Abstract
Knowledge of SGS in plants is vital to understand the ecological and evolutionary dynamics of populations and to plan conservation strategies. Some of the major factors that can affect spatial genetic structure (SGS) in plants are the level of gene flow, spatial arrangement and life stages of individuals within populations. Applying six highly variable microsatellite markers, we investigated the effect of these factors on spatial genetic structure selecting two natural populations of sycamore maple, which is an insect-pollinated, autotetraploid and an indigenous hardwood species in Germany and in other central European countries. The two study populations had different shapes (“compact” and “elongated”) and tree densities. Significant SGS extended to ~180 m in the elongated population and to ~35 m in the compact population. Juvenile plants of the compact population showed significant SGS up to 40 m. Estimate of Sp statistic in high-density population was almost double of that in the population with low density. Gene dispersal distance in the low-density population was about 9 times higher than in the population with high density. The similar level of significant SGS in both adult and juvenile plants suggested minimal or no effect of life stages of individuals on SGS in the sycamore maple population. The data presented in this study can provide guidelines for seed collection and to establish populations for the conservation and management of genetic resources of the species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Fine-scale spatial genetic structure (SGS) describes the distribution of genetic variants of individuals or groups over two-dimensional space (Epperson 1992). It provides insight into the dynamics of different ecological and evolutionary forces such as gene flow, genetic drift, inbreeding depression and natural selection, which shape the amount and distribution of genetic diversity within populations (Epperson 1993; Heywood 1991). The information about SGS is also vital for designing strategies for conservation of genetic resources in natural populations, for example, for selecting seed collection stands for ex-situ conservation of genetic resources (Finkeldey and Mátyás 2003). Furthermore, the lack of knowledge about the genetic structure within populations can lead to a biased assessment of other biological phenomena such as mating system and gene flow within plant populations (Ellstrand et al. 1978; Ritland 1985).
Genetic structuring of individuals within a continuous population can occur as a result of distance barriers for gene dispersal between sub-populations, known as “isolation-by-distance (IBD)” process (Wright 1943). Under the IBD process, limits to dispersal of pollen and seeds result in spatially non-random distribution of individuals, and the strength of SGS negatively correlates with the dispersal distance. Characterization of SGS can be performed using several autocorrelation methods based on the relationship between pairwise spatial and genetic distances (e.g., Loiselle et al. 1995; Smouse and Peakall 1999; Rousset 2000; Hardy 2003).
The genetic variation over short distances in sexually reproducing individuals may occur due to the combined effects of various factors, e.g., gene flow, natural selection, genetic drift and spatial arrangement of individuals (Endler 1977; Epperson 1993; Doligez et al. 1998). These factors are affected by the demography of a population such as population density and age. Reduced level of population density can increase inbreeding as a result of restricted gene flow between the fragments of populations, consequently creating clusters of individuals with similar genetic information. SGS has been found to be inversely correlated with adult tree density within populations (Vekemans and Hardy 2004).
Since plants are immobile, dispersal of pollen and seeds is spatially limited, and therefore, occurrence of SGS is common in trees with limited pollen and seed dispersal. However, SGS across plant species is not consistent (reviewed in Heywood 1991; Vekemans and Hardy 2004). Many studies in forest tree species reported no significant SGS as a result of extensive gene flow through pollen and seed (e.g., Epperson and Allard 1989; Knowles 1991). In contrast, other studies reported significant SGS within populations mainly because of restriction in gene flow through pollen and seed (e.g., Streiff et al. 1998; Vornam et al. 2004; Ng et al. 2006).
Evolutionary and ecological forces act throughout the life stages of populations; therefore, SGS can vary in different life stages (Epperson and Alvarez-Buylla 1997). An understanding of SGS in various life stages provides information about how ecological and evolutionary forces such as gene flow and selection act on various cohorts of plant species. Some studies that have taken the life stages into consideration reported greater significant SGS in early stages than in adults due to thinning or density-dependent mortality in the successive demographical stages (e.g., Epperson and Alvarez-Buylla 1997; Ng et al. 2004; Hardesty et al. 2005), while others reported increase in SGS in adults as compared to early stages as a result of low recruitment in the early stage, overlapping of generations in the adult cohort and increased effects of selection in the successive demographical stages (e.g., Kalisz et al. 2001; Latouche-Halle et al. 2003).
Sycamore maple (Acer pseudoplatanus L.) is an indigenous forest tree species in Germany and in other central European countries. It is a tall tree with a round and dense crown that reaches a height of about 40 m at the age of 150 years and can attain a diameter of 60–70 cm. It is one of the important species of the mixed montane forest zone in Central Europe and can grow from 300 m up to 2,000 m altitude (Spaethmann and Namvar 1985). Besides its common use for garden and landscape management, it is also important for forest management because of its ecological and economic value. The wood of the sycamore maple is of high quality and is used in many sectors of industry and handicraft.
Sycamore maple has been reported to be autotetraploid with chromosomes 2n = 4x = 52 (Darlington and Wylie 1955). Flowers are monoecious and most of them are morphologically hermaphroditic but functionally unisexual (De Jong 1976). On the basis of its morphological characteristics and flowering ecology, especially the structure and distribution of pollen kits on the pollen wall, it has been categorized as an insect- and wind-pollinated species (Hesse 1979). Seeds of sycamore maple are characterized as recalcitrant (Dickie et al. 1991), and they are primarily dispersed by wind (Greene and Johnson 1989). However, dispersal of seeds by water can also occur along the water bodies (http://www.dpi.vic.gov.au/dpi/vro/vrosite.nsf/pages/invasive_sycamore_maple).
Marker-based genetic studies in this species were initiated by Konnert (1992), who used three highly variable enzyme systems for clone identification. Furthermore, Konnert et al. (2001) investigated the inheritance pattern at 25 enzyme gene loci in single trees. Complexities in interpreting the zymograms of some of the enzyme systems due to the tetrasomic inheritance pattern of the species have been reported (Konnert 1992; Konnert et al. 2001). Investigation of chloroplast DNA markers in 19 populations of sycamore maple from different parts of Europe has led to the identification of 22 different haplotypes, and some unique haplotypes in both southern and central European populations were reported (Petit et al. 2003). However, so far, no studies have reported on the genetic diversity and structure of the species using highly variable molecular markers such as nuclear microsatellites.
The main aims of the present study are (1) to assess the SGS of sycamore maple in two natural populations with different spatial distribution patterns of trees and (2) to compare the SGS between adults and juveniles. Out of the two populations, one is compact and dense, while the other is more elongated and the trees are distributed in small clumps with lower density. Two populations with different densities in the same area were chosen to analyze potential differences in gene flow within natural populations of sycamore maple. We postulated that strong SGS can be found in the elongated population with lower population density and consisting of small groups of clumped trees due to more restricted gene flow as compared to the compact population. As a result of restricted seed dispersal, there is a possibility for early recruitment of juveniles close to each other with similar genetic information, and due to this effect, we expect to observe higher level of SGS in juveniles than in adults of sycamore maple.
We hope that the findings of this study will not only contribute to understanding SGS in adult and juveniles of sycamore maple, but can also help in effective planning for conservation and management of genetic resources within natural populations of the species. The results will also enhance our understanding about SGS of temperate broad-leaved trees.
Materials and methods
Sample collection
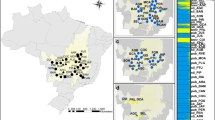
Sampling was carried out in two natural populations of sycamore maple, i.e., Södderich and Weisswassertal. Both populations are part of the Reinhausen forest district and are located near Göttingen, Germany (Södderich (longitude: 010° 01′ 52′′E; latitude: 51° 33′ 43′′ N; altitude: 317 m asl) and Weisswassertal (longitude: 010° 04′ 51′′E; latitude: 51° 34′ 28′′ N; altitude: 224 m asl). The shape of the Weisswassertal population is linear, and it is stretched about 2 km along the narrow valley of the Weisswassertal creek with small clumps of trees, while the Södderich population is compact and located on a flat area. The average slope of Weisswassertal population is about 3%. Total areas of the Södderich and Weisswassertal populations are about 3 and 65 ha, respectively. Density of reproductive individuals of sycamore maple in the Södderich was about 45, and in the Weisswassertal it was 1.26 trees per ha. The Weisswassertal population is isolated from the nearest sycamore population as it was surrounded by the populations of other species, while the Södderich is embedded within a large population of sycamore maple. The two populations are located about 2.50 km apart. They are mixed stands of beech (Fagus sylvatica), ash (Fraxinus execlsior), pedunculate oak (Quercus robur) and sycamore maple (Acer pseudoplatanus). Buds from 137 and 82 adult trees were collected from the Södderich and Weisswassertal populations, respectively. From the Södderich population, 112 juveniles were also sampled to compare the genetic diversity and SGS between adult and juveniles. The nearest juvenile of an estimated age of 2–5 years was sampled from each of the adult trees whenever it was available. Geographical positions of each individual tree of both populations and juveniles from the Södderich population were recorded by compass survey (Fig. 1).
DNA extraction and genotyping
DNA extraction and genotyping of six microsatellite markers were carried out following the protocols from Pandey et al. (2004) and Pandey (2005).
Statistical analysis
Genetic diversity
Since sycamore maple is an autotetraploid species, individual trees showed a maximum of four different alleles at a locus with the average number of alleles ranging from 1.5 to 2.65 per sample. Therefore, it was difficult to determine the exact number of copies of an allele present in a particular heterozygote. This made the interpretation of genotypes complex. Since we were not able to assign a correct genotype to the heterozygous individuals that had two or three bands, we scored each allele as “1” for presence and “0” for absence. Although we accept that this approach of scoring genotypes can have biased results of genetic diversity parameters, as a result of the complexity of the banding pattern this was the only applicable approach. Similar problems were also reported by many other authors (e.g., Samadi et al. 1999; Korbecka et al. 2003; Lian et al. 2001; Saltonstall 2003; Kosman and Leonard 2005), who developed or used microsatellite markers in polyploid species, and most of them scored the bands according to their presence or absence, i.e., as dominant markers, e.g., “1” for presence and “0” for absence of bands.
Spatial genetic structure (SGS)
Spatial genetic structures of the adult trees in the two populations, i.e., Södderich and Weisswassertal and the natural regeneration of the Södderich population, were estimated following the Loiselle et al. (1995) pairwise kinship coefficient (Fij) method using the computer program SPAGEDI (Hardy and Vekemans 2002). This method is based on the principle of probability of identity by descent of alleles in sampled genotypes in relation to their spatial location. The size of the distance class was based on the even sample size in each distance class. Statistical significance (95% upper and lower confidence intervals) of the observed pairwise kinship coefficient (Fij) was determined after 10,000 permutations.
Sp statistic and gene dispersal distance
In order to quantify SGS and to compute neighborhood sizes and gene dispersal distances in the two sycamore populations, Sp statistic was estimated as described in Vekemans and Hardy (2004). Using the Sp statistic, Wright’s neighborhood size (Nb) (Wright 1943, 1946) was also estimated. Nb is defined as Nb = 4 πDeσ 2g , where σ 2g is the axial variance of gene dispersal and De the effective population density. Estimate of gene dispersal distance (σg) was performed assuming ratios of effective to census density, De/D, of 0.1 and 0.5 (Vekemans and Hardy 2004; De-Lucas et al. 2009). Estimates of −^ bF and ^ F1 were carried out using SPAGEDHI program (Hardy and Vekemans 2002). Statistical significance (P ≤ 0.05) of the analyses was performed by permutation tests with 10,000 replications.
Results
Genetic diversity
The numbers of alleles observed at the six microsatellite loci in the Södderich and the Weisswassertal populations are presented in Table 1. The total number of alleles found in the Weisswassertal population (N a = 85) was slightly higher than in the Södderich population (N a = 80). Accordingly, the average number of alleles observed in the Weisswassertal population was 14.2 and 13.3 in the Södderich population. The number of alleles observed at microsatellite loci MAP-2, MAP-12 and MAP-33 was higher in the Weisswassertal as compared to the Södderich population. On the other hand, microsatellite loci MAP-9, MAP-12 and MAP-46 had more alleles in the Södderich than in the Weisswassertal population (Table 1). The average number of alleles in the natural regeneration was 11.50 with a total number of 69 alleles. The lower average number of alleles in juveniles could be due to a lower number of samples of juveniles than of adults.
Spatial genetic structure (SGS)
The results of SGS analyses in the Södderich and Weisswassertal populations are presented in Fig. 2 and Table 2. In the adult trees of the Södderich population, statistically significant positive SGS was observed up to ~35 m, while significant negative SGS was observed at the distance classes of 60–71, 83–96 and 194–227 m.
Correlograms (solid line) showing kinship coefficient (Fij) (Loiselle et al. 1995) of the Södderich and the Weisswassertal populations of sycamore maple, with 95% confidence regions (dotted lines), which were obtained after 10,000 permutations
Although the kinship coefficient (Fij = 0.025) in the Weisswassertal population is weaker than in the Södderich population, a significant positive SGS was found to extend up to ~180 m. A negative significant SGS was observed at the distance classes of 434–525 and 611–691 m.
The juveniles of the Södderich population showed a similar level of significant positive SGS (~40 m) as the adults. However, the level of positive SGS in terms of pairwise kinship coefficient (Fij) value was considerably higher in adult trees (Fij = 0.064) than in the juveniles (Fij = 0.036), resulting in a larger magnitude of significant SGS in adults.
Sp statistics and gene dispersal estimates
Results of Sp statistics and dispersal estimates in the two sycamore populations are shown in Table 3. Sp value in the Södderich (Sp = 0.023) was about 2 times higher than in the Weisswassertal population (Sp = 0.011). Whereas gene flow distance (σg) in the Weisswassertal was almost nine times higher (340–761 m) than in the Södderich population (39–88 m).
Discussion
Our study revealed that sycamore maple possesses significant SGS in the smaller to medium distance classes. The level of SGS was different in the two populations of sycamore maple with different forms and densities. In the Södderich population, clumping of individuals with similar genetic information up to ~35 m was observed, while in the Weisswassertal population, significant SGS extended up to ~180 m. The variation in the level of SGS in the Södderich and Weisswassertal populations of the sycamore maple could be the result of differences in densities of reproducing individuals. The Södderich population possesses a higher density of trees and has a more or less square form (Fig. 1) that may have facilitated higher levels of gene flow by promoting random mating among trees, whereas the Weisswassertal population is isolated from other sycamore maple populations and it is stretched along the narrow valley of the Weisswassertal creek with a lower density of reproducing trees. In agreement with our results, Vekemans and Hardy (2004) reviewed the SGS of 48 studies of five plant species and found an inverse correlation between density and the level of SGS. On the other hand, gene dispersal distances were found to be higher in populations with low density (Vekemans and Hardy 2004). A similar pattern was found also in this study, indicating that sycamore maple is capable of compensating at least partially the direct effect of density on SGS. The area and the shape of the populations may have also affected the observed differences in the level of SGS of the two populations because the chances of random mating among trees are higher in the population with small area and square shape as compared to the population with larger area and elongated shape. However, in the present study, it is not possible to distinguish which of the factors, density, area or shape, has higher effect on the observed differences in SGS of the two populations.
The spatial genetic structure of sycamore maple observed in the Södderich population is slightly more pronounced as compared to SGS (20–32 m) reported in three natural populations of sugar maple in northwestern Ontario, Canada (Perry and Knowles 1991). Since both species have a similar pollination biology and seed dispersal mechanisms, i.e., winged fruits that are dispersed by wind and gravity, similar SGS can be expected. However, the results of spatial genetic clustering of related individuals up to a distance of ~180 m observed in the Weisswassertal population are not comparable with the findings of Perry and Knowles (1991) as they used more or less compact populations in their studies. They explained that restricted gene flow due to limited pollen and seed dispersal contributed to the local clumping of individuals with similar genetic information. This factor is also a likely the reason for the observed SGS in sycamore maple. The authors are not aware of any published study concerning seed dispersal distances in natural populations of sycamore maple. However, seed dispersal is reported to be by wind (Greene and Johnson 1989) and water (http://www.dpi.vic.gov.au/DPI/Vro/vrosite.nsf/pages/invasive_sycamore_maple). The winged fruits may allow the distribution by wind over wider distances. Fruits can fly hundreds of meters in strong winds in open areas. However, in dense forest stands the fruits can strike other trees and shrubs in the course of flight. Due to this effect, seed dispersal by wind is expected to be much more restricted in dense population than in an open forest area (Greene and Johnson 1989). As the Weisswassertal population is more open than the Södderich population, clumping of trees along the small valley in the Weisswassertal population may have occured as a result of seed dispersal by water and wind.
SGS of tree species can also be affected by the impact of flowering phenology involved in the mating system (Young and Merriam 1994). Individual trees of sycamore maple are heterodichogamous, being either protogynous or protoandrous (De Jong 1976). Due to this characteristic, there is a higher probability of individuals with asynchronous flowering within populations. In this case, the spatial distribution of potential mates depends on the temporal distribution of mature male and female flowers, which may result in genetic correlation among half-sib progenies at some spatial scale and in a local clustering of individuals bearing similar genetic information. Young et al. (1993) and Young and Merriam (1994) have considered the above-mentioned mechanisms to be one of the possible causes of the spatial genetic structure observed in A. saccharum, which is also a heterodichogamous tree species.
Similar to the results of our study, other studies in tree species with gravity-dispersed seeds reported significant SGS on shorter distance classes (e.g., Chung and Epperson 2000; Schueler et al. 2006) and the results were explained by the occurrence of overlapping seed shadow and restricted gene flow within populations. However, other studies (e.g., Chung et al. 2000) reported non-significant SGS due to long range seed dispersal combined with high population density in populations. Lower levels of significant SGS (20–30 m) than in this study were reported in a wind-pollinated species with gravity-dispersed seeds, Fagus sylvatica (Vornam et al. 2004; Dounavi et al. 2010). However, Streiff et al. (1998) reported a slightly higher level of significant SGS (up to 40 m) than in this study in other wind-pollinated species with heavy seeds, Quercus robur and Q. petraea by using microsatellite and allozyme gene markers.
Schnabel et al. (1991) studied the genetic structure of autotetraploid (Maclura pomifera) and diploid (Gleditsia triacanthos) co-occurring tree species and found that the SGS was slightly lower in the autotetraploid as compared to the diploid species. The authors argued that the rate of heterozygosity losses in polyploid species due to selfing and mating among relatives is much slower than for diploids, which would slow down the development of SGS in polyploids. The argument is relevant to the low level of significant genetic structure observed in sycamore maple, which is also an autotetraploid tree species with tetrasomic inheritance (Luo et al. 2006). However, the effects of polyploidization may differ depending on the pre-existing systems of incompatibility or self-sterility.
Contrary to our expectation and the findings of many studies, which have reported either increased (e.g., Kalisz et al. 2001; Latouche-Halle et al. 2003) or decreased levels (e.g., Ng et al. 2004; Hardesty et al. 2005) of SGS from seedlings to adults, our results showed similar levels of SGS in both cohorts of sycamore maple. Most of the studies argued that the different intensity of selection to the different life stages is the main reason for observing various levels of SGS in different life stages. However, this explanation does not fit to the findings in sycamore maple as the juveniles, and adult trees of the same population showed a similar level of SGS and indicated either similar levels or no effect of selection on both adult and juveniles of the species.
Conservation and management implications
The analysis of SGS in the two populations of sycamore maple confirmed the existence of small- to medium-scale family structures (Södderich: ~35 m and Weisswassertal: ~180 m). The results are relevant for selecting individual trees for the purpose of seed collection in stands of the respective dimensions. In order to avoid the collection of seeds from individuals with similar genetic characteristics, the distance between seed trees should be kept above 35 m in compact populations like the Södderich population and above 180 m in the isolated populations with lower tree densities like the Weisswassertal population. This can be useful to maintain genetic diversity in seed material and reduce the possibility of inbreeding later in planted stands. The result is particularly important for the Södderich population, since the stand has been approved for the collection of reproductive material according to EU rules. Since the microsatellites markers used in this study are presumed to be neutral, the guidelines proposed here can be relevant to any areas irrespective of environmental variations.
References
Chung MG, Epperson BK (2000) Clonal and spatial genetic structure in Eurya emarginata (Theaceae). Heredity 84:170–177
Chung MG, Chung MY, Oh GS, Epperson BK (2000) Spatial genetic structure in a Neolitsea sericea population (Lauraceae). Heredity 85:490–497
Darlington CD, Wylie AP (1955) Chromosome atlas of flowering plants. George Allen and Unwin LTD, London
De Jong PC (1976) Flowering and sex expression in Acer L. A biosystematic study. Mededelingen Landbouwhogeschool Wageningen, Netherlands
De-Lucas AI, González-Martínez SC, Vendramin GG, Hidalgo E, Heuertz M (2009) Spatial genetic structure in continuous and fragmented populations of Pinus pinaster Aiton. Mol Ecol 18:4564–4576
Dickie JB, May K, Morris SVA, Titley SE (1991) The effects of desiccation on seed survival in Acer platanoides L. and Acer pseudoplatanus L. Seed Sci Res 1:149–162
Doligez A, Baril C, Joly HI (1998) Fine-scale spatial genetic structure with non-uniform distribution of individuals. Genetics 148:905–919
Dounavi A, Koutsias N, Ziehe M, Hattemer HH (2010) Spatial patterns and genetic structures within beech populations (Fagus sylvatica L.) of forked and non-forked individuals. Eur J Forest Res 129:1191–1202
Ellstrand NC, Torres AM, Levin DA (1978) Density and the rate of apparent outcrossing in Helianthus annus (Asteraceae). Syst Bot 3:403–407
Endler JA (1977) Geographic variation, speciation, and clines. Princeton Univ. Press, Princeton
Epperson BK (1992) Spatial structure of genetic variation within populations of forest trees. New For 6:257–278
Epperson BK (1993) Recent advances in correlation analysis of spatial patterns of genetic variation. Evol Biol 27:95–155
Epperson BK, Allard RW (1989) Spatial autocorrelation analysis of the distribution of genotypes within populations of lodgepole pine. Genetics 121:369–377
Epperson BK, Alvarez-Buylla E (1997) Spatial autocorrelation analysis of family structure in multiple life stages of Cecropia obtusifolia. Evolution 51:275–282
Finkeldey R, Mátyás G (2003) Genetic variation of oaks (Quercus spp.) in Switzerland. 3. Lack of impact of postglacial recolonization history on nuclear gene loci. Theor Appl Genet 106:346–352
Greene DE, Johnson EA (1989) A model of wind dispersal of winged or plumed seeds. Ecology 70:339–347
Hardesty BD, Dick CW, Kremer A et al (2005) Spatial genetic structure of Simarouba amara Aubl. (Simaroubaceae), a dioecious, animal-dispersed, Neotropical tree, on Barro Colorado Island, Panama. Heredity 95:290–297
Hardy OJ (2003) Estimation of pairwise relatedness between individuals and characterization of isolation-by-distance processes using dominant genetic markers. Mol Ecol 12:1577–1588
Hardy OJ, Vekemans X (2002) SPAGeDi: a versatile computer program to analyse spatial genetic structure at the individual or population levels. Mol Ecol Notes 2:618–620
Hesse M (1979) Ultrastruktur und Verteilung des Pollenkitts in der insekten- und windblütingen Gattung Acer (Aceraceae). Pl Syst Evol 131:277–289
Heywood JS (1991) Spatial analysis of genetic variation in plant populations. Annu Rev Ecol Syst 22:335–355
Kalisz S, Nason JD, Hanzawa FM et al (2001) Spatial population genetic structure in Trillium grandiflorum: the roles of dispersal, mating, history, and selection. Evolution 55:1560–1568
Knowles P (1991) Spatial genetic structure within two natural stands of black spruce (Picea mariana (Mill.) B. S. P.). Silvae Genet 40:13–19
Konnert M (1992) Isozymanalysen bei Bergahorn (Acer pseudoplatanus L.)-Klonidentifizierung. Biochemische Untersuchungen zur Genetik von Waldbaumpopulationen, Schriftenreihe der Landesanstalt für Forstwirtschaft Nordrhein-Westfalen. pp 77–84
Konnert M, Ruetz W, Fromm M (2001) Genetic variation in Acer pseudoplatanus L. Inheritance of isozyme variants. For Genet 8:25–37
Korbecka G, Vrieling K, Squirrell J et al (2003) Characterization of six microsatellite loci in Echium vulgare (Boraginaceae). Mol Ecol Notes 3:274–276
Kosman E, Leonard KJ (2005) Similarity coefficients for molecular markers in studies of genetic relationships between individuals for haploid, diploid, and polyploidy species. Mol Ecol 14:415–424
Latouche-Halle C, Ramboer A, Bandou E, Caron H, Kremer A (2003) Nuclear and chloroplast genetic structure indicate fine-scale spatial dynamics in a Neotropical tree population. Heredity 91:181–190
Lian C, Nara K, Nakaya H et al (2001) Development of microsatellite markers in polyploid Salix reinii. Mol Ecol Notes 1:160–161
Loiselle BA, Sork VL, Nason J, Graham C (1995) Spatial genetic structure of a tropical understory shrub, Psychotria officinalis (Rubiaceae). Am J Bot 82:1420–1425
Luo ZW, Zhang Z, Zhang RM, Pandey M, Gailing O, Hattemer HH, Finkeldey R (2006) Modeling population genetic data in autotetraploid species. Genetics 172:639–646
Ng KKS, Lee SL, Koh CL (2004) Spatial structure and genetic diversity of two tropical tree species with contrasting breeding systems and different ploidy levels. Mol Ecol 13:657–669
Ng KKS, Lee SL, Saw LG, Plotkin JB, Koh CL (2006) Spatial structure and genetic diversity of three tropical tree species with different habitat preferences within a natural forest. Tree Genet Genom 2:121–131
Pandey M (2005) Development of microsatellites in sycamore maple (Acer pseudoplatanus L.) and their application in population genetics. Ph. D. Dissertation. Georg-August University of Goettingen, Germany (available at: http://www.uni-goettingen.de/de/98152.html#pandey)
Pandey M, Gailing O, Fischer D, Hattemer R, Finkeldey HH (2004) Characterization of microsatellite markers in sycamore (Acer pseudoplatanus L.). Mol Ecol Notes 4:253–255
Perry DJ, Knowles P (1991) Spatial genetic structure within three sugar maple (Acer saccharum Marsh.) stands. Heredity 66:137–142
Petit RJ, Aguinagalde I, de Beaulieu J-L et al (2003) Glacial refugia: hotspots but not melting pots of genetic diversity. Science 300:1563–1565
Ritland K (1985) The genetic-mating structure of subdivided populations. I. Open-mating model. Theor Pop Biol 27:51–74
Rousset F (2000) Genetic differentiation between individuals. J Evol Biol 13:58–62
Saltonstall K (2003) Microsatellite variation within and among North American lineages of Phragmites australis. Mol Ecol 12:1689–1702
Samadi S, Mavárez J, Pointier JP et al (1999) Microsatellite and morphological analysis of population structure in the parthenogenetic freshwater snail Melanoides tuberculata: insights into the creation of clonal variability. Mol Ecol 8:1141–1153
Schnabel A, Laushman RH, Hamrick JL (1991) Comparative genetic structure of two co-occurring tree species, Maclura pomifera (Moraceae) and Gleditsia triacanthos (Leguminosae). Heredity 67:357–364
Schueler S, Tusch A, Scholz F (2006) Comparative analysis of the within-population genetic structure in wild cherry (Prunus avium L.) at the self-incompatibility locus and nuclear microsatellites. Mol Ecol 15:3231–3243
Smouse PE, Peakall R (1999) Spatial autocorrelation analysis of individual multiallele and multilocus genetic structure. Heredity 82:561–573
Spaethmann W, Namvar K (1985) Der Bergahorn und die Gattung Acer. Allg Forst 42:1126–1131
Streiff R, Labbe T, Bacilieri R et al (1998) Within-population genetic structure in Quercus robur L. and Quercus petraea (Matt.) Liebl. Assessed with isozymes and microsatellites. Mol Ecol 7:317–328
Vekemans X, Hardy OJ (2004) New insights from fine-scale spatial genetic structure analyses in plant populations. Mol Ecol 13:921–934
Vornam B, Decarli N, Gailing O (2004) Spatial distribution of genetic variation in a natural beech stand (Fagus sylvatica L.) based on microsatellite markers. Cons Genet 5:561–570
Wright S (1943) Isolation by distance. Genetics 28:114–138
Wright S (1946) Isolation by distance under diverse systems of mating. Genetics 31:39–59
Young AG, Merriam HG (1994) Effects of forest fragmentation on the spatial genetic structure of Acer saccharum Marsh. (Sugar maple) populations. Heredity 72:201–2008
Young AG, Merriam HG, Warwick SI (1993) The effect of forest fragmentation on genetic variation in Acer saccharum Marsh. (sugar maple) populations. Heredity 71:277–289
Acknowledgments
This research was financially supported by Deutsche Forschungsgemeinschaft under DFG-Project Number Ha 501/32-1. We gratefully acknowledge the technical assistance of Mr. Thomas Seliger and Ms. Olga Artes.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Additional information
Communicated by U. Berger.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Pandey, M., Gailing, O., Hattemer, H.H. et al. Fine-scale spatial genetic structure of sycamore maple (Acer pseudoplatanus L.). Eur J Forest Res 131, 739–746 (2012). https://doi.org/10.1007/s10342-011-0546-9
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10342-011-0546-9