Abstract
Background
Lymphovascular invasion (LVI) is a prerequisite step in breast cancer (BC) metastasis. We have previously identified wild-type isocitrate dehydrogenase 2 (IDH2) as a key putative driver of LVI. Thus, we explored the prognostic significance of IDH2 at transcriptome and protein expression levels in pre-invasive and invasive disease.
Methods
Utlising tissue microarrays from a large well annotated BC cohort including ductal carcinoma in situ and invasive breast cancer (IBC), IDH2 was assessed at the transcriptomic and proteomic level. The associations between clinicopathological factors including LVI status, prognosis and the expression of IDH2 were evaluated.
Results
In pure DCIS and IBC, high IDH2 protein expression was associated with features of aggressiveness including high nuclear grade, larger size, comedo necrosis and hormonal receptor negativity and LVI, higher grade, larger tumour size, high NPI, HER2 positivity, and hormonal receptor negativity, respectively. High expression of IDH2 either in mRNA or in protein levels was associated with poor patient’s outcome in both DCIS and IBC. Multivariate analysis revealed that IDH2 protein expression was an independent risk factor for shorter BC specific-survival.
Conclusion
Further functional studies to decipher the role of IDH2 and its mechanism of action as a driver of BC progression and LVI are warranted.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Breast cancer (BC) progression is a complex multifactorial process. However, the invasion machinery which is a critical step in progression from in situ to infiltrating tumour followed by distant metastasis remains unclear. Deciphering the transcriptomics and proteomics that govern the invasive and metastatic cascades of BC is essential in understanding the mechanisms of cancer progression. Lymphovascular Invasion (LVI) is an independent prognostic factor of poor outcome in invasive BC (IBC) and is a prerequisite for the tumour metastasis [1,2,3,4]. Understanding the molecular mechanisms underlying BC progression, and in particular LVI, and unveiling their driver molecular pathways could ultimately improve patient outcomes [5]. Through stringent bioinformatics analysis, we have previously interrogated transcriptomic datasets of IBC [Molecular Taxonomy of Breast Cancer International Consortium (METABRIC) and The Cancer Genome Atlas (TCGA)] for putative drivers of LVI [6]. Briefly, LVI positive and negative cases from these cohorts were subjected to a method of weighted average difference (WAD) and subsequently differentially expressed genes (DEGs) were identified based on WAD rankings [7]. Forty-two significantly overexpressed and 57 downregulated genes identified in the METABRIC cohort were identified and validated in the TCGA cohort [8]. Wild-type Isocitrate Dehydrogenase 2 (IDH2) was a highly expressed gene associated with presence of LVI [8]. Moreover, IDH2 was reported to be differentially expressed between recurrent and non-recurrent DCIS and associated with DCIS recurrence and progression to invasive disease [9, 10].
IDH2 is a member of the isocitrate dehydrogenase family that plays a key role in cellular metabolism and acts in the tricarboxylic acid (TCA) cycle as an NADP + consuming enzyme, producing NADPH. In the reverse TCA cycle, when IDH2 causes reductive carboxylation of 2-oxoglutarate (2-OG), it consumes cell during hypoxia to survive the lower glucose levels [11].
Cellular energy and biosynthetic intermediates are produced by the TCA cycle, which are upregulated in metastasised cancer cells. Glycolysis is also upregulated in cancer cells to produce biosynthetic intermediates and energy needed for cellular proliferation and survival. Circulating tumour cells are predisposed to anoikis as a result of impaired glucose uptake. Thus, the metastasised tumour cells evade anoikis by upregulation of the TCA cycle [12]. Previous studies have reported upregulation of wild-type IDH2 in lung cancer, ovarian cancer, endometrioid carcinoma, and advanced colorectal cancer [13, 14]. Gain of function mutations in IDH2 result in an increase in the oncometabolite 2-hydroxyglutarate (2-HG) which is believed to link aberrant metabolism and aberrant epigenetics and gene regulation in cancer [15]. A well-known function of mutant IDH2 has been demonstrated in cancers such as glioma, cholangiocarcinoma, and breast solid papillary carcinoma with reverse polarity (SPCRP) [16]. Wild-type IDH2 overexpression is an indicator of poor outcome in lung cancer through the stimulation of the Warburg effect to help the maintenance of cancer cells via activation of hypoxia inducible factor 1α (HIF1α) which supports tumour growth in hypoxic environments [17]. However, the role of wild-type IDH2 in BC is still unclear. In this study, we aimed to assess the expression of wild-type IDH2 in BC and evaluate its role in tumour progression, particularly LVI, and patient outcome.
Materials and methods
IHD2 protein expression
Study cohorts
Large well-characterised BC cohorts consisting of pure ductal carcinoma in situ (DCIS; n = 776) and invasive disease (IBC; n = 859) from patients presented between 1998 and 2006 at Nottingham City Hospital, Nottingham, United Kingdom (UK) as previously described [18, 19] were utilised in this study. Patients’ demographic data, tumours’ morphological features, treatment data including surgery, hormonal therapy, radiotherapy and chemotherapy were available for both cohorts. Patients receiving neoadjuvant therapy were excluded in this study. Oestrogen Receptor (ER), Progesterone Receptor (PgR), HER2 status and Ki67 data was available [20, 21]. As per previous publications [20, 22, 23], ER and PgR was defined as ≥ 1%. HER2 positivity was defined when ≥ 10% of tumour cells showed strong membranous staining (score + 3), where chromogenic in situ hybridisation technique (CISH) was used to assess the gene amplification status in borderline cases (+2). BC molecular subtypes (for both DCIS and IBC cohorts) including luminal A (ER+/HER2−; Ki67 < 10%), Luminal B (ER+/HER2−; Ki67⩾10%), HER2-positive class (HER2+ regardless of ER status), and TN (ER−, PgR− and HER2 −) were defined based on IHC profile. For further understanding the molecular interactions of IDH2, available data on basal phenotype (CK5, CK14, and EGFR), EMT related markers (E-cadherin, N-cadherin, P-cadherin, TGF beta, and TWIST2), and glutamine metabolism proteins (SLC1A5, SLC3A2, SLC7A5, GLS, ALDH18A1, ALDH4A1, PRODH) were included in this study as per previous publications [24,25,26,27,28].
The patient records were regularly updated including patient’s outcome and follow-up. Local recurrence free interval (LRFI) in DCIS was defined as the time (in months) between 6 months after the first DCIS surgical removal and the occurrence of ipsilateral local recurrence. Patients had close/positive surgical margins or presented with residual tumour tissue and undergoing re-excision surgery within the first 6 months were not considered as recurrence. Patients who developed contralateral breast event after initial diagnosis of DCIS were censored at the time of occurrence of the contralateral disease. For IBC, data on breast cancer-specific survival (BCSS) was defined as the period (in months) extending from the date of primary surgery to the time of death due to breast cancer, and time to distant metastasis (TTDM) was defined as the period (in months) from primary surgery to occurrence of first distant metastasis.
Immunohistochemistry (IHC)
Twenty full face BC tissue sections (including DCIS and IBC) based on different tumour grades, LVI status and histological type were stained using IHC to evaluate the pattern of IDH2 protein expression prior to staining of Tissue Microarrays (TMAs). TMAs were previously prepared using a TMA Grand Master® (3D HISTECH®, Budapest, Hungary) [19, 26].
Primary antibody specificity for the mouse monoclonal anti-wild-type IDH2 antibody (ab55271, Abcam, UK) was validated using Western blot. An array of breast cancer cell line lysates was used, which include MCF-10A, MCF7, and MB-MDA-231 (obtained from the American Type Culture Collection, Rockville, MD, USA). IDH2 antibody was used at a dilution of 1:500, which showed a single specific band at the predicted molecular weight of 47 kDa (Supplementary Fig. 1).
Antigen retrieval was performed based on the manufacturer’s recommendations (citrate buffer pH 6.0 at 1000 W for 20 min using microwave). Expression of IDH2 protein was assessed by IHC using the Novocastra Novolink™ Polymer Detection Systems kit (Code: RE7280-K, Leica, Biosystems, UK), where 4 µm sections were incubated for 60 min with mouse monoclonal IDH2 (dilution 1:500). Normal kidney tissue was used as a positive control, while a negative control was carried out by omitting the primary antibody.
Scoring of IDH2 expression
Assessment of IDH2 cytoplasmic expression was performed using the semi-quantitative Histochemical score (H-score), where staining intensity was multiplied by the percentage of representative cells in the tissue for each intensity, producing a range of values between 0 and 300 [29]. All non-representative cores (folded tissue during processing and staining or cores with only normal breast tissue or those containing < 15% tumour cells relative to the whole core area) were excluded from scoring. The scoring was performed by AA blinded to patients’ clinicopathological and outcome, with a subset of cores (~ 10%) scored independently by another scorer (MAA) to calculate the inter-observer concordance. The protein expression of IDH2 was dichotomised by cut-off points generated from X-tile bioinformatics software version 3.6.1 (Yale University, USA) based on BCSS in IBC and LRFI in DCIS. An H-score of 70 was the optimal cut-off value of IDH2 protein expression in IBC, while a H-score of 45 was used to dichotomise DCIS cases into high and low expression.
IDH2 transcriptomic analysis
Two datasets comprising the METABRIC (n = 1980) [6] and TCGA breast carcinoma (TCGA BRCA, n = 854) [30] were used to evaluate IDH2 mRNA expression. The median was used as cut-off to categorise mRNA expression levels into high and low subgroups. For further validation of the prognostic significance of IDH2 expression in BC, the prognostic analytical module within the publicly available online dataset of Breast Cancer Gene Expression Miner v4.0 (http://bcgenex.centregauducheau.fr) was used.
Statistical analysis
Statistical analysis was performed using SPSS, version 24 (Chicago, IL, USA). The interclass correlation coefficient (ICC) test was used to assess the concordance rate of the IDH2 scoring between both the observers. The association between IDH2 and various clinicopathological parameters in both cohorts was analysed using Chi-square test. Mann–Whitney test was used to compare between IDH2 expression in DCIS and IBC. The Spearman’s rank correlation coefficient was used to assess the correlation between IDH2 mRNA and protein expression levels in IBC.
Log rank test and Kaplan–Meier curves were used for univariate survival analysis. Analysis with recurrence in DCIS was carried out for the breast conservative surgery (BCS) treated group (and not for those treated by mastectomy). Cox regression model was used for multivariate analysis. For all tests, a two-tailed p value < 0.05 was considered as statistically significant.
Results
Patterns of IDH2 protein expression
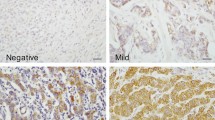
The full face tissue sections demonstrated an even staining of IDH2 expression within IBC and DCIS, indicating that TMA cores are representative for the whole tumour when assessing IDH2 expression. Adjacent normal breast terminal duct-lobular units showed occasional weak cytoplasmic staining. Occasional stained inflammatory cells and stromal fibroblasts were evident in a few cores. When present, IDH2 was expressed in the cytoplasm of epithelial tumour cells (Fig. 1). After exclusion of uninformative cores, 512 and 859 cases were available to evaluate the immunoreactivity in DCIS and IBC, respectively. There was a strong concordance between both observers in IDH2 immunoscoring (ICC = 0.855, p < 0.001). Distribution of IDH2 expression showed unimodal distribution. High IDH2 expression was observed in 49% and 59% of pure DCIS and IBC cases, respectively. IDH2 expression was higher in IBC than DCIS (Z = 9.5, p < 0.0001).
Photomicrographic images (× 40) for immunohistochemical protein expression of IDH2 in breast tissue microarray images; a normal breast terminal duct-lobular showing negative to faint staining, b negative expression in invasive breast carcinoma, c positive expression in invasive breast carcinoma, and d positive expression in DCIS
Significance of IDH2 protein expression
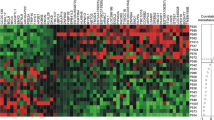
In pure DCIS, numerous clinicopathological parameters indicating poor DCIS prognosis were associated with high IDH2 expression (Table 1) including younger age at diagnosis, (p = 0.035), larger tumour size, higher nuclear grade (both p < 0.0001), presence of comedo type necrosis (p = 0.001), hormonal receptor negativity and HER2 positivity (all p < 0.0001). In IBC, high IDH2 protein expression was associated with positive LVI status (p = 0.002), high histological grade, high Nottingham Prognostic Index (NPI), hormonal receptor negativity, HER2 positivity (all p < 0.0001), large tumour size (p = 0.005), basal phenotype (p = 0.007) and high proliferation index (Ki67) (p = 0.008) (Table 2). There was positive correlation between IDH2 protein expression and the expression of EGFR (p < 0.0001), CK5 (p = 0.003), EMT markers including E-cadherin (p = 0.003), N-cadherin, P-cadherin and TWIST2 (all p < 0.0001). There was a positive correlation between IDH2 and enzymes within the glutamine-proline regulatory axis: GLS (p < 0.001), PRODH (p = 0.010), ALDH4A1 (p = 0.012), and solute transporters including SLC1A5 (p = 0.002), SLC3A2 and SLC7A5 (both; p < 0.001). Correlation matrix of IDH2 with other associated proteins in IBC is shown in Fig. 2.
Correlation matrix showing the association between IDH2 protein expression and with other biomarkers related to cellular proliferation, metabolism and epithelial mesenchymal transition. Positive correlation is displayed in blue colour and negative correlation in red colour. Colour intensity is proportional to the correlation coefficient
IDH2 protein expression and patient outcome
Survival analysis in DCIS revealed that higher IDH2 expression was correlated with shorter LRFI for all recurrences (HR = 2.4, 95% CI 1.3–4.5, p = 0.005, Fig. 3a) and a trend for association with shorter LRFI for invasive recurrences (HR = 1.9, 95% CI 0.9–4.4, p = 0.07 Fig. 3b) in patients treated solely with BCS without adjuvant radiotherapy. Multivariate survival analysis revealed that a high expression of IDH2 was an independent poor prognostic factor for DCIS recurrence (HR 2.0; 95% CI 1.1–3.9; p = 0.034) regardless of the other variables including patient age at diagnosis, DCIS size, nuclear grade, presence of comedo necrosis and surgical margin width (Table 3).
Comparable results were obtained in IBC where high IDH2 protein expression was associated with shorter BCSS (HR 1.6; 95% CI 1.2–2.2; p = 0.003; Fig. 4). High expression of IDH2 however was not associated with the distant metastasis (HR 1.1; 95% CI 0.9–1.4; p = 0.310; Supplementary Fig. 2). When the cohort was split into the intrinsic molecular subtypes, high expression of IDH2 protein in luminal B-like was associated with poor outcome in LVI positive tumours (HR 2.1; 95% CI 1.0–4.5; p = 0.044; Supplementary Fig. 3a). HER2 positivity was in borderline association with overexpression of IDH2 in LVI positive status (p = 0.062; Supplementary Fig. 3b), but not with TN subtype defined as ER−, PR−, HER2− (p = 0.221; Supplementary Fig. 3c).
In multivariate Cox regression analysis, high IDH2 protein expression was an independent predictor of shorter BCSS (HR 1.4; 95% CI 1.1–1.9; p = 0.042) regardless of the tumour grade, lymph node stage and nodal status (Table 4).
IDH2 mRNA expression
High IDH2 mRNA expression was observed in 50% of IBC in the METABRIC and TCGA datasets, respectively. There was a significant positive linear correlation between IDH2 protein expression and IDH2 mRNA expression (r = 0.240, p = 0.002) when tested in the Nottingham subset of the METABRIC cases (n = 288). Similar to IDH2 protein, in the METABRIC and TCGA datasets, overexpression of IDH2 mRNA was significantly associated with LVI-positivity (all; p = 0.001). In both datasets, high expression of IDH2 mRNA was associated with high histological grade and hormonal receptor negativity (both; p < 0.0001). Moreover, in the METABRIC cohort, high IDH2 mRNA levels were associated with large tumour size (p = 0.038), axillary lymph node positivity and HER2 positivity (all; p < 0.0001); (Table 5). In the METABRIC cohort, BCSS of patients with high IDH2 mRNA expression was significantly shorter than those with low expression (HR 1.38; 95% CI 1.2–1.6; p < 0.0001) (Fig. 5a), but not in TCGA dataset (HR = 1.0; 95% CI 0.7–1.5; p = 0.916); (Fig. 5b). External validation using the Gene Miner datasets (n = 4039) of IDH2 further supported that high expression of IDH2 mRNA was positively associated with shorter patient outcome (HR 1.26; 95% CI 1.1–1.4; p = 0.0002), Supplementary Fig. 4.
Discussion
Breast cancer represents a group of heterogeneous diseases that vary in their morphological, molecular, and clinical behaviour. This heterogeneity poses challenges in precise understanding of the biology of BC, and hence to define a personalised therapy approach [31].
Despite the breakthrough of the genetic and molecular analysis, the mechanisms underlying progression of breast in situ lesions into invasive disease, and those involved in distant metastasis and LVI are still to be defined. Wild-type IDH2 was previously described as a key factor in DCIS progression into invasive disease [9, 10], and controlling LVI [8]. However, the prognostic significance of IDH2 in breast cancer has not been described before. To the best of our knowledge, this is the first study addressing the role of wild-type IDH2 protein in BC using IHC and well annotated cohort of patients.
The current study included large cohorts of pre-invasive and IBC to assess the transcriptomic and proteomic expression of IDH2 expression and its correlation with the clinicopathological parameters and patients’ outcome. Our analysis of IDH2 expression in DCIS supported our hypothesis that this protein would be associated with features of high-risk DCIS. In addition, the poor prognostic significance of higher IDH2 expression was independent from other clinical and morphological features, and with a trend of association towards invasive recurrence and progression.
Similarly, results in IBC showed that a high IDH2 expression was associated with criteria of aggressive behaviour including LVI, larger tumour size, higher grade and poor NPI. This supports the results of previous studies which demonstrated that IDH2 is significantly associated with LVI [32]. High IDH2 expression was also an independent predictor of shorter BCSS either in proteomic or transcriptomic datasets. In METABRIC and bc-Exminer, datasets support the poor prognostic significance of high IDH2 expression.
Cellular energy produced by the TCA cycle is upregulated in highly proliferative and metastasised cancer cells. Energy metabolism and biosynthetic intermediates such as Alpha-ketoglutarate (αKG) produced through TCA from isocitrate by IDH2 is essential in tumour progression and metastasis. Wild-type IDH2 can reduce αKG and increase 2-HG production, in turn disrupting normal epigenetic regulation of transcription [15]. For example, in hypoxic condition in BC, IDH2 carboxylates α-KG from glutamine to citrate and elevates 2-HG levels, acting as an oncometabolite [33]. Moreover, when IDH2 reduces αKG, it can have an oncogenic impact on cellular differentiation [17]. αKG may have a role in the epithelial–mesenchymal transition (EMT), which is a key step during metastasis. It has been shown that blocking of αKG inhibits cellular transformation and cancerous cell invasion through transamination or reverse TCA cycle [12, 34]. Our data shows that IDH2 is associated with EMT proteins and factors including in cellular metabolism and energy production, which support its role in disease progression and metastasis. This study showed that high IDH2 protein was significantly associated with EGFR which can regulate epithelial mesenchymal transition (EMT), migration and invasion [35]. Moreover, this study also demonstrates that high N-cadherin is associated with high IDH2 protein. Previous studies elucidate that N-cadherin has a role in motility, invasion and metastasis by increasing MMP-9 production allowing tumour cells to penetrate the matrix barriers and to adhere to the endothelium. This may also increase the chance of tumour cells to enter and exit the vasculature [36]. Additionally, it has been reported that N-cadherin can induce EMT which plays a fundamental role in the invasion and metastasis of cancer cells [37]. Thus, the observed association between the high levels of IDH2 and N-cadherin explain the role of IDH2 in LVI as N-cadherin can promote two properties of the metastasised cell i.e. adhesion and invasion. IDH2 was also associated with TWIST2 EMT marker. EMT has a highly important role in cancer invasion and metastasis. In BC and lymph node metastases, Twist2 is overexpressed as it results in morphological transformation, downregulation of epithelial markers and upregulation of mesenchymal markers [38]. Thus, Interaction with EMT markers explains the role of IDH2 in DCIS progression into invasive disease and invasion of BC cells through the lymphovascular channels. EMT is a known mechanism attained by the breast carcinoma cells for basement membrane invasion and LVI [39]. Further mechanistic studies are highly warranted to understand the underlying molecular mechanisms.
Additionally, one of the major hallmarks of cancer is reprogramming energy metabolism [40]. High rates of glucose and glutamine are taken up by cancer cells to produce NADPH to survive and grow followed by decreasing the level of intracellular αKG [41]. Many studies provide evidence that oncogenes can alter cancer cell metabolism including glutamine transporters which were associated with high IDH2 in this study [26]. This may imply that IDH2 could potentially re-programme the metabolic pathways in BC and support tumour cells to survive and indicate its prognostic significance in BC. Our observations showed that high IDH2 was associated with high grade DCIS and IBC with a highly proliferative index. High expression of IDH2 was associated with EGFR, CK5 and CK14 which are connected to the highly proliferative basal phenotype [42, 43]. In this study, IDH2 was also associated with high Ki67 expression reflecting increased cellular proliferation [44]. It has been reported that IDH2 has a critical role in cell proliferation through the alteration of NADP levels. Beside the role of NADP as antioxidants, it has been reported that NADP has an important role in cell death. It links cell survival with biological properties such as energy metabolism and oxidative stress, which are factors that determine cell death [14]. Primary tumour cells must proliferate and invade the adjacent tissue to establish an invasion and metastasis cascades, which consists of many steps including basement membrane degradation and LVI. Proliferation lasts until the invasion of blood vessels or lymphatic channels occur. Tumour cells at this stage evade apoptosis and immune responses [45]. Thus, IDH2 may have an important role in cell proliferation, which is a perquisite step of the invasion and metastatic process, which can lead to the development of invasive cancer from a pre-invasive lesion, and for development of LVI and hence distant metastasis.
Furthermore, IDH2 mRNA and protein were also highly expressed in TNBC and HER2+ either in IBC or DCIS, in concordance with previous studies [5]. HER2+ tumours were more likely to have IDH2 protein which is perhaps unsurprising as high grade progressing DCIS and LVI are significantly associated with HER2 positivity [1]. It also has been shown that tumour microenvironment plays a major role in the HER2 signalling pathway, invasion and the development of LVI, which is a crucial step in metastasis [46]. IDH2 high expression might enhance HER2 signalling pathways that can have effect on the tumour microenvironment to support the growth of the tumour cell, stimulating invasion, LVI and metastasis in BC. The molecular subtypes differ from each other in their expression of claudin [47]. For example, low expression of the claudin tight junction protein and the high expression of proteins involved in EMT and cancer cell invasiveness were reported in TN compared to other subtypes [48]. This could explain the difference in the significance between these subtypes.
This study reveals that IDH2 is associated with poor prognostic characteristics and short-term survival outcomes in BC including higher local recurrence rate after diagnosis of DCIS or poor survival rate in IBC. Furthermore, a positive association of IDH2 and elevated levels of basal cytokeratin confers a poor prognosis. Basal cytokeratins are strongly associated with high histological grade, negative hormone status and worse patient outcome [39, 49]. Among subgroups, overexpression of IDH2 protein appears to play a particularly significant role in luminal B subtype which is in concordance with a recent study [50].
In conclusion high expression of IDH2 is an independent poor prognostic factor in BC. High expression of IDH2 may have a key role in BC progression from DCIS to IBC and in the development of LVI and metastasis. Further functional assessment of IDH2 in BC is warranted to detect its underlying mechanistic roles and its therapeutic potential.
Data availability
The authors confirm the data that has been used in this work is available on reasonable request.
References
Rakha EA, Martin S, Lee AHS, Morgan D, Pharoah PDP, Hodi Z, MacMillan D, Ellis IO (2012) The prognostic significance of lymphovascular invasion in invasive breast carcinoma. Cancer 118(15):3670–3680. https://doi.org/10.1002/cncr.26711
Zhang S, Zhang D, Yi S, Gong M, Lu C, Cai Y, Tang X, Zou L (2016) The relationship of lymphatic vessel density, lymphovascular invasion, and lymph node metastasis in breast cancer: a systematic review and meta-analysis. Oncotarget 8(2):2863–2873. https://doi.org/10.18632/oncotarget.13752
Liu YL, Saraf A, Lee SM, Zhong X, Hibshoosh H, Kalinsky K, Connolly EP (2016) Lymphovascular invasion is an independent predictor of survival in breast cancer after neoadjuvant chemotherapy. Breast Cancer Res Treat 157(3):555–564. https://doi.org/10.1007/s10549-016-3837-5
Mohammed RA, Martin SG, Mahmmod AM, Macmillan RD, Green AR, Paish EC, Ellis IO (2011) Objective assessment of lymphatic and blood vascular invasion in lymph node-negative breast carcinoma: findings from a large case series with long-term follow-up. J Pathol 223(3):358–365. https://doi.org/10.1002/path.2810
Aleskandarany M, Sonbul S, Mukherjee A, Rakha E (2015) Molecular mechanisms underlying lymphovascular invasion in invasive breast cancer. Pathobiology. https://doi.org/10.1159/000433583
Curtis C, Shah SP, Chin S-F, Turashvili G, Rueda OM, Dunning MJ, Speed D, Lynch AG, Samarajiwa S, Yuan Y, Gräf S, Ha G, Haffari G, Bashashati A, Russell R, McKinney S, Group M, Caldas C, Aparicio S, Curtis† C, Shah SP, Caldas C, Aparicio S, Brenton JD, Ellis I, Huntsman D, Pinder S, Purushotham A, Murphy L, Caldas C, Aparicio S, Caldas C, Bardwell H, Chin S-F, Curtis C, Ding Z, Gräf S, Jones L, Liu B, Lynch AG, Papatheodorou I, Sammut SJ, Wishart G, Aparicio S, Chia S, Gelmon K, Huntsman D, McKinney S, Speers C, Turashvili G, Watson P, Ellis I, Blamey R, Green A, Macmillan D, Rakha E, Purushotham A, Gillett C, Grigoriadis A, Pinder S, de Rinaldis E, Tutt A, Murphy L, Parisien M, Troup S, Caldas C, Chin S-F, Chan D, Fielding C, Maia A-T, McGuire S, Osborne M, Sayalero SM, Spiteri I, Hadfield J, Aparicio S, Turashvili G, Bell L, Chow K, Gale N, Huntsman D, Kovalik M, Ng Y, Prentice L, Caldas C, Tavaré S, Curtis C, Dunning MJ, Gräf S, Lynch AG, Rueda OM, Russell R, Samarajiwa S, Speed D, Markowetz F, Yuan Y, Brenton JD, Aparicio S, Shah SP, Bashashati A, Ha G, Haffari G, McKinney S, Langerød A, Green A, Provenzano E, Wishart G, Pinder S, Watson P, Markowetz F, Murphy L, Ellis I, Purushotham A, Børresen-Dale A-L, Brenton JD, Tavaré S, Caldas C, Aparicio S (2012) The genomic and transcriptomic architecture of 2,000 breast tumours reveals novel subgroups. Nature 486:346. https://doi.org/10.1038/nature10983. https://www.nature.com/articles/nature10983#supplementary-information
Kadota K, Nakai Y, Shimizu K (2008) A weighted average difference method for detecting differentially expressed genes from microarray data. Algorithms Mol Biol 3:8–8. https://doi.org/10.1186/1748-7188-3-8
Kurozumi S, Joseph C, Sonbul S, Alsaeed S, Kariri Y, Aljohani A, Raafat S, Alsaleem M, Ogden A, Johnston SJ, Aleskandarany MA, Fujii T, Shirabe K, Caldas C, Ashankyty I, Dalton L, Ellis IO, Desmedt C, Green AR, Mongan NP, Rakha EA (2019) A key genomic subtype associated with lymphovascular invasion in invasive breast cancer. Br J Cancer 120(12):1129–1136. https://doi.org/10.1038/s41416-019-0486-6
Gorringe KL, Hunter SM, Pang JM, Opeskin K, Hill P, Rowley SM, Choong DY, Thompson ER, Dobrovic A, Fox SB, Mann GB, Campbell IG (2015) Copy number analysis of ductal carcinoma in situ with and without recurrence. Mod Pathol 28(9):1174–1184. https://doi.org/10.1038/modpathol.2015.75
Hannemann J, Velds A, Halfwerk JB, Kreike B, Peterse JL, van de Vijver MJ (2006) Classification of ductal carcinoma in situ by gene expression profiling. Breast Cancer Res 8(5):R61. https://doi.org/10.1186/bcr1613
Smolkova K, Jezek P (2012) The role of mitochondrial NADPH-dependent isocitrate dehydrogenase in cancer cells. Int J Cell Biol 2012:273947. https://doi.org/10.1155/2012/273947
Teoh ST, Lunt SY (2018) Metabolism in cancer metastasis: bioenergetics, biosynthesis, and beyond. Wiley Interdiscip Rev Syst Biol Med. https://doi.org/10.1002/wsbm.1406
Bergaggio E, Piva R (2019) Wild-type IDH enzymes as actionable targets for cancer therapy. Cancers 11(4):563
Lv Q, Xing S, Li Z, Li J, Gong P, Xu X, Chang LE, Jin X, Gao F, Li W, Zhang G, Yang J, Zhang X (2012) Altered expression levels of IDH2 are involved in the development of colon cancer. Exp Ther Med 4(5):801–806. https://doi.org/10.3892/etm.2012.676
Raineri S, Mellor J (2018) IDH1: linking metabolism and epigenetics. Front Genet 9:493. https://doi.org/10.3389/fgene.2018.00493
Chiang S, Weigelt B, Wen H-C, Pareja F, Raghavendra A, Martelotto LG, Burke KA, Basili T, Li A, Geyer FC, Piscuoglio S, Ng CKY, Jungbluth AA, Balss J, Pusch S, Baker GM, Cole KS, von Deimling A, Batten JM, Marotti JD, Soh H-C, McCalip BL, Serrano J, Lim RS, Siziopikou KP, Lu S, Liu X, Hammour T, Brogi E, Snuderl M, Iafrate AJ, Reis-Filho JS, Schnitt SJ (2016) IDH2 mutations define a unique subtype of breast cancer with altered nuclear polarity. Can Res 76(24):7118. https://doi.org/10.1158/0008-5472.CAN-16-0298
Li J, He Y, Tan Z, Lu J, Li L, Song X, Shi F, Xie L, You S, Luo X, Li N, Li Y, Liu X, Tang M, Weng X, Yi W, Fan J, Zhou J, Qiang G, Qiu S, Wu W, Bode AM, Cao Y (2018) Wild-type IDH2 promotes the Warburg effect and tumor growth through HIF1alpha in lung cancer. Theranostics 8(15):4050–4061. https://doi.org/10.7150/thno.21524
Miligy IM, Gorringe KL, Toss MS, Al-Kawaz AA, Simpson P, Diez-Rodriguez M, Nolan CC, Ellis IO, Green AR, Rakha EA (2018) Thioredoxin-interacting protein is an independent risk stratifier for breast ductal carcinoma in situ. Mod Pathol 31(12):1807–1815. https://doi.org/10.1038/s41379-018-0086-7
Sonbul SN, Aleskandarany MA, Kurozumi S, Joseph C, Toss MS, Diez-Rodriguez M, Nolan CC, Mukherjee A, Martin S, Caldas C, Ellis IO, Green AR, Rakha EA (2018) Saccharomyces cerevisiae-like 1 (SEC14L1) is a prognostic factor in breast cancer associated with lymphovascular invasion. Mod Pathol 31(11):1675–1682. https://doi.org/10.1038/s41379-018-0092-9
Rakha EA, Pinder SE, Bartlett JMS, Ibrahim M, Starczynski J, Carder PJ, Provenzano E, Hanby A, Hales S, Lee AHS, Ellis IO, National Coordinating Committee for Breast P (2015) Updated UK recommendations for HER2 assessment in breast cancer. J Clin Pathol 68(2):93–99. https://doi.org/10.1136/jclinpath-2014-202571
Rakha EA, Agarwal D, Green AR, Ashankyty I, Ellis IO, Ball G, Alaskandarany MA (2017) Prognostic stratification of oestrogen receptor-positive HER2-negative lymph node-negative class of breast cancer. Histopathology 70(4):622–631. https://doi.org/10.1111/his.13108
Green AR, Aleskandarany MA, Agarwal D, Elsheikh S, Nolan CC, Diez-Rodriguez M, Macmillan RD, Ball GR, Caldas C, Madhusudan S, Ellis IO, Rakha EA (2016) MYC functions are specific in biological subtypes of breast cancer and confers resistance to endocrine therapy in luminal tumours. Br J Cancer 114(8):917–928. https://doi.org/10.1038/bjc.2016.46
Hammond MEH, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, Fitzgibbons PL, Francis G, Goldstein NS, Hayes M, Hicks DG, Lester S, Love R, Mangu PB, McShane L, Miller K, Osborne CK, Paik S, Perlmutter J, Rhodes A, Sasano H, Schwartz JN, Sweep FCG, Taube S, Torlakovic EE, Valenstein P, Viale G, Visscher D, Wheeler T, Williams RB, Wittliff JL, Wolff AC (2010) American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. Arch Pathol Lab Med 134(6):907–922. https://doi.org/10.1043/1543-2165-134.6.907
Abd El-Rehim DM, Ball G, Pinder SE, Rakha E, Paish C, Robertson JFR, Macmillan D, Blamey RW, Ellis IO (2005) High-throughput protein expression analysis using tissue microarray technology of a large well-characterised series identifies biologically distinct classes of breast cancer confirming recent cDNA expression analyses. Int J Cancer 116(3):340–350. https://doi.org/10.1002/ijc.21004
Aleskandarany MA, Rakha EA, Ahmed MA, Powe DG, Ellis IO, Green AR (2011) Clinicopathologic and molecular significance of phospho-Akt expression in early invasive breast cancer. Breast Cancer Res Treat 127(2):407–416. https://doi.org/10.1007/s10549-010-1012-y
El Ansari R, Craze ML, Miligy I, Diez-Rodriguez M, Nolan CC, Ellis IO, Rakha EA, Green AR (2018) The amino acid transporter SLC7A5 confers a poor prognosis in the highly proliferative breast cancer subtypes and is a key therapeutic target in luminal B tumours. Breast Cancer Res 20(1):21. https://doi.org/10.1186/s13058-018-0946-6
El Ansari R, Craze ML, Diez-Rodriguez M, Nolan CC, Ellis IO, Rakha EA, Green AR (2018) The multifunctional solute carrier 3A2 (SLC3A2) confers a poor prognosis in the highly proliferative breast cancer subtypes. Br J Cancer 118(8):1115–1122. https://doi.org/10.1038/s41416-018-0038-5
Craze ML, Cheung H, Jewa N, Coimbra NDM, Soria D, El-Ansari R, Aleskandarany MA, Wai Cheng K, Diez-Rodriguez M, Nolan CC, Ellis IO, Rakha EA, Green AR (2017) MYC regulation of glutamine–proline regulatory axis is key in luminal B breast cancer. Br J Cancer 118:258. https://doi.org/10.1038/bjc.2017.387
Matkowski R, Gisterek I, Halon A, Lacko A, Szewczyk K, Staszek U, Pudelko M, Szynglarewicz B, Szelachowska J, Zolnierek A, Kornafel J (2009) The prognostic role of tumor-infiltrating CD4 and CD8 T lymphocytes in breast cancer. Anticancer Res 29(7):2445–2451
Ciriello G, Gatza ML, Beck AH, Wilkerson MD, Rhie SK, Pastore A, Zhang H, McLellan M, Yau C, Kandoth C, Bowlby R, Shen H, Hayat S, Fieldhouse R, Lester SC, Tse GM, Factor RE, Collins LC, Allison KH, Chen YY, Jensen K, Johnson NB, Oesterreich S, Mills GB, Cherniack AD, Robertson G, Benz C, Sander C, Laird PW, Hoadley KA, King TA, Perou CM (2015) Comprehensive molecular portraits of invasive lobular breast cancer. Cell 163(2):506–519. https://doi.org/10.1016/j.cell.2015.09.033
Dawson SJ, Rueda OM, Aparicio S, Caldas C (2013) A new genome-driven integrated classification of breast cancer and its implications. EMBO J 32(5):617–628. https://doi.org/10.1038/emboj.2013.19
Kurozumi S, Joseph C, Sonbul S, Alsaeed S, Kariri Y, Aljohani A, Raafat S, Alsaleem M, Ogden A, Johnston SJ, Aleskandarany MA, Fujii T, Shirabe K, Caldas C, Ashankyty I, Dalton L, Ellis IO, Desmedt C, Green AR, Mongan NP, Rakha EA (2019) A key genomic subtype associated with lymphovascular invasion in invasive breast cancer. Br J Cancer. https://doi.org/10.1038/s41416-019-0486-6
Terunuma A, Putluri N, Mishra P, Mathé EA, Dorsey TH, Yi M, Wallace TA, Issaq HJ, Zhou M, Killian JK, Stevenson HS, Karoly ED, Chan K, Samanta S, Prieto D, Hsu TYT, Kurley SJ, Putluri V, Sonavane R, Edelman DC, Wulff J, Starks AM, Yang Y, Kittles RA, Yfantis HG, Lee DH, Ioffe OB, Schiff R, Stephens RM, Meltzer PS, Veenstra TD, Westbrook TF, Sreekumar A, Ambs S (2014) MYC-driven accumulation of 2-hydroxyglutarate is associated with breast cancer prognosis. J Clin Investig 124(1):398–412. https://doi.org/10.1172/JCI71180
Weinberg F, Hamanaka R, Wheaton WW, Weinberg S, Joseph J, Lopez M, Kalyanaraman B, Mutlu GM, Budinger GRS, Chandel NS (2010) Mitochondrial metabolism and ROS generation are essential for Kras-mediated tumorigenicity. Proc Natl Acad Sci USA 107(19):8788–8793. https://doi.org/10.1073/pnas.1003428107
Masuda H, Zhang D, Bartholomeusz C, Doihara H, Hortobagyi GN, Ueno NT (2012) Role of epidermal growth factor receptor in breast cancer. Breast Cancer Res Treat 136(2):331–345. https://doi.org/10.1007/s10549-012-2289-9
Hazan RB, Phillips GR, Qiao RF, Norton L, Aaronson SA (2000) Exogenous expression of N-cadherin in breast cancer cells induces cell migration, invasion, and metastasis. J Cell Biol 148(4):779–790. https://doi.org/10.1083/jcb.148.4.779
Nakajima S, Doi R, Toyoda E, Tsuji S, Wada M, Koizumi M, Tulachan SS, Ito D, Kami K, Mori T, Kawaguchi Y, Fujimoto K, Hosotani R, Imamura M (2004) N-Cadherin expression and epithelial-mesenchymal transition in pancreatic carcinoma. Clin Cancer Res 10(12):4125. https://doi.org/10.1158/1078-0432.CCR-0578-03
Fang X, Cai Y, Liu J, Wang Z, Wu Q, Zhang Z, Yang CJ, Yuan L, Ouyang G (2011) Twist2 contributes to breast cancer progression by promoting an epithelial–mesenchymal transition and cancer stem-like cell self-renewal. Oncogene 30:4707. https://doi.org/10.1038/onc.2011.181
Heerboth S, Housman G, Leary M, Longacre M, Byler S, Lapinska K, Willbanks A, Sarkar S (2015) EMT and tumor metastasis. Clin Transl Med 4(1):6. https://doi.org/10.1186/s40169-015-0048-3
Ward PS, Thompson CB (2012) Metabolic reprogramming: a cancer hallmark even warburg did not anticipate. Cancer Cell 21(3):297–308. https://doi.org/10.1016/j.ccr.2012.02.014
Cantor JR, Sabatini DM (2012) Cancer cell metabolism: one hallmark, many faces. Cancer Discov 2(10):881–898. https://doi.org/10.1158/2159-8290.Cd-12-0345
Laakso M, Loman N, Borg Å, Isola J (2005) Cytokeratin 5/14-positive breast cancer: true basal phenotype confined to BRCA1 tumors. Mod Pathol 18(10):1321–1328. https://doi.org/10.1038/modpathol.3800456
Badve S, Dabbs DJ, Schnitt SJ, Baehner FL, Decker T, Eusebi V, Fox SB, Ichihara S, Jacquemier J, Lakhani SR, Palacios J, Rakha EA, Richardson AL, Schmitt FC, Tan P-H, Tse GM, Weigelt B, Ellis IO, Reis-Filho JS (2010) Basal-like and triple-negative breast cancers: a critical review with an emphasis on the implications for pathologists and oncologists. Mod Pathol 24:157. https://doi.org/10.1038/modpathol.2010.200
Inwald EC, Klinkhammer-Schalke M, Hofstädter F, Zeman F, Koller M, Gerstenhauer M, Ortmann O (2013) Ki-67 is a prognostic parameter in breast cancer patients: results of a large population-based cohort of a cancer registry. Breast Cancer Res Treat 139(2):539–552. https://doi.org/10.1007/s10549-013-2560-8
Hunter KW, Crawford NPS, Alsarraj J (2008) Mechanisms of metastasis. Breast Cancer Res 10(Suppl 1):S2–S2. https://doi.org/10.1186/bcr1988
Banin-Hirata BK, de Oliveira CEC, Losi-Guembarovski R, Ozawa PMM, Vitiello GAF, de Almeida FC, Derossi DR, André ND, Watanabe MAE (2018) The prognostic value of regulatory T cells infiltration in HER2-enriched breast cancer microenvironment. Int Rev Immunol 37(3):144–150. https://doi.org/10.1080/08830185.2017.1401620
Dias K, Dvorkin-Gheva A, Hallett RM, Wu Y, Hassell J, Pond GR, Levine M, Whelan T, Bane AL (2017) Claudin-low breast cancer; clinical & pathological characteristics. PLoS ONE 12(1):e0168669–e0168669. https://doi.org/10.1371/journal.pone.0168669
Sabatier R, Finetti P, Guille A, Adelaide J, Chaffanet M, Viens P, Birnbaum D, Bertucci F (2014) Claudin-low breast cancers: clinical, pathological, molecular and prognostic characterization. Mol Cancer 13:228. https://doi.org/10.1186/1476-4598-13-228
Kordek R, Potemski P, Kusinska R, Pluciennik E, Bednarek A (2010) Basal keratin expression in breast cancer by quantification of mRNA and by immunohistochemistry. J Exp Clin Cancer Res 29:39. https://doi.org/10.1186/1756-9966-29-39
Liao G-S, Hsu H-M, Chu C-H, Hong Z-J, Fu C-Y, Chou Y-C, Golshan M, Dai M-S, Chen T-W, De-Chian C, Tsai W-C, Pan C-W, Hsu K-F, Kao E-N, Hsu Y-C, Chang T-H, Yu J-C (2018) Prognostic role of lymphovascular invasion and lymph node status among breast cancer subtypes. J Med Sci 38(2):54–61. https://doi.org/10.4103/jmedsci.jmedsci_105_17
Acknowledgements
Abrar Aljohani is supported and funded by Taif University, Kingdom of Saudi Arabia. We thank the Nottingham Health Science Biobank and Breast Cancer Now Tissue Bank for the provision of tissue samples.
Funding
This research was supported and funded by Taif University, Kingdom of Saudi Arabia.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
All authors declare that they have no conflict of interest.
Research involving human participants and/or animals
This article does not contain any studies with human participants or animals performed by any of the authors.
Informed consent
This work obtained ethics approval to use the human tissue samples by the North West – Greater Manchester Central Research Ethics Committee under the title Nottingham Health Science Biobank (NHSB), reference number 15/NW/0685. Informed consent was obtained from all individuals prior to surgery to use their tissue materials in research. This study was performed according to the REMARK guidelines for tumour prognostic studies.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Aljohani, A.I., Toss, M.S., Kurozumi, S. et al. The prognostic significance of wild-type isocitrate dehydrogenase 2 (IDH2) in breast cancer. Breast Cancer Res Treat 179, 79–90 (2020). https://doi.org/10.1007/s10549-019-05459-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10549-019-05459-7