, while weak concordant correlation was recorded with erythromycin resistance, using Kendall’s Tau correlation coefficient. On the other hand, Kendall’s Tau correlation coefficient indicated discordant correlation between spdC gene expression level and the AMR in the case of sulfamethoxazole/trimethoprim (weak correlation) and ciprofloxacin (moderate correlation). This correlation between spdC gene expression level and the AMR whether concordant or discordant, measured using Kendall’s Tau correlation coefficient, was insignificant (p > 0.05).
The median of spdC gene expression level was higher in resistant isolates, compared to susceptible isolates, to most of the tested antibiotics (vancomycin, linezolid, erythromycin, tetracycline, gentamicin, clindamycin, and chloramphenicol). In the case of sulfamethoxazole/trimethoprim and ciprofloxacin, lower median of spdC gene expression level was detected in resistant isolates. However, in all tested antibiotics, no significant difference was detectable between the median of spdC gene expression level in isolates with different resistance pattern, using the Wilcoxon rank-sum test (Supplementary Table 4).
Weak discordant correlation was recorded between methicillin resistance (cefoxitin resistance) phenotype and the spdC gene expression level, with higher median of spdC gene expression in MSSA isolates. However, no significant correlation was recorded between spdC gene expression level and MRSA phenotype using either Kendall’s Tau correlation coefficient (p = 0.62) or the Wilcoxon rank-sum test (p = 0.73).
Discussion
S. aureus expresses a vast array of virulence factors that impact disease progression and severity (Liu 2009). Antibiotic resistance and emergence of MDR isolates, in addition to unavailability of effective vaccine, complicate the treatment of S. aureus infections (Cheung et al. 2021). SpdC protein was described as a global regulator of S. aureus virulence that affects biofilm formation and pathogenesis in addition to altering resistance toward compounds targeting cell wall (Poupel et al. 2018).
The rates of antimicrobial susceptibility patterns and virulence in our isolates were similar to those recorded elsewhere. AMR was predominant among the tested isolates, where 81% of the isolates were MDR and about half of the isolates (49%) were MRSA. Similar rates of MRSA infection were reported previously in Egypt, Africa, and the Middle East (Elshimy et al. 2018; Elsayed et al. 2018; Zigmond et al. 2014), as well as in other regions worldwide (Diekema et al. 2019).
High rates of resistance to tested antibiotics such as penicillin, cefoxitin, tetracycline, clindamycin, erythromycin, gentamicin, and chloramphenicol were also detectable. Most of the isolates (81%) were penicillin-resistant; this penicillin resistance rate is slightly lower than those recorded in other studies in 2018 and 2020, where penicillin resistance rates exceeded 90% (Elsayed et al. 2018; Kashef et al. 2020; Manandhar et al. 2018). Similar observations regarding the increased susceptibility to penicillin in S. aureus were described previously (Butler-Laporte et al. 2018; Chabot et al. 2015). This might arise from the limited use of penicillin in treatment of S. aureus infections where restricting the use of a certain antimicrobial may help in restoring its activity (Baym et al. 2016; Maher et al. 2012).
Vancomycin and linezolid are considered among the last-line treatments of MRSA infections. Resistance to linezolid was detected in 9% of the tested isolates which was higher than the rates recorded previously for linezolid resistance in Egypt (Kashef et al. 2020) and other regions of the world (Gu et al. 2013; Quiles-Melero et al. 2013). Vancomycin resistance was detected in 31% of the tested isolates. This is a considerably high rate, where vancomycin is the drug of choice for MRSA treatment. Higher and lower rates of vancomycin resistance were recorded previously in Egypt (Al-Amery et al. 2019; Elsayed et al. 2018; Kashef et al. 2020; El Refai et al. 2014) and other areas (Alzolibani et al. 2012; Yilmaz and Aslantaş 2017). According to the meta-analysis carried by Wu et al. (2021), the highest prevalence of vancomycin-resistant S. aureus was reported in Africa (16%) with Nigeria having the highest prevalence rate (29%). In the USA, the threat of vancomycin resistance was reduced (CDC 2019).
Biofilm formation by S. aureus plays an important role in chronic disease progression and increases tolerance to antibiotics (Lister and Horswill 2014). All except four of the collected isolates were biofilm-forming (96%) which was similarly reported previously (Kashef et al. 2020; Omidi et al. 2020; Piechota et al. 2018). In addition, about 95% of the MDR isolates were biofilm-forming. The high rates of biofilm formation among MDR isolates were recorded in previous studies (Kwon et al. 2008). Similarly, we detected moderate number of isolates with the ability to produce other tested virulence factors as protease and hemolysin (71% and 79%, respectively).
All tested isolates carried spdC gene in their genome. This is in accordance with a previous study that confirmed the presence of spdC gene in all sequenced staphylococci isolates (Gründling et al. 2006). However, Yamaguchi et al. (2019) recently detected a vancomycin intermediate–resistant S. aureus strain that lacked spdC gene. In our study, we isolated 17 strains with intermediate vancomycin resistance and none of our strains lacked the spdC gene. Other mechanisms might be responsible for vancomycin intermediate resistance phenotype as single nucleotide polymorphisms in genes responsible for cell wall biosynthesis. The effect of spdC gene deletion on vancomycin resistance needs to be further confirmed.
Interestingly, there was no significant correlation between biofilm formation and the level of the spdC gene expression in our clinical isolates. The measured Kendall’s Tau correlation coefficient between spdC gene expression level and biofilm formation indicated moderate discordant correlation. This opposed the results reported earlier about highly reduced biofilm formation in spdC mutant (Poupel et al. 2018). No studies are available on the correlation between biofilm formation and spdC gene expression in clinical isolates.
This may indicate the indirect effect of SpdC on biofilm formation. Other regulators are documented to be implicated directly in controlling biofilm formation as WalKR, RNAIII, SarA, and SigB. SpdC negatively affects the expression of WalKR regulon which in turn is known to positively affect biofilm formation (Dubrac et al. 2007; Paharik and Horswill 2016; Poupel et al. 2018). Other studies are still required to elucidate the exact molecular mechanism for SpdC effect on biofilm formation.
Similar to the lack of significant correlation between spdC gene expression level and biofilm formation, we also failed to find any significant correlation between spdC gene expression level and extracellular protease activity in our isolates. Proteases are important virulence factors that can cleave and degrade several important host proteins, including the heavy chains of all human immunoglobulin classes, plasma proteinase inhibitor, and elastin. In addition, proteases also play a role in S. aureus invasiveness by degrading bacterial cell surface proteins responsible for bacterial adhesion such as protein A and fibronectin-binding protein (Karlsson and Arvidson 2002).
S. aureus produces a number of proteases. These include two cysteine proteases (staphopain A, ScpA; and staphopain B, SspB), a metalloprotease (aureolysin), a serine protease (V8 or SspA), and six serine-like proteases (Spls) that are SspA homologs (SplABCDEF) (Lehman et al. 2019). Synthesis of extracellular proteases is activated by the accessory gene regulator quorum-sensing system (Agr) and repressed by SarA protein (Lehman et al. 2019). SpdC was suggested to activate SplB and SplC (serine-like proteases B and C) and staphopain thiol protease (SspB) (Poupel et al. 2018). We reported discordant correlation between spdC gene expression level and the phenotypic protease activity. This may be caused by the predominant effect of one or more proteases over the others and/or the predominant effect of other regulators on proteases’ expression.
Also, no significant correlation between spdC expression level and delta-hemolysin activity was detectable; however, moderate discordant association was recorded. Poupel et al. (2018) reported the lack of SpdC effect on S. aureus delta-hemolysin gene. SpdC induces the expression of gamma-hemolysin; this may be due to the positive effect of SpdC on the expression of the TCS saeRS and its negative effect on the WalKR system (Poupel et al. 2018). However, delta-hemolysin production in S. aureus is mainly regulated by the Agr system (Divyakolu et al. 2019).
Similar to the results of Poupel et al. (2018) about the increased susceptibility to cell wall active agents in spdC mutant, we found a moderate concordant correlation between spdC gene expression level and the AMR especially in cell wall active agents as vancomycin and penicillin. However, this correlation was not significant. These agents are cell wall antibiotics that inhibit the late stages of cell wall biosynthesis (Sarkar et al. 2017). Only with methicillin (cefoxitin) resistance, discordant correlation was recorded.
The effect of SpdC on the susceptibility to antibiotics with targets other than cell wall biosynthesis was also tested in this study (ciprofloxacin, sulfamethoxazole/trimethoprim, chloramphenicol, erythromycin, clindamycin, tetracycline, gentamicin, and linezolid), where no significant correlation was detectable between their susceptibility pattern and spdC gene expression level. Although, there was moderate concordant correlation between spdC gene expression level and the AMR to these agents, except with ciprofloxacin and sulfamethoxazole/trimethoprim, where discordant correlation was detectable.
Collectively, we failed to record any significant correlation between spdC gene expression level and virulence or antimicrobial susceptibility in clinical S. aureus isolates. Similar observation regarding the lack of direct effect of spdC and other spd genes on gene transcription or translation was reported previously (Frankel et al. 2010). This may reflect the predominant effect of other virulence regulators in S. aureus as WalKR, Agr, SaeRS, SrrAB, ArlSR, and LytRS, in addition to SarA and SarA homologs (Pragman and Schlievert 2004; Bronner et al. 2003). However, these regulators might be partially affected by SpdC level indicated by the moderate correlation detected in most cases.
Previous studies have reported the predominant effect of one regulator over the other on various virulence factors such as the predominant negative effect of SarA over the positive effect of the Agr system on the protease activity (Karlsson and Arvidson 2002), also the dominant effect of Sae regulator over the effect of σB regulator on virulence gene expression in S. aureus during device-related infection (Goerke et al. 2005). Single nucleotide polymorphism in spdC gene might affect the level of different virulence regulators that are affected by SpdC.
The results of this study are preliminary, and more studies are still required to confirm this conclusion. This study has a limitation of the small number of isolates tested for spdC gene expression level (eight isolates). Testing this association on larger number of isolates is urgently required to confirm the results of this study together with the possible effect of the isolate source. spdC mutants need to be prepared from clinical isolates to confirm the role of SpdC on virulence and antimicrobial resistance. Also, the association between spdC gene expression level and the level of expression of other virulence genes needs to be tested. In addition, the exact mechanism of spdC effect on the expression and regulation of different virulence factors is required to be elucidated.
Conclusion
The increased rates of antibiotic resistance together with the prevalence of virulent strains in hospitals is alarming and urges for an effective infection control strategy. The previously reported role of SpdC protein as virulence regulator in S. aureus isolates needs further evaluation together with the determination of the predominant regulators for each virulence factor.
References
Al-Amery K, Elhariri M, Elsayed A, El-Moghazy G, Elhelw R, El-Mahallawy H, El Hariri M, Hamza D (2019) Vancomycin-resistant Staphylococcus aureus isolated from camel meat and slaughterhouse workers in Egypt. Antimicrob Resist Infect Control 8:129. https://doi.org/10.1186/s13756-019-0585-4
Alzolibani AA, Al RAA, Al SHA, Bilal JA, Ahmad MI, Bin SG (2012) Documentation of vancomycin-resistant Staphylococcus aureus (VRSA) among children with atopic dermatitis in Qassim region, Saudi Arabia. J Am Acad Dermatol 21:51–53. https://doi.org/10.1016/j.jaad.2011.11.683
Baym M, Stone LK, Kishony R (2016) Multidrug evolutionary strategies to reverse antibiotic resistance. Science 351(6268):aad3292. https://doi.org/10.1126/science.aad3292
Bleul L, Francois P, Wolz C (2022) Two-component systems of S. aureus: signaling and sensing mechanisms. Genes 13:34. https://doi.org/10.3390/genes13010034
Boswihi SS, Udo EE (2018) Methicillin-resistant Staphylococcus aureus : an update on the epidemiology, treatment options and infection control. Curr Med Res Pract 8:18–24. https://doi.org/10.1016/j.cmrp.2018.01.001
Bronner S, Monteil H, Prévost G (2003) Regulation of virulence determinants in Staphylococcus aureus: complexity and applications. FEMS Microbiol Rev 28:183–200. https://doi.org/10.1016/j.femsre.2003.09.003
Butler-Laporte G, Lee TC, Cheng MP (2018) Increasing rates of penicillin sensitivity in staphylococcus aureus. Antimicrob Agents Chemother 62(7):e00680-e718. https://doi.org/10.1128/AAC.00680-18
CDC (2011) Healthcare-associated infections. https://www.cdc.gov/hai/organisms/staph.html. Accessed on 16 September 2021
CDC (2019) Antibiotic resistance threats in the United States, 2019. U.S. Department of Health and Human Services, CDC, Atlanta, GA.
Chabot MR, Stefan MS, Friderici J, Schimmel J, Larioza J (2015) Reappearance and treatment of penicillin-susceptible Staphylococcus aureus in a tertiary medical centre. J Antimicrob Chemother 70:3353–3356. https://doi.org/10.1093/jac/dkv270
Cheung GYC, Bae JS, Otto M (2021) Pathogenicity and virulence of Staphylococcus aureus. Virulence 12:547–569. https://doi.org/10.1080/21505594.2021.187868816
Christensen GD, Simpson WA, Younger JJ, Baddour LM, Barrett FF, Melton DM, Beachey EH (1985) Adherence of coagulase-negative staphylococci to plastic tissue culture plates: a quantitative model for the adherence of staphylococci to medical devices. J Clin Microbiol 22:996–1006. https://doi.org/10.1128/jcm.22.6.996-1006.1985
CLSI (2012a) Performance standards for antimicrobial disk susceptibility tests: approved standard - eleventh edition. CLSI document M02-A11. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania, USA
CLSI (2012b) Methods for dilution antimicrobial susceptibility tests for bacteria that grow aerobically ; approved standard — ninth edition, CLSI document M07-A9. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania, USA.
CLSI (2017). Performance standards for antimicrobial susceptibility testing. 27th ed. Supplement M100. Clinical and Laboratory Standards Institute, Wayne, Pennsylvania, USA.
Diekema DJ, Pfaller MA, Shortridge D, Zervos M, Jones RN (2019) Twenty-year trends in antimicrobial susceptibilities among Staphylococcus aureus from the SENTRY Antimicrobial Surveillance Program. Open Forum Infect Dis 6:S47–S53. https://doi.org/10.1093/ofid/ofy270
Divyakolu S, Chikkala R, Ratnakar KS, Sritharan V (2019) Hemolysins of Staphylococcus aureus—an update on their biology, role in pathogenesis and as targets for anti-virulence therapy. Adv Infect Dis 09:80–104. https://doi.org/10.4236/aid.2019.92007
Dubrac S, Boneca IG, Poupel O, Msadek T (2007) New insights into the WalK/WalR (YycG/YycF) essential signal transduction pathway reveal a major role in controlling cell wall metabolism and biofilm formation in Staphylococcus aureus. J Bacteriol 189:8257–8269. https://doi.org/10.1128/JB.00645-07
Dubrac S, Bisicchia P, Devine KM, Msadek T (2008) A matter of life and death: cell wall homeostasis and the WalKR (YycGF) essential signal transduction pathway. Mol Microbiol 70:1307–1322. https://doi.org/10.1111/j.1365-2958.2008.06483
El Refai KH, Ghoniem E, El Hendawy G, Abdel Moteleb T, Hassan H (2014) Characterization of vancomycin-resistant Staphylococcus aureus in the National Liver Institute. Menoufia Med J 27:825–832. https://doi.org/10.4103/1110-2098.149802
Elsayed N, Ashour M, Amine AEK (2018) Vancomycin resistance among staphylococcus aureus isolates in a rural setting. Egypt. GERMS 8:134–139. https://doi.org/10.18683/germs.2018.1140
Elshimy R, Khattab RA, Zedan H, Hosny AE-DS, Elmorsy TH (2018) Study on prevalence and genetic discrimination of methicillin-resistant Staphylococcus aureus (MRSA) in Egyptian hospitals. African J Microbiol Res 12:629–646. https://doi.org/10.5897/ajmr2017.8742
Foster TJ (2017) Antibiotic resistance in Staphylococcus aureus. Current status and future prospects. FEMS Microbiol Rev 41:430–449. https://doi.org/10.1093/femsre/fux007
Frankel MB, Wojcik BM, Dedent AC, Missiakas DM, Schneewind O (2010) ABI domain-containing proteins contribute to surface protein display and cell division in Staphylococcus aureus. Mol Microbiol 78:238–252. https://doi.org/10.1111/j.1365-2958.2010.07334.x
Goerke C, Fluckiger U, Steinhuber A, Bisanzio V, Ulrich M, Bischoff M, Patti JM, Wolz C (2005) Role of Staphylococcus aureus global regulators sae and σB in virulence gene expression during device-related infection. Infect Immun 73:3415–3421. https://doi.org/10.1128/IAI.73.6.3415-3421.2005
Gründling A, Missiakas DM, Schneewind O (2006) Staphylococcus aureus mutants with increased lysostaphin resistance. J Bacteriol 188:6286–6297. https://doi.org/10.1128/JB.00457-06
Gu B, Kelesidis T, Tsiodras S, Hindler J, Humphries RM (2013) The emerging problem of linezolid-resistant Staphylococcus. J Antimicrob Chemother 68:4–11. https://doi.org/10.1093/jac/dks354
Karlsson A, Arvidson S (2002) Variation in extracellular protease production among clinical isolates of Staphylococcus aureus due to different levels of expression of the protease repressor sarA. Infect Immun 70:4239–4246. https://doi.org/10.1128/IAI.70.8.4239-4246.2002
Kashef MT, Saleh NM, Ramadan MA (2020) The antimicrobial activity of ciprofloxacin-loaded niosomes against ciprofloxacin-resistant and biofilm-forming Staphylococcus aureus. Infect Drug Resist 13:1619–1629. https://doi.org/10.2147/IDR.S249628
Kaur G, Balamurugan P, Vasudevan S, Jadav S, Princy SA (2017) Antimicrobial and antibiofilm potential of acyclic amines and diamines against multi-drug resistant Staphylococcus aureus. Front Microbiol 15(8):1767. https://doi.org/10.3389/fmicb.2017.01767
Kwon AS, Park GC, Ryu SY, Lim DH, Lim DY, Choi CH, Park Y, Lim Y (2008) Higher biofilm formation in multidrug-resistant clinical isolates of Staphylococcus aureus. Int J Antimicrob Agents 32:68–72. https://doi.org/10.1016/j.ijantimicag.2008.02.009
Lehman MK, Nuxoll AS, Yamada KJ, Kielian T, Carson SD, Feya PD (2019) Protease-mediated growth of staphylococcus aureus on host proteins is opp3 dependent. Mbio 10:e02553-e2618. https://doi.org/10.1128/mBio.02553-18
Lister JL, Horswill AR (2014) Staphylococcus aureus biofilms: recent developments in biofilm dispersal. Front Cell Infect Microbiol 4:178. https://doi.org/10.3389/fcimb.2014.00178
Liu GY (2009) Molecular pathogenesis of Staphylococcus aureus infection. Pediatr Res 65(5 Pt 2):71R-77R. https://doi.org/10.1203/PDR.0b013e31819dc44d
Magiorakos AP, Srinivasan A, Carey RB, Carmeli Y, Falagas ME, Giske CG, Harbarth S, Hindler JF, Kahlmeter G, Olsson-Liljequist B, Paterson DL, Rice LB, Stelling J, Struelens MJ, Vatopoulos A, Weber JT, Monnet DL (2012) Multidrug-resistant, extensively drug-resistant and pandrug-resistant bacteria: an international expert proposal for interim standard definitions for acquired resistance. Clin Microbiol Infect 18:268–281. https://doi.org/10.1111/j.1469-0691.2011.03570.x
Mahdally NH, George RF, Kashef MT, Al-ghobashy M, Murad FE, Attia AS (2021) Staquorsin : a novel Staphylococcus aureus Agr-mediated quorum sensing inhibitor impairing virulence in vivo without notable resistance development. Front Microbiol 12:700494. https://doi.org/10.3389/fmicb.2021.700494
Maher MC, Alemayehu W, Lakew T, Gaynor BD, Haug S, Cevallos V, Keenan JD, Lietman TM, Porco TC (2012) The fitness cost of antibiotic resistance in Streptococcus pneumoniae: insight from the field. PLoS ONE 7:e29407. https://doi.org/10.1371/journal.pone.0029407
Manandhar S, Singh A, Varma A, Pandey S, Shrivastava N (2018) Biofilm producing clinical staphylococcus aureus isolates augmented prevalence of antibiotic resistant cases in tertiary care hospitals of Nepal. Front Microbiol 9:2749. https://doi.org/10.3389/fmicb.2018.02749
Mesrati I, Saidani M, Jemili M, Ferjeni S, Slim A, Ben BIB (2018) Virulence determinants, biofilm production and antimicrobial susceptibility in Staphylococcus aureus causing device-associated infections in a Tunisian hospital. Int J Antimicrob Agents 52:922–929. https://doi.org/10.1016/j.ijantimicag.2018.05.004
Naimi HM, Rasekh H, Noori AZ, Bahaduri MA (2017) Determination of antimicrobial susceptibility patterns in Staphylococcus aureus strains recovered from patients at two main health facilities in Kabul. Afghanistan BMC Infect Dis 17:737. https://doi.org/10.1186/s12879-017-2844-4
Neuhäuser M (2011). In: Lovric M (ed) International encyclopedia of statistical science, 2011th edn. Springer, Berlin, Heidelberg, p 32
Nitulescu G, Margina D, Zanfirescu A, Olaru OT, Nitulescu GM (2021) Targeting bacterial sortases in search of anti-virulence therapies with low risk of resistance development. Pharm (Basel) 14(5):415. https://doi.org/10.3390/ph14050415
Omidi M, Firoozeh F, Saffari M, Sedaghat H, Zibaei M, Khaledi A (2020) Ability of biofilm production and molecular analysis of spa and ica genes among clinical isolates of methicillin-resistant Staphylococcus aureus. BMC Res Notes 13:19. https://doi.org/10.1186/s13104-020-4885-9
Paharik AE, Horswill AR (2016) The Staphylococcal biofilm: adhesins, regulation, and host response. Microbiol Spectr 4: https://doi.org/10.1128/microbiolspec.VMBF-0022-2015
Piechota M, Kot B, Frankowska-Maciejewska A, Gruzewska A, Woźniak-Kosek A (2018) Biofilm formation by methicillin-resistant and methicillin-sensitive Staphylococcus aureus strains from hospitalized patients in Poland. Biomed Res Int 2018:4657396. https://doi.org/10.1155/2018/4657396
Poupel O, Proux C, Jagla B, Msadek T, Dubrac S (2018) SpdC, a novel virulence factor, controls histidine kinase activity in Staphylococcus aureus. PLoS Pathog 14:e1006917. https://doi.org/10.1371/journal.ppat.1006917
Pragman AA, Schlievert PM (2004) Virulence regulation in Staphylococcus aureus: the need for in vivo analysis of virulence factor regulation. FEMS Immunol Med Microbiol 42:147–154. https://doi.org/10.1016/j.femsim.2004.05.005
Procop GW, Church DL, Hall GS, Janda WM (2017). In: Procop GW (ed) Koneman’s color atlas and textbook of diagnostic, 7th edn. Wolters Kluwer Health, Philadelphia, USA.
Puka L (2011) Kendall’s Tau. In: Lovric M. (ed) International encyclopedia of statistical science, 2011edn. Springer: Berlin, Heidelberg, pp. 14–32.
Quiblier C, Zinkernagel AS, Schuepbach RA, Berger-Bächi B, Senn MM (2011) Contribution of SecDF to Staphylococcus aureus resistance and expression of virulence factors. BMC Microbiol 11:72. https://doi.org/10.1186/1471-2180-11-72
Quiles-Melero I, Gómez-Gil R, Romero-Gómez MP, Sánchez-Díaz AM, De Pablos M, García-Rodriguez J, Gutiérrez A, Mingorance J (2013) Mechanisms of linezolid resistance among staphylococci in a tertiary hospital. J Clin Microbiol 51:998–1001. https://doi.org/10.1128/JCM.01598-12
Reddy PN, Srirama K, Dirisala VR (2017) An update on clinical burden, diagnostic tools, and therapeutic options of Staphylococcus aureus. Infect Dis Res Treat 10:117991611770399. https://doi.org/10.1177/1179916117703999
Sambrook J, Russell DW (2001) Molecular cloning : a laboratory manual, 3rd edn. Cold Spring Harbor Laboratory Press, Cold Spring Harbor, N.Y.
Sarkar P, Yarlagadda V, Ghosh C, Haldar J (2017) A review on cell wall synthesis inhibitors with an emphasis on glycopeptide antibiotics. Medchemcomm 8:516–533. https://doi.org/10.1039/c6md00585c
Tong SYC, Davis JS, Eichenberger E, Holland TL, Fowler VG (2015) Staphylococcus aureus infections: epidemiology, pathophysiology, clinical manifestations, and management. Clin Microbiol Rev 28:603–661. https://doi.org/10.1128/CMR.00134-14
Torlak E, Korkut E, Uncu AT, Şener Y (2017) Biofilm formation by Staphylococcus aureus isolates from a dental clinic in Konya, Turkey. J Infect Public Health 10:809–813. https://doi.org/10.1016/j.jiph.2017.01.004
Uhlemann A-C, Otto M, Lowy FD, DeLeo FR (2014) Evolution of community- and healthcare-associated methicillin-resistant Staphylococcus aureus. Infect Genet Evol 21:563–574. https://doi.org/10.1016/j.meegid.2013.04.030
van Hal SJ, Jensen SO, Vaska VL, Espedido BA, Paterson DL, Gosbell IB (2012) Predictors of mortality in staphylococcus aureus bacteremia. Clin Microbiol Rev 25:362–386. https://doi.org/10.1128/CMR.05022-11
World Health Organization (2017) WHO News. https://www.who.int/news/item/27-02-2017-who-publishes-list-of-bacteria-for-which-new-antibiotics-are-urgently-needed. Accessed on 16 may 2021
World Health Organization (2020) Antimicrobial resistance. https://www.who.int/news-room/fact-sheets/detail/antimicrobial-resistance. Accessed on 17 may 2021
Wu Q, Sabokroo N, Wang Y, Hashemian M, Karamollahi S, Kouhsari E (2021) Systematic review and meta-analysis of the epidemiology of vancomycin-resistance Staphylococcus aureus isolates. Antimicrob Resist Infect Control 10:101. https://doi.org/10.1186/s13756-021-00967-y
Yamaguchi T, Ando R, Matsumoto T, Ishii Y, Tateda K (2019) Association between cell growth and vancomycin resistance in clinical community-associated methicillin-resistant Staphylococcus aureus. Infect Drug Resist 12:2379–2390. https://doi.org/10.2147/IDR.S209591
Yilmaz EŞ, Aslantaş Ö (2017) Antimicrobial resistance and underlying mechanisms in Staphylococcus aureus isolates. Asian Pac J Trop Med 10:1059–1064. https://doi.org/10.1016/j.apjtm.2017.10.003
Zhang H, Zheng Y, Gao H, Xu P, Wang M, Li A, Miao M, Xie X, Deng Y, Zhou H, Du H (2016) Identification and characterization of Staphylococcus aureus strains with an incomplete hemolytic phenotype. Front Cell Infect Microbiol 6:146. https://doi.org/10.3389/fcimb.2016.00146
Zigmond J, Pecan L, Hájek P, Raghubir N, Omrani AS (2014) MRSA infection and colonization rates in Africa and Middle East: a systematic review & meta-analysis. Int J Infect Dis 21:391. https://doi.org/10.1016/j.ijid.2014.03.1227
Acknowledgements
The authors thank Dr. Salwa I. Bakr, the professor of clinical pathology and immunology, Faculty of Medicine, Ain Shams University, for her help in obtaining the clinical isolates and Dr. Mohsen N. Fakhry for help during the statistical analysis.
Funding
Open access funding provided by The Science, Technology & Innovation Funding Authority (STDF) in cooperation with The Egyptian Knowledge Bank (EKB).
Author information
Authors and Affiliations
Contributions
Conceptualization: Mona Kashef, Alaa El-Dien Hosny, and Mohammed Ramadan; investigation and methodology: Mayada Bakr and Mona Kashef; resources: Mayada Bakr and Mona Kashef; supervision: Mona Kashef, Alaa El-Dien Hosny, and Mohammed Ramadan; writing – original draft: Mayada Bakr; writing – review and editing: Mona Kashef and Mohammed Ramadan. All authors read and approved the final manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Bakr, M.E., Kashef, M.T., Hosny, A.ED.M.S. et al. Effect of spdC gene expression on virulence and antibiotic resistance in clinical Staphylococcus aureus isolates. Int Microbiol 25, 649–659 (2022). https://doi.org/10.1007/s10123-022-00249-6
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10123-022-00249-6