Abstract
The genetic structure of a population can provide important insights into animal movements at varying geographical scales. Individual and social behaviors, such as philopatry and dispersal, affect patterns of relatedness, age and sex structure, shaping the local genetic structure of populations. However, these fine scale patterns may not be detected within broader population genetic structure. Using SNP genotyping for pairwise relatedness estimates, we investigated the spatial and genetic structuring of 141 red foxes within south-central Sweden at two scales. First, we looked at broad scale population structuring among red foxes at the regional level. We then estimated pairwise relatedness values to evaluate the spatial and genetic structure of male, female and mixed sex pairs for patterns of philopatry and dispersal at a more localized scale. We found limited genetic differentiation at the regional scale. However, local investigations revealed patterns of female philopatry and male biased dispersal. There were significant differences in pairwise geographic distances between highly related same sex pairs with the average distance between related males, 37.8 km, being six times farther than that of related females, averaging 6.3 km. In summary, the low levels of genetic differentiation found in this study illustrates the mobility and dispersal ability of red foxes across scales. However, relatedness plays a strong role in the spatial organization of red foxes locally, ultimately contributing to male biased dispersal patterns.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Dispersal is a key driver of gene flow across space, linking populations both demographically and genetically (Clobert et al. 2001). Patterns of dispersal or connectivity arising from physical landscape features (Wright 1943; Balkenhol et al. 2015) or from behavioral traits such as natal philopatry or territoriality, can limit gene flow between groups of organisms, altering the genetic structure of a population (Macdonald 1983; Bowler and Benton 2005; van Dijk et al. 2015). Local gene flow patterns, where short distance movements, philopatric behavior or inbreeding avoidance influence patterns of relatedness, age and sex structure, can shape the genetic structure of populations considerably. However, such fine scale patterns may not be detected within broader population genetic structure (Rousset 2001).
Various genetic methods can provide important insights into animal movements at varying geographical scales by describing how individual behaviors, such as philopatry and dispersal, translates into gene flow (Prugnolle and De Meeus 2002). Advancements in the development of useful, low cost genetic markers (e.g. single nucleotide polymorphisms, SNPs), in combination with rigorous inference methods, offer an expanded framework for interpreting fine scale patterns, such as familial relationships, from genetic markers (Beaumont and Rannala 2004; Latch et al. 2006; Stauffer 2007; Stephens and Balding 2009; Banks and Peakall 2012). Similarly, the use of GPS telemetry for studying animal movements has dramatically increased, enhancing our ability to collect detailed movement data from wild and elusive, free-ranging animals (Cagnacci et al. 2010). Together, these tools can be used for investigation into factors jointly affecting animal dispersal, spatial and social organization, population dynamics and genetic structuring (Morales et al. 2010).
For many species, dispersal movements tend to be relatively short and long distance dispersal, while notable, is uncommon (Nathan 2005; Jordano 2017). In mammals, dispersal is often male-biased with females exhibiting philopatric behavior (Greenwood 1980). When examined at a finer spatial scale, this sex-biased dispersal can create different spatial patterns of relatedness between the sexes, with local relatedness higher among members of the more philopatric sex (Prugnolle and De Meeus 2002). In such philopatric systems, where closely related individuals settle within or close to the home range of their parents, high relatedness among individuals can lead to the emergence of kin structured populations (Kitchen et al. 2005; Croteau et al. 2010; Quaglietta et al. 2013).
The red fox, Vulpes vulpes, is a mid-sized canid with highly variable spatial and social behavior (von Schantz 1981; Doncaster and Macdonald 1991; Cavallini 1996). Red foxes have a geographic range that spans most of the northern hemisphere (Larivière and Pasitschniak-Arts 1996) and the potential to disperse over long distances (Colson et al. 2017; Walton et al. 2018). In most areas, red foxes form monogamous pairs or small family groups (von Schantz 1981; Cavallini 1996). These typically consist of a dominant, breeding pair and subordinate individuals, which are believed to be the offspring of one or both of the dominants (von Schantz 1981; Baker et al. 2004). However, kinship is rarely confirmed, and red foxes have been shown to exhibit polygynandry, multiple paternity and extra pair copulations (Baker et al. 2004) adding potential complexity to the kin structure of family groups.
Previous population genetic studies indicate that red foxes can exhibit a high degree of gene flow due to their dispersal ability (Gachot-Neveu et al. 2009; Teacher et al. 2011; Mullins et al. 2014; Atterby et al. 2015; Norén et al. 2015). However, studies of localized genetic patterns relating to philopatric behavior and dispersal between sub-populations is largely lacking for this species (but see Gachot-Neveu et al. 2009; Whiteside et al. 2011). Delineating patterns of familial clustering can ultimately lead to a better understanding of red fox social organization, spatial behavior and dispersal, as well as the mechanisms underlying such patterns, facilitating our understanding of the evolution of dispersal and group living in social mammals (Whiteside et al. 2011). Such data also lends itself to informing the ecology and management of red foxes (Quinn et al 2019) and the design and implementation of disease control programs such as oral vaccination programs (Zecchin et al. 2019). For instance, integrating genetic and movement data into management and/or monitoring programs can be helpful to understand dispersal and highlight corridors or barriers for movement and gene flow, which can aide in optimizing species control efforts or predicting the spread of diseases. This is particularly relevant to red foxes as they are considered a primary reservoir species for diseases such as rabies and canine distemper virus and as vectors for the spread of numerous other zoonotic parasites and diseases (Holmala and Kauhala 2006; Vervaeke et al. 2006; Atterby et al. 2015). Thus, such knowledge may lend vital information to wildlife managers and health authorities aiming to manage such an ecologically dynamic generalist species.
Here, we investigated the spatial and genetic structuring of red foxes within south-central Sweden at two different scales. First, we looked at regional scale genetic structuring among red foxes, where we expected gene flow to be high with panmixia occurring over a broad scale due to the high mobility of this species (Norén et al. 2015; Munro and Burg, 2017). We then evaluated the spatial and genetic structure of male, female and mixed sex pairs to compare patterns of philopatry and dispersal at a more localized scale. Here, we predicted that sex biased dispersal due to philopatric female behavior should lead to patterns of kin structuring within a population. The low levels of genetic differentiation found in this study illustrates high mobility and dispersal ability of red foxes across scales. However, relatedness plays a strong role in the spatial organization of red foxes locally, ultimately contributing to male biased dispersal patterns.
Material and methods
Study area and DNA sampling
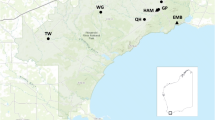
This study was conducted within a 13,000 km2 region in south-central Sweden (58°N-60°N; Fig. 1). Within this region, the northernmost fox trapping sites around Hedemora (60°16′N-15°59′E) and Grimsö (59°40′N-15°25′E), are separated from our local study area around Kolmården (58°40′N-16°22′E) by approximately 200 km (Hedemora is located ~ 70 km north of Grimsö and Grimsö is ~ 130 km north of Kolmården). The northern areas consists of a transitional border zone between boreonemoral forests in the south and boreal forests in the north, where the landscape is generally flat with mean altitude rising from 75 m a.s.l. in the south to 180 m a.s.l. in the north. Daily mean temperatures average 15 °C in summer to − 5 °C in winter. The ground is generally snow covered from late December to March. Locally, the study area around Kolmården encompasses approximately 1800 km2 within Södermanland County (see inset Fig. 1) representing a fragmented mosaic of productive agriculture areas, boreonemoral forests and scattered human settlements, representing a rural, anthropogenically modified landscape. Daily mean temperatures range from 20 °C in summer to − 5 °C in winter. Snow covers the ground irregularly from December to March.
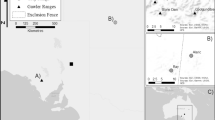
Spatial genetic structure of red foxes visualized by spatial principal component analyses (sPCA). The colors in the maps show the first three principal components only. Colors indicate genetic similarity (similar colors are more genetically similar individuals whereas contrasting colors are more genetically dissimilar) with panels showing different subsets of the population. Panel a represents all individuals included in the study. Panels b and c show local structuring of females and males from the Kolmården area, respectively. Overall, the population displays a high level of mixing, with limited structure, but, the overlapping similar color groupings in panel B indicate females have more localized structure than males. (Color figure online)
We collected genetic material (hair, tissue or blood) from red foxes that were live captured as part of a Global Positioning System (GPS) telemetry study during 2012–2019 (see Walton et al. 2017 for details of the capture and handling procedures). We also opportunistically collected red fox scat and tissue samples from deceased, unmarked red foxes found during field monitoring of GPS collared foxes. All live captured foxes were sexed, weighed and marked with plastic ear tags (Rototags, Dalton Supplies, Ltd.). Age of foxes was approximated, assuming a birth date of 15th April (Englund 1970), as either pup (< 6 months), sub-adult (6 months to < 1 year) or adult (> 1 year), using the date of capture in combination with the amount of tooth wear and coloration. All capture and handling protocols were approved by the Swedish Environmental Protection Board and the Swedish Animal Ethics Committee (permit numbers NV-03459–11, DNR 70–12, DNR 58–15, DNR 13–47).
DNA extraction and SNP genotyping
DNA from tissue, blood and hair was extracted using a Qiagen Symphony robot and the QiaSymphony DNA mini kit and QiaSymphony DNA investigator kit, respectively. Feces were manually extracted using Zymo's soil/fecal DNA kit. This kit was chosen after a review of eight different methods/kits (Supp. Table S1).
We genotyped individuals on a Fluidigm® Biomark™ microarray platform using the 96.96 dynamic array. The SNP (Single Nucleotide Polymorphism) assays were based on loci shown polymorphic in fox on Illumina’s® CanineHD™ 170 K Whole-Genome Genotyping BeadChip (marker references listed in Supp. Table S2). For the marker ascertainment we used a population of foxes from across Norway (N = 24) and Sweden (N = 24). The selected somatic markers were not deviating from Hardy Weinberg equilibria expectations and had a minor allele frequency exceeding 0.3. Additionally, we only included markers that clustered well for several DNA sources (tissue, blood, hair, and feces). In addition to 91 somatic markers, we used Sanger sequencing to design three markers that only amplified the Y- chromosome, and two mitochondrial markers for species recognition, for a total of 96 SNP markers.
All genotyping runs included three NTC (No Template Controls) and reference samples. Samples were run in duplicates for quality control (see Table 2). Samples that showed an amplification rate below 90% were discarded. For samples that amplified well (> 99%) the error rate fell below 1%. Basic marker statistics are presented in Supp. Table S2.
Genetic and spatial analyses
For analysis of spatial population structure and calculating pairwise relatedness distances described below, we used the last position of each GPS collared animal. All red foxes were captured in one of 13 baited wooden box traps in our study area. Given the identical trapping location for many foxes (range 1–15 individual foxes captured per trap) we chose to use the last GPS position as this provided a better representation of the area occupied by the animal (i.e. its home range) than the original trapping location. Additionally, it also allowed for animal movement over time, such as dispersal, to occur, whereas trapping location did not. For red foxes that were trapped but did not receive a collar (i.e. predominantly young foxes under < 6 months not meeting the necessary weight requirements), we used the GPS coordinates of the wooden box trap where the individual was captured and subsequently released to represent sample location. For scat or tissue, we used the coordinates taken directly at the site of field collection using a hand held GPS unit. For analyses at the regional and local levels we included only adult and subadult foxes, removing pup samples (n = 15) as pups still occupied their natal range, regardless of sex, potentially confounding results. For analyses of fine scale genetic structuring and patterns of kinship within a more localized area, we also removed individuals where we were not able to identify sex (scat samples, n = 4). In total, 126 red foxes contributed to the regional analyses and 103 to the local analyses.
To explore the genetic structure of red foxes we ran Genepop 4.2 (Raymond and Rousset 1995; Rousset 2008). In Genepop, we calculated basic marker statistics, such as expected heterozygosity (He), observed heterozygosity (Ho) and deviations from Hardy Weinberg equilibrium (HWE) (Supp. Table S2).
We then ran a spatial principal component analyses (sPCA) in the R package Adegenet (Jombart 2008) to visualize genetic differentiation geographically, both regionally and locally. The sPCA was run on the full dataset, as well as for subsets of male and female individuals from the southernmost trapping area around Kolmården, Sweden, to explore how distribution of genetic variation varied with sex locally. The K Nearest Neighbor algorithm was used with two neighbors for all runs (Jombart and Collins 2015). As the information content of the principal components (PCs) of each subset differed slightly, we included different numbers of components. The number of global PCs were 5, 5 and 4 and local PCs 3, 3 and 3 for all foxes, Kolmården females only, and Kolmården males only.
To determine kinship among individuals, we determined the relatedness coefficient (r) using R.3.3.2 (R-Core-Team 2018) and the R package ‘Related’(Wang 2011; Pew et al. 2015) between all individual pairs going back two generations. The r-value is estimated from similarities in the number of shared identical alleles, in relation to the population level allele frequencies. This allowed us to infer the most probable relationship between individuals, using a cutoff of 0.65. A first-degree pair is expected to show an r-value of approximately 0.5. Such relationships include the individual’s parents, full siblings, and offspring. A second degree relative will show an r-value of approximately 0.25, which includes the individual’s grandparents, grandchildren, aunts, uncles, nephews, nieces or half-siblings.
To determine if kin were spatially clustered at the local scale, we evaluated the pairwise mean geographic distances among all pairs of first degree relatives (r ≥ 0.40) for the different sex combinations: female-female, opposite sex, and male-male, using Welch two sample t-tests. We further examined how pairwise distances varied according to differing degrees of relatedness. For this, we categorized all red fox individuals according to their relatedness values where values of 0–0.10 and 0.10–0.20 indicated unrelated or distantly related individuals, 0.21–0.40 intermediate or second-degree relatives and ≥ 0.40 to be first-degree relatives.
Results
Genetic material collected from 180 red fox samples yielded 141 individual genotypes (54 females, 82 males, 5 unknown sex) of which 91 (32 females and 59 males) were fitted with GPS collars (Table 1). The mean amplification success exceeded 99% for tissue and blood extractions, whereas hair was lower at 96%. Feces showed highly variable amplification success, likely correlated to large differences in the degree of degradation in the field before the DNA could be collected. Error rates were low, but increased for samples with low amplification success. We therefore only included samples with an amplification success of 75% or more (Table 2). This allowed us to confidently separate individuals while simultaneously avoiding the inclusion of unique genotypes caused by genotyping errors (see Creel et al. 2003). Overall, the SNP markers showed high variability, with a mean minor allele frequency of 0.35 and the majority of them conformed to Hardy–Weinberg Equilibrium (HWE) expectations (Supp.Table S2). Before Bonferroni correction, 11 markers fell outside HWE, and after Bonferroni correction two markers remained significantly outside HWE. We kept all markers in the analyses as the analyses were not affected in any noticeable way with or without them.
Our sPCA analyses showed low levels of genetic differentiation when analyzing all individuals together at the regional level (Fig. 1a). Here, genetic similarity is indicated by similar colors and contrasting colors are more genetically dissimilar. Locally, the sPCA results of male and female groups showed females configured in tighter clustering of similar colors, whereas males exhibited more diffuse spatial structure (Fig. 1b and 1c). Geographic distances supported this patterning as well, with female kin being more spatially clustered at the local scale than males. Here, highly related female pairs showed the shortest average pairwise distances, whereas highly related male pairs showed the largest average distances. Opposite sex pairs showed intermediate distances (Fig. 2; Table 3). A t-test identified all pair types to be significantly different from all other pair types, with female pairs and male pairs being most different (t = 3.3661, df = 43.132, p-value = 0.00161). The difference in mean distance between opposite sex pairs and female-female pairs (t = -2.1794, df = 79.144, p-value = 0.03227) and opposite sex pairs and male-male pairs (t = 2.2642, df = 48.766, p-value = 0.02805, Table 3) were also significant. Examining the pairwise distances for all pair types according to their categorized relatedness values showed a similar pattern across all relatedness categories and all pair types (Table 4; Supp. Fig. S1).
Discussion
Our study highlights differing patterns of red fox genetic structure depending on spatial scale and sex. The limited genetic differentiation seen in the sPCA of red foxes at the regional level indicates high levels of gene flow, whereas patterns of relatedness revealed intersexual differences in spatial genetic structure at a local scale. Red foxes are a widespread omnivore showing high genetic diversity throughout their European range (Gachot-Neveu et al. 2009; Teacher et al. 2011; Kutschera et al. 2013; Galov et al. 2014; Atterby et al. 2015; Norén et al. 2015). The low levels of genetic differentiation found at the regional level follow expectations that red foxes are a mobile species and dispersal causes gene flow across large spatial scales (Peakall et al. 2003). Our results also do not indicate physical barriers to red fox movements within the regional scale of our study (Norén et al. 2015). Similarly, other canid species exhibiting dispersal over long distances also show limited phylogeographic structuring (e.g. grey wolf (Canis lupus), coyote (Canis latrans) and arctic fox (Vulpes lagopus) (Lehman and Wayne 1991; Vilà et al. 1999; Lai et al. 2017).
Locally, the geographic distances between highly related individuals showed spatial patterns indicating significant philopatric behavior among related females with larger geographic separation between related male foxes. The average distance between related male pairs, approximately 38 km, was over six times larger than the distance between related female pairs (6 km). This confirms our prediction that highly related females show kin clustering by occurring in closer proximity to related females compared to mixed sex or male red fox pairs. As restrictions to movement did not appear evident from the sPCA results at the regional scale, local scale movement patterns are probably mostly influenced by social barriers (Macdonald 1983).
In polygynous mating systems, such as that in red foxes, inbreeding avoidance (Pusey and Wolf 1996) may contribute to a sex biased dispersal pattern (Johnson and Gaines 1990). Dispersal may be a prerequisite for reproduction or a breeding territory vacancy (Greenwood 1980). Additionally, philopatric behavior, where offspring are allowed to settle and breed within or adjacent to their natal range, has been shown to create clustering of related individuals in the landscape (Benton and Bowler 2012). Evolutionarily, red fox females are thought to benefit more than males from being among kin through access to breeding sites (e.g. underground den systems), cooperative social interactions, knowledge of resource areas, or through inheritance of the home range (von Schantz 1981; Macdonald 1983; Baker et al. 1998). Thus, the genetic sub-structuring seen at the local scale is likely driven by male biased dispersal and female philopatry altering gene flow between groups, subsequently shaping local genetic patterns.
Mechanisms such as male biased dispersal and female philopatry do not work independently however, and genetic patterns can be expected to differ across populations where different dispersal strategies and demographic components such as survival and natality are at play (Kamler et al. 2013). For instance, management actions leading to high harvest or population control rates can create artificial source-sink systems (Pulliam 1988; McCullough 1996), that may disrupt the spatial dynamics and social organization of local populations (Maletzke et al. 2014). For example, locally high population turnover rates, either naturally occurring or as a result of anthropogenic actions can result in both increased philopatry and increased dispersal rates (Robinson et al. 2008). In other cases, disruption of family groups due to harvest could promote the association of unrelated females (Williams et al. 2008; Miller et al. 2010), resulting in an altered genetic structure. However, limited information exists on how carnivore populations respond to harvest or the effects of intense harvest on dispersal and social group dynamics, despite the relative value of such information for management and in modelling or managing outbreaks of disease (Robinson et al. 2008, Grear et al. 2010, Maletzke et al. 2014). Identifying relatedness underlying broader landscape genetic structure can offer important insights into how dispersal patterns can contribute to genetic structure locally. An understanding of life-history patterns, including dispersal and social structure is vital to understand the effects of management actions such as culling programs. It also has implications for models of disease transmission, and the need for consideration of social behaviors in conservation and management (Festa-Bianchet and Apollonio 2013).
This study highlights the use of SNP genotyping to provide detailed information on the relationships between individuals, in this case allowing for detection of different patterns of gene flow at the population and individual kin levels. The genetically identified patterns found in this study fit well with dispersal patterns found in other red fox movement studies (Storm et al. 1976; Allen and Sargeant 1993) and for mammals in general (Greenwood 1980). Additionally, a high degree of relatedness among individuals within a population can confound or may not be evident within broader population structure (Anderson et al. 2010; Norman et al. 2017). Here, our results indicate that some of the underlying genetic structure at the regional level may be due to familial clustering.
The power of genetic markers will likely prove valuable for future studies, providing new avenues for combining genetic methods with movement data and new tools for management. Studies using genetic methods should however, consider the scale they wish to address, as demographic and behavioral processes can operate at a range of scales to generate genetic structure (Anderson et al. 2010; van Dijk et al. 2015). For instance, a high level of short distance dispersal events will affect local spatial patterns of genetic differentiation (Rousset 2001) more than a low level of long dispersals (Nathan 2005). Thus, it is important to consider the scale at which such movements take place and the behavioral aspects that influence such movements.
Furthermore, populations are not closed. If we assume that our regional population of red foxes is a continuous population (as our regional results might suggest), it is noteworthy that fine scale genetic structuring can occur locally by social mechanisms not linked to the movement ability of the species. This suggests that movement capacity is not a prerequisite for fine scale genetic structuring to occur locally (Campbell and Strobeck 2006; Quaglietta et al. 2013) and that red foxes can be restricted by social barriers. However, limited attention has been given to examining intraspecific patterns of genetic structure at different spatial scales (Anderson et al. 2010), particularly for carnivores (Gompper 1996; Oehler and Litvaitis 1996). Many species in the order Carnivora show great flexibility in their social systems (Macdonald 1983) which can influence population genetic structuring at a very fine scale. Distinguishing between population structure and the underlying fine-scale social and kinship patterns that affect population genetic structure can ultimately lead to a more thorough understanding of the spatial, social and population dynamics of a species. Yet, these are still among some of the least understood attributes of both individual animals and populations (Sutherland et al. 2000; Nathan 2001; Kokko and López-Sepulcre 2006).
Two correlates of carnivore spacing behavior are sex and social system (Clobert et al. 2001). By combining animal movement data with patterns of genetic relatedness and allele frequencies in a population, we explored how dispersal contributes to the genetic and spatial organization of red foxes at regional and local scales. In summary, the levels of genetic similarity found in this study illustrates the dispersal ability of red foxes across scales, where locally, relatedness plays a strong role in the spatial organization of red foxes ultimately contributing to male biased dispersal patterns and restricted female movements. However, this fine scale structuring likely occurs by social mechanisms not related to the movement capacity of the species.
Data availability
Genetic data from this study is available as supplementary material. GPS data is available by request from the authors.
References
Allen SH, Sargeant AB (1993) Dispersal patterns of red foxes relative to population density. J of Wildl Manag 57:526–533
Anderson CD, Epperson BK, Fortin MJ, Holderegger R, James PM, Rosenberg MS, Scribner KT, Spear S (2010) Considering spatial and temporal scale in landscape-genetic studies of gene flow. Mol Ecol 19(17):3565–3575
Atterby H, Allnutt TR, MacNicoll AD, Jones EP, Smith GC (2015) Population genetic structure of the red fox (Vulpes vulpes) in the UK. Mammal Research 60(1):9–19
Baker PJ, Robertson CP, Funk SM, Harris S (1998) Potential fitness benefits of group living in the red fox. Vulpes vulpes Animal Behaviour 56(6):1411–1424
Baker PJ, Funk SM, Bruford MW, Harris S (2004) Polygynandry in a red fox population: implications for the evolution of group living in canids? Behav Ecol 15(5):766–778
Balkenhol N, Cushman S, Storfer A, Waits L (2015) Landscape genetics: concepts, methods, applications. Wiley, New Jersey, p 288
Banks SC, Peakall R (2012) Genetic spatial autocorrelation can readily detect sex-biased dispersal. Mol Ecol 21(9):2092–2105
Beaumont MA, Rannala B (2004) The Bayesian revolution in genetics. Nat Rev Genet 5(4):251–261
Benton TG, Bowler DE (2012) Linking dispersal to spatial dynamics. In: Benton TG, Bowler DE (eds) Dispersal ecology and evolution. Oxford University Press, Oxford, pp 251–265
Bowler DE, Benton TG (2005) Causes and consequences of animal dispersal strategies: relating individual behaviour to spatial dynamics. Biol Rev 80(2):205–225
Cagnacci F, Boitani L, Powell RA, Boyce MS (2010) Animal ecology meets GPS-based radiotelemetry: a perfect storm of opportunities and challenges. Phil. Trans. R. Soc. B 365(1550):2157–2162
Campbell V, Strobeck C (2006) Fine-scale genetic structure and dispersal in Canada lynx (Lynx canadensis) within Alberta Canada. Can J Zool 84(8):1112–1119
Cavallini P (1996) Variation in the social system of the red fox. Ethol Ecol Evol 8(4):323–342
Clobert J, Danchin E, Dhondt AA, Nichols J (2001) Dispersal. Oxford University Press, Oxford
Colson K, Smith JD, Hundertmark KJ (2017) St. Matthew Island colonized through multiple long-distance red fox (Vulpes vulpes) dispersal events. Can J Zool 95(8):607–609
Creel S, Spong G, Sands JL, Rotella J, Zeigle J, Joe L, Murphy KM, Smith D (2003) Population size estimation in Yellowstone wolves with error-prone noninvasive microsatellite genotypes. Mol Ecol 12(7):2003–2009
Croteau EK, Heist EJ, Nielsen CK (2010) Fine-scale population structure and sex-biased dispersal in bobcats (Lynx rufus) from southern Illinois. Can J Zool 88(6):536–545
Doncaster CP, Macdonald DW (1991) Drifting territoriality in the red fox Vulpes vulpes. J Anim Ecol 60:423–439
Festa-Bianchet M, Apollonio M (2013) Animal Behavior and Wildlife Conservation. Island Press, Washington
Gachot-Neveu H, Lefevre P, Roeder J-J, Henry C, Poulle M-L (2009) Genetic detection of sex-biased and age-biased dispersal in a population of wild carnivore, the red fox. Vulpes vulpes Zoological Science 26(2):145–152
Galov A, Sindičić M, Andreanszky T, Čurković S, Dežđek D, Slavica A, Hartl GB, Krueger B (2014) High genetic diversity and low population structure in red foxes (Vulpes vulpes) from Croatia. Mammalian Biology-Zeitschrift für Säugetierkunde 79(1):77–80
Gompper M (1996) Genetic relatedness among individuals within carnivore societies. In: Gittleman JL (ed) Carnivore behavior, ecology, and evolution 2. Cornell University Press, Ithaca, New York, pp 429–452
Grear DA, Samuel MD, Scribner KT, Weckworth BV, Langenberg JA (2010) Influence of genetic relatedness and spatial proximity on chronic wasting disease infection among female white-tailed deer. J Appl Ecol 47(3):532–540
Greenwood PJ (1980) Mating systems, philopatry and dispersal in birds and mammals. Anim Behav 28(4):1140–1162
Holmala K, Kauhala K (2006) Ecology of wildlife rabies in Europe. Mammal Review 36:17–36
Johnson ML, Gaines MS (1990) Evolution of dispersal: theoretical models and empirical tests using birds and mammals. Annu Rev Ecol Syst 21(1):449–480
Jombart T (2008) adegenet: a R package for the multivariate analysis of genetic markers. Bioinformatics 24(11):1403–1405
Jombart T, Collins C. 2015. A tutorial for discriminant analysis of principal components (DAPC) using adegenet 2.0. 0. London: Imperial College London, MRC Centre for Outbreak Analysis and Modelling.
Jordano P (2017) What is long-distance dispersal? And a taxonomy of dispersal events. J Ecol 105(1):75–84
Kamler JF, Gray MM, Oh A, Macdonald DW (2013) Genetic structure, spatial organization, and dispersal in two populations of bat-eared foxes. Ecology and Evolution 3(9):2892–2902
Kitchen AM, Gese EM, Waits LP, Karki SM, Schauster ER (2005) Genetic and spatial structure within a swift fox population. J Anim Ecol 74(6):1173–1181
Kokko H, López-Sepulcre A (2006) From individual dispersal to species ranges: perspectives for a changing world. Science 313(5788):789–791
Kutschera VE, Lecomte N, Janke A, Selva N, Sokolov AA, Haun T, Steyer K, Nowak C, Hailer F (2013) A range-wide synthesis and timeline for phylogeographic events in the red fox (Vulpes vulpes). BMC Evol Biol 13(1):114
Lai S, Bêty J, Berteaux D (2017) Movement tactics of a mobile predator in a meta-ecosystem with fluctuating resources: the arctic fox in the High Arctic. Oikos 126(7):937–947
Larivière S, Pasitschniak-Arts M (1996) Vulpes vulpes. Mammalian Species 537:1–11
Latch EK, Dharmarajan G, Glaubitz JC, Rhodes OE (2006) Relative performance of Bayesian clustering software for inferring population substructure and individual assignment at low levels of population differentiation. Conserv Genet 7(2):295–302
Lehman N, Wayne RK (1991) Analysis of coyote mitochondrial DNA genotype frequencies: estimation of the effective number of alleles. Genetics 128(2):405–416
Macdonald DW (1983) The ecology of carnivore social behaviour. Nature 301(5899):379
Maletzke BT, Wielgus R, Koehler GM, Swanson M, Cooley H, Alldredge JR (2014) Effects of hunting on cougar spatial organization. Ecology and Evolution 4(11):2178–2185
McCullough DR (1996) Spatially structured populations and harvest theory. J Wildl Manag 60:1–9
Miller BF, De Young RW, Campbell TA, Laseter BR, Ford WM, Miller KV (2010) Fine-scale genetic and social structuring in a central Appalachian white-tailed deer herd. J Mammal 91(3):681–689
Morales JM, Moorcroft PR, Matthiopoulos J, Frair JL, Kie JG, Powell RA, Merrill EH, Haydon DT (2010) Building the bridge between animal movement and population dynamics. Philosophical Transactions of the Royal Society B: Biological Sciences 365(1550):2289–2301
Mullins J, McDevitt AD, Kowalczyk R, Ruczyńska I, Górny M, Wójcik JM (2014) The influence of habitat structure on genetic differentiation in red fox populations in north-eastern Poland. Acta Theriologica 59(3):367–376
Munro KJ, Burg TM (2017) A review of historical and contemporary processes affecting population genetic structure of Southern Ocean seabirds. Emu-Austral Ornithology 117(1):4–18
Nathan R (2001) The challenges of studying dispersal. Trends Ecol Evol 16(9):481–483
Nathan R (2005) Long-distance dispersal research: building a network of yellow brick roads. Divers Distrib 11(2):125–130
Norén K, Statham MJ, Ågren EO, Isomursu M, Flagstad Ø, Eide NE, Berg TBG, Bech-Sanderhoff L, Sacks BN (2015) Genetic footprints reveal geographic patterns of expansion in Fennoscandian red foxes. Glob Change Biol 21(9):3299–3312
Norman AJ, Stronen AV, Fuglstad G-A, Ruiz-Gonzalez A, Kindberg J, Street NR, Spong G (2017) Landscape relatedness: detecting contemporary fine-scale spatial structure in wild populations. Landscape Ecol 32(1):181–194
Oehler JD, Litvaitis JA (1996) The role of spatial scale in understanding responses of medium-sized carnivores to forest fragmentation. Can J Zool 74(11):2070–2079
Peakall R, Ruibal M, Lindenmayer DB (2003) Spatial autocorrelation analysis offers new insights into gene flow in the Australian bush rat. Rattus fuscipes Evolution 57(5):1182–1195
Pew J, Wang J, Muir P, Frasier T. 2015. related: related: an R package for analyzing pairwise relatedness data based on codominant molecular markers. R package version 1.
Prugnolle F, De Meeus T (2002) Inferring sex-biased dispersal from population genetic tools: a review. Heredity 88(3):161
Pulliam HR (1988) Sources, sinks, and population regulation. Am Nat 132(5):652–661
Pusey A, Wolf M (1996) Inbreeding avoidance in animals. Trends Ecol Evol 11(5):201–206
Quaglietta L, Fonseca VC, Hájková P, Mira A, Boitani L (2013) Fine-scale population genetic structure and short-range sex-biased dispersal in a solitary carnivore, Lutra lutra. J Mammal 94(3):561–571
Quinn CB, Alden PB, Sacks BN (2019) Noninvasive sampling reveals short-term genetic rescue in an insular red fox population. J Hered 110(5):559–576
Raymond M, Rousset F (1995) GENEPOP v.l.2: population genetics software for exact tests and ecumenicism. J Hered 86(3):248–249
R-Core-Team. 2018. R: A language and environment for statistical computing.
Robinson HS, Wielgus RB, Cooley HS, Cooley SW (2008) Sink populations in carnivore management: cougar demography and immigration in a hunted population. Ecol Appl 18(4):1028–1037
Rousset F (2001) Genetic approaches to the estimation of dispersal rates. In: Clobert J, Danchin E, Dhondt AA, Nichols JD (eds) Dispersal. Oxford University Press, Oxford, pp 18–28
Rousset F (2008) genepop’007: a complete re-implementation of the genepop software for Windows and Linux. Molecular ecology resources 8(1):103–106
Stauffer HB (2007) Contemporary Bayesian and frequentist statistical research methods for natural resource scientists. Wiley, New Jersey
Stephens M, Balding DJ (2009) Bayesian statistical methods for genetic association studies. Nat Rev Genet 10(10):681
Storm G, Andrews RD, Phillips RL, Bishop RA, Siniff DB, Tester JR (1976) Morphology, reproduction, dispersal, and mortality of midwestern red fox populations. Wildlife Monographs 49:3–82
Sutherland GD, Harestad AS, Price K, Lertzman KP (2000) Scaling of natal dispersal distances in terrestrial birds and mammals. Conservation Ecology. https://doi.org/10.5751/ES-00184-040116
Teacher AG, Thomas JA, Barnes I (2011) Modern and ancient red fox (Vulpes vulpes) in Europe show an unusual lack of geographical and temporal structuring, and differing responses within the carnivores to historical climatic change. BMC Evol Biol 11(1):214
van Dijk RE, Covas R, Doutrelant C, Spottiswoode CN, Hatchwell BJ (2015) Fine-scale genetic structure reflects sex-specific dispersal strategies in a population of sociable weavers (Philetairus socius). Mol Ecol 24(16):4296–4311
Vervaeke M, van der Giessen J, Brochier B, Losson B, Jordaens K, Verhagen R, de Lezenne CC, Teunis P (2006) Spatial spreading of Echinococcus multilocularis in Red foxes (Vulpes vulpes) across nation borders in Western Europe. Preventive Veterinary Medicine 76(3–4):137–150
Vilà C, Amorim IR, Leonard JA, Posada D, Castroviejo J, Petrucci-Fonseca F, Crandall KA, Ellegren H, Wayne RK (1999) Mitochondrial DNA phylogeography and population history of the grey wolf Canis lupus. Mol Ecol 8(12):2089–2103
von Schantz T (1981) Female cooperation, male competition, and dispersal in the red fox Vulpes vulpes. Oikos. https://doi.org/10.2307/3544073
Walton Z, Samelius G, Odden M, Willebrand T (2017) Variation in home range size of red foxes Vulpes vulpes along a gradient of productivity and human landscape alteration. PloS one 12(4):e0175291
Walton Z, Samelius G, Odden M, Willebrand T (2018) Long-distance dispersal in red foxes Vulpes vulpes revealed by GPS tracking. Eur J Wildl Res 64(6):64
Wang J (2011) COANCESTRY: a program for simulating, estimating and analysing relatedness and inbreeding coefficients. Mol Ecol Resour 11(1):141–145
Whiteside HM, Dawson DA, Soulsbury CD, Harris S (2011) Mother knows best: dominant females determine offspring dispersal in red foxes (Vulpes vulpes). PLoS ONE 6(7):e22145
Williams SC, DeNicola AJ, Ortega IM (2008) Behavioral responses of white-tailed deer subjected to lethal management. Can J Zool 86:1358–1366
Wright S (1943) Isolation by distance. Genetics 28(2):114
Zecchin B, De Nardi M, Nouvellet P, Vernesi C, Babbucci M, Crestanello B, Donnelly CA (2019) Genetic and spatial characterization of the red fox (Vulpes vulpes) population in the area stretching between the Eastern and Dinaric Alps and its relationship with rabies and canine distemper dynamics. PLoS ONE 14(3):e0213515
Acknowledgements
We gratefully acknowledge Bo Söderberg, Eric Ringaby, Lars Jäderberg, and Kent Sköld for their expert assistance in field capturing and collaring of red foxes. We additionally owe a debt of gratitude to Helena Königsson for her invaluable expertise and laboratory analyses. The Swedish Environmental Protection Board and the Swedish Hunters Association provided funding for this study.
Funding
Open Access funding provided by Inland Norway University Of Applied Sciences.
Author information
Authors and Affiliations
Contributions
ZW, MH, KO and GS contributed to the study conception and design. Material preparation, data collection and analysis were performed by ZW, GS, MH, AN. Funding for this study was acquired by TW, GS, MO, KO, GS. The first draft of the manuscript was written by ZW and MH and all authors commented on and revised previous versions of the manuscript. All authors read and approved the final manuscript
Corresponding author
Ethics declarations
Conflict of interest
The authors declare that they have no conflict of interest.
Research using human and animal consent
All animal capture and handling protocols were approved by the Swedish Environmental Protection Board and the Swedish Animal Ethics Committee (permit numbers NV-03459–11, DNR 70–12, DNR 58–15, DNR 13–47).
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Supplementary Information
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Walton, Z., Hagenlund, M., Østbye, K. et al. Moving far, staying close: red fox dispersal patterns revealed by SNP genotyping. Conserv Genet 22, 249–257 (2021). https://doi.org/10.1007/s10592-021-01332-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10592-021-01332-7