Abstract
The geography of the Black Hills region of South Dakota and Wyoming may limit connectivity for many species. For species with large energetic demands and large home ranges or species at low densities this can create viability concerns. Carnivores in this region, such as cougars (Puma concolor), have the additive effect of natural and human-induced mortality; this may act to decrease long-term viability. In this study we set out to explore genetic diversity among cougar populations in the Black Hills and surrounding areas. Specifically, our objectives were to first compare genetic variation and effective number of breeders of cougars in the Black Hills during three harvest regimes: pre (2003–2006), moderate (2007–2010), and heavy (2011–2013), to determine if harvest impacted genetic variation. Second, we compared genetic structure of the Black Hills cougar population with cougar populations in neighboring eastern Wyoming and North Dakota. Using 20 microsatellite loci, we conducted genetic analysis on DNA samples from cougars in the Black Hills (n = 675), North Dakota (n = 113), and eastern Wyoming (n = 62) collected from 2001–2013. Here we report that the Black Hills cougar population maintained genetic variation over the three time periods. Our substructure analysis suggests that the maintenance of genetic variation was due to immigration from eastern Wyoming and possibly North Dakota.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
By the early 1900s, the North American cougar (Puma concolor) population had been displaced from two-thirds of its historic range (Young and Goldman 1946). Populations have since increased, and currently occupy the western United States and part of southern Florida (Beier 1991; Logan and Sweanor 2001). Several studies have suggested habitat barriers (i.e., fragmentation and habitat degradation) can limit gene flow resulting in distinct genetic structure among some cougar populations (Ernest et al. 2003; Mcrae et al. 2005; Loxterman 2011; Andreasen et al. 2012; Holbrook et al. 2012). In contrast, others have argued immigration/emigration can maintain genetic variation (Biek et al. 2006; Anderson et al. 2004) and can limit population subdivision in wild cougar populations (Sinclair et al. 2001; Anderson et al. 2004).
Advances in DNA analyses have enabled genetics to play an increasingly important role in the conservation and management of many wildlife populations (DeYoung and Honeycutt 2005; Culver and Schwartz 2011). Genetic data can allow for estimates of effective population size (Ne), dispersal, inbreeding, and gene flow (Culver and Schwartz 2011), and recently has been applied to many wide-ranging carnivore populations (e.g., Spong et al. 2000; Haag et al. 2010; Coster and Kovach 2012; Croteau et al. 2012). The additive effects of human-induced mortality to some North American cougar populations may affect the genetic viability of these populations; particularly those in unique geographic regions, i.e., the Black Hills National Forest.
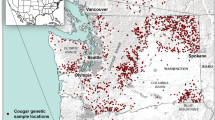
Historically, cougars were found throughout South Dakota (Young and Goldman 1946), but were thought to have been eradicated by the early 1900s (South Dakota Mountain Lion Management Plan 2010). The Black Hills complex is relatively small (Fig. 1) with an estimated 6703 km2 of suitable cougar habitat (Fecske 2003), and is largely surrounded by inhospitable cougar habitat (Anderson et al. 2004), i.e., the Northern Great Plains. During the late 1980s to early 1990s, reports of cougar sightings in the Black Hills increased; some of which were eventually verified by state biologists (South Dakota Mountain Lion Management Plan 2010). By 2002, there was an estimated population size of 127–149 cougars inhabiting the Black Hills (Fecske 2003). The cougar population in the Black Hills likely resulted from immigrants from neighboring Wyoming populations, which either founded the current population, or augmented a small remnant population that persisted (Fecske 2003).
Initial genetic data suggested that the Black Hills were founded by immigrants from north-central and southeastern Wyoming, and that sustained gene flow had occurred between regions (Anderson et al. 2004). By 2006, a concomitant study showed that cougars in the Black Hills had maintained genetic variation, and the population was saturated (Thompson 2009) with a conservative modeled population estimate of about 250 individuals (J. A. Jenks, South Dakota State University, unpublished data). As a result, South Dakota Department of Game, Fish and Parks (SDGFP) initiated its first cougar harvest season in 2005 and harvest limits in the Black Hills Fire Protection Area have increased steadily from 2005–2013 (Fig. 2).
The semi-isolated nature of the Black Hills could make the Black Hills cougar population prone to deleterious effects from potential inbreeding and increased genetic drift (Culver and Schwartz 2011). Prolonged and intense harvest has the potential to result in a greater loss of genetic variation and alteration of population structure (Allendorf et al. 2008). Thus, the objectives of this study were to: assess and compare genetic variation (i.e., heterozygosity, allelic richness, effective population size) of cougars in the Black Hills during three different harvest regimes: pre (2001–2006), moderate (2007–2010), and heavy (2011–2013), and examine the impact of increased harvest on population substructure. Additionally, while previous and current telemetry data verify cougar dispersal out of the Black Hills into other cougar populations (e.g., Wyoming, Montana, Nebraska, and North Dakota; Thompson 2009; Thompson and Jenks 2010), data confirming immigration into the Black Hills remains limited. Our final objective was to compare genetic structure of Black Hills cougars to that of neighboring cougar populations in North Dakota and eastern Wyoming, and conduct population assignment tests to determine evidence of immigration.
Materials and methods
Study area
We conducted our study in the Black Hills National Forest and surrounding lands (Fig. 1), located in western South Dakota and northeastern Wyoming. The Black Hills represent the eastern most extension of the Rocky Mountains (Froiland 1990), encompassing approximately 8400 km2 (Fecske et al. 2004). Maximum elevation in the Blacks Hills is 2207 m above mean sea level, with topography consisting of rock outcrops, rolling hills, steep ridges, canyonlands, and gulches (Froiland 1990). The most abundant tree species occurring in the Black Hills is ponderosa pine (Pinus ponderosa). White spruce (Picea glauca), aspen (Populus tremuloides), and birch (Betula spp.) trees also are prevalent at higher elevations; burr oak (Quercus macrocarpa) draws occur at lower elevations (Larson and Johnson 1999). Prey species available to cougars include: white-tailed deer (Odocoileus virginianus), mule deer (O. hemionus), elk (Cervus elaphus), bighorn sheep (Ovis canadensis), mountain goat (Oreamnos americanus), porcupine (Erethizon dorsatum) as well as a variety of small mammal and domestic livestock species. The cougar is the apex predator in the Black Hills, occurring sympatrically with bobcat (Lynx rufus) and coyote (Canis latrans). The Black Hills is surrounded by the Northern Great Plains. The closest breeding cougar populations occur in the Pine Ridge region, Nebraska (48 km southeast), Badlands, North Dakota (120 km north/northwest), Laramie Range, Wyoming (160 km southwest), and the Bighorn Mountains, Wyoming (200 km west; Thompson and Jenks 2010).
Capture methods
From 1 January 2003–1 January 2013, we captured cougars primarily with the use of trained hounds; however, we also utilized walk-in live traps, foot-hold traps, and leg-hold traps (Logan et al. 1999). We immobilized cougars with a mixture of telazol (5.0 mg/kg) and xylazine (1.0 mg/kg; Kreeger and Armeno 2007) based on estimated live animal body weight via dart rifle (Dan-Inject, Børkop, Denmark, EU). We weighed, measured, sexed, and estimated age of cougars by tooth wear and pelage characteristics (Anderson and Lindzey 2000). We collected blood samples for genetic analysis from all captured cougars and fitted them with VHF (Telonics MOD-500 (NH), Inc., Mesa, Arizona, USA) or GPS (ATS G2110E, Inc. Isanti, Minnesota, USA; Northstar D-cell, King George, Virginia, USA) radio collars. Immobilized cougars were reversed with 0.125 mg/kg yohimbine, released on site, and observed from a distance to ensure safe recovery. We collected tissue samples from cougars within the Black Hills, South Dakota, from harvest and non-harvest related mortalities. We collaborated with the Wyoming Game and Fish Department to obtain tissue samples collected during 2011–2013 biopsy darting and harvest from cougars in the Wyoming portion of the Black Hills, as well from other eastern regions of Wyoming (i.e., Casper, Sheridan, and Laramie). Additionally, we collaborated with the North Dakota Game and Fish Department to obtain tissue samples collected from 2003–2012 from harvest, non–harvest related mortalities, and live captures. All animal capture procedures were approved by the South Dakota State University Animal Care and Use Committee (Approval number 11-078A) and followed recommendations of the American Society of Mammalogists (Sikes and Gannon 2011).
Genetic analysis
DNA extraction and genetic analysis was conducted at the National Genomics Center for Wildlife and Fish Conservation, United States Forest Service, Rocky Mountain Research Station (Missoula, Montana, USA). Samples were analyzed at 20 microsatellite loci used in previous studies on cougars (Menotti-Raymond and O’Brien 1995; Menotti-Raymond et al. 1999): Fca43, Fca57, Fca77, Fca90, Fca96, Fac132, Fca559, Fca176, Fca35, Lc109, Fca391, Fca08, Fca30, Fca82, Fca149, PcoA208, PcoB10, PcoC112, PcoB210, PcoC108. Genomic DNA from blood and tissue was extracted with the Dneasy Tissue Kit (Qiagen, Inc., Valencia, CA, USA). Polymerase chain reaction (PCR) volume (10 μl) contained 1.0 ml DNA, 1× reaction buffer (Applied Biosystems, Foster City, CA, USA), 2.0 mM MgCl2, 200 mM of each dNTP, 1 mM reverse primer, 1 mM dye-labeled forward primer, 1.5 mg/ml BSA, and 1U Taq polymerase (Applied Biosystems). The PCR profile was 94 °C/5 min, ([94 °C/1 min, 55 °C/1 min, 72 °C/30 s] × 36 cycles). The resultant products were visualized on a LI-COR DNA analyzer (LI-COR Biotechnology). We tested for genotyping error using program DROPOUT (McKelvey and Schwartz 2005) following Schwartz et al. (2006).
We grouped individuals into cohorts based on birth year, which were subsequently categorized under the appropriate harvest regime (pre, moderate, and heavy). Cougars can breed as early as 2 years of age; thus, we used a 2 year sliding window approach to avoid the inclusion of parents and offspring in the same cohort. We calculated observed (HO) and expected (HE) heterozygosity, effective alleles (Ae), and tested for deviations from Hardy–Weinberg equilibrium (HWE) using GenAlEx 6.5 (Peakall and Smouse 2006, 2012) and Genepop (Raymond and Rousset 1995; Rousset 2008). To account for variation in sample size we used HPRARE 1.0 (Kalinowski 2005), which uses rarefied measures to calculate allelic richness (Ar). Most methods used to estimate effective population size (Ne) assume discrete generations, which can cause severe bias for species with overlapping generations (Luikart et al. 2010), such as cougars. However, if a single cohort is sampled, effective population size can be estimated as effective number of breeders (Schwartz et al. 1998; Waples 2005). Therefore, we estimated effective population size as the effective number of breeding individuals (from here on referred to as NB) genetically contributing to the population (Waples and Teel 1990; Waples 2005). We calculated NB for each cohort using a bias correction method based on linkage disequilibrium (Hill 1981; Waples 2006; Waples and Do 2010), as implemented by NeEstimator V2 (Do et al. 2014). Statistical analysis among cohorts and harvest regimes was determined using an ANOVA in the R statistical program (R Core Team 2015).
We examined population structure between Black Hills, eastern Wyoming, and North Dakota cougar populations from 2003–2013 using STRUCTURE (Pritchard et al. 2000). We used Geneclass2 (Piry et al. 2004) and STRUCTURE (Pritchard et al. 2000) to conduct population assignment tests to determine immigration between the Black Hills and North Dakota cougar populations using samples from 2001–2013. Finally, we assessed overall genetic variation and population structure for cougars in the Black Hills, eastern Wyoming (i.e., Casper, Sheridan, and Laramie) and North Dakota using individuals from the same birth period (2011–2013) to alleviate temporal effects. We used the R statistical program (R Core Team 2015) diveRsity to determine FST values between the three populations, and performed Fisher’s exact test between the Black Hills and eastern Wyoming populations to test for panmixia (Keenan et al. 2013).
Geneclass2 assigns/excludes individuals using predefined population subdivision based on different genetic assignment criteria (Piry et al. 2004). We employed the frequency based method (Paetkau et al. 1995) using the Monte Carlo resampling method with 1000 simulated individuals and an alpha of 0.01 (Paetkau et al. 2004). STRUCTURE uses both allele frequency and a Bayesian model-based clustering method to infer population structure on the basis of genotypes (Pritchard et al. 2000). To infer population structure, we excluded available geographic information, and assumed individuals had mixed ancestry (admixture model) and correlated allele frequencies. We set both burn-in periods to 10,000 and evaluated 1–10 possible genetic clusters (K), with three iterations. To select the appropriate number of genetic clusters (K), we used Structure Harvester, which collates STRUCTURE results (Earl and vonHoldt 2012) to determine the most suitable K using the Evanno method (Evanno et al. 2005).
Results
We analyzed 675 unique genotypes from Black Hills cougars (pre-harvest: n = 288; moderate harvest: n = 289; and heavy harvest: n = 98), 113 unique genotypes from North Dakota cougars, and 62 unique genotypes from eastern Wyoming cougars at 20 microsatellite loci. We observed no significant difference in observed (P = 0.473) or expected (P = 0.886) heterozygosity levels, allelic richness (P = 0.764), or effective alleles (P = 0.745) in cougars in the Black Hills under different harvest regimes (Table 1). Significant deviations from HWE occurred at PcoB10 (P < 0.05) during the pre-harvest regime, Fca90 (P < 0.05), Fca149 (P < 0.05), and Pco108 (P < 0.05) during the moderate harvest regime, and at Bco210 (P < 0.05) during the heavy harvest regime. We found effective population (NB) size ranged from a low of 38 (28–55; 95 % CI) pre-harvest and a high of 98 (70–150; 95 % CI) moderate harvest, before declining to 62 (42–106; 95 % CI) heavy harvest (Table 2).
STRUCTURE analysis for Black Hills, eastern Wyoming, and North Dakota resulted in a K = 3 (mean Ln P(K) = −37,526.6; Fig. 3), when excluding prior population information. Plotting these results geographically indicated there was more genetic similarity between Black Hills and eastern Wyoming cougars, in comparison to North Dakota cougars. Hierarchical analysis using samples only from the Black Hills and eastern Wyoming resulted in a K = 2 (mean Ln P(K) = −31,844.8; Fig. 4) with no obvious geographic structuring. STRUCTURE analysis for Black Hills and North Dakota populations resulted in a K = 2 (mean Ln P(K) = −36,200.6; Fig. 5), but suggested geographic structuring into a Black Hills and a North Dakota group. We used these results to confirm immigration of cougars (n = 2) into the Black Hills from North Dakota, as well as emigration out of the Black Hills (n = 6) into North Dakota, based on 98 % ancestry in the Black Hills cluster (Table 3). Results from Geneclass2 allowed assignment of two cougars to North Dakota that were originally sampled from the Black Hills cougar population; these individuals were the same as those identified by STRUCTURE. However, we were only able to assign four cougars (of the six identified by STRUCTURE) to the Black Hills population that were originally sampled from North Dakota, based on a >85 % probability (Table 3).
Population structure of Black Hills (red and blue), eastern Wyoming (red and blue), and North Dakota (green) cougars inferred by STRUCTURE, K = 3. Populations are divided by a vertical black line. Each individual is represented by a single column, where the color(s) of the column represent degree of similarity to each population
Population structure of Black Hills (red and green) and eastern Wyoming (red and green), cougars inferred by STRUCTURE, K = 2. Populations are divided by a vertical black line. Each individual is represented by a single column, where the color(s) of the column represent degree of similarity to each population
Genetic variation between the three populations was comparable (Table 4), and we observed no significant difference in observed (P = 0.605) or expected (P = 0.617) heterozygosity, allelic richness (P = 0.261), or effective alleles (P = 0.296). While we found no significant deviations from HWE in the Wyoming cougar population, we found deviations from HWE at Fca132 (P < 0.05), Fca391 (P < 0.05), and PcoA208 (P < 0.05) in the North Dakota cougar population. There were 96 alleles in Black Hills, 64 alleles in North Dakota, and 86 alleles in the eastern Wyoming cougar population, as well as alleles unique to the Black Hills (n = 2), North Dakota (n = 3), and eastern Wyoming population (n = 6). Overall, FST between Black Hills and North Dakota was 0.10 (0.07–0.13; 95 % CI), 0.07 (0.04–0.09; 95 % CI) between North Dakota and eastern Wyoming, and 0.03 (0.02–0.04; 95 % CI) between Black Hills and eastern Wyoming (Table 5); thus, indicating population differentiation, as confidence intervals do not incorporate zero. Fisher’s exact test between Black Hills and eastern Wyoming revealed a significant difference (P < 0.05) at all but 6 loci.
Discussion
Our results indicate that cougars in the Black Hills have maintained genetic variability despite the semi-isolated nature of the region, years of increasing harvest, and harvest pressure from both South Dakota and Wyoming. Multiple measures of genetic variation (i.e., heterozygosity and allelic richness), confirm that Black Hills cougars have maintained genetic variability since the pre-harvest period (Thompson 2009), though we acknowledge that simulations would provide further insight on the processes observed. We found no significant difference in heterozygosity and allelic richness in our cohort analysis across the three harvest regimes: pre–harvest, moderate harvest, and heavy harvest (Table 1), despite the populations’ recent decrease in size from its highest estimate of approximately 250 individuals (Fig. 6). Observed heterozygosity for Black Hills cougars was similar to that of other cougar populations in the western United States, i.e., Texas, Wyoming, Colorado, Utah, Arizona, New Mexico, and Idaho (HO 0.52–0.61; Walker et al. 2000; Anderson et al. 2004; Mcrae et al. 2005; Loxterman 2011; Holbrook et al. 2012), and paralleled other studies (Sinclair et al. 2001; Anderson et al. 2004) that found no evidence of population subdivision. Though measures of genetic variation indicate this population has retained genetic viability, we recognize that unlike demographic effects, which are immediate and more recognizable, genetic effects occur over many generations and therefore, may not be apparent for several generations (Harris et al. 2012).
Loss of genetic diversity has been documented in some wild populations following decreases in population size and/or Ne [African elephant, Loxodonta africana africana, Whitehouse and Harley (2001); mountain goat, Oreamnos americanus, Ortego et al. (2011); Saimaa ringed seal, Phoca hispida saimensis, Valtonen et al. (2012); Far eastern leopard, Panthera pardus orientalis, Sugimoto et al. (2014)]. Other studies of wild populations found either no reduction in genetic diversity [coyote, Canis latrans, Williams et al. (2003); Kerguelen mouflon, Ovis aries, Kaeuffer et al. (2007)], or occasionally even an increase in diversity that resulted from increased immigration [great reed warblers, Acrocephalus arundinaceus, Hansson et al. (2000)]. Williams et al. (2003) found dispersal/immigration preserved genetic variation in a coyote population despite decades of intensive removal. Our cohort estimates of NB (Table 2; Fig. 6) reflected the changes in estimated population size corresponding to the harvest regimes, with lower estimates during the pre- and heavy harvest regimes, and higher estimates during the moderate harvest regime. Although NB decreased during the heavy harvest regime, estimates were similar if not higher than pre–harvest NB estimates. Thus, it is likely that a combination of dispersal and immigration and relatively stable NB, has resulted in genetic maintenance in Black Hills cougars across the three harvest regimes. However, we believe that additional genetic monitoring is warranted to monitor the recent decline observed in NB to ensure this is not a continuing trend.
Wright (1978) suggested that an FST value between 0 and 0.05 indicates little genetic differentiation; based on FST (Table 5), South Dakota and eastern Wyoming cougar populations were more closely related in comparison to North Dakota cougars. Furthermore, STRUCTURE results (Fig. 3) and the similarity in genetic variability at 20 microsatellite loci (Table 3) between South Dakota and eastern Wyoming cougars would suggest that these populations may be acting as a large population as postulated by Anderson et al. (2004). STRUCTURE analysis between Black Hills and eastern Wyoming also depicted little genetic differentiation (Fig. 4). Though Fisher’s exact test revealed statistical significance at 14 loci, rejecting panmixia; this is likely a reflection of the number of unique alleles in the Wyoming cougar population (n = 6) than in the Black Hills population (n = 2). However, a more thorough assessment of the population structure of Wyoming cougars using a larger sample size may provide more insight.
Although Black Hills cougars likely colonized the North Dakota Badlands, there seems to be genetic differentiation between the two Dakota populations, as evidenced by genetic variability results (Tables 4 and 5) and population assignment tests (Table 3). This differentiation in the North Dakota population likely resulted from genetic drift within and/or immigration from other cougar populations, such as Montana, which has been documented (Wilkens 2014). In addition, we were able to assign individual cougars to either North Dakota or South Dakota populations with relatively high probabilities (>85 %); confirming both immigration into the Black Hills from neighboring cougar populations, and emigration from the Black Hills into North Dakota. Our results also suggest eastern Wyoming and South Dakota experience a higher frequency of cougar movements (emigration and immigration) compared to the North Dakota population.
Regardless of habitat barriers that may limit dispersal movements among populations (McRae et al. 2005), such constraints are not preventing cougar dispersal movements out of or into the Black Hills (Thompson and Jenks 2010). Several radio marked individuals from this project were documented dispersing from the Black Hills into North Dakota, Wyoming, and Montana. Additionally, we documented immigration into the South Dakota portion of the Black Hills from an ear-tagged Wyoming cougar. Population immigration likely has sustained the genetic variability of Black Hills cougars, as dispersal facilitates transfer of genetic material, thereby maintaining genetically healthy populations (Logan and Sweanor 2001; Sinclair et al. 2001; Anderson et al. 2004; Biek et al. 2006). Moreover, no clinical signs of inbreeding depression (e.g., crooked tails, cowlicks) have been documented in cougars in the Black Hills (Thompson 2009; Jansen 2011; Juarez 2014).
Conservation and management implications
Genetic assessment of the Black Hills cougar population revealed that the population has maintained genetic viability likely as a result of compensatory emigration/immigration, during years of increased harvest. However, we recommend continued genetic monitoring of cougars in the Black Hills every 3–5 years given that genetic changes occur on an evolutionary time scale (i.e., over many generations; Harris et al. 2012). The cougar populations evaluated in this study may not show genetic effects of harvest pressure and decreasing population size for several generations. As cougars in the Black Hills experience harvest pressure from both South Dakota and Wyoming, continued genetic monitoring is warranted (Schwartz et al. 2007). Furthermore, because cougars in North Dakota and other regions in Wyoming also are experiencing harvest, continued assessment of genetic variation between the three neighboring cougar populations will be important. Dispersal of Black Hills cougars has been documented to North Dakota, Wyoming, Montana, Nebraska, Oklahoma, and Minnesota (Thompson 2009; Thompson and Jenks 2010), which illustrates the difficulty of managing cougars within agency boundaries. Therefore, because cougars do not adhere to state boundaries we also recommend a large scale cougar database, which could provide an effective means for collaboration among management agencies to continue to successfully maintain healthy populations of the North American cougar.
References
Allendorf FW, England PR, Luikart G, Ritchie PA, Ryman N (2008) Genetic effects of harvest on wild animal populations. Trends Ecol Evol 23:327–337
Anderson CR Jr, Lindzey FG (2000) A guide to estimating mountain lion age classes. Wyoming Cooperative Fish and Wildlife Research Unit, Laramie
Anderson CR Jr, Lindzey FG, McDonald DB (2004) Genetic structure of cougar populations across the Wyoming Basin: metapopulation or megapopulation. J Mammal 85:1207–1214
Andreasen AM, Stewart KM, Longland WS, Beckmann JP, Forister ML (2012) Identification of source-sink dynamics in mountain lions of the Great Basin. Mol Ecol 21:5689–5701
Beier P (1991) Cougar attacks on humans in the United States and Canada. Wildl Soc Bull 19:403–412
Biek RA, Akamine N, Schwartz MK, Ruth TK, Murphy KM, Poss M (2006) Genetic consequences of sex-biased dispersal in a solitary carnivore: yellowstone cougars. Biol Lett 2:312–316
Coster SS, Kovach AI (2012) Anthropogenic influences on the spatial genetic structure of black Bears. Conserv Genet 13:1247–1257
Croteau EK, Heist EJ, Nielsen CK, Hutchinson JR, Hellgren EC (2012) Microsatellites and mitochondrial DNA reveal regional population structure in bobcats (Lynx rufus) of North America. Conserv Genet 13:1637–1651
Culver M, Schwartz MK (2011) Conservation genetics and cougar management. In: Jenks JA (ed) Managing cougars in North America. Jack H Berryman Institute, Utah State University, Logan, pp 91–109
DeYoung RW, Honeycutt RL (2005) The molecular toolbox: genetic techniques in wildlife ecology and management. J Wildl Manag 69:1362–1384
Do C, Waples RS, Peel D, Macbeth GM, Tillett BJ, Ovenden JR (2014) NeEstimator V2: re-implementation of software for the estimation of contemporary effective population size (Ne) from genetic data. Mol Ecol Resour 14:209–214
Earl DA, vonHoldt BM (2012) STRUCTURE HARVESTER: a website and program for visualizing STRUCTURE output and implementing the Evanno method. Conserv Genet Resour 4:359–361. doi:10.1007/s12686-011-9548-7
Ernest HB, Boyce WM, Bleich VC, May B, Stiver SJ, Torres SG (2003) Genetic structure of mountain lion (Puma concolor) populations in California. Conserv Genet 4:353–366
Evanno G, Regnaut S, Goudet J (2005) Detecting the number of clusters of individuals using the software STRUCTURE: a simulation study. Mol Ecol 14:2611–2620
Fecske DM (2003) Distribution and abundance of American martins and cougars in the Black Hills of South Dakota and Wyoming. Disseration, South Dakota State University
Fecske DM, Jenks JA, Lindzey FG (2004) Mortality of an adult cougar due to a forest fire. Prairie Naturalist 36:137–140
Froiland SG (1990) Natural history of the Black Hills and Badlands. The Center for Western Studies, Augustana College, Sioux Falls
Haag T, Santos AS, Sana A, Morato RG, Cullen L Jr, Crawshaw JR, De Angelo C, Di Bitetti MS, Salzano FM, Eizirik E (2010) The effect of habitat fragmentation on the genetic structure of a top predator: loss of diversity and high differentiation among remnant populations of Atlantic Forest jaguars (Panthera onca). Mol Ecol 19:4906–4921
Hansson B, Bensch S, Hasselquist D, Lillandt BG, Wennerberg L, Von Schantz T (2000) Increase of genetic variation over time in a recently founded population of great reed warblers (Acrocephalus arundinaceus) revealed by microsatellites and DNA fingerprinting. Mol Ecol 9:1529–1538
Harris RB, Wall WA, Allendorf FW (2012) Genetic consequences of hunting: what do we know and what should we do? Wildl Soc Bull 30:634–643
Hill WG (1981) Estimation of effective population size from data on linkage disequilibrium. Genet Res 38:209–216
Holbrook JD, De Young RW, Janecka JE, Tewes ME, Honeycutt RL, Young JH (2012) Genetic diversity, population structure, and movements of mountain lions (Puma concolor) in Texas. J Mammal 93:989–1000
Jansen BD (2011) Anthropogenic factors affecting mountain lions in the Black Hills of South Dakota. Disseration, South Dakota State University
Juarez RL (2014) Evaluating methods to estimate population size and assessing temporal genetic varition of cougars in the Black Hills. Thesis, South Dakota State University
Kaeuffer R, Coltman D, Chapuis J, Pontier D, Réale D (2007) Unexpected heterozygosity in an island mouflon population founded by a single pair of individuals. Proc Biol Sci 274:527–533
Kalinowski ST (2005) HP-Rare: a computer program for performing rarefaction on measures of allelic diversity. Mol Ecol Notes 5:187–189
Keenan K, McGinnity P, Cross TF, Crozier WW, Prodöhl PA (2013) diveRsity: an R package for the estimation of population genetics parameters and their associated errors. Methods Ecol Evol 4:782–788
Kreeger TJ, Armeno JM (2007) Handbook of wildlife chemical immobilization, 3rd edn. Self-published, Laramie
Larson GE, Johnson JR (1999) Plants of the Black Hills and Bear Lodge Mountains: a field guide with color photographs. A South Dakota State University, College of Agriculture and Biological Sciences Book. Fenske Media Corporation, Rapid City
Logan KA, Sweanor L (2001) Desert puma: evolutionary ecology and conservation of an enduring carnivore. Island Press, Washington, DC
Logan KA, Sweanor LL, Smith JF, Hornocker MG (1999) Capturing pumas with foot-hold snares. Wildl Soc Bull 27:201–208
Loxterman JL (2011) Fine scale population structure of pumas in the Intermountain ‘West’. Conserv Genet 12:1049–1059
Luikart G, Ryman N, Tallmon DA, Schwartz MK, Allendorf FW (2010) Estimation of census and effective population sizes: the increasing usefulness of DNA-based approaches. Conserv Genet 11:355–373
McKelvey KS, Schwartz MK (2005) DROPOUT: a program to identify problem loci and samples for noninvasive genetic samples in a capture-mark-recapture framework. Mol Ecol Notes 5:716–718
McRae BH, Beier P, Dewald LE, Huynh LY, Keim P (2005) Habitat barriers limit gene flow and illuminate historical events in a wide-ranging carnivore, the American puma. Mol Ecol 14:1965–1977
Menotti-Raymond MA, O’Brien SJ (1995) Evolutionary conservation of ten microsatellite loci in four species of Felidae. J Hered 86:319–322
Menotti-Raymond MA, David VA, Lyons LA, Shäffer AA, Tomlin JF, Hutton MK, O’Brien SJ (1999) A genetic linkage map of microsatellites of the domestic cat (Felis catus). Genomics 57:9–23
Ortego J, Yannic G, Shafer AA, Mainguy J, Festa-Bianchet M, Coltman DW, Côté SD (2011) Temporal dynamics of genetic variability in a mountain goat (Oreamnos americanus) population. Mol Ecol 20:1601–1611
Paetkau D, Calvert W, Stirling I, Strobeck C (1995) Microsatellite analysis of population structure in Canadian polar bears. Mol Ecol 4:347–354
Paetkau D, Slade R, Burden M, Estoup A (2004) Genetic assignment methods for the direct, real-time estimation of migration rate: a simulation based exploration of accuracy and power. Mol Ecol 13:55–65
Peakall R, Smouse PE (2006) GENALEX 6: genetic analysis in Excel. Population genetic software for teaching and research. Mol Ecol Notes 6:288–295
Peakall R, Smouse PE (2012) GenAlEx 6.5: genetic analysis in Excel. Population genetic software for teaching and research-an update. Bioinformatics 28:2537–2539
Piry S, Alapetite A, Cornuet JM, Paetkau D, Baudouin L, Estoup A (2004) GeneClass2: a software for genetic assignment and first-generation migrant detection. J Hered 95:536–539
Pritchard JK, Stephens M, Donnelly P (2000) Inference of population structure using multilocus genotype data. Genetics 155:945–959
R Core Team (2015) R: A language and environment for statistical computing. R Foundation for Statistical Computing,Vienna, Austria. URL http://www.R-project.org/
Raymond M, Rousset F (1995) GENEPOP (version 1.2): population genetics software for exact tests and ecumenicism. J Hered 86:248–249
Rousset F (2008) GENEPOP’007: a complete re-implementation of the GENEPOP software for Windows and Linux. Mol Ecol Resour 8:103–106
Schwartz MK, Tallmon DA, Luikart G (1998) Review of DNA-based census and effective population size estimators. Anim Conserv 1:293–299
Schwartz MK, Cushman SA, McKelvey KS, Hayden J, Engkjer C (2006) Detecting genotyping errors and describing American black bear movement in northern Idaho. Ursus 17:138–148
Schwartz MK, Luikart GH, Waples RS (2007) Genetic monitoring as a promising tool for conservation and management. Trends Ecol Evol 22:25–33
Sikes RS, Gannon WL (2011) Guidelines of the American Society of Mammalogists for the use of wild animals in research. J Mammal 92:235–253
Sinclair EA, Swenson EL, Wolfe ML, Choate DC, Bates B, Crandall KA (2001) Gene flow estimates in Utah’s cougars imply management beyond Utah. Anim Conserv 4:257–264
Spong G, Johansson M, Björklund M (2000) High genetic variation in leopards indicates large and long-term stable effective population size. Mol Ecol 9:1773–1782
Sugimoto T, Aramilev VV, Kerley LL, Nagata J, Miquelle DG, McCullough DR (2014) Noninvasive genetic analyses for estimating population size and genetic diversity of the remaining Far Eastern leopard (Panthera pardus orientalis) population. Conserv Genet 15:521–532
Thompson DJ (2009) Population demographics of cougars in the Black Hills: survival, dispersal, morphometry, genetic structure, and associated interactions with density dependence. Dissertation, South Dakota State University
Thompson DJ, Jenks JA (2010) Dispersal movements of subadult cougars from the Black Hills: the notions of range expansion and recolonization. Ecosphere 1:1–11
Valtonen M, Palo JU, Ruokonen M, Kunnasranta M, Nyman T (2012) Spatial and temporal variation in genetic diversity of an endangered freshwater seal. Conserv Genet 13:1231–1245
Walker CW, Harveson LA, Pittman MT, Tewes ME, Honeycutt RL (2000) Microsatellite variation in two populations of mountain lions (Puma concolor) in Texas. Southwest Nat 45:196–203
Waples RS (2005) Genetic estimates of contemporary effective population size: to what time periods do the estimates apply? Mol Ecol 14:3335–3352
Waples RS (2006) A bias correction for estimates of effective population size based on linkage disequilibrium at unlinked loci. Conserv Genet 7:167–184
Waples RS, Do C (2010) Linkage disequilibrium estimates of contemporary Ne using highly variable genetic markers: a largely untapped resource for applied conservation and evolution. Evol Appl 3:244–262
Waples RS, Teel DJ (1990) Conservation genetics of pacific salmon: temporal changes in allele frequency. Conserv Biol 4:144–156
Whitehouse AM, Harley EH (2001) Post-bottleneck genetic diversity of elephant populations in South Africa, revealed using microsatellite analysis. Mol Ecol 10:2139–2149
Wilkens D (2014) Ecology of mountain lions (Puma concolor) in the North Dakota Badlands: population dynamics and prey use. Thesis, South Dakota State University
Williams C, Blejwas K, Johnston J, Jaeger M (2003) Temporal genetic variation in a coyote (Canis latrans) population experiencing high turnover. J Mammal 84:177–184
Wright S (1978) Evolution and the genetics of populations, Vol 4: Variability within and among natural populations. University of Chicago Press, Chicago
Young SP, Goldman EA (1946) The Puma: mysterious American cat. Dover Publications, Inc., New York
Acknowledgments
Financial support for this project was funded by Federal Aid to Wildlife Restoration under project W-75-R (Study No. 7545) administered through the South Dakota Department of Game, Fish and Parks. We thank J. Alexander, S. Griffin, J. Broecher, L. Meduna, A. Rhoe, B. Tycz, M. Gottlob, B. Parr, L. Wiechmann, J. Kanta, A. Lindbloom, T. Berdan, J. Felio, K. Cudmore, and T. Solano for assistance with cougar captures. We thank all residence and property owners throughout the Black Hills for allowing access to their property. We thank K. Morgan, C. Engkjer, K. McKelvey, and J. Tucker for assistance with genetics analysis. We thank the Wyoming Game and Fish and North Dakota Game and Fish departments for providing additional DNA samples. We also thank the Civil Air Patrol pilots G. Hewett, L. Becht, R. Deweese, C. Goodrich, and M. Beason for assistance with flight time and aerial telemetry.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Juarez, R.L., Schwartz, M.K., Pilgrim, K.L. et al. Assessing temporal genetic variation in a cougar population: influence of harvest and neighboring populations. Conserv Genet 17, 379–388 (2016). https://doi.org/10.1007/s10592-015-0790-5
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-015-0790-5