Abstract
Invasions of non-native species are of great concern as they have a devastating impact on native biodiversity and can also affect the economy of a region. Multiple introductions in several locations of a new range greatly promote the success of non-native species. The raccoon (Procyon lotor) is an omnivore whose native distribution extends from southern Canada to Panama. It has been successfully introduced in many European countries. We examined the microsatellite and mitochondrial diversity of raccoon populations in Central Europe (Germany, Poland, and Czech Republic) in order to determine their introduction sources and pathways as well as the factors affecting genetic structure in this invasive species. We found low diversity of the mtDNA control region and moderate diversity of microsatellite markers. Raccoon showed three hierarchic levels of genetic structure which separate at different levels sampled from Czech Republic, Germany and raccoon inhabiting two different habitats in Poland. In Poland the raccoon population was established through migration from Germany to Czech Republic. Analysis of the intensity of migration between two different habitat types indicated source-sink dynamics in the Polish populations of raccoons. Our results confirm the high intensity of the raccoon invasion in Central Europe and point to specific measures needed as part of an effective management strategy.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
There has been a continuous increase in invasions by non-native species due to accidental transport by humans, deliberate introduction, or the international trade in plants and animals (DAISIE 2009). It is of great concern, as invasive species often have a devastating impact on native biodiversity and may have great economic costs. Despite the global scale of this process, involving a large number of plant and animal species over a huge geographical range, the processes underlying the success of invasive species are insufficiently understood.
The two critical stages of invasion by non-native species are establishment (introduction) and spread in the new region. Introduction, usually of a small number of individuals, should cause a bottleneck, low genetic diversity and therefore reduced adaptability to the new environment. Indeed, some populations of introduced species go extinct or persist only in the small area of introduction (Lockwood et al. 2007). However, many non-native species have been introduced in large numbers (high propagule pressure) through several separate events, and in those cases their genetic diversity can be even higher than in a single population of the native range (Dlugosch and Parker 2008; Zalewski et al. 2010). High propagule pressure may increase the likelihood that the introduced species will establish and spread. Introduced individuals may originate from a few genetically divergent sources in the native range or from distant lineages; such admixture of individuals of distinct origin may increase their genetic and adaptive variation and increase their vigour (Facon et al. 2008; Kolbe et al. 2007; Lucek et al. 2010). Clearly the features of a particular introduction, and especially multiple introductions in various locations of the new range, greatly affect the success of non-native species.
The spread of non-native species in a new range consists of major dispersal events and often is linked to high gene flow between areas. This pattern leads to the assumption that populations of an invasive species in a new range should share a fairly uniform genetic structure. During expansion, however, genetic variation should become partitioned among populations via random genetic drift and/or adaptation to local conditions (natural selection). Differentiation via drift takes time, but local adaption to environmental conditions should be very rapid, especially in species inhabiting highly diverse habitats (generalists). If, during an introduction, animals of different origins are released in different locations and during expansion undergo genetic admixture upon secondary contact in the new range, this may additionally facilitate the emergence of adaptive evolutionarily novel traits and thereby accelerate expansion (Seehausen 2004; Keller and Taylor 2010). To mitigate the introduction and spread of non-native species and to develop effective management strategies, we need information about introduction pathways, multiple introduction events and sources of origin, as well as the factors affecting the genetic structure of invasive species (Simberloff 2003, Abdelkrim et al. 2005).
The raccoon Procyon lotor is a carnivore whose native distribution extends from southern Canada to Panama (Zeveloff 2002). Twenty-five subspecies have been recognised in North America (Zeveloff 2002); this is driven by the high genetic variation of this species across its native range. In the eastern and central United States, 76 mtDNA were found among 311 samples (Cullingham et al. 2008a, b). Haplotypes grouped in three main lineages were found, although the main haplotype groups corresponded only roughly to subspecies designations (Cullingham et al. 2008a, b). In its native range the raccoon exhibits high plasticity in habitat selection and in feeding habits. It occupies a great variety of habitat types, including wetlands, marshes, forests, farmland and urban areas, and feeds on a large variety of food including plant material, vertebrates, invertebrates and rubbish (Zeveloff 2002). As an omnivorous predator, the raccoon can have a severely damaging impact on birds (especially waterfowl) and reptiles (Ratnaswamy et al. 1997; Hartman et al. 1997; Hartman and Eastman 1999; Zeveloff 2002; Elis et al. 2007). The raccoon is considered one of the most chronic nuisance animals in North America; it can damage fields and garden plants, and can cause damage or other problems around houses (Conovery 1987; Beasley and Rhodes 2008). It carries rabies, canine distemper and many parasites, including Baylisascaris procyonis; which presents a threat to public health (Zeveloff 2002; Cote et al. 2012). Management and population control of this species are now of great concern in North America (Rosatte et al. 2007; Beasley and Rhodes 2008; Rosatte et al. 2010).
Raccoons were successfully introduced in Europe, both deliberately and accidentally (escape from fur farms or breeding centres) in the 1930s (1934 in Germany, 1936 in Belarus; Lutz 1984; Chesnokov 1989). In Germany, the natural spread of raccoons was limited in the first stage of invasion, and 20 years after introduction, the population was still confined to a relatively small area (Lutz 1984). Then its expansion accelerated. In the following years raccoons colonised new areas, and recently as many as a million raccoons were estimated to be living in Germany, with the number steadily climbing (Hohmann 2001). Since 1954 the raccoon has been a game species in Germany, with an annual bag of more than 50,000 in 2010 (Vos et al. 2012). Hunting has not prevented its expansion to other countries, however. Fifty years after the first effective introduction, its distribution range extends beyond Germany into the Netherlands, France, Switzerland, Austria and Poland (Lutz 1984; Bartoszewicz et al. 2008). Besides the main route of entry from Germany, in many European countries the raccoon was and still is commonly bred in captivity, mainly by private owners, for the pet trade and wildlife collections. Such pet raccoons have escaped or been released into the wild, and have been encountered sporadically in isolated locations or have established small local populations, as in France (1966) and Spain (2001) (Leger 1999; Garcia et al. 2012). Over recent years the raccoon has also been encountered in Belgium, Luxembourg, Hungary, the Czech Republic and Serbia (Mitchell-Jones et al. 1999; Červeny et al. 2001; Schley et al. 2001; Milenkovic 2003), but data on the origins of these populations are lacking: did the raccoons of a given population originate through expansion from Germany or from a separate introduction?
Raccoon introduction presents a potential threat to the many protected native species raccoons may prey upon (Bartoszewicz et al. 2008; Garcia et al. 2012), as well as an important health threat to animals and humans from the pathogens raccoons may host (Park et al. 2000; Hohmann et al. 2002, Popiołek et al. 2011). To reduce these risks and curb the raccoon invasion we need to know the current pathways of introduction, potential secondary contact zones, and the genetic structure of populations. Development of an effective management strategy requires extensive knowledge of the genetic structure and variability of the species. In this paper we analysed the microsatellite and mitochondrial diversity of the raccoon, an invasive species in Central Europe, to determine its introduction sources and pathways as well as the factors affecting its genetic structure.
Materials and methods
Study area and sampling methods
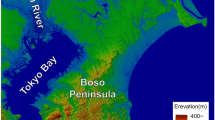
We collected 174 samples from raccoons between 2007 and 2012 (Table 1). Hair and tissue samples were taken from road-kill or from raccoons we live-trapped in “Warta Mouth” National Park and adjacent area. Additional raccoon muscle tissue samples were obtained from hunters culling raccoons as part of their game management activity in Poland, Germany and the Czech Republic (Fig. 1). All tissue samples were stored at −20 °C prior to DNA extraction.
Raccoon tissue samples were collected from Western Poland, Eastern Germany, Oberlausitzer Heide Biosphere Reserve and Saxony region and the Czech Republic, west of the city of Přerov. Three additional samples came from other sites: near Berlin, Germany (no. 1), northwest of Pol1 (no. 2), and Małopolska Province in southern Poland (no. 3) (Fig. 1). In Poland samples were collected in two different habitat types. “Warta Mouth” National Park (Pol1) is located near the German border (52°34′N, 14°43′E). The national park, a protected area since 1984, covers 80.7 km2 of the lower Warta River valley and 20 km of the river itself. It is dominated by wetlands, scrub willow, marshes, meadows and pastures. Only 1 % of the area is covered by forest. Remaining samples from Poland (Pol2) were collected about 30 km south of national park. That area is dry woodland dominated by beech (Fagus silvatica), oak (Quercus spp.) and pine (Pinus sylvestris), with only a few small rivers and water bodies. Study site Ger1 is northwest of the city of Dresden. The area is densely populated, with a number of human settlements and a dense road system. Temperate broadleaf forest or woodland cover is interspersed with arable fields, many water bodies and rivers. Study site Ger2 is in the Oberlausitzer Heide Biosphere Reserve (51°19′N, 14°32′E), characterised by high habitat diversity, with heathland, temperate broadleaf forest and woodlands, and many fishponds, rivers and streams. The area is moderately populated and intensively used for agriculture and forestry. Study site CzR is in the eastern Czech Republic west of the city of Přerov, with numerous fish ponds, canals and small water bodies.
DNA isolation, microsatellite genotyping and mitochondrial DNA amplification and sequencing
DNA was extracted from dried ear fragments or ethanol-preserved tissue using the NucleoSpin Tissue Kit (Macherey and Nagel, Dueren, Germany) according to the manufacturer’s protocol. Individual samples were genotyped at 19 microsatellite loci (Cullingham et al. 2006; Fike et al. 2007) in 6 multiplex reactions. All reactions were performed with Qiagen Mastermix (Qiagen Ltd., Crawley, UK) with one primer of each pair fluorescently dyed. Amplification products were resolved on an ABI PRISM 3130xl Genetic Analyzer (Applied Biosystems, Foster City, USA) and sized with internal lane standard LIZ 500 using Genemapper v. 4.0 (Applied Biosystems, Foster City CA, USA).
A subset of 60 samples equally representing all sampled regions was selected for mitochondrial DNA fragment amplification using primers PLO-CRL1 and L15997 (Cullingham et al. 2008a, b), which amplify the most variable fragment of the raccoon mitochondrial control region. The obtained 600 bp fragment gave 467 bp of clear sequence. All samples were sequenced in both directions on an ABI PRISM 3130xl. Sequences were checked manually and aligned with the ClustalX algorithm in Bioedit 7.0.5.3 (Hall 1999). All the newly obtained haplotypes have been deposited in GenBank.
Genetic analysis
Mitochondrial DNA
DnaSP 5.0 (Librado and Rozas 2009) was used to determine the number of haplotypes (h) and variable sites (S), and to calculate haplotype (Hd) and nucleotide diversity (π). To infer the origin of haplotypes detected in the studied populations, we made a phylogenetic analysis using raccoon haplotypes detected in the original range of the species (Cullingham et al. 2008a, b). We used Arlequin 3.01 (Excoffier and Laval 2005) to construct minimum spanning network presenting haplotype relationships, the network was subsequently visualized in HapStar 0.5 (Teacher and Griffiths 2011).
Microsatellite diversity indices
Tests of linkage disequilibrium between all pairs of loci were implemented in GENEPOP v. 3.4 (Raymond and Rousset 1995) with 10,000 permutations. The associated probability values were corrected for multiple comparisons by applying the Bonferroni adjustment for p = 0.05 significance level. The same software was used to test for deviation from Hardy–Weinberg equilibrium of microsatellite genotype frequencies. We used FSTAT 2.9.3 (Goudet 1995) to estimate genetic diversity parameters: number of alleles (NA), observed and expected heterozygosity (Ho, HE) and inbreeding coefficient (FIS). To assure that any departures from Hardy–Weinberg equilibrium were not an effect of null alleles and that the heterozygosity deficits were evenly distributed across loci we performed the jackknifing procedure in FSTAT 2.9.3, removing one locus at a time and checking the contribution of the remaining loci to the FIS value.
Population structure
To infer the raccoon population structure of the whole sampled region we used a Bayesian model-based clustering method implemented in STRUCTURE 2.3.3 (Pritchard et al. 2000). We used an admixture model and correlated allele frequencies with LOCPRIOR model as a population structure of recently introduced, invasive species is expected to be weak. This model uses prior information on sampling locations, but it do not tend to find structure when it is not present in the data set (Falush et al. 2003, Hubisz et al. 2009). Ten independent runs were performed for each K value from K = 1 to 10. Each run had 100,000 iterations with a burn-in of 100,000 iterations. To infer the number of genetic clusters in our data set, we used the ΔK method of Evanno et al. (2005) which calculates the rate of change in the log-probability of the data between successive K values (ΔK) to determine the most likely number of clusters with STRUCTURE HARVESTER (Earl and vonHoldt 2012). To obtain an average individual’s genotype proportions across runs we used CLUMPP (Jakobsson and Rosenberg 2007). We assigned each individual to the group for which its inferred ancestry was higher than 0.5 (more than a half of genome assigned to the same group). We then applied the procedure of Coulon et al. (2008) to find all levels of hierarchical structure that exist among raccoons in sampled area. As ΔK method detects the uppermost level of structure (Evanno et al. 2005), we repeated the analysis on each K detected on the previous run. The process was repeated until the most probable number of K was 1. As ΔK method cannot find the true K = 1 we also tested whether lnP(D) was maximum for K = 1. All subsequent population genetic analyses were performed on STRUCTURE-defined genetic groups. To compare the level of differentiation among genetic groups we computed pairwise FST using FSTAT 2.9.3. Analyses of molecular variance (AMOVA) were performed using Arlequin 3.01 to test the genetic relationships among populations grouped by country of origin, to check for multiple introductions within different countries.
To assess the effect of genetic isolation-by-distance (IBD) we tested the correlations between genetic distance and geographic distance (calculated as straight-line-distance, Euclidean distance) between all pairs of individual raccoons. Genetic distances between pairs of individuals were calculated by a method implemented in GenAlEx v. 6.4 (Peakall and Smouse 2006), and the Mantel test was performed with significance based on 10,000 matrix permutations.
Within-population microsatellite diversity and demographic history
Genetic diversity was measured for all STRUCTURE-defined genetic groups. Mean number of alleles (NA), allelic richness (AR), observed and expected heterozygosity (HO and HE) and inbreeding coefficient (FIS) were estimated using FSTAT 2.9.3. We used LDNE (Waples and Do 2006) to estimate the effective population sizes of previously identified clusters. The model assumes closed populations and eliminates and controls for the possible bias of small sample size. We set 0.02 as the lowest allele frequency and 95 % as the confidence interval.
We used BOTTLENECK 1.2.02 to detect any recent bottleneck and expansion in each population (Cornuet and Luikart 1997). An excess of observed gene diversity relative to expected gene diversity for the number of alleles detected in a sample may indicate population decline. Conversely, a deficit of observed gene diversity may indicate that the population is growing. Two mutation models considered appropriate for microsatellites were applied: the strict stepwise mutation model (SMM) and a two-phase model (TPM). For the TPM we used a model that includes 90 % single-step mutations and 10 % multiple-step mutations. Significant deviations of observed heterozygosity over all loci were tested using the nonparametric Wilcoxon test.
Estimating rates of migration between Polish raccoon populations
To evaluate the direction and extent of recent migrations between genetic clusters identified by STRUCTURE among the raccoons sampled in Poland, we applied the approach described by Holderegger and Gugerli (2012). We could apply this procedure only for raccoons sampled in Poland where our sampling was extensive and we had exact geographic coordinates for each sample. First we used TESS 2.3.1 (Chen et al. 2007) to define the population genetic structure of those individuals across the landscape prior to migration. Similarly to STRUCTURE, Tess applies an MCMC algorithm but also incorporates the geographic coordinates of samples as prior information. We ran Tess with the no-admixture model first, to estimate the upper limit of the number of different clusters (Durand et al. 2009). We set a burn-in period of 100,000 iterations followed by sampling of 200,000 iterations, and tested for the number of clusters from K = 2 to 5 with 10 replicates of each K. As the most likely number of clusters we chose the one that matches the plateau of the deviance information criterion (DIC) curve. Then we performed 10 replicates of Tess analysis with the admixture model, applying the same parameters as before for the most likely K identified. We used CLUMPP (Jakobsson and Rosenberg 2007) to average an individual’s genotype proportions across 10 independent runs. Each individual was assigned to the genetic cluster for which its inferred ancestry was higher than 50 %. The next step was to use the cluster assignment in Tess to test for the rates of migration in BIMR (Faubet and Gaggiotti 2008). As BIMR assumes that sampling took place before migration, we used the clusters identified by Tess software rather than geographically defined groups. BIMR uses an MCMC algorithm, which allows for departure from Hardy–Weinberg equilibrium by using population-specific inbreeding coefficients, assumes migration-drift equilibrium in the prior generation, and calculates asymmetrical migration rates between pairs of populations during the last generation. The program uses the F-model which assumes that the admixture took place before the last generation of migration. That allows the program to characterise the rates of migration even between populations that are weakly differentiated. We set 95 % highest posterior density intervals (HPDI) and 25–45 % acceptance rates after 10 replicate runs to ensure convergence, with a burn-in and sample size of 100,000 and a thinning interval of 50 iterations. To extract parameter estimates, we chose the run with the lowest assignment component of total deviance (Dassign).
Results
Mitochondrial DNA variation
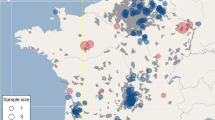
Partial sequences of the mitochondrial control region obtained for 72 samples representing all regions equally revealed four different haplotypes. Three of them (PLO2, GenBank acc. no. EF030393; PLO13, EF0303345; PLO16, EF0303348) were previously identified in a study of the phylogenetic relationships of raccoon populations in the original range of the species (Cullingham et al. 2008a, b) and one was newly detected (PL1, KC565878). Among those four haplotypes were ten segregating sites; haplotype diversity was 0.48 and nucleotide diversity 0.0076. Three haplotypes were detected in raccoons from Germany, two in the samples from Poland, and one from the Czech material. Haplotype PLO2 was the most frequent; it occurred in 68.05 % of the sampled individuals. Haplotype PLO16 was found in 19.4 %, PL1 in 6.9 % and PLO13 in 5 % of the individuals. Figure 1 presents the distribution of haplotypes among the sampled populations.
The minimum spanning network confirmed the division of raccoon mitochondrial haplotypes to three distinct phylogenetic lineages (Fig. 2). Haplotype PLO2 was placed in lineage III, and haplotypes PLO 13, PLO16 and PL1 in phylogenetic lineage II according to Cullingham et al. (2008a, b).
Microsatellite diversity
Among the 19 tested microsatellite loci we found six (PLO3-117F, PLO-M15F, PLO-M3F, PLO-M2R, PLOT4, PLOT9) with strong Hardy–Weinberg disequilibrium according to Fisher’s exact test, probably due to nonamplifying alleles; those loci were excluded from further analyses. The exact test for linkage disequilibrium among the remaining loci indicated no significant deviations for either comparison (p > 0.05). At the analysed loci, we found from 3 to 20 alleles on average. At all loci the observed heterozygosity was consistently lower than expected heterozygosity, resulting in slightly positive FIS values for all loci (Supplementary materials, Table S1). Heterozygote deficits were distributed evenly over the sampled loci. In the jackknifing procedure, removing one locus at a time, all loci contributed similarly to the given FIS (Supplementary materials, Table S2), including the three loci with significant FIS; thus we can conclude that the deviations from HWE are independent of null alleles but are connected to a population-level effect. Multilocus FIS was 0.116.
Population structure
On the basis of repeated STRUCTURE runs we detected three levels of hierarchical structure. At each round of the analysis the most probable K value based on the ΔK method was 2 (Table 1). Then we ran subsequent runs of STRUCTURE for samples assigned to different clusters based on average individual genotype proportion q > 0.5. When maximum lnP (D) indicated the most probable K value was one, we treated this cluster as the bottom level of structuring and performed subsequent runs for remaining samples. The results of the average individual’s genotype proportions for three consecutive runs of STRUCTURE are presented on Fig. 3. As the result of the first run all individuals from the Czech Republic apart of three were clustered in separate cluster (Fig. 3; (Level I) the light grey colour, cluster 1) and all others were grouped together in second cluster (cluster 1a). One individual sampled in southern Poland was grouped within a cluster 1a, even though its mitochondrial haplotype was shared with the Czech population. In the second run, we did not find additional substructure among individuals from cluster 1 but for remaining individuals again two groups were detected (Fig. 3; Level II). Most individuals grouped according to their sampling locality. Individuals sampled in Poland, but outside “Warta Mouth” National Park (Pol2) and in Germany (Ger1 and Ger2) grouped mostly in cluster 2a while cluster 2 consisted of individuals sampled in Poland in “Warta Mouth” National Park area, although individuals assigned to opposite cluster were also detected. According to this result, individuals assigned to cluster 2 were treated as a separate genetic group despite its geographic proximity to other samples from Poland. Again, we did not detect additional level of structuring among individuals assigned at level II to cluster 2. In the third round of analysis [Fig. 3; (Level III)], performed for individuals assigned to cluster 2a in the second run there was a visible structure between samples from Poland (Pol2) and Germany, but with a number of individuals assigned to opposite clusters. According to this result individuals were assigned to cluster 3 and cluster 4. The individual sampled in southern Poland, as well as three individuals sampled in the Czech Republic that were assigned to cluster 1a in the first round of analysis remained in cluster 4 (mainly individuals from Germany). When performing a separate STRUCTURE run on individuals assigned to clusters 3 and 4 no additional structuring was detected. On the basis of repeated STRUCTURE runs we detected three levels of hierarchical structure. At each round of the analysis the most probable K value based on the ΔK method was 2 (Table 1). Then we made further runs of STRUCTURE for samples assigned to different clusters based on average individual genotype proportion q > 0.5. When maximum lnP (D) indicated the most probable K value was one, we treated this cluster as the bottom level of structuring and performed subsequent runs for remaining samples. The results of the average individual’s genotype proportions for three consecutive runs of STRUCTURE are presented in Fig. 3. In the first run all individuals from the Czech Republic except three were placed in a separate cluster (Fig. 3; Level I, light grey, cluster 1) and all others were grouped in a second cluster (cluster 1a). One individual sampled in southern Poland was grouped in cluster 1a even though its mitochondrial haplotype was shared with the Czech population. In the second run we did not find additional substructure among the individuals from cluster 1 but for the remaining individuals two groups were again detected (Fig. 3; Level II). Most individuals grouped according to their sampling locality. Individuals sampled in Poland but outside Warta Mouth National Park (Pol2) and in Germany (Ger1 and Ger2) grouped mostly in cluster 2a, while cluster 2 consisted of individuals sampled in Poland in the Warta Mouth National Park area, although individuals assigned to the opposite cluster were also detected. According to this result, individuals assigned to cluster 2 were treated as a separate genetic group despite its geographic proximity to other samples from Poland. Again, we did not detect an additional level of structuring among individuals assigned at level II to cluster 2. In the third round of analysis (Fig. 3; Level III), performed for individuals assigned to cluster 2a in the second run, there was visible structure between the samples from Poland (Pol2) and Germany, but with a number of individuals assigned to opposite clusters. According to this result, individuals were assigned to cluster 3 and cluster 4. The individual sampled in southern Poland and the three individuals sampled in the Czech Republic that were assigned to cluster 1a in the first round of analysis remained in cluster 4 (mainly individuals from Germany). A separate STRUCTURE run on individuals assigned to clusters 3 and 4 did not detect additional structuring.
Average individual membership in genetic groups assessed in three consecutive runs of STRUCTURE revealing three levels of hierarchical structure observed in the dataset. Details described in Materials and method section. I First run performed for all individuals sampled, sampling locations indicated at the top and marked with dashed lines. II Second run of STRUCTURE, for all individuals assigned to dark grey cluster with individual q > 0.5 in the previous run. III Third run of STRUCTURE, for all individuals assigned to dark grey cluster with individual q > 0.5 in the previous run. Separate runs of STRUCTURE were performed for all individuals assigned to clusters shaded light grey in runs I, II and III. All resulted in K = 1. According to this result, individuals were grouped in clusters 1, 2, 3 and 4, and this grouping was used for subsequent population genetic analysis
Pairwise comparisons of FST values between STRUCTURE-defined genetic groups (Table 2) showed significant differentiation between all of them (p < 0.00001). Differentiation was the highest between the cluster 1, consisting of individuals from the Czech Republic, and all other individuals (0.169–0.229). Differentiation among the other clusters ranged from 0.033 to 0.048. AMOVA revealed only 9.1 % variation among raccoon populations grouped by country of origin, and the variance was not significant. Variance between populations within countries was 3.4 % (p = 0.000) and 87.5 % (p = 0.000) of variance was due to within-population diversity. Isolation-by-distance analysis (Mantel test) showed a significant but weak positive relationship between geographic and genetic distance (Fig. 4).
Within-population diversity
Genetic diversity indices were estimated for each STRUCTURE defined cluster (Table 3). According to allelic richness values corrected for sample size, diversity was the lowest in the cluster 1 formed from individuals from the Czech Republic. The remaining populations had similar and moderate values of genetic diversity. Cluster 1 had also the lowest effective population size. No recent bottlenecks were confirmed for any population although the SMM model indicated significant heterozygote deficiency in clusters 2 (Pol1), and 3 (Pol2) suggesting recent population expansion (Tables 4, 5).
Rates of migration between sites in Poland
Tess analysis of individuals originating from Poland, for which geographic coordinates were known, identified two different clusters corresponding to those obtained from the STRUCTURE. We identified two immigrant individuals from “Warta Mouth” National Park outside Park and five individuals that migrated in opposite direction. For BIMR analysis those migrants were placed with the populations of origin identified by Tess. Probability estimates of ancestral origin over the last generation estimated with BIMR indicated asymmetric migration rates between two clusters. Mean migration rates were consistent across all 10 BIMR runs. The run with the lowest Bayesian deviance (Dassign) indicated mean 0.281 emigration (expressed as a proportion of population that emigrated within the last generation) from “Warta Mouth” National Park (95 % HDPI 0.069–0.507, mode 0.278) and mean 0.033 immigration to the park (95 % HDPI 0.005–0.442, mode = 0.008).
Discussion
In this study we used both mitochondrial and microsatellite (nuclear) markers to obtain data from an extensive sampling of raccoons in Poland, Germany and the Czech Republic in order to understand the genetic differentiation patterns of this invasive introduced species. These are the first results on raccoon genetic variation in Central Europe, the main introduced range in Europe. We found relatively high genetic diversity of microsatellite markers but reduced genetic diversity of mitochondrial markers, an effect of the low number of individuals introduced in Europe. Despite this fact, intensive expansion of this species from Germany as well as the Czech Republic has been observed.
Mitochondrial diversity
Our analysis of the mitochondrial diversity of raccoons from Poland, Germany and the Czech Republic indicates that the species became a successful invader despite strong reduction of its genetic diversity. We found only four distinct mitochondrial haplotypes, in contrast to results from the native range, where 76 haplotypes were found among 311 samples. Other European populations of raccoons also exhibit low mitochondrial diversity. In Spain only two haplotypes were found among 58 individuals (Alda et al. 2013), and in central and eastern Germany only six haplotypes were detected in a sample of 193 raccoons (Frantz et al. 2013). The haplotypes reported in our study overlap with those detected in the German study. PLO2 was the most common haplotype in our study, in line with findings from Frantz et al. (2013). In their work the PLO2 haplotype was separated into PLO2a and PLO2b, as they analysed a slightly longer fragment. Haplotype PLO2a occurred in the Hesse region (central Germany), and PLO2b was widespread in the former East Germany. Unfortunately we could not distinguish those two haplotypes, but the occurrence of one of them in eastern Germany is in accord with our result, as we detected PLO2 in Poland and in eastern Germany (Saxony). In the same region we also found haplotype PLO16, similar to Frantz et al. (2013). Haplotype PL1, found in our study, and denoted as PLO110 (longer sequence) in the German study, was found in central eastern Germany. Haplotype PLO 13, which we found only in the Czech population and one sample from southern Poland, was detected in western Germany by Frantz et al. (2013). The absence of this haplotype in all other populations sampled in this study suggests a separate invasion route of raccoons from Germany to the Czech Republic and further to southern Poland. In general, the good agreement between these two studies strongly supports the German origin of the Polish and Czech raccoon populations.
From the mitochondrial data we cannot identify the origin of the introduced populations. The haplotype that characterises the Czech population is common and widespread across the native range in eastern North America (Cullingham et al. 2008a). The same applies to the haplotypes found in the German and Polish populations. The PLO2 haplotype is the most common and widespread in the native range. Similarly, the other two haplotypes detected in this study are spread throughout the native range, as the population structure of raccoons in North America is the result of intensive artificial exchange of individuals between regions (Cullingham et al. 2008a, b).
Population structure and within-population diversity
Data analysis showed the presence of different genetic clusters in the studied area, confirmed by both mitochondrial and nuclear markers and probably attributable to two different invasion routes in the studied region. The Czech Republic group was clearly distinguished from all others by STRUCTURE clustering and FST pairwise comparisons. In view of its having the lowest values for heterozygosity, allelic richness and effective population size, it was probably established from fewer individuals than were the populations of the other sites. Interestingly, three individuals from the Czech Republic and the only raccoon sampled in southern Poland (Fig. 1, point 3) also possessed the PLO13 haplotype but shared ancestry in its nuclear genome with raccoons from Germany and not the Czech population. This might be the result of a few generations of backcrossing with raccoons that shared ancestry with those from northern Poland and Germany, despite its origin from the Czech maternal lineage. On the other hand, this finding also indicates a separate invasion route, from western Germany to the Czech Republic. We did not sample individuals from western Germany so we cannot exclude the possibility that individuals from the Czech Republic would cluster with them in nuclear genome analyses. We suspect, however, that in southern Poland there is a zone of contact between individuals from two different invasion routes. It seems a likely scenario, given the short time elapsed since raccoons were introduced in Poland. This would be important from the management point of view, as such a mixed population would be expected to exhibit high levels of genetic diversity and high invasive potential, as has been suggested for intraspecific hybrids of non-native species (Facon et al. 2008; Kolbe et al. 2007; Lucek et al. 2010). Confirmation of this scenario requires extensive sampling of raccoons in that region.
Apart from the Czech population, we found an additional level of substructure among the remaining samples. The genetic structure of raccoons from other locations is detectable but weak in terms of FST, but consecutive runs of STRUCTURE revealed a clear division between raccoons inhabiting Warta Mouth National Park and the others from Poland and Germany. The clustering together of individuals from Poland and Germany in the second round of analysis shows their common origin. Both clusters detected in Poland were fixed for the most common mitochondrial haplotype, PLO2, which was also found in samples from Germany. What is more, the samples from both Polish clusters showed lower allelic richness than those from Germany. This would support the scenario of raccoon invasion from Germany to western Poland, as we observe a diversity gradient such as that seen when a population from a nearby source area disperses into an unoccupied area by a gradual stepping-stone process. The genetic variation present in the source population will progressively decline with the distance to newly established sites (Le Corre and Kremer 1998). Weak differentiation indicating a strong and recent invasion was also confirmed by AMOVA, as the differentiation between regions hosting populations from each country was low and non-significant. The weak but significant IBD pattern also confirms intensive colonisation and high mobility of individuals.
The consistent pattern of heterozygote deficiency across all markers as well as the positive FIS values across populations suggest the Wahlund effect, possibly resulting from mixing of individuals from different sources, or else inbreeding within populations established from highly related individuals. The haplotype composition and low effective population size, combined with the high density of raccoons in Germany (Hohmann 2001), rather argue for interbreeding of related individuals and against multiple sources of introduction of individual populations. The level of intrapopulation genetic diversity, expressed as expected heterozygosity values and allelic richness, are comparable with those reported for raccoon populations in the native species range (Cullingham et al. 2006, 2008a, Root et al. 2009, Hauver et al. 2010, Santonastaso et al. 2012). Four of the loci analysed in this study were also used to reveal the genetic diversity of raccoons in the native range. The number of alleles for those loci in individuals sampled in the studied region is similar, or in two cases even higher than in native populations (Table S3, Supplementary material). A similar level of population genetic diversity was also found in invasive populations of raccoon in Spain (Alda et al. 2013). A higher number of alleles in an introduced population can be a result of mixing individuals from different source populations. As mentioned before, the only population exhibiting lower diversity is the one from the Czech Republic, but the number of alleles for all analysed individuals is higher than the number of alleles when individuals from the Czech Republic are excluded. This would also support the separate origin of the Czech population.
The model-based assignment analyses, both those from STRUCTURE and those from Tess incorporating geographic location of samples, disclosed two different clusters among the raccoons sampled in Poland despite their close geographic proximity. Although geographically close, those raccoons have very different habitats. Individuals grouped in cluster 2 were sampled mainly in the “Warta Mouth” National Park where habitat is dominated by wetlands, scrub willow, marshes and meadows. The raccoons grouped in cluster 3 inhabit a much drier area outside the park covered by coniferous and deciduous forest. In analysing the asymmetrical effective movement between these two contrasting habitats we detected almost ten times higher emigration from the Park than immigration to it, although the HDPI values overlapped. What is more, the low FST between these two groups indicates a substantial level of overall gene flow between them and probably prevents detection of directional migrations. On the other hand, the sharp division between raccoons sampled in Warta Mouth National Park and outside it could suggest source-sink dynamics between these two subpopulations, related to raccoon density in these two habitats. Raccoons inhabit the floodplains of the lower Warta River valley (in the Park) at much higher density, and they have smaller home ranges; the raccoons inhabiting the forest have much larger home ranges, suggesting their density is lower there (Bartoszewicz et al. 2008). Such optimal habitats where raccoon density is very high could host source populations for further colonisation. This means that the management strategy should concentrate especially on such areas, to reduce the rate of invasion to other places. Those signs of strong and intensive expansion of the raccoon in Poland are supported by the results of the BOTTLENECK test, showing a significant heterozygote deficiency in clusters 2 and 3 formed from Polish individuals.
Despite extensive expansion of the raccoon in Central Europe, associated with long-distance dispersal and intensive gene flow between different areas, the population of raccoons is structured. In the new range of an invasive species its genetic structure is often well developed (Zeisset and Beebee 2003; Herborg et al. 2007; Rollins et al. 2009), and sometimes invasive species exhibit higher structure than native populations (Marrs et al. 2008; Gillis et al. 2009); however patterns of genetic differentiation are not uniform (e.g. Schrey et al. 2011). The genetic structure of invasive species is developed by multiple introductions to multiple locations (e.g. connected with the distribution of farms from which the feral populations originated) as well as by geographical barriers (e.g. mountain ranges) and genetic drift in newly established populations (Lecis et al. 2008; Zalewski et al. 2009, 2010; Inoue et al. 2012). Our results suggest that genetic structure can also develop by invasion and adaptation of alien species to different habitats. Genetic diversification may have further implications for the native species. By exploiting different habitats, raccoons are likely to exert a different kind of predation pressure on their prey, altering both the size and dynamics of the prey population and in turn altering the ecosystem.
Here we demonstrated that an introduced species, the raccoon, lost much of its mtDNA diversity upon introduction but maintained most of the nuclear diversity present in the native population. This finding testifies to the high adaptive ability of the raccoon and points to the very high risk that a few escapees from farms or a few animals released by private owners will establish a new population. Admixture upon secondary contact between raccoons from separate introductions may further accelerate expansion. Further expansion poses a great threat to native species, and to human health as well, as raccoons carry many diseases and parasites (Popiołek et al. 2011; Beltran-Beck et al. 2012). The risk of disease transmission would be high because raccoons are very likely to exploit anthropogenic habitats (Bartoszewicz et al. 2008; Garcia et al. 2012). Therefore management should above all aim to prevent their establishment at new locations. It should also aim to reduce the opportunity for admixture resulting from secondary contact between populations from various introductions, and to prevent accidental or deliberate introduction of ranch animals to an already established population, which would pose the risk of increasing the genetic diversity of the invasive species.
References
Abdelkrim J, Pascal M, Calmet C, Samadi S (2005) Importance of assessing population genetic structure before eradication of invasive species: examples from insular Norway rat populations. Conserv Biol 19:1509–1518
Alda F, Ruiz-López MJ, García FJ, Gompper ME, Eggert LS, García JT (2013) Genetic evidence for multiple introduction events of raccoons (Procyon lotor) in Spain. Biol Invasion 15:687–698
Bartoszewicz M, Okarma H, Zalewski A, Szczesna J (2008) Ecology of the raccoon (Procyon lotor) from western Poland. Ann Zool Fenn 45:291–298
Beasley JC, Rhodes OE (2008) Relation between raccoon abundance and crop damage. Hum Wildl Confl 2:248–259
Beltran-Beck B, Garcia FJ, Gortazar C (2012) Raccoons in Europe: disease hazards due to the establishment of an invasive species. Eur J Wildl Res 58:5–15
Červeny J, Andĕra M, Koubek P, Homolka M, Toman A (2001) Recently expanding mammal species in the Czech republic: distribution, abundance and legal status. Beiträge zur Jagd-und Wildforschung 26:111–125
Chen C, Durand E, Forbes F, François O (2007) Bayesian clustering algorithms ascertaining spatial population structure: a new computer program and a comparison study. Mol Ecol Notes 7:747–756
Chesnokov NI (1989) Dzikije zywotnyje meniajut adriesa. Mysl, Moskwa
Conovery MR (1987) Reducing raccoon and birds damage to small corn plots. Wildl Soc Bull 15:268–272
Cornuet JM, Luikart G (1997) Description and power analysis of two tests for detecting recent population bottlenecks from allele frequency data. Genetics 144:2001–2014
Cote H, Garant D, Robert K, Mainguy J, Pelletier F (2012) Genetic structure and rabies spread potential in raccoons: the role of landscape barriers and sex-biased dispersal. Evol Appl 5:393–404
Coulon A, Fritzpatrik JW, Bowman R, Stith BM, Makarewich CA, Stenzler LM, Lovette IJ (2008) Congruent population structure inferred from dispersal behaviour and intensive genetic surveys of the threatened Florida scrub-jay (Aphelocoma cœrulescens). Mol Ecol 17:1685–1701
Cullingham CI, Kyle CJ, White BN (2006) Isolation, characterization and multiplex genotyping of raccoon tetranucleotide microsatellite loci. Mol Ecol Notes 6:1030–1032
Cullingham CI, Kyle CJ, Pond BA, White BN (2008a) Genetic structure of raccoons in Eastern North America based on mtDNA: implications for subspecies designation and rabies disease dynamics. Can J Zool 86:947–958
Cullingham CI, Pond BA, Kyle CJ, Rees EE, Rosatte RC, White BN (2008b) Combining direct and indirect genetic methods to estimate dispersal for informing wildlife disease management decisions. Mol Ecol 17:4874–4886
DAISIE (2009) Handbook of alien species in Europe. Series: invading nature. Springer series in invasion ecology. Volume 3. Springer, Dordrecht
Dlugosch KM, Parker IM (2008) Founding events in species invasions: genetic variation, adaptive evolution, and the role of multiple introductions. Mol Ecol 17:431–449
Durand E, Jay F, Gaggiotti OE, François O (2009) Spatial inference of admixture proportions and secondary contact zones. Mol Biol Evol 26:1963–1973
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361
Elis JC, Shulman MJ, Jessop H, Suomala R, Morris SR, Seng V, Wagner M, Mach K (2007) Impact of raccoons on breeding success in large colonies of Great Black Gulls and Herring Gulls. Waterbirds 30:375–383
Evanno et al (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Excoffier LG, Laval SS (2005) Arlequin ver. 3.0: an integrated software package for population genetics data analysis. Evol Bioinform Online 1:47–50
Facon B, Pointier JP, Jarne P, Sarda V, David P (2008) High genetic variance in life-history strategies within invasive populations by way of multiple introductions. Curr Biol 18:363–367
Falush D, Stephens M, Pritchard JK (2003) Inference of population structure using multilocus genotype data: linked loci and correlated allele frequencies. Genetics 164:1567–1587
Faubet P, Gaggiotti OE (2008) A new Bayesian method to identify the environmental factors that influence recent migration. Genetics 178:1491–1504
Fike JA, Drauch AM, Beasley JC, Dharmarajan G, Rhodes OE (2007) Development of 14 multiplexed microsatellite loci for raccoons Procyon lotor. Mol Ecol Notes 7:525–527
Frantz AC, Heddergott M, Lang J, Schulze C, Ansorge H, Runge M, Braune S, Michler F-U, Wittstatt U, Hoffmann L, Hohmann U, Michler BA, Van Den Berge K, Horsburgh GJ (2013) Limited mitochondrial DNA diversity is indicative of a small number of founders of the German raccoon (Procyon lotor) population. Eur J Wildl Res. doi:10.1007/s10344-013-0719-6
Garcia JT, Garcia FJ, Alda F, Gonzalez JL, Aramburu MJ, Cortes Y, Prieto B, Pliego B, Perez M, Herrera J, Garcia-Roman L (2012) Recent invasion and status of the raccoon (Procyon lotor) in Spain. Biol Invasions 14:1305–1310
Gillis NK, Walters LJ, Fernandes FC, Hoffman EA (2009) Higher genetic diversity in introduced than in native populations of the mussel Mytella charruana: evidence of population admixture at introduction sites. Divers Distrib 15:784–795
Goudet J (1995) FSTAT (Version1.2): a computer program to calculate F-statistics. J Hered 86:485–486
Hall TA (1999) BioEdit: a user-friendly biological sequence alignment editor and analysis program for Windows 95/98/NT. Nucleic Acids Symp Ser 41:95–98
Hartman LH, Eastman DS (1999) Distribution of introduced raccoons Procyon lotor on the Queen Charlotte Islands: implications for burrow-nesting seabirds. Biol Conserv 88:1–13
Hartman LH, Gaston AJ, Eastman DS (1997) Raccoon predation on ancient murrelets on East Limestone Island, British Columbia. J Wildl Manag 61:377–388
Hauver SA, Gehrt SD, Prange S, Dubach J (2010) Behavioral and genetic aspects of the raccoon mating system. J Mamm 91:749–757
Herborg LM, Weetman D, Van Oosterhout C, Hanfling B (2007) Genetic population structure and contemporary dispersal patterns of a recent European invader, the Chinese mitten crab, Eriocheir sinensis. Mol Ecol 16:231–242
Hohmann U (2001) Stand und Perspektiven der Erforschung des Waschbaren in Deutschland. Beiträge zur Jagd-und Wildforschung 26:181–186
Hohmann U, Voig S, Andreas U (2002) Raccoons take the offensive. A current assessment. In: Kowarik I, Starfinger U (Eds) Biologische Invasionen, Neobiota 1: pp 191–192
Holderegger R, Gugerli F (2012) Where do you come from, where do you go? Directional migration rates in landscape genetics. Mol Ecol 21:5640–5642
Hubisz M, Falush D, Stephens M, Pritchard J (2009) Inferring weak population structure with the assistance of sample group information. Mol Ecol Resour 9:1322–1332
Inoue T, Kaneko Y, Yamazaki K, Anezaki T, Yachimori S, Ochiai K, Lin LK, Pei KJC, Chen YJ, Chang SW, Masuda R (2012) Genetic population structure of the masked palm civet Paguma larvata (Carnivora: Viverridae) in Japan, revealed from analysis of newly identified compound microsatellites. Conserv Genet 13:1095–1107
Jakobsson M, Rosenberg N (2007) CLUMPP: a cluster matching and permutation program for dealing with label switching and multimodality in analysis of population structure. Bioinformatics 23:1801–1806
Keller SR, Taylor DR (2010) Genomic admixture increases fitness during a biological invasion. J Evolut Biol 23:1720–1731
Kolbe JJ, Glor RE, Schettino LR, Lara AC, Larson A, Losos JB (2007) Multiple sources, admixture, and genetic variation in introduced Anolis lizard populations. Conserv Biol 21:1612–1625
Le Corre V, Kremer A (1998) Cumulative effects of founding events during colonization on genetic diversity and differentiation in an island and stepping-stone model. J Evol Biol 11:495–512
Lecis R, Ferrando A, Ruiz-Olmo I, Manas S, Domingo-Roura X (2008) Population genetic structure and distribution of introduced American mink (Mustela vison) in Spain, based on microsatellite variation. Conserv Genet 9:1149–1161
Leger F (1999) Le Raton laveur en France. Bull Mens ONC 241:16–37
Librado P, Rozas J (2009) DnaSP v5: a software for comprehensive analysis of DNA polymorphism data. Bioinformatics 25:1451–1452
Lockwood JL, Hoopes MF, Marchetti MP (2007) Invasion ecology, 1st edn. Blackwell, Oxford, pp 1–304
Lucek K, Roy D, Bezault E, Sivasundar A, Seehausen O (2010) Hybridization between distant lineages increases adaptive variation during a biological invasion: stickleback in Switzerland. Mol Ecol 19:3995–4011
Lutz W (1984) The distribution of the raccoon (Procyon lotor, Linne 1758) in Central Europe. Z Jagdwiss 30:218–228
Marrs RA, Sforza R, Hufbauer RA (2008) When invasion increases population genetic structure: a study with Centaurea diffusa. Biol Invasions 10:561–572
Milenkovic DC (2003) The first record of the free-ranging raccoon (Procyon lotor Linnaeus, 1758) in Yugoslavia. Mamm Biol 6:116–117
Mitchell-Jones AJ et al (1999) The atlas of European mammals. T&A D Poyser Natural History, London
Park SY, Glaser C, Murray WJ, Kazacos KR, Rowley HA, Fredrick DR, Bass N (2000) Raccoon roundworm (Baylisascaris procyonis) encephalitis: Case report and field investigation. Pediatrics 106:E56
Peakall R, Smouse PE (2006) GenAlEx 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Popiołek M, Szczesna-Staskiewicz J, Bartoszewicz M, Okarma H, Smalec B, Zalewski A (2011) Helminth parasites of an introduced invasive carnivore species, the raccoon (Procyon lotor L.), from the Warta Mouth National Park (Poland). J Parasitol 97:357–360
Pritchard JK, Stevens M, Donnely P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
Ratnaswamy MJ, Warren RJ, Kramer MT, Adam MD (1997) Comparisons of lethal and nonlethal techniques to reduce raccoon depredation of sea turtle nests. J Wildl Manag 61:368–376
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rollins LA, Woolnough AP, Wilton AN, Sinclair R, Sherwin WB (2009) Invasive species can’t cover their tracks: using microsatellites to assist management of starling (Sturnus vulgaris) populations in Western Australia. Mol Ecol 18:1560–1573
Root JJ, Puskas RB, Fischer JW, Swope CB, Neubaum MA, Reeder SA, Piaggio AJ (2009) Landscape genetics of raccoons (Procyon lotor) associated with ridges and valleys of Pennsylvania: implications for oral rabies vaccination programs. Vector Borne Zoonotic Dis 9:583–588
Rosatte R, Sobey K, Donovan D, Allan M, Bruce L, Buchanan T, Davies C (2007) Raccoon density and movements after population reduction to control rabies. J Wildl Manag 71:2373–2378
Rosatte R, Ryckman M, Ing K, Proceviat S, Allan M, Bruce L, Donovan D, Davies JC (2010) Density, movements, and survival of raccoons in Ontario, Canada: implications for disease spread and management. J Mamm 91:122–135
Santonastaso TT, Dubach J, Hauver SA, Graser WH, Gehrt SD (2012) Microsatellite analysis of raccoon (Procyon lotor) population structure across an extensive metropolitan landscape. J Mamm 93:447–455
Schley L et al (2001) Neuburger und Heimkehre unte den Wildtieren Luxemburgs. Beitr Jagd-Wildforschung 26:141–154
Schrey AW, Grispo M, Awad M et al (2011) Broad-scale latitudinal patterns of genetic diversity among native European and introduced house sparrow (Passer domesticus) populations. Mol Ecol 20:1133–1143
Seehausen O (2004) Hybridization and adaptive radiation. Trends Ecol Evol 19:198–207
Simberloff D (2003) How much information on population biology is needed to manage introduced species? Conserv Biol 17:83–92
Teacher AGF, Griffiths DJ (2011) HapStar: automated haplotype network layout and visualization. Mol Ecol Notes 11:151–153
Vos A, Ortmann S, Kretzschmar AS, Kohnemann B, Michler F (2012) The raccoon (Procyon lotor) as potential rabies reservoir species in Germany: a risk assessment. Berl Munch Tierarztl Wochenschr 125:228–235
Waples RS, Do C (2006) LdNe: a program for estimating effective population size from data on linkage disequilibrium. Mol Ecol Notes 8:753–756
Zalewski A, Piertney SB, Zalewska H, Lambin X (2009) Landscape barriers reduce gene flow in an invasive carnivore: geographical and local genetic structure of American mink in Scotland. Mol Ecol 18:1601–1615
Zalewski A, Michalska-Parda A, Bartoszewicz M, Kozakiewicz M, Brzeziński M (2010) Multiple introductions determine the genetic structure of an invasive species population: American mink Neovison vison in Poland. Biol Conserv 143:1355–1363
Zeisset I, Beebee TJC (2003) Population genetics of a successful invader: the marsh frog Rana ridibunda in Britain. Mol Ecol 12:639–646
Zeveloff SI (2002) Raccoons a natural history. Smithsonian Institution Press, Washington
Acknowledgments
We are grateful to Dr. M. Popiołek, Dr. H. Baacke, Landesjagdverband Sachsen e.V. (Dresden, Germany) and the staff of “Warta Mouth” National Park for help in collecting raccoon samples. Michael Jacobs line-edited the manuscript. This study was supported under research project no. NN309 135535, funded by the Polish Ministry of Science and Higher Education.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution License which permits any use, distribution, and reproduction in any medium, provided the original author(s) and the source are credited.
About this article
Cite this article
Biedrzycka, A., Zalewski, A., Bartoszewicz, M. et al. The genetic structure of raccoon introduced in Central Europe reflects multiple invasion pathways. Biol Invasions 16, 1611–1625 (2014). https://doi.org/10.1007/s10530-013-0595-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10530-013-0595-8