Abstract
The objective of this study was to evaluate the genotype by environment interaction using the additive main effects and multiplicative interaction model for seeds yield of pea cultivars grown in Poland. Twelve pea (Pisum sativum L.) cultivars: Bohun, Boruta, Cysterski, Ezop, Kavalir, Lasso, Medal, Santana, Tarchalska, Terno, Wenus and Zekon were evaluated in 20 environments (ten locations in 2 years). The experiment was laid out as randomized complete block design with three replicates. Seeds yield ranged from 26.10 dt ha−1 (for Wenus in Radostowo 2011) to 79.73 dt ha−1 (for Lasso in Słupia 2010), with an average of 50.70 dt ha−1. AMMI analyses revealed significant genotype and environmental effects as well as genotype-by-environment interaction with respect to seeds yield. In the analysis of variance, 89.19% of the total seeds yield variation was explained by environment, 1.65% by differences between genotypes, and 8.33% by GE interaction. The cultivar Terno is the highest stability. The cultivar Tarchalska is recommended for further inclusion in the breeding program because its stability and the highest averages of seeds yield.
Similar content being viewed by others
Explore related subjects
Discover the latest articles, news and stories from top researchers in related subjects.Avoid common mistakes on your manuscript.
Introduction
Poland has favourable agro-ecological conditions for the cultivation of peas for seeds. This species is characterized by high fertility, but at the same time significant variability of yield level. It also has a high protein content in seeds and the ability to bind atmospheric nitrogen, which is of both ecological and economic importance. Peas also play an important role in crop rotation as a plant that interrupts the frequent succession of cereals. Due to its short growing season it is a valuable forecrop for winter plant species. The relatively deep root system of peas and the large mass of post-harvest residues enrich the soil with nitrogen and organic matter. Peas also play an important role in the plant production systems where chemicals are not used or only used to a limited extent. Pea seeds can be used as human food and animal feed. In some countries, pea seeds are almost entirely used as a component of the human diet or are mainly used as a raw material for the production of high-protein concentrates. According to Bilalis et al. (2015), Davidsson et al. (2001), due to their nutritional value, peas may be considered as an alternative source of protein for soya in the countries where it is not a natural species or where soya beans of that species cannot be used due to allergic reactions or intolerance. Pea seeds contain about 22–24% total protein, 1.5% fat, 55% nitrogen-free extract compounds, and 6–8% crude fibre. They are an important source of lysine amino acid, whose content in 1 kg of seeds is 3–4 times higher than in cereal grains.
Pea is often described as a species adapted to a wide type of environmental conditions (e.g., Annicchiarico and Iannucci 2002) many studies have showed a significant influence of environment and genotype-by-environment (GE) interaction on phenotypic performance of seeds yield and other agronomically important traits (Moot and McNeil 1995; Acikgoz et al. 2009; Tolessa et al. 2013). An important goal of pea breeding has always been to maximize productivity by increasing seed yield. Obtaining as much knowledge as possible about the inheritance of the seed yield will allow acceleration of the genetic progress of this species. This trait is very complex and its expression depends on factors such as: genetics, environment and GE (Tan et al. 2012). The genotype response to diverse environmental conditions during the growing season determines the seed yield.
Among the main methods used to evaluate GE interaction, the following can be highlighted: (1) Eberhart–Russel (1966); (2) additive main effects and multiplicative interaction (AMMI) (Gollob 1968; Gabriel 1971; Mandel 1971; Gauch 1988, 2006; Zobel et al. 1988; van Eeuwijk 1995; Dias and Krzanowski 2003); and (3) mixed model (Henderson 1975; Smith et al. 2005; Resende 2007; Piepho et al. 2008). The literature shows several studies on the use of these three methods to assess GE interaction in different crops (Piepho 1994; Annicchiarico 1997; Yang et al. 2009; Malosetti et al. 2013).
The objective of this study was to assess genotype by environment interaction for seeds yield in pea (Pisum sativum L.) grown in Poland by the AMMI model. The AMMI method combines the traditional analysis of variance for the main effects of genotype and environment as well as the principal component analysis (PCA) with multiplicative parameters into single analysis (Zobel et al. 1988).
Materials and methods
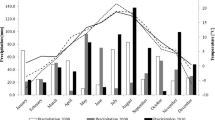
Material for the studies covered 12 pea (Pisum sativum L.) cultivars: Bohun, Boruta, Cysterski, Ezop, Kavalir, Lasso, Medal, Santana, Tarchalska, Terno, Wenus and Zekon. All 12 cultivars were in the official catalogue of varieties—The Polish National List (NLI) published by The Research Centre for Cultivar Testing (COBORU) at 2011. Field experiments were carried out over two years: 2010–2011 at ten locations: Bezek (51° 12′ 06″ N, 23° 16′ 06″ E), Chrząstowo (53° 09′ 52″ N, 17° 35′ 02″ E), Cicibór Duży (52° 04′ 39″ N 23° 06′ 48″ E), Głębokie (52° 38′ 41″ N, 18° 26′ 18″ E), Kawęczyn (52° 10′ 09″ N, 20° 20′ 45″ E), Krzyżewo (53° 01′ 34″ N, 22° 45′ 29″ E), Pawłowice (50° 27′ 14″ N, 18° 29′ 28″ E), Słupia (50° 36′ 03″ N, 19° 58′ 28″ E), Sulejów (51° 21′ 10″ N, 19° 53′ 05″ E) and Radostowo (53° 59′ 27″ N, 18° 43′ 59″ E). During each year, the experiment with genotypes (cultivars) was carried out in a randomized block design, with three replications. Peas were most frequently sown in the first decade of April. Harvesting took place in the last decade of July. The pH of soil, depending on the year and the experimental station, ranged from 6.2 to 7.2 pH. Cereals (winter or spring wheat) were the most common pre-crop. The seeds were treated with fungicide and Nitragin before sowing. In all experiments, after sowing, the soil herbicide was sprayed, and in case of weeds, a second spraying on dicotyledonous or monocotyledonous weeds was applied. Two insecticide sprays were used in all pest control experiments, and in case of pest occurrence, a third spraying was performed. The following fertilizers were applied: P2O5—from 30 to 60 kg ha−1, K2O—from 70 to 110 kg ha−1, N—from 20 to 30 kg ha−1. The meteorological conditions in Bezek, Cicibór duży, Chrząstowo, Głębokie, Kawęczyn, Krzyżewo, Pawłowice, Radostowo, Słupia and Sulejów during the vegetation season of pea in 2010, 2011 and over an extended period were presented in Table 1.
A two-way fixed effect model was fitted to determine the magnitude of the main effects of variation and their interaction on seeds yield. Least-squares means were simultaneously produced for the AMMI model. The model first fits additive effects for the main effects of genotypes (G) and environments (E) followed by multiplicative effects for GE interaction by principal component analysis. The AMMI model (Gauch and Zobel 1990; Nowosad et al. 2016) is given by:
where yge is the seeds yield mean of genotype g in environment e, μ is the grand mean, αg is the genotypic mean deviations, βe is the environmental mean deviations, N is the number of PCA axis retained in the adjusted model, λn is the eigenvalue of the PCA axis n, γgn is the genotype score for PCA axis n, δen is the score eigenvector for PCA axis n, Qge is the residual, including AMMI noise and pooled experimental error. Expected distribution of Qge is normal.
The AMMI stability value (ASV) was used to compare the stability of genotypes as described by Purchase et al. (2000) as follows:
where SS is the sum of squares, IPCA1 and IPCA2 are the first and the second interaction principal component axes, respectively; and the IPCA1 and IPCA2 scores were the genotypic scores in the AMMI model. ASV is the distance from zero in a two-dimensional scatterplot of IPCA1 scores against IPCA2 scores. Since the IPCA1 score contributes more to GE sum of square, it has to be weighted by the proportional difference between IPCA1 and IPCA2 scores to compensate for the relative contribution of IPCA1 and IPCA2 total GE sum of squares. The higher the IPCA score, either negative or positive, the more specifically adapted a genotype is to certain environments. Lower ASV score indicate a more stable genotype across environments.
Genotype selection index (GSI) was calculated for each genotype which incorporates both mean seeds yield and ASV index in single criteria (GSIi) as (Farshadfar and Sutka 2003; Bocianowski et al. 2019a)
where GSIi is genotype selection index for ith genotype, RYi is rank of mean seeds yield for ith genotype, RASVi is rank for the AMMI stability value for the ith genotype.
The relationships between the seeds yield observed in different environments were assessed based on Pearson’s correlation coefficients. All the analyses were conducted using the GenStat v. 18 statistical software package.
Results
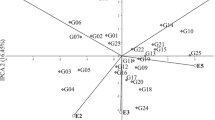
In the analysis of variance, the sum of squares for environments main effect represented 89.19% of the total, and this factor had the highest effect on seeds yield. The differences between genotypes explained 1.65% of the total seeds yield variation, while the effects of GE interaction explained 8.33% (Table 2). Values for the three principal components were also highly significant (Table 2). The three principal components of GE interaction accounted jointly for 68.99% of the whole effect it had on the variation of seeds yield. The first principal component (IPCA 1) accounted for 35.39% of the variation caused by interaction, IPCA 2 accounted for 18.03% while IPCA 3 accounted for 15.57% (Fig. 1 and Table 2).
Biplot for genotype by environment interaction of pea (Pisum sativum L.) cultivars in 20 environments, showing the effects of primary and secondary components (IPCA1 and IPCA2, respectively); Be Bezek, Ch Chrząstowo, Ci Cicibór Duży, Gł Głębokie, Ka Kawęczyn, Kr Krzyżewo, Pa Pawłowice, Sł Słupia, Su Sulejów, Ra Radostowo
The results of field trials demonstrated the impact of weather conditions, environment and genotypes on the seeds yield of pea genotypes. The seeds yield of the tested genotypes varied from 26.10 dt ha−1 (for Wenus in Radostowo 2011) to 79.73 dt ha−1 (for Lasso in Słupia 2010), throughout the 20 environments, with an average of 50.70 dt ha−1 (Table 3). The reason of low yields in Radostowo in the 2011 was small amount of rainfall in June (36.2 mm) and theirs very unproportionate spread as well as high amount of precipitation in July (114 mm) what caused strong lodging of pea’s cultivars. Very profitable rainfall was observed in period of vegetation in Słupia at 2010 (V—197.8 mm, VI—78.5 mm, VII—166.9 mm) as well as they profitable distribution (Table 1). The cultivar Tarchalska had the highest average seeds yield (54.10 dt ha−1), and the cultivar Zekon had the lowest (48.82 dt ha−1). The average seeds yield per environment also varied from 33.73 dt ha−1 in Radostowo 2011, to 73.80 dt ha−1 in Słupia 2010.
The stability of genotypes, environments, and specific GE interactions were showed in the AMMI1 biplot (Fig. 1). Among the tested genotypes, cultivar Santanta had the highest IPCA1 value of 1.860, while the smallest value of IPCA1 was − 2.578 for cultivar Wenus (Fig. 1). Among the tested environments, the smallest IPCA1 value was observed in Głębokie 2011 (− 4.582), while the highest value of IPCA1 was 1.604 in Radostowo 2011 (Fig. 1 and Table 3). The reaction to environmental changes as weather conditions, agronomic factors as well as stress has been described as the stability of the genotype. In the conducted research, the variability of this component resulted from climatic conditions.
The stability of tested genotypes can be evaluated according to biplot for seeds yield (Fig. 2). The cultivars Cysterski and Santanta interacted positively with the Głębokie 2010, Pawłowice 2010 and Cicibór Duży 2010, but negatively with the Głębokie 2011 and Słupia 2010 (Figs. 1, 2). The cultivars Bohun, Ezop, Tarchalska, Terno and Wenus interacted positively with the Głębokie 2011 and Radostowo 2010, but negatively with the Radostowo 2011 and Pawłowice 2011. The analysis showed that some genotypes have high adaptation; however, most of them have specific adaptability. AMMI stability values (ASV) revealed variations in seeds yield stability among the 12 genotypes (Table 3). A stable variety is defined as one with ASV value close to zero. Consequently, the cultivars Terno and Zekon with ASV of, respectively, 0.494 and 0.959 were the most stable, while the cultivar Wenus (ASV = 5.061) was the least stable (Table 3). Cultivar Kavalir, with average seeds yield of 50.63 dt ha−1 very close to the general mean of 50.70 dt ha−1, is distinguished on the biplot (Fig. 2). Generally, all cultivars were average seeds yield close to general mean, but with different adaptations (Figs. 1, 2): Bohun, Ezop, Tarchalska and Wenus showed specific adaptation to the conditions in Głębokie 2011, and Boruta, Cystersko, Kavalir, Lasso, Medal and Santana showed specific adaptation to the conditions in Pawłowice (both years), Zekon showed specific adaptation to the conditions in Bezek 2011, and Terno shows the highest stability. The cultivar Tarchalska has the best genotype selection index, equal to 7 (Table 3). However, the cultivars Cysterski, Santana and Wenus have the worst genotype selection index (19).
The positively statistically significant (α = 0.05) correlation coefficients for seed yield were observed for following pair of environments: Sł10-Ka10 (r = 0.60), Su10-Ch10 (0.59), Su10-Pa10 (0.62), Ch11-Ka10 (0.67), Ch11-Su10 (0.78), Ci11-Sł10 (0.59), Gł11-Sł10 (0.63), Gł11-Ra10 (0.65), Ka10-Ka11 (0.61), Ka11-Su10 (0,78), Ka11-Ch11 (0.65), Pa10-Kr11 (0.76), Pa10-Pa11 (0.71), Pa11-Kr11 (0.81), and Be10-Su11 (0.61) (Fig. 3). However, negative correlation coefficients were observed for seed yield in Ci10 and Sł11 (− 0.59) as well as Gł10 and Sł11 (− 0.60) (Fig. 3).
Discussion
The seeds yield in pea (Pisum sativum L.) is a trait determined by multiple genes that cause change in the performance of genotypes depending on the cultivation environment. The plant’s morphological characteristics are influenced by abiotic environmental factors such as soil structure, availability of oxygen and nutrients, soil moisture, temperature, and biotic factors such as plant genetic features and occurring soil microorganisms (Hodge et al. 2001; Rich and Watt 2013). High diversity of fungal communities may be beneficial for crop production, especially high diversity of AM fungi may be an important factor in maintaining productivity and stability in sustainable agricultural ecosystems (Kjøller and Rosendahl 2001;Unterseher et al. 2011). In this study, the three sources of variation were highly significant. Similar results for seeds yield in pea obtained Krajewski et al. (2012) and Tolessa et al. (2013). Krajewski et al. (2012) obtained significant GE interaction for seeds yield of two populations of lines of the F5 generation (110 and 101 lines, respectively) grown in Poznań and Wiatrowo, Poland. Tolessa et al. (2013) analysed 14 pea genotypes in 16 environments of Ethiopia. Boros and Sawicki (1997) showed that high yielding varieties are characterized by lower yield stability than average. Dore et al. (1998) report that the yield of a narrow-leaved Solara variety in a relatively small area of France ranged from 3.07 to 7.09 t ha−1 in 3 years of research. The basic cause of this state is considered to be variable humidity conditions prevailing during the growing season. They affect the cast and all elements of the yielding structure, the length of subsequent growing stages of vegetation, the height and lodging of plants. Multi-location experiments the AMMI biplot allows to visualize the main genotype effect in different environments (Bocianowski et al. 2019a). The AMMI model was used to study many species (Abakemal et al. 2016; Edwards 2016; Nowosad et al. 2017, 2018; Bocianowski et al. 2019d). Indicating GE’s interaction and other interactions is possible using the AMMI model. This model ensures an increase in the correctness of estimation (Bocianowski et al. 2018, 2019b). The use of AMMI allows the division of genotypes or other factors into groups based on similarity as well as characterization and detection of trends depending on the environment. It provides a very efficient tool for extracting genotypes or varieties that are characterized by greater efficiency in diverse environments. The use of such a strategy makes it possible to obtain more information from the GE interaction. The seeds yield expression in Poland is the most influenced trait by genotype and environment main effects as well as GE interaction. AMMI analyzes allowing for the estimation of the effect of genotype interaction in each environment, thus enable the identification of genotypes most accurately matched to the environmental conditions. Significant GE interaction of seeds yield was indicated with AMMI analysis application.
High genotypes stability is linked with the AMMI stability value. Determination of the main effect of the genotype, the environment, and the most meaningful GE interactions could be assessed based on the AMMI results displayed on GE biplot. The AMMI models are able to measure the weight of the environments, the genotypes and their interactions using a value that measure genotype stability in all environments taking into account the seeds yield. The observation and analysis of GE interaction in multi-environment yield trials is very important for evaluation, selection and recommendation of crop cultivars. AMMI models are widely used in plant breeding of the most popular crops such as wheat (Sardouei-Nasab et al. 2019), maize (Branković-Radojčić et al. 2018; Bernardo Júnior et al. 2018; Bocianowski et al. 2019c), rice (Inabangan-Asilo et al. 2019), cassava (Fotso et al. 2018), peanuts (Patil et al. 2018), cotton (Riaz et al. 2019), and willow (Fabio et al. 2017).
References
Abakemal D, Shimelis H, Derera J (2016) Genotype-by-environment interaction and yield stability of quality protein maize hybrids developed from tropical-highland adapted inbred lines. Euphytica 209:757–769
Acikgoz E, Ustun A, Gul I, Anlarsal E, Tekeli AS, Nizam I, Avcıoglu R, Geren H, Cakmakci S, Aydinoglu B, Yucel C, Avci M, Acar Z, Ayan I, Uzun A, Bilgili U, Sincik M, Yavuz M (2009) Genotype × environment interaction and stability analysis for dry matter and seed yield in field pea (Pisum sativum L.). Span J Agric Res 7(1):96–106
Annicchiarico P (1997) Joint regression vs. AMMI analysis of genotype–environment interactions for cereals in Italy. Euphytica 94:53–62
Annicchiarico P, Iannucci A (2002) Adaptation strategy, germplasm type and adaptive traits for field pea improvement in Italy based on variety responses across climatically contrasting environments. Field Crop Res 108(2):133–142
Bernardo Júnior LAY, da Silva CP, de Oliveira LA, Nuvunga JJ, Pires LPM, Von Pinho RG, Balestre M (2018) AMMI bayesian models to study stability and adaptability in maize. Agron J 110(5):1765–1776
Bilalis D, Karkanis A, Angelopoulou F, Travlos I, Antoniadis A, Ntatsi G, Lazaridi E, Savvas D (2015) Effect of organic and mineral fertilization on root growth and mycorrhizal colonization of pea crops (Pisum sativum L.). Bull UASVM Hortic 72(2):288–294
Bocianowski J, Szulc P, Nowosad K (2018) Soil tillage methods by years interaction for dry matter of plant yield of maize (Zea mays L.) using additive main effects and multiplicative interaction model. J Integr Agric 17(12):2836–2839
Bocianowski J, Niemann J, Nowosad K (2019a) Genotype-by-environment interaction for seed quality traits in interspecific cross-derived Brassica lines using additive main effects and multiplicative interaction model. Euphytica 215:7
Bocianowski J, Nowosad K, Szulc P (2019b) Soil tillage methods by years interaction for harvest index of maize (Zea mays L.) using additive main effects and multiplicative interaction model. Acta Agr Scand Sect B-Soil Plant Sci 69(1):75–81
Bocianowski J, Nowosad K, Tomkowiak A (2019c) Genotype–environment interaction for seed yield of maize hybrids and lines using the AMMI model. Maydica 64:M 13
Bocianowski J, Warzecha T, Nowosad K, Bathelt R (2019d) Genotype by environment interaction using AMMI model and estimation of additive and epistasis gene effects for 1000-kernel weight in spring barley (Hordeum vulgare L.). J Appl Genet 60(2):127–135
Boros L, Sawicki J (1997) Ocena wybranych odmian i form w kolekcji grochu siewnego (Pisum sativum L.). Cz. II. Stabilność plonowania i współzależność cech. Zesz Probl Post Nauk Roln 446:107–112 [in Polish]
Branković-Radojčić D, Babić V, Girek Z, Živanović T, Radojčic A, Filipović M, Srdić J (2018) Evaluation of maize grain yield and yield stability by AMMI analysis. Genetika 50(3):1067–1080
Davidsson L, Dimitriou T, Walczyk T, Hurrel RF (2001) Iron absorption from experimental infant formulas based on pea (Pisum sativum)-protein isolate: the effect of phytic acid and ascorbic acid. Bt J Nutr 85:59–63
Dias CT, Krzanowski WJ (2003) Model selection and cross validation in additive main effects and multiplicative interaction models. Crop Sci 43:865–873
Dore T, Meynard JM, Sebillotte M (1998) The role of grain number, nitrogen nutrition and stem number in limiting pea crop (Pisum sativum) yields under agricultural conditions. Eur J Agron 8:29–37
Eberhart SA, Russel WA (1966) Stability parameters for comparing varieties. Crop Sci 6:36–40
Edwards JW (2016) Genotype × environment interaction for plant density response in maize (Zea mays L.). Crop Sci 56:1493–1505
Fabio ES, Volk TA, Miller RO, Serapiglia MJ, Gauch HG, Van Rees KCJ, Hangs RD, Amichev BY, Kuzovkina YA, Labrecque M, Johnson GA, Ewy RG, Kling GJ, Smart LB (2017) Genotype × environment interaction analysis of North American shrub willow yield trials confirms superior performance of triploid hybrids. GCB Bioenergy 9(2):445–459
Farshadfar E, Sutka J (2003) Locating QTLs controlling adaptation in wheat using AMMI model. Cereal Res Commun 31:249–256
Fotso AK, Hanna R, Kulakow P, Parkes E, Iluebbey P, Ngome FA, Suh C, Massussi J, Choutnji I, Wirnkar VL (2018) AMMI analysis of cassava response to contrasting environments: case study of genotype by environment effect on pests and diseases, root yield, and carotenoids content in Cameroon. Euphytica 214:155
Gabriel KR (1971) The biplot graphic display of matrices with application to principal component analysis. Biometrika 58:453–467
Gauch HG (1988) Model selection and validation for yield trials with interaction. Biometrics 44:705–715
Gauch HG (2006) Statistical analysis of yield trials by AMMI and GGE. Crop Sci 46:1488–1500
Gauch HG, Zobel RW (1990) Imputing missing yield trial data. Theor Appl Genet 79:753–761
Gollob HF (1968) A statistical model which combines features of factor analytic and analysis of variance techniques. Psychometrika 33:73–115
Henderson CR (1975) Best linear estimation and prediction under a selection model. Biometrics 31:423–447
Hodge A, Campbell CD, Fitter AH (2001) An arbuscular mycorrhizal fungus accelerates decomposition and acquires nitrogen directly from organic material. Nature 413:297–299
Inabangan-Asilo MA, Swamy BPM, Amparado AF, Descalsota-Empleo GIL, Arocena EC, Reinke R (2019) Stability and G × E analysis of zinc-biofortified rice genotypes evaluated in diverse environments. Euphytica 215:61
Kjøller R, Rosendahl S (2001) Molecular diversity of glomalean (arbuscular mycorrhizal) fungi determined as distinct Glomus specific DNA sequences from roots of field grown peas. Mycol Res 105:1027–1032
Krajewski P, Bocianowski J, Gawłowska M, Kaczmarek Z, Pniewski T, Święcicki W, Wolko B (2012) QTL for yield componenets and protein content: a multienvironment study of two pea (Pisum sativum L.) populations. Euphytica 183:323–336
Malosetti M, Ribaut JM, van Eeuwijk FA (2013) The statistical analysis of multi-environment data: modeling genotype-by-environment interaction and its genetic basis. Front Physiol 4:1–17
Mandel J (1971) A new analysis of variance model for non-additive data. Technometrics 13:1–18
Moot DJ, McNeil DL (1995) Yield components, harvest index and plant type in relation to yield differences in field pea genotypes. Euphytica 86(1):30–40
Nowosad K, Liersch A, Popławska W, Bocianowski J (2016) Genotype by environment interaction for seed yield in rapeseed (Brassica napus L.) using additive main effects and multiplicative interaction model. Euphytica 208:187–194
Nowosad K, Liersch A, Poplawska W, Bocianowski J (2017) Genotype by environment interaction for oil content in winter oilseed rape (Brassica napus L.) using additive main effects and multiplicative interaction model. Indian J Genet Plant Breed 77(2):293–297
Nowosad K, Tratwal A, Bocianowski J (2018) Genotype by environment interaction for grain yield in spring barley using additive main effects and multiplicative interaction model. Cereal Res Commun 46(4):729–738
Patil AS, Hedvat I, Levy Y, Galili S, Hovav R (2018) Genotype-by-environment effects on the performance of recombinant inbred lines of Virginia-type peanut. Euphytica 214:83
Piepho HP (1994) Best linear unbiased prediction (BLUP) for regional yield trials: a comparison to additive main effects and multiplicative interaction (AMMI) analysis. Theor Appl Genet 89:647–654
Piepho HP, Möhring J, Melchinger AE, Büchse A (2008) BLUP for phenotypic selection in plant breeding and variety testing. Euphytica 161:209–228
Purchase JL, Hatting H, van Deventer CS (2000) Genotype × environment interaction of winter wheat (Triticum aestivum L.) in South Africa: II. Stability analysis of yield performance. S Afr J Plant Soil 17:101–107
Resende MDV (2007) Estimaçao e prediçao em modelos lineares mistos. In: Resende MDV (ed) Matemática e estatística na análise de experimentos e no melhoramento genético. Embrapa Florestas, Colombo, pp 101–170
Riaz M, Farooq J, Ahmed S, Amin M, Chattha WS, Ayoub M, Kainth RA (2019) Stability analysis of different cotton genotypes under normal and water-deficit conditions. J Integr Agric 18(6):1257–1265
Rich SM, Watt M (2013) Soil conditions and cereal root system architecture: review and considerations for linking Darwin and Weaver. J Exp Bot 64:1193–1208
Sardouei-Nasab S, Mohammadi-Nejad G, Nakhoda B (2019) Yield stability in bread wheat germplasm across drought stress and non-stress conditions. Agron J 111(1):175–181
Smith AB, Cullis BR, Thompson R (2005) The analysis of crop cultivar breeding and evaluation trials: an overview of current mixed model approaches. J Agric Sci 143:449–462
Tan M, Koc A, Dumlu Gul Z (2012) Morphological characteristics and seed yield of east Anatolian local forage pea (Pisum sativum ssp. arvense L.) ecotypes. Turk J Field Crops 17(1):24–30
Tolessa TT, Keneni G, Sefera T, Jarso M, Bekele Y (2013) Genotype × environment interaction and performance stability for grain yield in field pea (Pisum sativum L.) genotypes. Int J Plant Breed 7(2):116–123
Unterseher M, Jumpponen A, Öpik M, Tedersoo L, Moora M, Dormann CF, Schnittler M (2011) Species abundance distributions and richness estimations in fungal metagenomics-lessons learned from community ecology. Mol Ecol 20:275–285
van Eeuwijk FA (1995) Linear and bilinear models for the analysis of multi-environment trials: I. An inventory of models. Euphytica 84:1–7
Yang RC, Crossa J, Cornelius PL, Burgueno J (2009) Biplot analysis of genotype × environment interaction: proceed with caution. Crop Sci 49:1564–1576
Zobel RW, Wright MJ, Gauch HG (1988) Statistical analysis of yield trial. Agron J 80:388–393
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Conflict of interest
Authors declare that they have no conflict of interest.
Ethical approval
This article does not contain any studies with human participants or animals performed by any of the authors.
Additional information
Publisher's Note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Bocianowski, J., Księżak, J. & Nowosad, K. Genotype by environment interaction for seeds yield in pea (Pisum sativum L.) using additive main effects and multiplicative interaction model. Euphytica 215, 191 (2019). https://doi.org/10.1007/s10681-019-2515-1
Received:
Accepted:
Published:
DOI: https://doi.org/10.1007/s10681-019-2515-1