Abstract
Planctomycetes is a fascinating phylum of mostly aquatic bacteria, not only due to the environmental importance in global carbon and nitrogen cycles, but also because of a unique cell biology. Their lifestyle and metabolic capabilities are not well explored, which motivated us to study the role of Planctomycetes in biofilms on marine biotic surfaces. Here, we describe the novel strain Pan54T which was isolated from algae in a hydrothermal area close to the volcanic island Panarea in the Tyrrhenian Sea, north of Sicily in Italy. The strain grew best at pH 9.0 and 26 °C and showed typical characteristics of planctomycetal bacteria, e.g. division by polar budding, formation of aggregates and presence of stalks and crateriform structures. Phylogenetically, the strain belongs to the genus Rubinisphaera. Our analysis suggests that Pan54T represents a novel species of this genus, for which we propose the name Rubinisphaera italica sp. nov. We suggest Pan54T (= DSM 29369 = LMG 29789) as the type strain of the novel species.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Together with Verrucomicrobia, Lentisphaerae, Kirimatiellaeota and Chlamydiae, Planctomycetes form the medically and biotechnologically relevant PVC superphylum (Spring et al. 2016; Wagner and Horn 2006; Devos and Ward 2014). In the past, Planctomycetes were postulated as the missing link between bacteria and eukaryotes (Devos and Reynaud 2010) beyond the bacterial cell plan (Devos et al. 2013; Fuerst and Sagulenko 2011). This finding was based on proposed exceptional planctomycetal features, such as, lack of peptidoglycan (König et al. 1984), a compartmentalised cell plan (Lindsay et al. 1997), a nucleus-like structure (Fuerst and Webb 1991) and performance of endocytosis (Lonhienne et al. 2010). Further investigation of the planctomycetal physiology and morphology based on the advent of novel techniques changed this picture (Jogler et al. 2011; Jogler and Jogler 2013; Rivas-Marin et al. 2016). In particular, presence of peptidoglycan in some Planctomycetes was confirmed (Jeske et al. 2015; van Teeseling et al. 2015) and thus the cell plan of Planctomycetes was reinterpreted to be Gram-negative (Boedeker et al. 2017; Devos 2014a, b). But still, Planctomycetes remain exceptional and enigmatic in comparison to well-characterised canonical bacteria. They e.g. divide unusually, either by budding, binary fission or even a combination of both (Wiegand et al. 2019) and lack canonical divisome proteins including the otherwise universal FtsZ (Jogler et al. 2012; Pilhofer et al. 2008).
Many Planctomycetes survive in oligotrophic environments, such as seawater, by utilising a range of high-molecular-weight sugars derived from algae (Jeske et al. 2013; Lachnit et al. 2013) after attaching to these nutrient-rich surfaces (Bengtsson et al. 2012; Bondoso et al. 2015, 2014, 2017; Lage and Bondoso 2014; Vollmers et al. 2017). In this context, a specialised morphology including the unique pili-forming crateriform structures and an extremely enlarged periplasm might be involved in the uptake and cleavage of such polymeric compounds (Boedeker et al. 2017).
It is frequently observed that Planctomycetes are highly abundant in biofilms on nutrient-rich marine surfaces (Bengtsson and Øvreås 2010; Kohn et al. 2019b). This is astonishing when considering their moderate growth rates compared to faster-growing competitors in this ecological niche (Frank et al. 2014; Wiegand et al. 2018). It is thus likely that Planctomycetes are ‘talented’ producers of secondary metabolites, which could mediate (symbiotic) interactions with algae or act as antibiotics (Jeske et al. 2013). Gene clusters involved in small molecule production were predicted in planctomycetal genomes, which substantiates Planctomycetes as a promising source of such bioactive compounds (Graca et al. 2016; Wiegand et al. 2019).
Taken together, Planctomycetes are amongst the most maverick of all bacteria known thus far (Wiegand et al. 2018). We are steadily aiming to expand the collection of known Planctomycetes and recently presented 79 novel strains in an overview article (Wiegand et al. 2019). Here, we introduce and validly describe Pan54T, a novel planctomycetal strain that was isolated close to the volcanic island Panarea, and describe its morphology, physiology and phylogeny. The sampling location Panarea is located in the Tyrrhenian Sea, has an area of 3.3 km2 and is the second smallest of the Aeolian islands. The island itself is only a small part of a sub-marine edifice in form of a truncated cone with an eastern protrusion with its base being 1500 m below sea level (Gabbianelli et al. 1990). The entire cone has a diameter of 23 km and an area of 460 km2 and several thermal springs are located in proximity to the island. In the surroundings of Panarea areas with increased temperatures, higher levels of nutrients including nitrogen and sulfur sources are present, which motivated us to choose this geographical location as a valuable source of so far unknown species of prokaryotes (Gugliandolo et al. 2015).
Material and methods
Cultivation conditions and isolation
For strain isolation and cultivation M1H NAG ASW medium was used. For medium preparation 0.25 g peptone (Bacto), 0.25 g yeast extract (Bacto), 2.38 g (4-(2-hydroxyethyl)-1-piperazineethanesulfonic acid) (HEPES) (10 mM), 250 mL artificial sea water (ASW) and 20 mL sterile-filtered Hutner’s basal salt solution were mixed in a final volume of 973 mL double distilled water. The pH was adjusted to 7.5 using 5 M KOH and the solution was autoclaved for 20 min at 121 °C. After cooling, the following solutions were added aseptically: 1 mL of 25% (w/v) glucose, 5 mL vitamin solution, 1 mL trace element solution and 20 mL of a stock solution with 50 g/L N-acetyl glucosamine (NAG). ASW contained 46.94 g/L NaCl, 7.84 g/L Na2SO4, 21.28 g/L MgCl2 × 6 H2O, 2.86 g/L CaCl2 × 2 H2O, 0.384 g/L NaHCO3, 1.384 g/L KCl, 0.192 g/L KBr, 0.052 g/L H3BO3, 0.08 g/L SrCl2 × 6 H2O and 0.006 g/L NaF and was freshly prepared before addition to the base solution. Hutner’s basal salt solution was prepared by first dissolving 10 g nitrilotriacetic acid (NTA) in 700 mL double distilled water and adjusting the pH to 7.2 using 5 M KOH. After that, the following compounds were added: 29.7 g MgSO4 × 7 H2O, 3.34 g CaCl2 × 2 H2O, 0.01267 g Na2MoO4 × 2 H2O, 0.099 g FeSO4 × 7 H2O and 50 mL metal salt solution 44. The solution was filled up to 1 L, sterilised by filtering and stored at 4 °C. Metal salt solution 44 consisted of 250 mg/L Na2-EDTA, 1095 mg/L ZnSO4 × 7 H2O, 500 mg/L FeSO4 × 7 H2O, 154 mg/L MnSO4 x H2O, 39.5 mg/L CuSO4 × 5 H2O, 20.3 mg/L CoCl2 × 6 H2O and 17.7 mg/L Na2B4O7 × 10 H2O. In the first step, EDTA was dissolved and, if required, a few drops of concentrated H2SO4 were added to retard precipitation of heavy metal ions. Metal salt solution 44 was sterilised by filtration and stored at 4 °C. Vitamin solution contained per liter: 10 mg p-aminobenzoic acid, 4 mg biotin, 20 mg pyridoxine hydrochloride, 10 mg thiamine hydrochloride, 10 mg Ca-pantothenate, 4 mg folic acid, 10 mg riboflavin, 10 mg nicotinamide and 0.2 mg vitamin B12. p-Aminobenzoic acid was dissolved first and the solution was sterilised by filtration and stored in the dark at 4 °C. The trace element solution containing 1.5 g/L Na-nitrilotriacetate, 500 mg/L MnSO4 x H2O, 100 mg/L FeSO4 × 7 H2O, 100 mg/L Co(NO3)2 × 6 H2O, 100 mg/L ZnCl2, 50 mg/L NiCl2 × 6 H2O, 50 mg/L H2SeO3, 10 mg/L CuSO4 × 5 H2O, 10 mg/L AlK(SO4)2 × 12 H2O, 10 mg/L H3BO3, 10 mg/L NaMoO4 × 2 H2O and 10 mg/L Na2WO4 × 2 H2O was sterilised by filtration and stored in the dark at 4 °C.
Strain Pan54T was isolated on the 10th of September 2013 from an algal surface in hydrothermal area 26 (sampling site 38.6392 N, 15.1051 E) close to the island Panarea in the north of Sicily, Italy. Algae leaves were initially washed with 0.5 × artificial sea water (ASW) and placed on M1H ASW solid medium (lacking NAG) containing 8 g/L gelrite additionally supplemented with 20 mg/L cycloheximide, 1000 mg/L streptomycin and 200 mg/L ampicillin. Two different pH values (6.5 and 8.0) were tested and plates were cultivated at 20 °C for 2–3 weeks. Isolated colonies were then streaked on a new plate and maintained in liquid M1H NAG ASW medium. Initial amplification and sequencing of the 16S rRNA gene was performed as previously described (Rast et al. 2017).
Deposition of genomic data
Genome data (acc. no. SJPG00000000) and the 16S rRNA gene sequence (acc. no. MK554545) were deposited in the GenBank database.
Light microscopy
Phase contrast (Phaco) analyses were performed employing a Nikon Eclipse Ti inverted microscope with a Nikon DS-Ri2 camera (blue LED). Specimens were immobilised in MatTek glass bottom dishes (35 mm, No. 1.5) employing a 1% agarose cushion (Will et al. 2018). Images were analysed using the Nikon NIS-Elements software (version 4.3). To determine the cell size, at least 100 representative cells were counted manually (Annotations and Measurements, NIS-Elements) or by using the NIS-Elements semi-automated object count tool (smooth: 4×, clean: 4×, fill holes: on, separate: 4×).
Electron microscopy
For field emission scanning electron microscopy (FESEM) bacteria were fixed in 1% (v/v) formaldehyde in HEPES buffer (3 mM HEPES, 0.3 mM CaCl2, 0.3 mM MgCl2, 2.7 mM sucrose, pH 6.9) for 1 h on ice and washed once employing the same buffer (Rast et al. 2017). Cover slips with a diameter of 12 mm were coated with a poly-l-lysine solution (Sigma-Aldrich) for 10 min, washed in distilled water and air-dried. 50 µL of the fixed bacteria solution was placed on a cover slip and allowed to settle for 10 min. Cover slips were then fixed in 1% glutaraldehyde in TE buffer (20 mM TRIS, 1 mM EDTA, pH 6.9) for 5 min at room temperature and subsequently washed twice with TE buffer before dehydrating in a graded series of acetone (10, 30, 50, 70, 90, 100%) on ice for 10 min at each concentration. Samples from the 100% acetone step were brought to room temperature before placing them in fresh 100% acetone. Samples were then subjected to critical-point drying with liquid CO2 (CPD 300, Leica). Dried samples were covered with a gold/palladium (80/20) film by sputter coating (SCD 500, Bal-Tec) before examination in a field emission scanning electron microscope (Zeiss Merlin) using the Everhart Thornley HESE2 detector and the inlens SE detector in a 25:75 ratio at an acceleration voltage of 5 kV. Transmission electron microscopy (TEM) was performed as described before (Kohn et al. 2016).
Physiological analyses
For determination of the pH optimum 100 mM HEPES was used for cultivations at pH 7.0, 7.5 and 8.0. For cultivation at pH 5.0 and 6.0 HEPES was replaced by 100 mM 2-(N-morpholino)ethanesulfonic acid (MES), whereas 100 mM N-cyclohexyl-2-aminoethanesulfonic acid (CHES) served as a buffering agent at pH 9.0 and 10.0. Cultivations for determination of the pH optimum were performed at 28 °C. For determination of the temperature optimum Pan54T was cultivated in M1H NAG ASW medium at pH 7.5 at different temperatures ranging from 10 to 40 °C. Fatty acid composition of Pan54T was analysed based on a protocol described previously (Kohn et al. 2016).
Phylogenetic analyses
The genome of strain Pan54T was published previously (Wiegand et al. 2019) and is available from GenBank under acc. no. SJPG00000000. The GenBank acc. no. of the 16S rRNA gene is MK554545. 16S rRNA gene phylogeny was computed for strain Pan54T, the type strains of all described planctomycetal species (May 2019) and all isolates recently published (Wiegand et al. 2019). The 16S rRNA gene sequences were aligned with SINA (Pruesse et al. 2012). The phylogenetic analysis was done employing a maximum likelihood (ML) approach with 1,000 bootstraps, the nucleotide substitution model GTR, gamma distribution and estimation of proportion of invariable sites (GTRGAMMAI option) (Stamatakis 2014). Three 16S rRNA genes of bacterial strains from the PVC superphylum served as outgroup. The rpoB nucleotide sequences (encoding the RNA polymerase β-subunit) were taken from publicly available genome annotations and the sequence identities were determined as described previously (Bondoso et al. 2013) with Clustal Omega (Sievers et al. 2011) alignment and matrix calculation upon extracting only those parts of the sequence that would have been sequenced with the described primer set. The average nucleotide identity (ANI) was calculated using OrthoANI (Lee et al. 2016). The average amino acid identity (AAI) was gained with the aai.rb script of the enveomics collection (Rodriguez-R and Konstantinidis 2016). The percentage of conserved proteins (POCP) was calculated as described before (Qin et al. 2014).
Results and discussion
Phylogenetic analysis
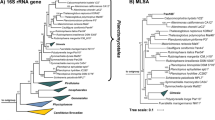
In our phylogenetic analysis Pan54T appears in a monophyletic clade with its closest relative Rubinisphaera brasiliensis DSM 5305T (Fig. 1). R. brasiliensis DSM 5305T was originally isolated from a water sample of Lagoa Vermelha, a salt pit near Rio de Janeiro, Brasil, initially described as “Planctomyces brasiliensis” in 1989 and was later reclassified (Scheuner et al. 2014; Schlesner 1989). R. brasiliensis is currently the only validly described species within the genus Rubinisphaera. Pan54T and R. brasiliensis DSM 5305T share a 16S rRNA gene sequence identity of 96.2%. This value is below the threshold of 98.7% for a novel species (Stackebrandt and Ebers 2006), but above the threshold for a novel planctomycetal genus of 94.5% (Yarza et al. 2014). Both strains share 76.5% rpoB gene identity, which is below the 96.3% cutoff proposed to distinguish between different planctomycetal species and above the threshold for novel genera of 72% sequence identity (Bondoso et al. 2013). Average nucleotide identity (ANI) between both strains is 69.7% and distinctly below the species threshold of 95 - 96% for ANIb (ANI calculated with the BLAST algorithm) (Kim et al. 2014). The average amino acid identity (AAI) between Pan54T and its relative R. brasiliensis DSM 5305T is 58.1% and thus fits in the range for species belonging to the same bacterial genus (Rodriguez et al. 2018). Finally, the percentage of conserved proteins (POCP) of 62.3% between both strains also indicates that they belong to the same genus, as a prokaryotic genus is proposed to be formed by a group of species with pairwise POCP values higher than 50% (Qin et al. 2014). Close relatives apart from R. brasiliensis DSM 5305T include Planctomicrobium piriforme P3T (Kulichevskaya et al. 2015), Gimesia maris DSM 8797T (Scheuner et al. 2014), Fuerstiella marisgermanici NH11T (formerly designated “Fuerstia marisgermanicae”) (Kohn et al. 2019a, 2016) and species of the Planctopirus genus (Fig. 1).
Phylogenetic analysis. The phylogenetic tree highlighting the position of Pan54T is depicted. 16S rRNA gene phylogeny was computed using the maximum likelihood method. Bootstrap values after 1000 resamplings (in %) are given at the nodes. The outgroup consists of three 16S rRNA genes from the PVC superphylum. The “Pirellula clade” includes species of the genera Rhodopirellula, Rubripirellula, Roseimaritima, Mariniblastus, Pirellula and Blastopirellula. The “Thermophilic clade” comprises the genera Thermostilla, Thermogutta and Thermopirellula
Morphological, physiological and biochemical analyses
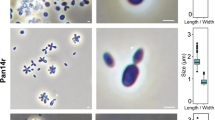
For a morphological characterization, Pan54T cells were harvested during the exponential growth phase. Pan54T cells were found to be pear-shaped (1.6 ± 0.2 µm × 0.8 ± 0.1 µm) (Fig. 2a–c) and form strong aggregates and biofilms. Cells have a textured surface and contain evenly distributed crateriform structures (Fig. 2d, e). Pan54T divides by polar budding (Fig. 2a). Daughter cells have the same shape as mother cells. Thin sections of Pan54T cells show a condensed nucleoid and cytoplasmic invaginations (Fig. 2f, g). Colonies are white indicating a lack of carotenoids as pigmenting compounds. Detailed information on morphology, locomotion and cell division is summarised in Table 1. Pan54T has a very similar morphology as R. brasiliensis. In the direct comparison Pan54T has a slightly elongated shape and crateriform structures appear to be more pronounced. Similar to R. brasiliensis rosette formation was observed rather than formation of larger aggregates (Scheuner et al. 2014). The colony colour of non-pigmented Pan54T differs from the yellow to orange pigmentation of R. brasiliensis.
Microscopy images and cell size plot of Pan54T. Pictures from light microscopy (LM) (a, b), scanning electron microscopy (SEM) (d, e) and transmission electron microscopy (TEM) (f, g) are shown. The scale bars are 1 µm. For determination of the cell size (c) at least 100 representative cells were counted manually or by using a semi-automated object count tool during scanning electron microscopy
In physiological experiments, Pan54T grew at a temperature range of 14–27 °C (Fig. 3a) and a pH range of 6.0–10.0 (Fig. 3b), but failed to grow at 30 °C or above (Fig. 3a). The optimal conditions turned out to be pH 9.0 and 26 °C. The observed temperature optimum of Pan54T is lower compared to R. brasiliensis (30–33 °C) (Schlesner 1989). R. brasiliensis even grows at temperatures of 37 °C (Schlesner 1989), while Pan54T failed to grow at 30 °C or higher. Taken together, growth of Pan54T is mesophilic and slightly alkaliphilic. A maximal growth rate of 0.039 h−1 was observed under the given conditions corresponding to a generation time of 18 h (Fig. 3). This value is in the range of 0.01–0.09 h−1 (generation times of 8–70 h), which we typically observed for planctomycetal strains isolated and characterised in our lab so far.
Major fatty acids of Pan54T include summed feature 3 (palmitoleic acid (16:1 ω7c), 15:0 iso 2-OH) (57%), palmitic acid (16:0) (36%) and cis-vaccenic acid (18:1 ω7c) (5%) (Table 2). The composition is similar to R. brasiliensis in which 16:0, 16:1 and 18:1 were also found as the major fatty acids (Scheuner et al. 2014). However, the composition appears to be more restricted to these three in Pan54T as other fatty acid were only detected in traces (< 0.5%) in this strain.
Genomic characteristics
The genome of Pan54T has a size of 6.7 Mb and is slightly larger compared to R. brasiliensis (6.0 Mb), whereas the GC content is lower (48.8% for Pan54T, 56.4% for R. brasiliensis). Relevant genome characteristics are summarised in Table 1. Automated gene prediction and annotation identified 5275 putative protein-encoding genes, of which 42% (2229 genes) are annotated as hypothetical proteins. The calculated values correspond to 787 protein-coding genes per Mb and a coding density of 85%. Except for differences in GC content no striking differences between genome features of Pan54T and R. brasiliensis DSM 5305T were observed (Table 1). The number of 16S rRNA in both strains is identical and number of tRNAs is very similar.
Conclusion
Based on the data collected during strain characterisation, Pan54T represents a novel species within the genus Rubinisphaera. We propose the name Rubinisphaera italica sp. nov. and present Pan54T as the type strain of the species.
Emended description of the genus Rubinisphaera Scheuner et al. (2014)
The description of the genus Rubinisphaera given previously (Scheuner et al. 2014), with the following modification: The GC content is between 48 and 57%.
Description of Rubinisphaera italica sp. nov.
Rubinisphaera italica (i.ta'li.ca. L. fem. adj. italica of Italy; corresponding to the isolation of the strain from Italy). Cells are pear-shaped (length: 1.6 ± 0.2 µm, width: 0.8 ± 0.1 µm), form aggregates and divide by polar budding. Cells grow at ranges of 14–27 °C (optimum 26 °C) and pH 6.0–10.0 (optimum 9.0). Colonies are white. The genome (acc. no. SJPG00000000) and 16S rDNA sequence (acc. no. MK554545) are available from the GenBank database. The genome has a GC content of 48.8% and is 6.70 Mb in length. The proposed type strain is Pan54T (DSM 29369 = LMG 29789) isolated from an algal surface at a hydrothermal area close to Panarea, Italy.
References
Bengtsson MM, Øvreås L (2010) Planctomycetes dominate biofilms on surfaces of the kelp Laminaria hyperborea. BMC Microbiol 10:261
Bengtsson MM, Sjøtun K, Lanzén A, Øvreås L (2012) Bacterial diversity in relation to secondary production and succession on surfaces of the kelp Laminaria hyperborea. ISME J 6:2188–2198
Boedeker C, Schuler M, Reintjes G, Jeske O, van Teeseling MC, Jogler M, Rast P, Borchert D, Devos DP, Kucklick M, Schaffer M, Kolter R, van Niftrik L, Engelmann S, Amann R, Rohde M, Engelhardt H, Jogler C (2017) Determining the bacterial cell biology of Planctomycetes. Nat Commun 8:14853
Bondoso J, Albuquerque L, Nobre MF, Lobo-da-Cunha A, da Costa MS, Lage OM (2015) Roseimaritima ulvae gen. nov., sp. nov. and Rubripirellula obstinata gen. nov., sp. nov. two novel planctomycetes isolated from the epiphytic community of macroalgae. Syst Appl Microbiol 38:8–15
Bondoso J, Balague V, Gasol JM, Lage OM (2014) Community composition of the Planctomycetes associated with different macroalgae. FEMS Microbiol Ecol 88:445–456
Bondoso J, Godoy-Vitorino F, Balague V, Gasol JM, Harder J, Lage OM (2017) Epiphytic Planctomycetes communities associated with three main groups of macroalgae. FEMS Microbiol Ecol 93(3):fiw255
Bondoso J, Harder J, Lage OM (2013) rpoB gene as a novel molecular marker to infer phylogeny in Planctomycetales. Antonie Van Leeuwenhoek 104:477–488
Devos DP (2014a) PVC bacteria: variation of, but not exception to, the Gram-negative cell plan. Trends Microbiol 22:14–20
Devos DP (2014b) Re-interpretation of the evidence for the PVC cell plan supports a Gram-negative origin. Antonie Van Leeuwenhoek 105:271–274
Devos DP, Jogler C, Fuerst JA (2013) The 1st EMBO workshop on PVC bacteria-Planctomycetes-Verrucomicrobia-Chlamydiae superphylum: exceptions to the bacterial definition? Antonie Van Leeuwenhoek 104:443–449
Devos DP, Reynaud EG (2010) Evolution. Intermediate steps. Science 330:1187–1188
Devos DP, Ward NL (2014) Mind the PVCs. Environ Microbiol 16:1217–1221
Frank O, Michael V, Pauker O, Boedeker C, Jogler C, Rohde M, Petersen J (2014) Plasmid curing and the loss of grip—the 65-kb replicon of Phaeobacter inhibens DSM 17395 is required for biofilm formation, motility and the colonization of marine algae. Syst Appl Microbiol 38:120–127
Fuerst JA, Sagulenko E (2011) Beyond the bacterium: planctomycetes challenge our concepts of microbial structure and function. Nat Rev Microbiol 9:403–413
Fuerst JA, Webb RI (1991) Membrane-bounded nucleoid in the eubacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 88:8184–8188
Gabbianelli G, Gillot P, Lanzafame G, Romagnoli C, Rossi P (1990) Tectonic and volcanic evolution of Panarea (Aeolian islands, Italy). Mar Geol 92:313–326
Graca AP, Calisto R, Lage OM (2016) Planctomycetes as novel source of bioactive molecules. Front Microbiol 7:1241
Gugliandolo C, Lentini V, Bunk B, Overmann J, Italiano F, Maugeri TL (2015) Changes in prokaryotic community composition accompanying a pronounced temperature shift of a shallow marine thermal brine pool (Panarea Island, Italy). Extremophiles 19:547–559
Jeske O, Jogler M, Petersen J, Sikorski J, Jogler C (2013) From genome mining to phenotypic microarrays: Planctomycetes as source for novel bioactive molecules. Antonie Van Leeuwenhoek 104:551–567
Jeske O, Schüler M, Schumann P, Schneider A, Boedeker C, Jogler M, Bollschweiler D, Rohde M, Mayer C, Engelhardt H, Spring S, Jogler C (2015) Planctomycetes do possess a peptidoglycan cell wall. Nat Commun 6:7116
Jogler C, Glöckner FO, Kolter R (2011) Characterization of Planctomyces limnophilus and development of genetic tools for its manipulation establish it as a model species for the phylum Planctomycetes. Appl Environ Microbiol 77:5826–5829
Jogler C, Waldmann J, Huang X, Jogler M, Glöckner FO, Mascher T, Kolter R (2012) Identification of proteins likely to be involved in morphogenesis, cell division, and signal transduction in Planctomycetes by comparative genomics. J Bacteriol 194:6419–6430
Jogler M, Jogler C (2013) Towards the development of genetic tools for Planctomycetes. In: Fuerst JA (ed) Planctomycetes: cell structure, origins and biology. Springer, Berlin, pp 141–164
Kim M, Oh H-S, Park S-C, Chun J (2014) Towards a taxonomic coherence between average nucleotide identity and 16S rRNA gene sequence similarity for species demarcation of prokaryotes. Int J Syst Evol Microbiol 64:346–351
Kohn T, Heuer A, Jogler M, Vollmers J, Boedeker C, Bunk B, Rast P, Borchert D, Glöckner I, Freese HM (2019a) Corrigendum: Fuerstia marisgermanicae gen. nov., sp. nov., an unusual member of the phylum Planctomycetes from the German Wadden Sea. Front Microbiol 10:1029
Kohn T, Wiegand S, Boedeker C, Rast P, Heuer A, Jetten MSM, Schüler M, Becker S, Rohde C, Müller R-W, Brümmer F, Rohde M, Engelhardt H, Jogler M, Jogler C (2019b) Planctopirus ephydatiae, a novel Planctomycete isolated from a freshwater sponge. Syst Appl Microbiol. https://doi.org/10.1016/j.syapm.2019.126022
Kohn T, Heuer A, Jogler M, Vollmers J, Boedeker C, Bunk B, Rast P, Borchert D, Glöckner I, Freese HM, Klenk HP, Overmann J, Kaster AK, Wiegand S, Rohde M, Jogler C (2016) Fuerstia marisgermanicae gen. nov., sp. nov., an unusual member of the phylum Planctomycetes from the German Wadden Sea. Front Microbiol 7:2079
König E, Schlesner H, Hirsch P (1984) Cell wall studies on budding bacteria of the Planctomyces/Pasteuria group and on a Prosthecomicrobium sp. Arch Microbiol 138:200–205
Kulichevskaya IS, Ivanova AA, Detkova EN, Rijpstra WIC, Damste JSS, Dedysh SN (2015) Planctomicrobium piriforme gen. nov., sp. nov., a stalked planctomycete from a littoral wetland of a boreal lake. Int J Syst Evol Microbiol 65:1659–1665
Lachnit T, Fischer M, Kunzel S, Baines JF, Harder T (2013) Compounds associated with algal surfaces mediate epiphytic colonization of the marine macroalga Fucus vesiculosus. FEMS Microbiol Ecol 84:411–420
Lage OM, Bondoso J (2014) Planctomycetes and macroalgae, a striking association. Front Microbiol 5:267
Lee I, Ouk Kim Y, Park SC, Chun J (2016) OrthoANI: An improved algorithm and software for calculating average nucleotide identity. Int J Syst Evol Microbiol 66:1100–1103
Lindsay MR, Webb RI, Fuerst JA (1997) Pirellulosomes: a new type of membrane-bounded cell compartment in planctomycete bacteria of the genus Pirellula. Microbiol 143:739–748
Lonhienne TG, Sagulenko E, Webb RI, Lee KC, Franke J, Devos DP, Nouwens A, Carroll BJ, Fuerst JA (2010) Endocytosis-like protein uptake in the bacterium Gemmata obscuriglobus. Proc Natl Acad Sci USA 107:12883–12888
Pilhofer M, Rappl K, Eckl C, Bauer AP, Ludwig W, Schleifer KH, Petroni G (2008) Characterization and evolution of cell division and cell wall synthesis genes in the bacterial phyla Verrucomicrobia, Lentisphaerae, Chlamydiae, and Planctomycetes and phylogenetic comparison with rRNA genes. J Bacteriol 190:3192–3202
Pruesse E, Peplies J, Glöckner FO (2012) SINA: accurate high-throughput multiple sequence alignment of ribosomal RNA genes. Bioinformatics 28:1823–1829
Qin Q-L, Xie B-B, Zhang X-Y, Chen X-L, Zhou B-C, Zhou J, Oren A, Zhang Y-Z (2014) A proposed genus boundary for the prokaryotes based on genomic insights. J Bacteriol 196:2210–2215
Rast P, Glöckner I, Boedeker C, Jeske O, Wiegand S, Reinhardt R, Schumann P, Rohde M, Spring S, Glöckner FO (2017) Three novel species with peptidoglycan cell walls form the new genus Lacunisphaera gen. nov. in the family Opitutaceae of the verrucomicrobial subdivision 4. Front Microbiol 8:202
Rivas-Marin E, Canosa I, Santero E, Devos DP (2016) Development of genetic tools for the manipulation of the Planctomycetes. Front Microbiol 7:914
Rodriguez-R LM, Konstantinidis KT (2016) The enveomics collection: a toolbox for specialized analyses of microbial genomes and metagenomes. PeerJ Preprints 4:e1900v1
Rodriguez RL, Gunturu S, Harvey WT, Rossello-Mora R, Tiedje JM, Cole JR, Konstantinidis KT (2018) The Microbial Genomes Atlas (MiGA) webserver: taxonomic and gene diversity analysis of Archaea and Bacteria at the whole genome level. Nucleic Acids Res 46:W282–W288
Scheuner C, Tindall BJ, Lu M, Nolan M, Lapidus A, Cheng, J-F, Goodwin L, Pitluck S, Huntemann M, Liolios K (2014) Complete genome sequence of Planctomyces brasiliensis type strain (DSM 5305T), phylogenomic analysis and reclassification of Planctomycetes including the descriptions of Gimesia gen. nov., Planctopirus gen. nov. and Rubinisphaera gen. nov. and emended descriptions of the order Planctomycetales and the family Planctomycetaceae. Stand Gen Sci 9:10
Schlesner H (1989) Planctomyces brasiliensis sp. nov., a halotolerant bacterium from a salt pit. Syst Appl Microbiol 12:159–161
Sievers F, Wilm A, Dineen D, Gibson TJ, Karplus K, Li W, Lopez R, McWilliam H, Remmert M, Söding J (2011) Fast, scalable generation of high-quality protein multiple sequence alignments using Clustal Omega. Mol Syst Biol 7:539
Spring S, Bunk B, Sproer C, Schumann P, Rohde M, Tindall BJ, Klenk HP (2016) Characterization of the first cultured representative of Verrucomicrobia subdivision 5 indicates the proposal of a novel phylum. ISME J 10:2801–2816
Stackebrandt E, Ebers J (2006) Taxonomic parameter revisited: tarnished gold standards. Microbiol Today 33:152–155
Stamatakis A (2014) RAxML version 8: a tool for phylogenetic analysis and post-analysis of large phylogenies. Bioinformatics 30:1312–1313
van Teeseling MC, Mesman RJ, Kuru E, Espaillat A, Cava F, Brun YV, VanNieuwenhze MS, Kartal B, van Niftrik L (2015) Anammox Planctomycetes have a peptidoglycan cell wall. Nat Commun 6:6878
Vollmers J, Frentrup M, Rast P, Jogler C, Kaster AK (2017) Untangling genomes of novel planctomycetal and verrucomicrobial species from Monterey Bay kelp forest metagenomes by refined binning. Front Microbiol 8:472
Wagner M, Horn M (2006) The Planctomycetes, Verrucomicrobia, Chlamydiae and sister phyla comprise a superphylum with biotechnological and medical relevance. Curr Opin Biotechnol 17:241–249
Wiegand S, Jogler M, Jogler C (2018) On the maverick Planctomycetes. FEMS Microbiol Rev 42:739–760
Wiegand S, Jogler M, Boedeker C, Pinto D, Vollmers J, Rivas-Marín E, Kohn T, Peeters SH, Heuer A, Rast P, Oberbeckmann S, Bunk B, Jeske O, Meyerdierks A, Storesund JE, Kallscheuer N, Lücker S, Lage OM, Pohl T, Merkel BJ, Hornburger P, Müller R-W, Brümmer F, Labrenz M, Spormann AM, Op den Camp HJM, Overmann J, Amann R, Jetten MSM, Mascher T, Medema MH, Devos DP, Kaster A-K, Øvreås L, Rohde M, Galperin MY, Jogler C (2019) Cultivation and functional characterization of 79 planctomycetes uncovers their unique biology. Nat Microbiol. https://doi.org/10.1038/s41564-019-0588-1
Will SE, Henke P, Boedeker C, Huang S, Brinkmann H, Rohde M, Jarek M, Friedl T, Seufert S, Schumacher M (2018) Day and night: metabolic profiles and evolutionary relationships of six axenic non-marine cyanobacteria. Gen Biol Evol 11:270–294
Yarza P, Yilmaz P, Pruesse E, Glöckner FO, Ludwig W, Schleifer KH, Whitman WB, Euzeby J, Amann R, Rossello-Mora R (2014) Uniting the classification of cultured and uncultured bacteria and archaea using 16S rRNA gene sequences. Nat Rev Microbiol 12:635–645
Acknowledgements
Part of this work was funded by the Deutsche Forschungsgemeinschaft (DFG) Grant No. KA 4967/1–1 and JO 893/4–1, Grant ALWOP.308 of the Nederlandse Organisatie voor Wetenschappelijk Onderzoek (NWO) grant and SIAM (Soehngen Institute for Anaerobic Microbiology) Grant No. 024002002. We thank the Scientific Diving Center of Bergakademie Freiberg (Germany) as well as Thomas Pohl, Peter Hornburger and all participants of the 2013 Panarea Expedition for support during sampling. We thank Ina Schleicher for skillful technical assistance. Lastly, we would like to thank Brian Tindall and Regine Fähnrich as well as the BCCM/LMG Bacteria collection for on-going support during strain deposition.
Author information
Authors and Affiliations
Contributions
NK wrote the manuscript, analyzed the data and prepared the figures, SW and MJ performed the genomic and phylogenetic analysis, AH isolated the strain and performed the initial strain cultivation and deposition, SHP and CB performed the light microscopic analysis, MSMJ contributed to text preparation and revised the manuscript, MR performed the electron microscopic analysis, CJ and MJ took the samples in Panarea, Italy. CJ supervised the study. All authors read and approved the final version of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Ethical statement
This article does not contain any studies with animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Kallscheuer, N., Jogler, M., Wiegand, S. et al. Rubinisphaera italica sp. nov. isolated from a hydrothermal area in the Tyrrhenian Sea close to the volcanic island Panarea. Antonie van Leeuwenhoek 113, 1727–1736 (2020). https://doi.org/10.1007/s10482-019-01329-w
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10482-019-01329-w