Abstract
Here we show that verrucomicrobial community structure and abundance are extremely sensitive to changes in chemical factors linked to soil fertility. Terminal restriction fragment length polymorphism fingerprint and real-time quantitative PCR assay were used to analyze changes in verrucomicrobial communities associated with contrasting soil nutrient conditions in tropical regions. In case study Model I (“Slash-and-burn deforestation”) the verrucomicrobial community structures revealed disparate patterns in nutrient-enriched soils after slash-and-burn deforestation and natural nutrient-poor soils under an adjacent primary forest in the Amazonia (R = 0.819, P = 0.002). The relative proportion of Verrucomicrobia declined in response to increased soil fertility after slash-and-burn deforestation, accounting on average, for 4 and 2 % of the total bacterial signal, in natural nutrient-poor forest soils and nutrient-enriched deforested soils, respectively. In case study Model II (“Management practices for sugarcane”) disparate patterns were revealed in sugarcane rhizosphere sampled on optimal and deficient soil fertility for sugarcane (R = 0.786, P = 0.002). Verrucomicrobial community abundance in sugarcane rhizosphere was negatively correlated with soil fertility, accounting for 2 and 5 % of the total bacterial signal, under optimal and deficient soil fertility conditions for sugarcane, respectively. In nutrient-enriched soils, verrucomicrobial community structures were related to soil factors linked to soil fertility, such as total nitrogen, phosphorus, potassium and sum of bases, i.e., the sum of calcium, magnesium and potassium contents. We conclude that community structure and abundance represent important ecological aspects in soil verrucomicrobial communities for tracking the changes in chemical factors linked to soil fertility under tropical environmental conditions.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Members of the phylum Verrucomicrobia have been shown to make up 1–10 % of the total bacterial 16S rRNA in soils (Buckley and Schmidt 2001, 2003; Lupatini et al. 2013; Navarrete et al. 2015a, b). Bergmann et al. (2011) recognized the dominance of Verrucomicrobia in soil bacterial communities across a range of biomes in Antarctica, Europe, and the Americas. The broad distribution of Verrucomicrobia in soils suggests that they are important members of terrestrial ecosystems (Buckley and Schmidt 2001; Felske and Akkermans 1998). Despite the fact that members of this phylum have typically been recalcitrant to cultivation methods (Janssen et al. 1997, 2002; Janssen 1998; Joseph et al. 2003; Davis et al. 2005), new strategies for isolation, as well as novel methods for detection of sought-after microorganisms on solid media, have yielded more Verrucomicrobia isolates (Stevenson et al. 2004; Sangwan et al. 2005; Pol et al. 2007; Islam et al. 2008). As a consequence, genomic and physiological characterizations of Verrucomicrobia isolates have contributed to the knowledge of their biology and ecology (Isanapong et al. 2012; Wertz et al. 2012). However, most information on the ecology of Verrucomicrobia is revealed by the correlations of 16S rRNA gene abundance with environmental parameters, such as soil moisture and soil physicochemical factors (Buckley and Schmidt 2001; Jordaan and Bezuidenhout 2013; Pan et al. 2014).
The culture-independent approach based on the direct recovery of bacterial 16S rRNA genes from tropical soils has revealed the occurrence of Verrucomicrobia from different Brazilian biomes: Amazon (Borneman and Triplett 1997; Kim et al. 2007; Navarrete et al. 2010, 2015b), Atlantic forest (Bruce et al. 2010), Cerrado (Quirino et al. 2009) and Pampa (Lupatini et al. 2013). However, the information acquired is still not sufficient as a systematic identification of taxa responding to the alterations in soil chemical factors. In soils from the Amazon region, Verrucomicrobia were present at different abundance under diverse soil management practices, opening the possibilities to investigate verrucomicrobial community as bioindicator of tropical soil management effects.
Biological processes, species or communities can serve as successful bioindicators (Holt and Miller 2011). A new perspective in microbial ecology has emerged due to the progresses of molecular biology, allowing the interpretation of ecological aspects by culture-independent approaches. Microbial community structure, including the number of species and the relative abundance of species, has been assessed by molecular fingerprinting techniques (Burlage 1998). Microbial community abundance, an ecological concept referring to the relative representation of a community in a particular ecosystem, can be estimated using quantitative molecular approaches (Smith and Osborn 2009). These ecological aspects of microbial communities can be statistically related to environmental parameters, such as soil chemical factors (Jesus et al. 2009; Kuramae et al. 2010, 2012; Navarrete et al. 2013; Pan et al. 2014).
In the present study, we examined changes in the verrucomicrobial community associated with high soil fertility after slash-and-burning deforestation in the Amazonia (Model I), and decline in soil fertility associated with different management practices for sugarcane (Model II). For this purpose, the verrucomicrobial community structure was assessed by terminal restriction fragment polymorphism (T-RFLP) analysis, and the verrucomicrobial community abundance was estimated by real-time quantitative PCR (qPCR). The explicit relationship between the verrucomicrobial community structure and soil chemical factors was examined by multivariate statistical analyses.
Materials and methods
Soil was sampled in two contrasting soil nutrient content situations: Model I (Slash-and-burn deforestation): soil samples were collected in nutrient-enriched soils after slash-and-burn deforestation and natural nutrient-poor soils under adjacent primary forest in three discontinuous areas in the Amazonian region. Model II (Management practices for sugarcane): soil samples were collected from sugarcane rhizosphere with optimal and deficient soil nutrients in a greenhouse mesocosm experiment. Fieldwork was conducted under legal authorization (SISBIO 4845833).
Model I: slash-and-burn deforestation
Study sites and soil sampling
Soil samples were collected in three discontinuous areas located in the Southeastern Brazilian Amazon, State of Mato Grosso, Brazil: area 1 (15°11′45″S and 59°03′31″W), area 2 (14°21′38″S and 57°21′27″W) and area 3 (13°21′57″S and 54°54′24″W) described previously (Navarrete et al. 2015b). The three discontinuous sampling areas were considered replicates, and soil samples were collected from deforested sites identified just after forest clearing and adjacent primary forest sites exactly as described in Navarrete et al. (2015b). Samples were transported to the laboratory under ice and stored at −20 °C until processing within 72 h after sampling. Soil fertility properties were determined previously for the same soil samples used in this study by Navarrete et al. (2015a, b) (Supplementary Table 1).
Soil DNA isolation and verrucomicrobial 16S rDNA T-RFLP fingerprinting analysis
DNA was extracted from 250 mg soil samples from deforested and forest sites using the Power Soil DNA Isolation Kit (Mo Bio Laboratories Inc., Carlsbad, CA, USA), according to the manufacturer’s instructions. DNA extraction was performed in duplicate for each soil sample and quantified using Qubit dsDNA BR Assay Kit (Invitrogen, Carlsbad, CA, USA). DNA concentrations were adjusted to 20 ng µl−1 and stored at −20 °C until use. T-RFLP analysis was used to characterize the verrucomicrobial community structure in soils from deforested sites and forest sites. Verrucomicrobial 16S rRNA gene fragments were amplified in duplicate 25 μl reactions using the primers VMB537f (O’Farrell and Janssen 1999) and 1378r (Heuer et al. 1997). The forward primer was labeled with hexacarboxifluorescein at the 5′end. Each 25 μl reaction mixture contained 2.5 μl of reaction buffer 10× (Invitrogen, Carslad, CA, USA), 0.75 μl of MgCl2 (50 mM), 0.5 μl of each primer (10 µM), 0.2 U of Platinum Taq DNA Polymerase (Invitrogen), 0.5 μl of each dNTP (1 mM), 10 ng of bovine serum albumin (BSA; 10 mg ml−1) and 10 ng of template DNA. The following conditions were chosen for amplification after optimization: initial denaturation for 5 min at 95 °C; 35 cycles of 1 min at 95 °C, 30 s at 60 °C, and 1.5 min at 72 °C; and final extension for 10 min at 72 °C. The duplicate hexacarboxifluorescein-labeled PCR products for each sample were pooled and purified using GFX™ PCR DNA and Gel Band Purification Kit (Amersham Pharmacia Biotech, NJ, USA) after analysis by gel electrophoresis. Purified products were split into three tubes (175 ng in each tube) and digested in separate 15 μl-reactions with 10 U of the restriction enzymes AluI, MspI and HhaI (Invitrogen, Carlsbad, CA, USA) for 3 h at 37 °C. Fluorescently labeled terminal restriction fragments (TRF) were separated and detected using an ABI PRISM 3100 Genetic Analyzer capillary sequencer (Applied Biosystems, Foster City, CA, USA). Before injection, the samples were precipitated using sodium acetate/EDTA, and denatured in the presence of 10 μl formamide and 0.25 μl GS-500 ROX size standard (Applied Biosystems, Foster City, CA, USA). The TRF patterns were evaluated using Peak Scanner (Applied Biosystems) and T-REX (http://trex.biohpc.org/) software. TRFs were defined by aligning peaks using clustering threshold (range specified equal to 1.0). TRFs <50 bp or contributing to 0.5 % of the total TRF signal were excluded. Matrices (concatenated matrix with the three enzymes) for presence/absence were analyzed using CANOCO 4.5 (ter Braak and Šmilauer 2002) to generate ordination of T-RFLP patterns by principal component analysis (PCA). A distance matrix (Jaccard metric) was constructed from presence/absence data. This similarity matrix was used for ANOSIM statistics to investigate differences in soil verrucomicrobial community structure inhabiting nutrient-enriched soils after slash-and-burn deforestation in the Amazonia and natural nutrient-poor soils under adjacent primary forest. The magnitude of R indicates the degree of separation between two communities, with a score of ‘1’ indicating complete separation and ‘0’ indicating no separation. Calculation of similarity coefficient and ANOSIM were carried out using Primer six (version 6.1.5, Primer-E Ltd., Plymouth, UK). The explicit relationship between the verrucomicrobial community structure and soil chemical factors was examined by constrained ordination generated by a redundancy analysis (RDA) performed using CANOCO 4.5 (ter Braak and Šmilauer 2002).
Quantitative real-time PCR assay
Quantitative real-time PCR (qPCR) using the 16S rRNA marker gene was performed to assess the abundance of the verrucomicrobial and total bacterial communities using the same soil DNA samples as those for T-RFLP fingerprinting. As standards, 16S rRNA gene amplicons from Verrucomicrobia spinosum (DSMZ 4136) and a bacterial clone from an environmental sample were obtained by PCR using primers pA-F (5′-AGAGTTTGATCCTGGCTCAG-3′) (Edwards et al. 1989) and 1378R (5′-CGGTGTGTACAAGGCCCGGGAAGG-3′) (Heuer et al. 1997), purified (QIA-quick PCR purification kit, Qiagen, Venlo, the Netherlands) and ligated into the pGEM-T vector (Promega, Leiden, the Netherlands). Ligation products were transformed with E. coli JM109 competent cells (Promega, Leiden, the Netherlands). The presence or absence of the insert was determined by PCR using SP6 and T7 primers, and plasmid DNA was isolated (QIAprep Spin Miniprep Kit, Qiagen, Venlo, the Netherlands) from appropriate clones (i.e., belonging to the desired target group). DNA standard curves were generated by producing a dilution series of 104–108 copies µl−1 using duplicate ten-fold dilutions of isolated plasmid DNA. For qPCR of 16S rRNA gene fragments from Verrucomicrobia and from total bacteria, we used the following primer pairs: Ver53/Eub518 and Eub338/Eub518, respectively (Lane 1991; Muyzer et al. 1993; Stevenson et al. 2004). Each 25 µl reaction contained 12.5 µl QPCR SYBR green 2× reaction mix (Abgene, Epsom, UK), 1.25 µl of each primer (30 µM), 2.5 µl of bovine serum albumin (BSA; 10 mg ml−1) and 50 ng of template DNA. All mixes were made using a CAS-1200 pipetting robot (Corbett Research, Sydney, Australia). PCR conditions for Verrucomicrobia and total bacteria were described by Fierer et al. (2005) with the modification of annealing temperature (60 °C) and forward primer (Ver53) in the case of Verrucomicrobia. PCR amplifications and product quantification were performed using the Rotor-Gene 3000 (Corbett Research, Sydney, Australia). Melting curve analysis of amplicons was conducted to confirm that fluorescent signals originated from specific amplicons and not from primer-dimers or other artifacts. DNA samples were processed using five replicate reactions for each of six sampling sites in field. Automated analysis of PCR amplicon quality (for example, PCR baseline subtraction, Ct-threshold setting to the linear amplification phase) and quantity was performed with ROTOR-GENE 6 software (Corbett Research, Sydney, Australia). Statistical analyses of qPCR data were performed using the STATISTICA 10 package (StatSoft Inc, Tulsa, OK, USA). One-way ANOVA was used to determine significance of the differences among all soil samples. The comparison of soil samples was based on post hoc analysis using Tukey’s HSD test.
Model II: management practices for sugarcane
Experimental design, treatments and soil sampling
Sugarcane plants (Saccharum spp.) were grown in a greenhouse mesocosm experiment as described previously (Navarrete et al. 2015a). Six treatments and three replications were used in a completely randomized design. Mineral fertilizer was applied in the form of urea (450 g N kg−1) at a rate of 60 kg N ha−1 in treatments containing nitrogen fertilizer. Vinasse, a liquid residue of the ethanol distillation, was applied to the soil at a rate of 0.06 L kg−1 (120 m3 ha−1) in treatments containing vinasse as fertilizer. The experiment consisted of two conditions of soil-surface straw coverage: dry and chopped leaves from adult sugarcane plants straw (10 t ha−1) and uncovered surface. Accordingly, the experiment included the following treatments: N, nitrogen fertilizer; N + S, N fertilizer and straw; N + V, N and vinasse as fertilizers; N + V + S, N and V as fertilizers and straw; C, excluding any N, V fertilizer and straw (control); and C + S, excluding any N and V fertilizer and including straw. Ten sugarcane plants were grown in each mesocosm. Sugarcane plants were removed in pairs from each mesocosm at 50 and 150 days after planting and soil fertilization on the optimal and deficient soil fertility conditions for sugarcane, respectively, according to van Raij et al. (1996). Plants were healthy at 50 days after planting and they presented nutrient deficiency symptoms at 150 days after planting. Roots and associated soil were transported to the laboratory on ice and then processed to obtain the rhizosphere soil. The roots were shaken to remove the loose soil, and the tightly attached soil including small aggregates (< 0.5 cm) was used for DNA extraction. The fertility status of the soil on optimal and deficient soil nutrients in each experimental treatment was determined previously by Navarrete et al. (2015a) (Supplementary Table 2).
DNA isolation, verrucomicrobial 16S rDNA T-RFLP fingerprinting and quantitative real-time PCR assay
Soil DNA was extracted using the same conditions used for Model I. T-RFLP analysis was used to characterize the verrucomicrobial community structure in sugarcane rhizosphere soils from healthy and nutrient deficient symptom plants collected on the optimal and deficient soil fertility for sugarcane, respectively. T-RFLP fingerprinting of verrucomicrobial 16S rRNA genes, and qPCR analysis of verrucomicrobial and bacterial 16S rRNA genes were performed as described previously for Model I.
Results
Model I: slash-and-burn deforestation
PCA ordination based on verrucomicrobial T-RFLP data from restriction profiles generated by enzymes AluI, MspI and HhaI revealed distinct groups for nutrient-enriched soils after slash-and-burn deforestation and natural nutrient-poor soils under adjacent primary forest (Fig. 1). This grouping pattern was confirmed by a significant R-value (R = 0.819, P = 0.002) by analysis of similarity (ANOSIM) of the presence and absence of terminal restriction fragments in T-RFLP profiles. Soil samples from deforested sites were distinctly grouped in the ordination (R = 0.610, P = 0.003). However, verrucomicrobial community structure did not group according to the sampling sites in forest soils (R = 0.173, P = 0.003).
Principal component analysis plot based on the structure of soil verrucomicrobial communities as determined by T-RFLP analysis in nutrient-enriched soils after slash-and-burn forest clearing and natural nutrient-poor soils in an adjacent primary forest in three discontinuous areas after forest clearing and burning. Symbols refer to individual replicates (A, B, C, D and E) of the following sampling sites: open squares forest site located at Area 1 (FS1), open diamonds forest site located at Area 2 (FS2), open circle forest site located at Area 3 (FS3), black squares deforested site located at Area 1 (DS1), black diamonds deforested site located at Area 2 (DS2), black circle deforested site located at Area 3 (DS3)
The relationship between soil chemical characteristics and verrucomicrobial community structures revealed by T-RFLP fingerprinting for nutrient-enriched soils after slash-and-burn deforestation (deforested soils) and natural nutrient-poor soils (forest soils) were calculated by redundancy analysis (RDA). A total of 64.3 % of all variation was explained by the first two RDA axes (Fig. 2). RDA results showed that community structures from deforested soils were related to soil chemical factors linked to soil fertility, such as total nitrogen (Ntot), phosphorus (P), potassium (K), calcium (Ca), magnesium (Mg), sum of bases (the sum of Ca, Mg and K), and soil pH. Verrucomicrobial community structures from forest soils were related to manganese (Mn), copper (Cu), iron (Fe) and organic matter (OM) content as well as cation exchange capacity (CEC) and potential acidity (H + Al) (Fig. 2).
Constrained ordination diagram for sample plots (deforested and forest soil samples) in the first two redundancy analysis (RDA) axes based on the soil chemical characteristics of the different sampling sites and their relationship with the verrucomicrobial T-RFLP generated by restriction with enzymes AluI, MspI and HhaI. Symbols refer to individual replicates (A, B, C, D and E) of the following sampling sites: open squares forest site located at Area 1 (FS1), open diamonds forest site located at Area 2 (FS2), open circle forest site located at Area 3 (FS3), black squares deforested site located at Area 1 (DS1), black diamonds deforested site located at Area 2 (DS2), black circle deforested site located at Area 3 (DS3)
Real-time quantitative PCR revealed significant differences (P < 0.05) in 16S rRNA gene copies between deforested soils and forest soils with regard to the relative and absolute proportions of the verrucomicrobial community in each of three replicates of sampling area (Table 1). Verrucomicrobia accounted, on average, for 4 and 2 % of the total bacterial signal in the soil samples from forest and deforested sites, respectively.
Model II: management practices for sugarcane
PCA ordination of the T-RFLP profiles generated by enzymes AluI, MspI and HhaI revealed distinct groups of soil Verrucomicrobia under optimal and deficient soil fertility for sugarcane cultivation (Fig. 3). Disparate grouping was confirmed by a significant R-value (R = 0.786, P = 0.002) verified by ANOSIM based on the presence and absence of terminal restriction fragments in T-RFLP profiles. Verrucomicrobial community structure did not group according to the experimental treatments (R = 0.244, P = 0.002).
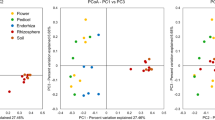
RDA showed relationships between soil chemical characteristics and verrucomicrobial community structures for sugarcane rhizosphere soil collected at optimal and deficient soil fertility for sugarcane. A total of 31.8 % of all variation was explained by the first two RDA axes (Fig. 4). Based on this multivariate analysis, the verrucomicrobial community structures from sugarcane rhizosphere receiving the optimal soil fertility for sugarcane were related to total N, P, K, Ca, Mn, Cu, Fe and OM contents, and soil pH and CEC (Fig. 4). In turn, verrucomicrobial community structures from sugarcane rhizosphere collected in nutrient-deficient soils were related to Zn and Mg contents as well as H + Al (Fig. 4).
Constrained ordination diagram for sample plots (sugarcane rhizosphere soil samples collected on optimal and deficient soil fertility for sugarcane) in the first two redundancy analysis (RDA) axes based on the soil chemical characteristics of the different soil treatments and their relationship with the verrucomicrobial T-RFLP data from restriction profiles generated by enzymes AluI, MspI and HhaI
ANOVA analyses were carried out on qPCR data targeting 16S rRNA gene abundances for total bacteria, as well as Verrucomicrobia (Table 2). These analyses did not show significant differences for total bacteria abundance between soil samples collected at the optimal and deficient soil fertility for sugarcane in any experimental treatments. However, the total number of 16S rRNA gene copies of rhizosphere Verrucomicrobia was significantly different between these two soil fertility conditions (Table 2). The proportion of the bacterial community that could be attributed to the phylum Verrucomicrobia, as based upon 16S rRNA gene copy detection, varied among soil samples from different experimental treatments, and it accounted, on average, for 2 and 5 % of the total bacterial signal, for soil samples collections taken from the optimal and deficient fertility soil, respectively (Table 2).
Discussion
Soils of the Amazon ecosystem are extremely low in all nutrients (Aubert and Tavernier 1972), characterized by low natural fertility, high exchangeable aluminium saturation, aluminium toxicity, K deficiency, high P fixation and low CEC (Cochrane and Sanchez 1982; Mendonça-Santos et al. 2006). Cultivation of acid soils in the Amazon is preceded by cutting and removing the economically important trees and burning the remaining aerial biomass (Fujisaka et al. 1996). Certini (2005) revised the effects of fire on properties of forest soils and reported that slash-and-burn deforestation contributes to the availability of micro- and macronutrients in the soil by releasing nutrients from OM. The resulting changes in soil nutrient availability in turn affects soil C and N dynamics by accelerating OM decomposition (Certini 2005). Navarrete et al. (2015b) reported short-term increase in soil fertility after slash-and-burn deforestation in Amazonia based on chemical analysis of the soil fertility status for the same soil samples used in the present study. The authors showed that slash-and-burn forest clearing in Amazonia decreased soil organic matter (OM) content and factors linked to soil acidity, and raised soil pH, base saturation, and the concentration of exchangeable bases. Nye and Greenland (1960) proposed the ‘nutrient-rich ash’ hypothesis to explain the observed increase in soil nutrient availability after slash-and-burn deforestation. The increased availability of nutrients from ash incorporation and soil OM combustion lead to increased soil fertility, with effects on soil bacterial communities composition and their potential functions (Navarrete et al. 2015b).
The distribution of the Verrucomicrobia phylum in soils (including plant rhizosphere soils) is variable and apparently extremely sensitive to changes in the environment (Kielak et al. 2008; Bruce et al. 2010; Pan et al. 2014; Navarrete et al. 2015a, b). Buckley and Schmidt (2001) showed that the distribution of rRNA from Verrucomicrobia in soil was affected by environmental characteristics that change in relation to time, soil history, and soil depth, and revealed that a statistically significant amount of the variation in verrucomicrobial 16S rRNA genes abundance can be explained by changes in soil moisture content together with other soil factors. In the present study, we showed that the verrucomicrobial community structure and abundance were affected by soil chemical factors linked to soil fertility, such as total N, P, K and sum of bases, i.e., the sum of calcium, magnesium and K contents.
Wertz et al. (2012) considered that Verrucomicrobia may exert a great impact with regard to nitrogen availability in certain ecosystems, including oligotrophic environments. This consideration was based on genomic data from TAV2, which is a member of the Opitutaceae from the gut of the termite Reticulitermes flavipes, and genomic data from the as-yet-unpublished Verrucomicrobium strain DG1235. As termite diets have a low N content, Verrucomicrobia may contribute the N pool within the gut ecosystem. Isanapong et al. (2013) showed some Verrucomicrobia seem to be well adapted to environments with low nitrogen availability by carrying genes for biosynthesis of amino acids and transcribing of those genes in situ. In our soils, higher verrucomicrobial community abundance was related to low N availability.
Huang et al. (2012) reported a decrease in abundance of Verrucomicrobia after increases in available nitrogen, phosphorus and potassium, and soil OM resulting from cotton straw application in soil. Pitombo et al. (2015) also found a negative effect of straw application in sugarcane on abundance of members of Verrucomicrobia. A negative effect of K content on abundance of Verrucomicrobia was found in grassland soil with yearly addition of inorganic fertilizers in a 54-year experiment (Pan et al. 2014) and sugarcane-cultivated soil with addition of vinasse in a short-term experiment (Navarrete et al. 2015a). Lima et al. (2014) showed that K also negatively affected Verrucomicrobia in Amazonian Dark Earth, one of the most fertile soils in the world. In addition, Mendes et al. (2015) reported lower abundance of Verrucomicrobia in deforested, agricultural and pasture soils than primary forest soils in Brazilian Amazonia, where the K content was higher than undisturbed soils. Our findings on verrucomicrobial community structure and abundance corroborate the previous findings that higher verrucomicrobial community abundance is associated with soils containing lower P and K contents.
Members of the phylum Verrucomicrobia have been shown to be present in varying plant-soil ecosystems (Chow et al. 2002; DeAngelis et al. 2009; Kielak et al. 2008; Nunes da Rocha et al. 2011, 2013; Sanguin et al. 2006; Zul et al. 2007). However, to date, there is no information about the verrucomicrobial community inhabiting the sugarcane rhizosphere soil. Nunes da Rocha et al. (2013) found specialized groups within several subdivisions of Verrucomicrobia that are leek rhizosphere competent. According to the authors, the lifestyles of these specialized groups of Verrucomicrobia with the plant may suggest that these bacteria either interact with the plants themselves or with other microbial communities associated with plants. Our results extend this current understanding of verrucomicrobial communities that associate with plants and suggest that these bacteria also interact with the soil chemical factors of the root-free soil surrounding of sugarcane plant roots, especially those related to soil fertility. The verrucomicrobial community structures from sugarcane rhizosphere given the optimal soil fertility for sugarcane were related to total N, P, K, Ca, Mn, Cu, Fe and OM contents. With regard to the relationship between Verrucomicrobia and soil OM content, the generally oligotrophic phylum Verrucomicrobia may be highly dependent on C availability due to a slow-growing life strategy (Bergmann et al. 2011; He et al. 2012). In this sense, although the effects of sugarcane crop residue on soil biological properties are expected to be more pronounced over long time periods (Graham et al. 2002), our results showed a tendency for increased abundance of Verrucomicrobia in soil with straw blanket coverage.
In our study, forest soils and sugarcane-cultivated soils on deficient fertility are considered to be oligotrophic since they contain low levels of nutrients and deforested soils and sugarcane-cultivated soils on optimal fertility as copiotrophic environments due to increase in nutrient availability after slash-and-burn forest clearing and fertilization, which increases nutrient availability (Carbonetto et al. 2014; Navarrete et al. 2015b). Under this assumption, bacteria in forest soils and sugarcane-cultivated soils on deficient fertility are expected to be k-selected and to present low growth rates and very efficient nutrient uptake systems with higher substrate affinities. In contrast, bacteria in deforested soils and sugarcane-cultivated soils on optimal fertility are expected to be r-selected and to have higher rates of activity per biomass unit, higher turnover rates and faster growth rates. Our results showed a trend toward these statements since a reduction in the abundance of Verrucomicrobia, a bacterial taxon with oligotrophic characteristics (Carbonetto et al. 2014; Navarrete et al. 2015b), was detected in deforested soils and sugarcane-cultivated soils on optimal fertility. Moreover, this is in agreement with recent findings that confirm a correlation between Verrucomicrobia abundance patterns and conditions of limited nutrient availability in tropical soil (Navarrete et al. 2015b). Culture-independent analysis of soil microbial communities revealed Verrucomicrobia as a bacterial phylum adapted to low substrate concentrations in soil (Noll et al. 2005). High-throughput sequencing of the same soil DNA samples used in this study revealed a high abundance of Verrucomicrobia at the phylum and class (Spartobacteria) levels in Amazon forest soils (Navarrete et al. 2015b), where the fertility is naturally low and maintained through litter nutrient cycling under high moisture condition (Fearnside 2005).
Based on our case studies (Models I and II), a clear link has been established between soil verrucomicrobial community structure and abundance and soil fertility status. Verrucomicrobial communities in the studied soils are consistent with an oligotrophic lifestyle, revealing decreased abundance and disparate community structure pattern on nutrient-enriched soils in comparison to nutrient-deficient soils. In conclusion, the community structure and abundance represent important ecological aspects in soil verrucomicrobial communities for tracking the changes in chemical factors linked to soil fertility under slash-and-burn deforestation and management practices for sugarcane.
References
Aubert G, Tavernier R (1972) Soil survey. In: Soils of the humid tropics. Committee on tropical soils. National Academy of Sciences, Washington DC, pp 17–46
Bergmann GT, Bates ST, Eilers KG, Lauber CL, Caporaso JG, Walters WA, Knight R, Fierer N (2011) The under-recognized dominance of Verrucomicrobia in soil bacterial communities. Soil Biol Biochem 43:1450–1455
Borneman J, Triplett EW (1997) Molecular microbial diversity in soils from Eastern Amazonia: evidence for unusual microorganisms and microbial population shifts associated with deforestation. Appl Environ Microbiol 63:2647–2653
Bruce T, Martinez IB, Neto OM, Vicente ACP, Kruger RH, Thompson FL (2010) Bacterial community diversity in the Brazilian Atlantic forest soils. Microb Ecol 60:840–849
Buckley DH, Schmidt TM (2001) Environmental factors influencing the distribution of rRNA from Verrucomicrobia in soil. FEMS Microbiol Ecol 35:105–112
Buckley DH, Schmidt TM (2003) Diversity and dynamics of microbial communities in soils from agro-ecosystems. Environ Microbiol 5:441–452
Burlage RS (1998) Molecular techniques. In: Burlage RS, Atlas R, Stahl D, Geesey G, Sayler G (eds) Techniques in microbial ecology. Oxford University Press, New York, pp 289–336
Carbonetto B, Rascovan N, Álvarez R, Mentaberry A, Vázquez MP (2014) Structure, composition and metagenomic profile of soil microbiomes associated to agricultural land use and tillage systems in Argentine Pampas. PLoS One 9(6):e99949
Certini G (2005) Effects of fire on properties of forest soils: a review. Oecologia 143:1–10
Chow ML, Radomski CC, McDermott JM, Davies J, Axelrood PE (2002) Molecular characterization of bacterial diversity in Lodgepole pine (Pinus contora) rhizosphere soils from British Columbia forest soils differing in disturbance and geographic source. FEMS Microbiol Ecol 42:347–357
Cochrane T, Sanchez P (1982) Land resource, soil and their management in the Amazon region: a state of knowledge report. In: Hecht S (ed) Amazon: agriculture and land use. Ciat, Cali, pp 137–209
Davis KER, Joseph SJ, Janssen PH (2005) Effects of growth medium, inoculum size, and incubation time on culturability and isolation of soil bacteria. Appl Environ Microbiol 71:826–834
DeAngelis KM, Brodie EL, DeSantis TZ, Andersen GL, Lindow SE, Firestone MK (2009) Selective progressive response of soil microbial community to wild oat roots. ISME J 3:168–178
Edwards U, Rogall T, Blocker H, Emde M, Bottger EC (1989) Isolation and direct complete nucleotide determination of entire genes—characterization of a gene coding for 16S ribosomal RNA. Nucleic Acid Res 17:7843–7853
Fearnside PM (2005) Desmatamento na Amazônia brasileira: História, índices e consequências. Megadiversidade 1:113–122
Felske A, Akkermans ADL (1998) Prominent occurrence of ribosomes from an uncultured bacterium of the Verrucomicrobiales cluster in grassland soils. Lett Appl Microbiol 26:219–223
Fierer N, Jackson JA, Vilgalys R, Jackson RB (2005) Assessment of soil microbial community structure by use of taxon-specific quantitative PCR assays. Appl Environ Microbiol 71:4117–4120
Fujisaka S, Bell W, Thomas N, Hurtado L, Crawford E (1996) Slash-and-burn agriculture, conversion to pasture, and deforestation in two Brazilian Amazon colonies. Agric Ecosyst Environ 59:115–130
Graham MH, Haynes RJ, Meyer JH (2002) Soil organic matter content and quality: effects of fertilizer applications, burning and trash retention on a long-term sugarcane experiment in South Africa. Soil Biol Biochem 34:93–102
He ZL, Piceno Y, Deng Y, Xu MY, Lu ZM, DeSantis T, Andersen G, Hobbie SE, Reich PB, Zhou JZ (2012) The phylogenetic composition and structure of soil microbial communities shifts in response to elevated carbon dioxide. ISME J 6:259–272
Heuer H, Krsek M, Baker P, Smalla K, Wellington EMH (1997) Analysis of actinomycete communities by specific amplification of genes encoding 16S rRNA and gel-electrophoretic separation in denaturing gradients. Appl Environ Microbiol 63:3233–3241
Holt EA, Miller SW (2011) Bioindicators: using organisms to measure environmental impacts. Nat Educ Knowl 3:8
Huang W, Bai Z, Hoefel D, Hu Q, Lv Q, Zhuang G, Xu S, Qi H, Zhang H (2012) Effects of cotton straw amendment on soil fertility and microbial communities. Front Environ Sci Eng 6:336–349
Isanapong J, Goodwin L, Bruce D, Chen A, Detter C, Han J, Han CS, Held B, Huntemann M, Ivanova N, Land ML, Mavromatis K, Nolan M, Pati A, Pennacchio L, Pitluck S, Szeto E, Tapia R, Woyke T, Rodrigues JLM (2012) High-quality draft genome sequence of the Opitutaceae bacterium strain TAV1, a symbiont of the wood-feeding termite Reticulitermes flavipes. J Bacteriol 194:2744–2745
Isanapong J, Hambright WS, Willis AG, Boonmee A, Callister SJ, Burnum KE, Paša-Tolić L, Nicora CD, Wertz JT, Schmidt TM, Rodrigues JLM (2013) Development of an ecophysiological model for Diplosphaera colotermitum TAV2, a termite hindgut Verrucomicrobium. ISME J 7:1803–1813
Islam T, Jensen S, Reigstad LJ, Larsen O, Birkeland N-K (2008) Methane oxidation at 55 °C and pH 2 by a thermoacidophilic bacterium belonging to the Verrucomicrobia phylum. Proc Nat Acad Sci USA 105:300–304
Janssen PH (1998) Pathway of Glucose catabolism by strain VeGlc2, an anaerobe belonging to the Verrucomicrobiales lineage of bacterial descent. Appl Environ Microbiol 64:4830–4833
Janssen PH, Schuhmann A, Moerschel E, Rainey FA (1997) Novel anaerobic ultramicrobacteria belonging to the Verrucomicrobiales lineage of bacterial descent isolated by dilution culture from anoxic rice paddy soil. Appl Environ Microbiol 63:1382–1388
Janssen PH, Yates PS, Grinton BE, Taylor PM, Sait M (2002) Improved culturability of soil bacteria and isolation in pure culture of novel members of the divisions Acidobacteria, Actinobacteria, Proteobacteria, and Verrucomicrobia. Appl Environ Microbiol 68:2391–2396
Jesus EC, Marsh TL, Tiedje JM, Moreira FMS (2009) Changes in land use alter the structure of bacterial communities in Western Amazon soils. ISME J 3:1004–1011
Jordaan K, Bezuidenhout CC (2013) The impact of physico-chemical water quality parameters on bacterial diversity in the Vaal River, South Africa. Afr J Oral Health Sci 39:385–395
Joseph SJ, Hugenholtz P, Sangwan P, Osborne CA, Janssen PH (2003) Laboratory cultivation of widespread and previously uncultured soil bacteria. Appl Environ Microbiol 69:7210–7215
Kielak A, Pijl AS, van Veen JA, Kowalchuk GA (2008) Differences in vegetation composition and plant species identity lead to only minor changes in soil-borne microbial communities in a former arable field. FEMS Microbiol Ecol 63:372–382
Kim JS, Sparovek G, Longo RM, De Melo WJ, Crowley D (2007) Bacteria diversity of terra preta and pristine forest soil from the western Amazon. Soil Biol Biochem 39:684–690
Kuramae EE, Gamper HA, Yergeau E, Piceno YM, Brodie EL, DeSantis TZ, Andersen GL, van Veen JA, Kowalchuk GA (2010) Microbial secondary succession in a chronosequence of chalk grasslands. ISME J 4:711–715
Kuramae EE, Yergeau E, Wong LC, Pijl AS, van Veen JA, Kowalchuk GA (2012) Soil characteristics more strongly influence soil bacterial communities than land-use type. FEMS Microbiol Ecol 79:12–24
Lane D (1991) 16S/23S rRNA sequencing. In: Stackebrandt E, Goodfellow M (eds) Nucleic acid techniques in bacterial systematics. Wiley, West Sussex, pp 115–175
Lima AB, Cannavan FS, Navarrete AA, Teixeira WG, Kuramae EE, Tsai SM (2014) Amazonian Dark Earth and plant species from the Amazon region contribute to shape rhizosphere bacterial communities. Microb Ecol. doi:10.1007/s00248-014-0472-8
Lupatini M, Suleiman AKA, Jacques RJS, Antoniolli ZI, Kuramae EE, Camargo FAO, Roesch LFW (2013) Soil-borne bacterial structure and diversity does not reflect community activity in Pampa biome. PLoS One 8:e76465
Mendes LW, Tsai SM, Navarrete AA, de Hollander M, van Veen JA, Kuramae EE (2015) Soil-borne microbiome: linking diversity to function. Microb Ecol. doi:10.1007/s00248-014-0559-2
Mendonça-Santos ML, dos Santos HG, Coelho MR, Bernardi ACC, Machado PLOA, Manzatto CV, Fidalgo ECC (2006) Soil and land use in the Brazilian Amazon. In: Moreira FMS, Siqueira JO, Brussaard L (eds) Soil biodiversity in Amazonian and other Brazilian ecosystems. CAB International, Wallingford, pp 13–42
Muyzer G, de Wall EC, Uitterlinden AG (1993) Profiling of complex microbial populations by denaturing gradient gel electrophoresis analysis of polymerase chain reaction-amplified genes coding for 16S rRNA. Appl Environ Microbiol 59:695–700
Navarrete AA, Cannavan FS, Taketani RG, Tsai SM (2010) A molecular survey of the diversity of microbial communities in different Amazonian agricultural model systems. Diversity 2:787–809
Navarrete AA, Kuramae EE, de Hollander M, Pijl AS, van Veen JA, Tsai SM (2013) Acidobacterial community responses to agricultural management of soybean in Amazon forest soils. FEMS Microbiol Ecol 83:607–621
Navarrete AA, Diniz TR, Braga LPP, Silva GGZ, Franchini JC, Rossetto R, Edwards RA, Tsai SM (2015a) Multi-analytical approach reveals potential microbial indicators in soil for sugarcane model systems. PLoS One 10:e0129765
Navarrete AA, Tsai SM, Mendes LW, Faust K, de Hollander M, Cassman N, van Veen JA, Kuramae EE (2015b) Soil microbiome responses to the short-term effects of Amazonian deforestation. Mol Ecol 24:2433–2448
Noll M, Matthies D, Frenzel P, Derakshani M, Liesack W (2005) Succession of bacterial community structure and diversity in a paddy soil oxygen gradient. Environ Microbiol 7:382–395
Nunes da Rocha U, Van Elsas JD, Van Overbeek LS (2011) Verrucomicrobia subdivision 1 strains display a difference in the colonization of the leek (Allium porrum) rhizosphere. FEMS Microbiol Ecol 78:297–303
Nunes da Rocha U, Plugge CM, George I, Van Elsas JD, Van Overbeek LS (2013) The rhizosphere selects for particular groups of Acidobacteria and Verrucomicrobia. PLoS One 8:e82443
Nye P, Greenland D (1960) The soil under shifting cultivation. Technical communication no. 51. Commonwealth Bureau of Soils, Harpenden, 156 pp
O’Farrell KA, Janssen PH (1999) Detection of Verrucomicrobia in a pasture soil by PCR-mediated amplification of 16S rRNA genes. Appl Environ Microbiol 65:4280–4284
Pan Y, Cassman N, de Hollander M, Mendes LW, Korevaar H, Geerts RHEM, van Veen JA, Kuramae EE (2014) Impact of long term N, P, K and NPK fertilization on the composition and potential functions of the bacterial community in grassland soil. FEMS Microbiol Ecol 90:195–205
Pitombo LM, Carmo JB, Cantarella H, Rossetto R, Hollander M, López MV, Kuramae EE (2015) Exploring soil microbial 16S rRNA sequence data to increase carbon yield and nitrogen efficiency of a bioenergy crop. Glob Change Biol Bioenergy. doi:10.1111/gcbb.12284
Pol A, Heijmans K, Harhangi HR, Tedesco D, Jetten MSM, Op den Camp HJM (2007) Methanotrophy below pH 1 by a new Verrucomicrobia species. Nature 450:874–878
Quirino BF, Pappas GJ, Tagliaferro AC, Collevattia RG, Neto EL, Silva MRSS, Bustamante MMC, Krüger RH (2009) Molecular phylogenetic diversity of bacteria associated with soil of the savanna-like Cerrado vegetation. Microbiol Res 164:59–70
Sanguin H, Remenant B, Dechesne A, Thioulouse J, Vogel TM, Nesme X, Moenne-Loccoz Y, Grundman G (2006) Potential of a 16S rRNA-based taxonomic microarray for analyzing the rhizopshere effects of maize on Agrobacterium spp. and bacterial communities. Appl Environ Microbiol 72:4302–4312
Sangwan P, Kovac S, Davis KER, Sait M, Janssen PH (2005) Detection and cultivation of soil Verrucomicrobia. Appl Environ Microbiol 71:8402–8410
Smith CJ, Osborn AM (2009) Advantages and limitations of quantitative PCR (Q-PCR)-based approaches in microbial ecology. FEMS Microbiol Ecol 67:6–20
Stevenson BS, Eichorst SA, Wertz JT, Schmidt TM, Breznak JA (2004) New strategies for cultivation and detection of previously uncultured microbes. Appl Environ Microbiol 70:4748–4755
ter Braak CJF, Šmilauer P (2002) CANOCO reference manual and CanoDraw for windows user’s guide: software for canonical community ordination (version 4.5). Microcomputer Power, Ithaca. www.canoco.com
van Raij B, Cantarella H, Quaggio JA, Furlani AMC (eds) (1996) Recomendações de adubação e calagem para o Estado de São Paulo, 2nd edn. Instituto Agronômico, IAC, Campinas, Boletim técnico 100
Wertz JT, Kim E, Breznak JA, Schmidt TM, Rodrigues JLM (2012) Genomic and physiological characterization of the Verrucomicrobia isolate Diplosphaera colitermitum gen. nov., sp. nov., reveals microaerophily and Nitrogen fixation genes. Appl Environ Microbiol 78:1544–1555
Zul D, Denzel S, Kotz A, Overmann J (2007) Effects of plant biomass, plant diversity, and water content on bacterial communities in soil lysimeters: implications for the determinants of bacterial diversity. Appl Environ Microbiol 73:6916–6929
Acknowledgments
This study was supported by a grant from Fundação de Amparo à Pesquisa do Estado de São Paulo (FAPESP—2012/13321-7 and 2011/51749-6), Coordenação de Aperfeiçoamento de Pessoal de Nível Superior (CAPES/Wageningen—2238/10-1), Conselho Nacional de Desenvolvimento Científico (CNPq—147861/2013-6, 152084/2011-8 and 485801/2011-6), and BE-Basic 008.002.005. Publication number 5876 of the NIOO-KNAW, Netherlands Institute of Ecology.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Navarrete, A.A., Soares, T., Rossetto, R. et al. Verrucomicrobial community structure and abundance as indicators for changes in chemical factors linked to soil fertility. Antonie van Leeuwenhoek 108, 741–752 (2015). https://doi.org/10.1007/s10482-015-0530-3
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-015-0530-3