Abstract
DNA repair is an essential agent in cancer development, progression, prognosis, and response to therapy. We have adapted a cellular repair assay based on the formamidopyrimidine DNA glycosylase (Fpg)-modified comet assay to assess DNA repair kinetics. The removal of oxidized nucleobases over time (0–480 min) was analyzed in peripheral blood mononuclear cells (PBMCs) and 8 cell lines. DNA damage was induced by exposure to either Ro19-8022 plus visible light or potassium bromate (KBrO3). The initial amount of damage induced by Ro 19–8022 plus light varied between cell lines, and this was apparently associated with the rate of repair. However, the amount of DNA damage induced by KBrO3 varied less between cell types, so we used this agent to study the kinetics of DNA repair. We found an early phase of ca. 60 min with fast removal of Fpg-sensitive sites, followed by slower removal over the following 7 h. In conclusion, adjusting the initial damage at T0 to an equal level can be achieved by the use of KBrO3, which allows for accurate analysis of subsequent cellular DNA repair kinetics in the first hour after exposure.
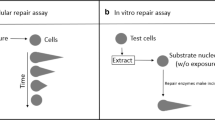
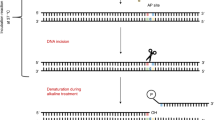
Graphical Abstract

Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Humans are constantly exposed to environmental and occupational hazards that can cause DNA damage. This damaged DNA is removed by DNA repair mechanisms, thus maintaining genome integrity and preventing mutations. Therefore, DNA repair capacity may be a useful biomarker when studying cancer development, progression, prognosis, and response to clinical therapy. To assess inter-individual differences in DNA repair activity, the comet assay (single-cell gel electrophoresis) is a suitable, robust, and sensitive method. Ostling and Johanson as well as Singh et al., in the original papers describing the comet assay for DNA breaks, also applied it to DNA repair, studying the decrease of ionizing radiation-induced strand breaks over time (Ostling and Johanson 1984; Singh et al. 1988). A similar approach can be used for other types of DNA damage that can be detected by the comet assay. The standard comet assay was modified to reveal oxidized bases by incubating the nucleoids after lysis with the bacterial enzyme endonuclease III (Endo III) to convert oxidized pyrimidines to breaks(Collins et al. 1993), or with formamidopyrimidine DNA glycosylase (Fpg) to detect oxidized purines (Dusinska and Collins 1996). Using H2O2 to induce DNA damage in HeLa cells and human lymphocytes, removal of strand breaks and oxidized bases was followed over time; differences in DNA repair kinetics were found between the human cell line HeLa and freshly isolated human peripheral blood mononuclear cells (PBMCs) (Collins et al. 1995).
The comet assay has already been applied in clinical settings, for example, in studying radiotherapy susceptibility and individualized health risk assessment. Palyvoda et al. (2003) estimated the cellular repair rates of healthy donors and of patients with squamous cell carcinoma of the head and neck (SCCHN) before radiotherapy, by measuring residual strand breaks at six time intervals from 0 min to 3 h after treating lymphocytes with γ-irridation (3 Gy dose) on ice, in order to predict treatment outcome. Ocolotobiche et al. (2021) used this assay to identify patients at higher risk of radiotherapy side effects, although they found no significant difference between patients and healthy volunteers. Valdiglesias et al. (2020) evaluated and compared the suitability of different blood preparations (frozen PBMCs, fresh blood, and frozen blood) isolated from a healthy population and treated with different challenging agents (bleomycin, methyl methane sulfonate (MMS), and UV light) in the cellular repair assay to assess the response to DNA damage. Various DNA-damaging agents can be used to induce different types of lesions in DNA for studying DNA repair activity with the comet assay, namely, photosensitizer Ro 19–8022 plus visible light, potassium bromate (KBrO3), MMS, UVC, and benzo(a)pyrene diol epoxide (BPDE). The DNA of cells treated with Ro 19–8022 plus light or KBrO3 contains oxidized purines, mainly 8-oxoguanine (8-oxoG), and MMS induces DNA alkylations, substrates for base excision repair (BER). UVC and BPDE induce pyrimidine dimers and bulky adducts, respectively, and are repaired by nucleotide excision repair (NER) (Collins et al. 2002; Speit et al. 2004; Ballmaier and Epe 2006; Luan et al. 2007; Camenisch and Naegeli 2009; Hašplová et al. 2012; Goto et al. 2015). In the current study, we focused on cellular repair after a challenge with Ro 19–8022 plus light, or KBrO3. Until now, there have been few studies measuring DNA repair after KBrO3 exposure (Kumar et al. 2022; Parlanti et al. 2012; Platel et al. 2011); in these papers, KBrO3 was used to induce 8-oxoG in DNA repair gene-deficient or silenced cell lines. Although the mechanism of KBrO3-induced DNA damage is not yet fully understood, it seems to provide a useful alternative approach to studying BER of oxidized DNA.
The cellular repair assay can accurately monitor repair kinetics, but some aspects of the assay need to be standardized before it can be used in clinical settings or human biomonitoring, For instance, it is not known whether the level of initial DNA damage affects the rate of repair. It is important to distinguish between the prevention of damage by antioxidant defenses and the removal of damage by DNA repair (Collins and Azqueta 2012; Collins 2014). Our aim in this paper was to identify factors affecting the rate of repair of oxidized bases and to propose a reliable approach to the comparative study of DNA repair rates in different cell lines that could also be of use in studying inter-individual variations in DNA repair in humans.
Materials and methods
Chemicals
RMPI-1640 medium, fetal bovine serum, 100 U/ml penicillin, 100 µg/ml streptomycin, trypsin–EDTA 1 × , SYBR Gold, and chemicals and reagents used for the comet assay were purchased from Sigma Aldrich (Heidelberg, Germany); McCoy’s 5 a Medium was purchased from Cytiva AS; Eagle’s Minimum Essential Medium and Dulbecco’s Modified Eagle Medium (DMEM) were purchased from Biowest AS. Trypan blue solution (0.4%) was purchased from Invitrogen Company (Thermo Fisher Scientific). Lymphoprep was purchased from Fresenius Kabi Norge As.
Cells
The following cell lines were used: HCT-116 (human colorectal carcinoma cell line), LNCaP (human prostate adenocarcinoma cell line), TK-6 (human lymphoblastoid cell line), LLC-pk1 (porcine kidney cell), V79-4 (Chinese hamster lung fibroblast cell line), MCF-7 (human breast cancer cell line), Caco2 (human colorectal adenocarcinoma cell line), HeLa cells (human cervical cell line), and HepG2 (human hepatocellular carcinoma cell line). All cell lines were grown in the appropriate medium, according to the protocol provided by the ATCC; they were incubated at 37 °C in a humidified incubator with a 5% CO2 atmosphere.
PBMCs were collected from 5 non-smoking, healthy volunteers (25–35 years) under the approval of the Regional Ethical Committee Southeast Norway. The venous blood (sampled in a vacutainer tube with EDTA as anticoagulant) was diluted in a 15 ml plastic tube at a volume ratio of 1:1 with sterile PBS, underlayed with the same volume of Lymphoprep, and centrifuged at 250 × g for 30 min at 4 °C with the brake off. PBMCs were isolated from the interface between PBS and Lymphoprep, washed with PBS, centrifuged (250 × g, 5 min at 4 °C), and resuspended in 1 ml sterile PBS.
Exposure of cells to Ro 19–8022, or KBrO 3
Non-adherent cells (PBMCs and TK-6) were suspended in RMPI-1640 with 10% fetal bovine serum and placed into Petri dishes at a concentration of 2.5 × 105 cells/ml. Adherent cells were seeded into 24-well plates and allowed to grow to 70–85% confluence and were then detached with 1 × trypsin–EDTA, washed with PBS, centrifuged (250 × g, 5 min at 4 °C), and resuspended in 1 ml sterile PBS. Five hundred microliters of cell suspension was kept as control, and the remaining cells (in a Petri dish) were placed on ice, treated with 1 µM Ro 19–8022 (a gift from F. Hoffmann-La Roche), and irradiated with visible light (33 cm from a 500 W tungsten halogen source) for 5 min.
In later experiments, non-adherent cells were exposed to a range of concentrations of KBrO3 between 0 and 100 mM, in appropriate cell medium for 1 h at 37 °C. After treatment, cells were washed with PBS, centrifuged (250 × g, 5 min at 4 °C), and resuspended in 1 ml sterile PBS. Adherent cells were seeded into 24-well plates and allowed to grow to 70–85% confluence, exposed to the same range of concentrations of KBrO3 as suspension cells in appropriate cell medium for 1 h at 37 °C after treatment, centrifuged, and resuspended in 1 ml sterile PBS.
Cytotoxicity test
To assess the Ro 19–8022 plus light or KBrO3-induced cytotoxicity, Trypan blue tests were conducted in parallel with the comet assay before and after exposure. In all cases, viability was higher than 80% (Table S1).
Alkaline comet assay for strand breaks and alkali-labile sites
After treatment with Ro 19–8022 plus light or KBrO3, cell suspensions were mixed with 0.7% low melting point agarose (LMPA), and 50 µL was placed on a glass slide pre-coated with 1% normal melting agarose. Gels were set at 4 °C, and the embedded cells were lysed at 4 °C overnight (lysis buffer: 2.5 M NaCl, 100 mM Na2EDTA, 10 mM Tris base, pH = 10, and 1% Triton X-100 added just before use). Slides were then placed in an alkaline electrophoresis solution (0.3 M NaOH, 1 mM Na2EDTA, pH > 12) for 20 min at 4 °C for unwinding and electrophoresed in the same solution for 20 min at a voltage gradient of 0.8 V/cm across the platform in a horizontal electrophoresis chamber (Bio-Rad, Richmond, CA, USA). Finally, the slides were rinsed once with PBS (1 × , pH = 7.4), twice in distilled water, and left to dry. For scoring, slides were stained with 1 µM SYBR™ Gold at the recommended 10,000 × dilution for 30 min in the dark and then rinsed twice in distilled water.
The Comet IV semi-automated image analysis system was used to evaluate 50 comets per gel. The percentage of DNA in the tail (% tail DNA) was the descriptor used, and the median value of % tail DNA from 100 comets was used to measure DNA damage for each condition.
Fpg-modified comet assay
Fpg was produced by Norgenotech AS, Norway, and was the same enzyme (made in one batch) as used by the European Comet Assay Validation Group (ECVAG) (Møller et al. 2010). Aliquots were diluted tenfold with Fpg reaction buffer (40 mM HEPES, 0.1 M KCl, 0.5 mM Na2EDTA, 0.2 mg/mL BSA, pH = 8) with the addition of 10% glycerol and stored at − 80 °C. For each experiment, an aliquot was diluted with 30 ml of Fpg reaction buffer, reaching a final dilution of 60,000 times from the original crude preparation (0.5 µg/ml total protein). The slides were then placed on a plastic rack, 50 µl Fpg solution or the reaction buffer was added to each gel, and a 22 × 22 mm coverslip was placed on top. Then, the rack was transferred to a pre-heated moist box and placed in an incubator for 1 h at 37 °C. After incubation, the slides were placed at 4 °C in a cold room to stop the Fpg reaction. The coverslips were removed, and all slides were transferred to the electrophoresis tank; subsequent steps were as for the standard comet assay for strand breaks. Net Fpg-sensitive sites were estimated by subtracting % tail DNA with buffer incubation only from % tail DNA with Fpg incubation.
Cellular repair assay
After treatment with Ro 19–8022 plus light or KBrO3, suspension cells were placed in a T75 flask with cell-specific medium at 37 °C in a humidified incubator with a 5% CO2 atmosphere to allow for DNA repair. Cells were sampled at 0, 15, 30, 45, 60, and 120 min and collected by centrifugation (250 × g, 5 min at 4 °C). Adherent cells were detached by 1 × trypsin–EDTA at 0, 15, 30, 45, 60, and 120 min after exposure, and collected by centrifugation (250 × g, 5 min at 4 °C). (Additional time points were included for incubations with KBrO3, namely, 240 and 480 min.) Cells were resuspended in 1 ml sterile PBS mixed with 0.7% LMPA to prepare gels and analyzed with the comet assay as described.
H 2 O 2 resistance assay
To assess antioxidant status, cells were exposed to H2O2 (0 µM, 12.5 µM, 25 µM, 50 µM, 100 µM) on ice for 5 min and subsequently washed and resuspended in sterile PBS. Cell suspensions were mixed with 0.7% LMPA to prepare gels, which were then processed with the comet assay for strand breaks. This approach of measuring H2O2 resistance has shown meaningful variation among samples of lymphocytes from individuals, with lower resistance in smokers, and increased resistance after taking a vitamin C supplement (Panayiotidis and Collins 1997).
Data analysis
Statistical analysis was conducted using IBM SPSS Statistics 21.0 and Excel. The normality of data distribution of all parameters was tested by the Kolmogorov–Smirnov test, then using ANOVA with Dunnett’s post hoc test for differences between groups, and Student’s t-test was used for differences between interval 0–60 min and interval 60–480 min. In case the data did not fit a normal distribution, the Kruskal–Wallis test and Mann–Whitney U test were used. All statistical tests were conducted with the confidence level set at 95% (P = 0.05). To study the rate of DNA repair, the half-life of DNA damage was estimated assuming first-order kinetics using the formula: \({t}_{(\frac{1}{2})}=\left(-Ln\left(2\right)*t\right)/(Ln\left(\frac{{N}_{t}}{{N}_{0}}\right))\) (t = time; Nt = damage at time point t; N0 = damage at start of measurement) (Petrucci et al. 1997).
Results
Removal of DNA damage after treatment of cells with Ro 19–8022 plus light
Cell lines as well as PBMCs from healthy volunteers were treated with 1 µM Ro 19–8022 plus light to introduce DNA base oxidation damage, after which the cells were monitored for the removal of DNA damage (i.e., DNA repair) at several time intervals between 0 and 120 min (Fig. 1a). Removal of net Fpg-sensitive sites was seen in all cell lines except MCF-7 and HepG2. It is notable that the initial damage (Fpg-sensitive sites) varied widely between cell lines, ranging between 19.8% tail DNA in MCF-7 and 69.3% tail DNA in HCT-116. We reason that it is important to start with the same level of DNA damage to make a valid comparison between cells, because as shown in Fig. 2a, the DNA damage removal over a period of 120 min was significantly related to the initial DNA damage (R2 = 0.86, P < 0.001), consistent with the law of mass action. Significant levels of removal were found in most of the cell lines, V79-4 (15 min), LLC-pk1, LNCaP (30 min), PBMCs (45 min), HCT-116, TK-6 (60 min); no significant removal was observed in MCF-7 and HepG2.
Removal of DNA damage in eight cell lines plus PBMCs treated with a 1 µM Ro 19–8022 plus light for 5 min or b 10 mM KBrO3 for 1 h. After treatment, the cells were incubated to allow repair of the damage in appropriate culture medium at 37 °C for time intervals as indicated. The removal of oxidized bases was monitored using the Fpg-modified comet assay. Data are shown as the mean of median values of three repeat experiments. In the case of PBMCs, there were 2 repeat experiments with each of the 5 samples
A Association between relative decrease of net Fpg-sensitive sites induced by Ro 19–8022 plus 5 min light over the first 120 min of incubation and initial level of net Fpg-sensitive sites induced by this treatment. b Association between relative decrease of net Fpg-sensitive sites induced by KBrO3 over the first 120 min of incubation and initial level of net Fpg-sensitive sites induced by KBrO3
Kinetics of removal of DNA damage after treatment of cells with Ro 19–8022 + light or with KBrO 3
Although photosensitizer Ro 19–8022 is frequently used to induce oxidized bases in DNA, finding a suitable concentration of this chemical to produce the same initial DNA damage in all cell lines was a challenge; for instance, doubling the concentration of Ro 19–8022 applied to MCF-7 and V79-4 cells (both cell lines with low initial levels of DNA damage) did not significantly increase DNA damage levels.
We therefore tested KBrO3 as an alternative DNA-damaging chemical, which produces more consistent levels of DNA damage (data not shown). Similar levels of damage (around 60–70% tail DNA) were induced by 10 mM KBrO3 in the 8 cell types (PBMCs being the exception, with only 35.8% net Fpg-sensitive sites at time zero). The removal of damage (i.e. DNA repair) was monitored for 480 min (Fig. 1b). Some cell lines were able to efficiently repair the KBrO3-induced DNA damage reaching background levels already after 120 min, whereas in other cell lines, DNA damage was still significantly higher in the treated cells after 120 min when compared to background levels (Fig. S1) in that same cell type. The percentage removal of damage after 120 min showed no significant correlation with the initial DNA damage induced (R2 = 0.005, P > 0.05) (Fig. 2b).
Initial DNA damage (net Fpg-sensitive sites) related to H 2 O 2 sensitivity of cells
The antioxidant status of cells was assessed by their resistance to oxidation by H2O2; a low level of DNA breaks induced by incubation with H2O2 at 4 °C reflects high antioxidant status. As shown in Fig. 3a, the induction of DNA damage by H2O2 significantly correlates with the initial level of net Fpg-sensitive sites caused by Ro 19–8022 plus light (R2 = 0.54, P < 0.05)—indicating that the level of net Fpg-sensitive sites induced by Ro 19–8022 is influenced by the cellular antioxidant status. No such association exists between antioxidant status and net Fpg-sensitive sites induced by KBrO3 (R2 = 0.11, P > 0.05) (Fig. 3b).
Timing of repair analyses
The question now arises, how best to express the rate of DNA repair? From a biochemical point of view, the initial rate would be the best measure, but the half-time of removal of damage, t (1/2), is another potentially useful parameter (Collins 2014). In an attempt to select the most appropriate measure, we first studied the overall removal of DNA damage by taking the average of all cell lines (Fig. 4). To test if DNA repair is following first-order kinetics, the natural logarithm of DNA damage (average of all cell lines) was plotted versus time, because if this graph is linear and has a negative slope, the reaction is considered to be first-order. Two phases of DNA repair could be distinguished with a linear part in the first 60 min and a linear part in the following period, representing a phase of fast repair and a subsequent period of slow repair (see Fig. 4). On the basis of this observation, t (1/2) was calculated for each cell line for 2 different intervals, namely, between 0 and 60 min and between 60 and 480 min (Table 1). On average, the t (1/2) was tenfold lower in the first 60 min compared to the period from 60 to 480 min. However, there were large differences between cell types varying between 2.4-fold lower repair in the second phase compared to the first phase in V79-4 cells (P = 0.03), to 26.5-fold differences in HeLa cells (P = 0.01). Interestingly, a lower repair rate in the first phase was accompanied by a higher repair rate (thus lower t (1/2)) in the second phase. This may indicate that if more damage remains after 60 min, then higher repair is necessary for the cell to continue the removal of DNA damage. Since this remaining DNA damage at t = 60 min cannot be controlled, we suggest that only the interval between 0 and 60 min will reliably reflect DNA repair capacity. Nevertheless, the later phase of DNA repair may still be of interest.
Discussion
The cellular repair assay based on the comet assay is a straightforward method for assessing DNA repair activity by measuring the kinetics of DNA damage removal over time. Indeed, Valdiglesias et al. (2020) published an extensive protocol to apply this assay in human biomonitoring by testing differently processed blood samples, including whole blood cells and frozen and fresh PBMCs with different DNA damage-inducing agents. Although this approach has already been applied in human biomonitoring studies, it has not been systematically validated, and various aspects of the assay still need to be addressed before it can be used as a reliable measure of repair rate. Therefore, we further optimized this assay for studying cellular DNA repair kinetics, with a focus on the evaluation of BER, by exposing cells to compounds that induce 8-oxoG, namely, Ro 19–8022 and KBrO3. 8-OxoG was subsequently detected using the Fpg-comet assay. The use of Fpg is thought to increase the specificity of the assay for measuring 8-OxoG. However, other types of DNA lesions such as ring-opened purine lesions are also detected, and therefore, the repair kinetics in the current study will not solely reflect the repair of 8-oxoG. The specificity of the assay could be further improved by using the mammalian equivalent of Fpg, 8-oxoguanine DNA glycosylase (human form – hOGG1), and MUTYH glycosylase (Vodicka et al. 2020) for detecting the remaining levels of 8-oxoG with higher precision. Replacing Fpg with other DNA glycosylases would make this assay also fit the study of repair of other types of lesions; for instance, Muruzabal et al. (2020) described the use of hAAG to study the removal of alkylation DNA damage.
Additionally, the type of exposure can improve the specificity; exposure to Ro 19–8022 plus light induces more lesions than just 8-oxoG, and therefore, the repair kinetics reflect the combination of repair by different glycosylases and different types of lesions. Our study showed that exposure to KBrO3 resulted in less variation in the level of DNA damage, consistent with conclusions from the recent inter-laboratory ring trial (Møller et al. 2020). Moreover, cell-free experiments indicated that the involvement of hydroxyl radicals and singlet oxygen in the lesions produced by KBrO3 can be excluded, and data were consistent with a radical mechanism involving bromine radicals (Ballmaier and Epe 1995). As a result, the net Fpg-sensitive sites induced by KBrO3 may be less affected by the antioxidant capacity of cells when compared to Ro 19–8022 plus light. Indeed, in the current study, we showed that the initial level of net Fpg-sensitive sites at T0 after exposure to Ro 19–8022 plus light was related to the antioxidant capacity of cells, whereas after exposure to KBrO3, it was not (Fig. 2). However, it should be stated here that the damaging properties of KBrO3 actually need glutathione (GSH), which is also an important intracellular antioxidant. In this study, we assessed the antioxidant capacity of cells by a short incubation (5 min) with hydrogen peroxide, subsequently measuring the number of strand breaks with the comet assay. This method has previously been successfully used to study the interaction between antioxidant status and genotoxicity (Davies et al. 2001).
Cells have complex molecular mechanisms in response to xenobiotic stress. Redox regulation is an essential mechanism for regulating cellular processes, and there is a balance between ROS (reactive oxygen species) formation and antioxidant defense. Loss of this balance causes excessive DNA oxidation, which could induce DNA repair activity as a “back-up” system. Therefore, as well as studying the association between antioxidant defense and the initial damage induced by 1 µM Ro 19–8022 plus light, we looked at the association between the amount of DNA damage at T0 and repair activity. For instance, some cell types (MCF-7 and LncaP) have relatively low initial damage at T0 after exposure to Ro 19–8022 plus light (possibly due to a higher antioxidant capacity) and, as a result, showed limited DNA repair activity. These cells may, however, still have the capacity to repair more if more damage had been induced. Therefore, the initial amount of damage seemed to be an important determinant of the DNA repair measurement. In other words, to improve the comparison between cell lines or cells from different individuals in human biomonitoring using the cellular DNA repair assay, it is important to induce similar levels of DNA damage at T0. To reach similar initial amounts of DNA damage in the cell lines, we first adjusted doses of Ro 19–8022 plus light (data not shown), but it seemed impossible to achieve this goal. Therefore, another DNA-challenging agent was needed. The ring trial performed by the hCOMET COST Action suggested that KBrO3 could replace Ro 19–8022 as a positive control for the Fpg-modified comet assay (Møller et al. 2020a). So far, KBrO3 has been used in only a few studies with the Fpg-modified comet assay. TK-6 cells and THP-1 cells were treated with between 1 and 5 mM KBrO3 for 1–3 h (Møller et al. 2020; Muruzabal et al. 2020; Platel et al. 2011); Kumar et al. (2022) treated U2OS cells with 0–20 mM KbrO3 for 1 h to induce 8-oxoG. The cell viability of U2OS wild type in 10 mM KbrO3 is 80%. In our experiments, we exposed cells to slightly higher levels of KBrO3 (10 mM), compared to published concentrations, in order to reach similar levels of damage in all cell types.
The direct DNA damage after exposure to KBrO3 at T0 is assessed after 1 h of incubation with high concentrations of KBrO3. In that period, cells may react by upregulating their antioxidant ability, for instance by re-synthesis of GSH or the expression of antioxidant enzymes such as SOD, TrX, and catalase, but DNA damage is induced nonetheless. As a result, the DNA repair pathways can be activated, and the full DNA repair capacity can be studied after 1 h. However, this early induction of DNA repair may interfere with having a reliable measurement at T0, because the formation and removal of DNA damage occur simultaneously. Still, after transformation of the data by the natural logarithm, an association was observed that reached linearity, indicating that T0 can still be used to reliably calculate the half-life over the period of 0–60 min. After 60 min, the DNA repair activity was observed to slow down, resulting in a larger t (1/2). Since the amount of DNA damage at T60 can thus not be controlled, we suggest that when comparing cell lines or individuals, only the early phase will reliably reflect inter-individual differences in DNA repair. There have been previous reports of biphasic repair of single-strand breaks (SSBs) (Furuno et al. 1979; Sossou et al. 2005), double-strand breaks (DSBs) (Dolling et al. 1998; Shibata and Jeggo 2020; Stamato et al. 1993; Wlodek and Hittelman 1987; Wu et al. 2002), or both SSBs and DSBs (Olive 1998; V. Calini et al. 2002) induced by ionizing radiation or other genotoxic agents. It is not known at this moment what these two phases represent. There are several possibilities: Some DNA lesions may be more difficult to reach for the DNA repair enzymes since the structure of chromatin could have an important influence on DNA repair kinetics (Wheeler and Wierowski 1983). High levels of DNA damage will result in high repair activity, in agreement with the Law of Mass Action, which states that the rate of a reaction R is equal to the concentration of reactant ([A]) multiplied by a rate of constant (k1), R = k1[A]. Therefore, low reactant levels (adduct) will have low reaction (repair) rates and decline as the reactant is used (Kumar et al. 2022; Parlanti et al. 2012).
We have shown significant differences in BER rates among an arbitrarily selected group of cell lines plus a sample of PBMCs. It remains to be seen whether comparable variation exists between PBMCs from different individuals. PBMCs are widely used in human biomonitoring studies, but in theory, other cell types can also be used. PBMCs circulate the whole body and are well-suited for exposure studies in environmental and occupational health research (Esteves et al. 2020; Koppen et al. 2018; Milić et al. 2019). PBMCs are easily accessible and not complicated to handle compared with other cell types. Of course, DNA repair can be different in PBMCs when compared to internal organs. However, while significant correlations were found between DNA repair in organs and PBMCs (Herrera et al. 2009; Slyskova et al. 2012), more work needs to be done in understanding the use of surrogate tissues. Other surrogates used in human biomonitoring include epithelial cells from the eye, tear duct, buccal, or nasal cells to measure DNA damage (Rojas et al. 2014; Russo et al. 2020), but these were not yet used for studying DNA repair.
In conclusion, our study, using the Fpg-modified comet assay to follow the removal of DNA base damage, demonstrates the difference in cellular repair kinetics shown by two different 8-oxoG-inducing agents in various cell lines. Using KBrO3 to induce DNA base oxidation resulted in similar initial damage levels in all cell lines, avoiding the complication of varying initial damage apparently owing to differing antioxidant status that is seen with Ro 19–8022. Moreover, t (1/2) over the first hour after exposure could be an optimal indicator for measuring the DNA repair ability of 8-oxoG by BER.
Data availability
All data generated or analyzed during this study are included in this published article.
References
Ballmaier D, Epe B. Oxidative DNA damage induced by potassium bromate under cell-free conditions and in mammalian cells. Carcinogenesis. 1995. https://doi.org/10.1093/carcin/16.2.335.
Ballmaier D, Epe B. DNA damage by bromate: mechanism and consequences. Toxicology. 2006;221(2–3):166–71. https://doi.org/10.1016/j.tox.2006.01.009.
Calini V, Urani C, Camatin M. Comet assay evaluation of DNA single- and double-strand breaks induction and repair in C3H10T1/2 cells. Cell Biol Toxicol. 2002;18:369–79. https://doi.org/10.1023/A:1020811522100.
Camenisch U, Naegeli H. Role of DNA repair in the protection against genotoxic stress. Mol, Clin Environ Toxicol. 2009;99:111–50.
Collins AR, Azqueta A. DNA repair as a biomarker in human biomonitoring studies; further applications of the comet assay. Mutation Research - Fundamental and Molecular Mechanisms of Mutagenesis. 2012;736(1–2):122–9. https://doi.org/10.1016/j.mrfmmm.2011.03.005.
Collins AR. Measuring oxidative damage to DNA and its repair with the comet assay. Biochim Biophys Acta Gen Subj. 2014;1840(2):794–800. https://doi.org/10.1016/j.bbagen.2013.04.022.
Collins AR, Duthie SJ, Dobson VL. Direct enzymic detection of endogenous oxidative base damage in human lymphocyte DNA. Carcinogenesis. 1993;14(9):1733–5. https://doi.org/10.1016/0921-8777(94)00043-6.
Collins AR, Ai-guo M, Duthie SJ. The kinetics of repair of oxidative DNA damage (strand breaks and oxidised pyrimidines) in human cells. Mutat Res-DNA Repair. 1995;336(1):69–77.
Collins AR, Gedik C, Wood S, White A, Dubois J, Duez P, et al. Inter-laboratory validation of procedures for measuring 8-oxo-7,8-dihydroguanine/8-oxo-7,8-dihydro-2′-deoxyguanosine in DNA. Free Radic Res. 2002;36(3):239–45. https://doi.org/10.1080/10715760290019246.
Davies RJH, Collins AR, Horvathovat E. Biochemical and biomedical aspects of oxidative modification Irish area section organized and edited by oxidative DNA damage, antioxidants and DNA repair: applications of the comet assay. 2001. Available from: http://www.rri.sari.ac.uk/escodd
Dolling J-AD, Boreham DL, Brown REJ, Raaphorst GP. Modulation of radiation-induced strand break repair by cisplatin in mammalian cells. International journal of radiation biology. 1998: 61–69. https://doi.org/10.1080/095530098141735
Dusinska M, Collins A. Detection of oxidised purines and UV-induced photoproducts in DNA of single cells, by inclusion of lesion-specific enzymes in the comet assay. Thirteenth Scandinavian Cell Toxicol Congress. 1996. https://doi.org/10.1177/0261192996024003.
Esteves F, Amaro R, Silva S, Sánchez-Flores M, Teixeira JP, Costa C. The impact of comet assay data normalization in human biomonitoring studies outcomes. Toxicol Letter. 2020;332(6):56–64. https://doi.org/10.1016/j.toxlet.2020.06.024.
Furuno I, Yada T, Matsudaira H, Maruyama T. Induction and repair of DNA Strand breaks in cultured mammalian cells following fast neutron irradiation. Int J Radiat Biol. 1979;36(6):639–48. https://doi.org/10.1080/09553007914551471.
Goto N, Bazar G, Kovacs Z, Kunisada M, Morita H, Kizaki S, et al. Detection of UV-induced cyclobutane pyrimidine dimers by near-infrared spectroscopy and aquaphotomics. Sci Rep Nat Publishing Group. 2015;5(1):11808. https://doi.org/10.1038/srep11808.
Hašplová K, Hudecová A, Magdolénová Z, Bjøras M, Gálová EŠ, Miadoková E, et al. DNA alkylation lesions and their repair in human cells: modification of the comet assay with 3-methyladenine DNA glycosylase (AlkD). Toxicol Letter. 2012;208(1):76–81. https://doi.org/10.1016/j.toxlet.2011.10.005.
Herrera M, Dominguez G, Garcia JM, Peña C, Jimenez C, Silva J, et al. Differences in repair of DNA cross-links between lymphocytes and epithelial tumor cells from colon cancer patients measured in vitro with the comet assay. Clin Cancer Res. 2009;15(17):5466–72. https://doi.org/10.1158/1078-0432.CCR-08-3268.
Koppen G, De Prins S, Jacobs A, Nelen V, Schoeters G, Langie SAS. The comet assay in human biomonitoring: cryopreservation of whole blood and comparison with isolated mononuclear cells. Mutagenesis. 2018;33(1):41–7. https://doi.org/10.1093/mutage/gex034.
Kumar N, Theil AF, Roginskaya V, Ali Y, Calderon M, Watkins SC, et al. Global and transcription-coupled repair of 8-oxoG is initiated by nucleotide excision repair proteins. Nat Commun. Nature Research; 2022;13(1). https://doi.org/10.1038/s41467-022-28642-9
Luan Y, Suzuki T, Palanisamy R, Takashima Y, Sakamoto H, Sakuraba M, et al. Potassium bromate treatment predominantly causes large deletions, but not GC > TA transversion in human cells. Mutat Res - Fundam Mol Mech Mutagen. 2007;619(1–2):113–23. https://doi.org/10.1016/j.mrfmmm.2007.02.029.
Milić M, Ožvald I, Vinković Vrček I, Vučić Lovrenčić M, Oreščanin V, Bonassi S, et al. Alkaline comet assay results on fresh and one-year frozen whole blood in small volume without cryo-protection in a group of people with different health status. Mutat Res Genet Toxicol Environ Mutagen. 2019;843(8):3–10. https://doi.org/10.1016/j.mrgentox.2019.03.009.
Møller P, Muruzabal D, Bakuradze T, Richling E, Bankoglu EE, Stopper H, et al. Potassium bromate as positive assay control for the Fpg-modified comet assay. Mutagenesis. 2020;35(4):341–8. https://doi.org/10.1093/mutage/geaa011.
Møller P, Möller L, Godschalk RWL, Jones GDD. Assessment and reduction of comet assay variation in relation to DNA damage: studies from the European Comet Assay Validation Group. Mutagenesis. 2010. 109–11. https://doi.org/10.1093/mutage/gep067
Muruzabal D, Sanz-Serrano J, Sauvaigo S, Gützkow KB, López de Cerain A, Vettorazzi A, et al. Novel approach for the detection of alkylated bases using the enzyme-modified comet assay. Toxicol Letter. 2020;330(4):108–17. https://doi.org/10.1016/j.toxlet.2020.04.021.
Ocolotobiche EE, Dauder RM, Güerci AM. Radiosensitivity of radiotherapy patients: the effect of individual DNA repair capacity. Mutat Res/Gen Toxicol Environ Mutagen. 2021;867(6):503371. https://doi.org/10.1016/j.mrgentox.2021.503371.
Olive PL. The role of DNA single- and double-strand breaks in cell killing by ionizing radiation. Radiat Res. 1998;150(5s):S42-51. https://doi.org/10.2307/3579807.
Ostling O, Johanson KJ. Microelectrophoretic study of radiation-induced DNA damages in individual mammalian cells. Biochem Biophys Res Commun. 1984;123(1):291–8. https://doi.org/10.1016/0006-291X(84)90411-X.
Palyvoda O, Polańska J, Wygoda A, Rzeszowska-Wolny J. DNA damage and repair in lymphocytes of normal individuals and cancer patients studies by the comet assay and micronucleus tests. Acta Biochim Pol. 2003;50(1):181–90. https://doi.org/10.18388/abp.2003_3725.
Panayiotidis M, Collins AR. Ex vivo assessment of lymphocyte antioxidant status using the comet assay. Free Radical Research. 1997;27(5):533–7. https://doi.org/10.3109/10715769709065793.
Parlanti E, Derrico M, Degan P, Calcagnile A, Zijno A, van der Pluijm I, et al. The cross talk between pathways in the repair of 8-oxo-7,8-dihydroguanine in mouse and human cells. Free Radic Biol Med. 2012;53(11):2171–7. https://doi.org/10.1016/j.freeradbiomed.2012.08.593.
Petrucci RH, Herring FG, Madura JD, Bissonnette C. General chemistry : principles and modern applications;1997.
Platel A, Nesslany F, Gervais V, Claude N, Marzin D. Study of oxidative DNA damage in TK6 human lymphoblastoid cells by use of the thymidine kinase gene-mutation assay and the in vitro modified comet assay: determination of no-observed-genotoxic-effect-levels. Mutat Res/Gen Toxicol Environ Mutagen. 2011;726(2):151–9. https://doi.org/10.1016/j.mrgentox.2009.06.006.
Rojas E, Lorenzo Y, Haug K, Nicolaissen B, Valverde M. Epithelial cells as alternative human biomatrices for comet assay. Front Genet. 2014;5(11):1–1. https://doi.org/10.3389/fgene.2014.00386.
Russo C, Acito M, Fatigoni C, Villarini M, Moretti M. B-comet assay (Comet assay on buccal cells) for the evaluation of primary DNA damage in human biomonitoring studies. Int J Environ Res Public Health. 2020;17(24):1–14. https://doi.org/10.3390/ijerph17249234.
Singh NP, Mccoy MT, Tice RR, Schneider EL. A simple technique for quantitation of low levels of DNA damage in individual cells. Exp Cell Res. 1988. https://doi.org/10.1016/0014-4827(88)90265-0.
Shibata A, Jeggo PA. Roles for 53BP1 in the repair of radiation-induced DNA double strand breaks. DNA Repair (amst). 2020. https://doi.org/10.1016/j.dnarep.2020.102915.
Slyskova J, Korenkova V, Collins AR, Prochazka P, Vodickova L, Svec J, et al. Functional, genetic, and epigenetic aspects of base and nucleotide excision repair in colorectal carcinomas. Clin Cancer Res. 2012;18(21):5878–87. https://doi.org/10.1158/1078-0432.CCR-12-1380.
Sossou M, Flohr-Beckhaus C, Schulz I, Daboussi F, Epe B, Radicella JP. APE1 overexpression in XRCC1-deficient cells complements the defective repair of oxidative single strand breaks but increases genomic instability. Nucleic Acids Res. 2005;33(1):298–306. https://doi.org/10.1093/nar/gki173.
Speit G, Schütz P, Bonzheim I, Trenz K, Hoffmann H. Sensitivity of the FPG protein towards alkylation damage in the comet assay. Toxicol Letter. 2004;146(2):151–8. https://doi.org/10.1016/j.toxlet.2003.09.010.
Stamato T, Guerriero S, Denko N. Two methods for assaying DNA double-strand break repair in mammalian cells by asymmetric field inversion gel electrophoresis. Radiat Res. 1993. https://doi.org/10.2307/3578257.
Valdiglesias V, Sánchez-Flores M, Fernández-Bertólez N, Au W, Pásaro E, Laffon B. Expanded usage of the Challenge-Comet assay as a DNA repair biomarker in human populations: protocols for fresh and cryopreserved blood samples, and for different challenge agents. Arch Toxicol. 2020;94(12):4219–28. https://doi.org/10.1007/s00204-020-02881-5.
Vodicka P, Urbanova M, Makovicky P, Tomasova K, Kroupa M, Stetina R, et al. Oxidative damage in sporadic colorectal cancer: Molecular mapping of base excision repair glycosylases in colorectal cancer patients. Int J Mol Sci. 2020;21(7):2473. https://doi.org/10.3390/ijms21072473.
Wheeler KT, Wierowski JV. DNA repair kinetics in irradiated undifferentiated and terminally differentiated cells. Radiat Environ Biophys. 1983;22(1):3–19. https://doi.org/10.1007/BF01323757.
Wlodek D, Hittelman WN. The repair of double-strand DNA breaks correlates with radiosensitivity of L5178Y-S and L5178Y-R cells. Radiat Res. 1987. https://doi.org/10.2307/3577085.
Wu J, Yin M-B, Hapke G, Aroly K´, Oth T´, Rustum YM. Induction of biphasic DNA double strand breaks and activation of multiple repair protein complexes by DNA topoisomerase I drug 7-ethyl-10-hydroxy-camptothecin. 2002. https://doi.org/10.1124/mol.61.4.742.
Acknowledgements
We would like to thank Kristine Bjerve Gützkow and Ann-Karin Hardie Olsen for sending us some cell lines as a gift.
Funding
The work was supported by the following grants: the European Union’s Horizon 2020 research and innovation training networks (ITN, aDDress, Marie-Skłodowska-Curie grant agreement no.812829); TEPCAN project granted in the program “Applied research” under the Norwegian Financial Mechanisms 2014–2021/POLNOR 2019 (EEA and Norway Grants), and by STIM4 + project granted in the program “Applied research” under the Norwegian Financial Mechanisms 2014–2021, Contract No. 14/2020, Project RO-NO-2019–0540.
The funding agencies were not involved in the design of the study, the collection, analysis, and interpretation of data, and in writing the manuscript.
Author information
Authors and Affiliations
Contributions
Roger Godschalk, Andrew Collins, Frederik-Jan van Schooten, and Sergey Shaposhnikov conceived the project; Roger Godschalk, Andrew Collins, Gunnar Brunborg, Sergey Shaposhnikov, and Anne Lene Nordengen critically assessed the results and the manuscript; Congying Zheng, Andrew Collins, and Sergey Shaposhnikov designed in vitro toxicity experiments, which were performed by Congying Zheng with technical input from Anne Lene Nordengen; Congying Zheng wrote the manuscript with input from the other authors; Congying Zheng performed the statistical analysis. All authors read and approved the final submitted manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing interests.
Ethics approval
The manuscript does not contain clinical studies or patient data.
Consent to participate
The manuscript does not contain clinical studies or patient data.
Consent for publication
Not applicable.
Conflict of interest
The authors declare no competing interests.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Zheng, C., Collins, A., Brunborg, G. et al. Assay conditions for estimating differences in base excision repair activity with Fpg-modified comet assay. Cell Biol Toxicol 39, 2775–2786 (2023). https://doi.org/10.1007/s10565-023-09801-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10565-023-09801-0