Abstract
A highly informative set of 16 microsatellite markers was used to fingerprint 695 apple accessions from eight Dutch collections. Among the total sample, 475 different genotypes were distinguished based on multi-locus microsatellite variation, revealing a potential redundancy within the total sample of 32%. The majority of redundancies were found between collections, rather than within collections. No single collection covered the total observed diversity well, as each collection consisted of about 50% of unique accessions. These findings reflected the fact that most collection holders focus on common Dutch varieties, as well as on region-specific diversity. Based on the diversity patterns observed, maintenance of genetic resources by a network of co-operating collection holders, rather than by collecting the total diversity in a single collection appears to be an efficient approach. Comparison of microsatellite and passport data showed that for many accessions the marker data did not provide support for the registered variety names. Verification of accessions showed that discrepancies between passport and molecular data were largely due to documentation and phenotypic determination errors. With the help of the marker data the varietal names of 45 accessions could be corrected. Microsatellite genotyping of apple appears to be an efficient tool in the management of collections and in variety identification. The development of a marker database was considered relevant as a reference instrument in variety identification and as a source of information about thus far unexplored diversity that could be of interest in the development of new apple varieties.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Cultivated apple (Malus × domestica Borkh.) is one of the most important fruit crops grown in temperate zones worldwide (Janick et al. 1996). Various wild Malus species, in particular M. sieversii (Ledeb.) M. Roemer, are thought to form the ancestors of cultivated apple, although the genetic background of apple is still not fully understood (Janick et al. 1996; Gharghani et al. 2009). Cultivated apple is generally believed to have its centre of diversity and domestication in Central Asia, from which it has spread in various directions (Gharghani et al. 2009). Apple cultivation in Southern Europe dates back to prehistoric times. It is generally believed that the Romans were responsible for the introduction of apple in North-western Europe. Like other countries in the sub-region, the Netherlands has a rich history of apple cultivation with a large diversity developed in the country throughout the last centuries. Already in the eighteenth century, Knoop (1758) described approximately 300 apple varieties that occurred in the Netherlands.
Over 7,000 apple varieties known across the globe have been reported in American publications from 1804 to 1904 (Ragan 1926), whereas nowadays the world’s production is based on a limited number of cultivars (Hokanson et al. 2001). Also in the Netherlands, a wide diversity of apple varieties was grown as small orchards of standard trees on almost each farm, castle and monastery until halfway into the twentieth century. After the Second World War, mixed farm types increasingly developed into specialized cattle or fruit farms. Orchards originally intended for self-support with diverse standard trees and a life cycle of up to 80 years, were increasingly replaced by high density plantations of spindle trees in large blocks with only a limited number of varieties per orchard. The uniform, high density plantations with a life cycle of up to 15 years facilitated mechanization, easier fruit harvesting and other preferred cultural practices. In the same period, as a result of increased breeding activities worldwide, new varieties better adapted to these new practices were developed that gradually replaced the old local varieties.
In the Netherlands, several non-governmental organizations (NGOs) have developed collections of old apple varieties and other fruit crops for the purpose of protecting the national cultural heritage. These NGOs can be found in all parts of the country, and have largely focused on regional diversity. In addition, a collection of old Dutch apple varieties is maintained by the Centre for Genetic Resources, the Netherlands (CGN). This latter collection has its origin in the 1970s when traditional varieties were collected that were no longer used in commercial fruit production (Blommers 1982). Because the collection holders in the country to a large extent operate independently from each other, particularly regarding collection composition, the diversity that is actually conserved at the national level is largely unknown. A complicating factor in determining this diversity is the lack of an adequate documentation system for some NGOs and questions on the assigned variety name for many conserved trees. In the case of old cultivars, often variety names have to be assigned to trees a posteriori, which forms quite a challenge if based on phenotypic examination, particularly when the large number of potential identities is considered.
Nowadays, molecular marker techniques are widely applied in germplasm characterization in order to assist and complement phenotypic assessments (Bretting and Widrlechner 1995). Their main advantage is that variation can be measured directly at the DNA level, which makes these techniques insensitive to environmental influences. Over the last decades, a large variety of different techniques has emerged (Spooner et al. 2005), including the use of microsatellites that are codominantly inherited and that are generally found to be highly informative and robust markers. Microsatellites are DNA fragments consisting of short repeat motifs of 1–6 base pairs that can be amplified by the Polymerase Chain Reaction (PCR). Polymerase slippage during DNA replication, or slipped strand mispairing, is considered the main cause of variation in the number of repeat units of a microsatellite, resulting in length polymorphisms that are detected by gel electrophoresis (Queller et al. 1993; Jarne and Lagoda 1996). A drawback of microsatellites is that high development costs are involved if sequence information of their flanking regions is not available for PCR primer design. However, appropriate microsatellites have already been determined for many crops.
For apple, several hundreds of microsatellite markers have been developed and positioned on a genetic linkage map (Gianfranceschi et al. 1998; Liebhard et al. 2002; Silfverberg-Dilworth et al. 2006). In apple, microsatellites are usually found to be highly polymorphic and, thus, have proven to be a powerful marker system for diversity studies (Hokanson et al. 2001; Coart et al. 2003; Gharghani et al. 2009) and for variety discrimination (Guilford et al. 1997; Hokanson et al. 1998; Goulão and Oliveira 2001). Microsatellites have also been used to monitor variation in genes for important traits, such as resistance to apple scab (Vinatzer et al. 2004).
Here we used a set of 16 published microsatellite markers to fingerprint nearly 700 hundred apple accessions from various Dutch collections. Specifically, our study aimed at increasing insight in the diversity that is conserved at the collection level and nation-wide, and to explore the usefulness of microsatellite markers for the purpose of variety identification.
Materials and methods
Study material
In 2004, Dutch collections were selected based on their size and their geographical location in the Netherlands (Table 1). The collection of the ‘POMologische Vereniging Noord-Holland’ actually consists of three, separately managed collections at different locations that therefore were treated as separate collections throughout the study. The entire collection of CGN was sampled, supplemented with wild material from Kazakhstan and a number of rootstocks that at the time of the study had not been formally included in the collection. Selections were made from the seven other collections, mainly on the basis of a presumed Dutch origin and their importance for apple cultivation in the Netherlands in the past or at present. In order to enable variety identification, part of the study material was selected because of their presumed occurrence in more than one collection. However, for efficiency reasons, the inclusion of large groups of accessions with identical documented name was avoided.
DNA extraction
In spring 2004, from each selected tree, approximately 100 mg of young leaf tissue was collected in Eppendorf tubes that were immediately frozen in liquid nitrogen. The samples were stored at −80°C upon return to the laboratory until further processing. Tissue samples were freeze-dried overnight and ground mechanically into a powder using a Retch shaking mill. Total genomic DNA was isolated using a combination of the methods of Fulton et al. (1995) and the DNeasy 96 Plant Kit (Qiagen, Westburg, the Netherlands).
Microsatellite analysis
Available DNA samples of the varieties ‘Discovery’, ‘Golden Delicious’ and ‘Prima’, provided by Plant Research International (Wageningen, the Netherlands), were added to the study material and used as references throughout the microsatellite analyses. In previous studies these varieties had already been used as references in order to calibrate marker sizes between studies (e.g. Koopman et al. 2007). The varieties ‘Discovery’ and ‘Prima’ are also part of the reference set used in the European project HiDRAS that aims to identify genetic factors controlling apple fruit quality (Gianfranceschi and Soglio 2004).
A set of 16 microsatellites was selected on the basis of their technical qualities, reported level of variability, genome coverage, linkage to known trait loci and use in previous or ongoing studies (Table 2). Details about the markers, such as primer sequences and exact map locations can be found on the website of the HiDRAS project (www.hidras.unimi.it/index.html) and in the references listed in Table 2. Microsatellite analyses basically followed the procedures described in Van Treuren et al. (2008) and individual markers were combined in four multiplexes (Table 3). Multiplexing was carried out prior to PCR, except for microsatellites CH02c06, CH02d08, Hi07f01 and SdSSR that were amplified separately and were combined with the other multiplex components after PCR. Fluorescently labelled PCR products were separated by capillary electrophoresis using an ABI Prism 3700 DNA analyzer (Applied Biosystems). Fragment sizes and peak areas were determined using the software package GENESCAN (release 1.1 3700 software, Applied Biosystems) and further processed with the software package Genotyper (version 3.5 NT, Perkin-Elmer).
Data analysis
Observed DNA fragments were recorded by the number of base pairs correct to two decimal places. Subsequently, a frequency distribution of the fragment sizes correct to one decimal place was made for each microsatellite locus in order to convert the recorded values to discrete alleles. In most cases, the fragments could be assigned unambiguously to discrete alleles, even when the data indicated allele size differences of only a single base pair (Fig. 1).
Microsatellite data were transformed into binary scores in order to calculate similarity values between all pairs of samples according to the methods of Jaccard (1908). Similarity values were used to visualize genetic relationships among the accessions by a UPGMA (unweighted pair group mean with arithmetic averaging) cluster analyses carried out using NTSYS-pc (Rohlf 1993). The data were checked for potential scoring errors at individual loci by generating pseudo replicate datasets each containing all but one of the original marker loci (jack-knifing). In case jack-knifing resulted in matching of samples that appeared dissimilar based on the total set of markers, the data of the microsatellite underlying the difference in results were re-examined.
The potential presence of null alleles was disregarded. As a consequence, homozygosity was assumed when samples showed only a single allele for a microsatellite locus. In order to obtain estimates of the level of diversity detected by individual microsatellites, allele frequencies in the total sample were used to calculate PIC (Polymorphic Information Content) values according to the methods of Botstein et al. (1980). Allele frequencies were also used to estimate the resolving power of the entire data set. For that purpose, the probability of observing identical multilocus microsatellite genotypes by chance was estimated (Hokanson et al. 1998). For a random mating population of a diploid organism, and assuming Hardy–Weinberg equilibrium and independent segregation among loci, this probability can be estimated by:
where m denotes the number of loci, n the number of alleles at locus a and p a,i the frequency of the ith allele at locus a.
The microsatellite data were used to quantify genotypic and allelic diversity for the total set of nearly 700 accessions and for each of the collections separately. In order to correlate the microsatellite data and the registered accession names, two separate duplication analyses were carried out. Firstly, potential duplication groups, hereafter denoted as ‘passport duplicates’ were constructed based on identical or similar registered accession names, or on known synonymy. Subsequently, the molecular data were used to determine relationships. Secondly, duplication groups, hereafter denoted as ‘marker duplicates’ were constructed based on identical microsatellite profiles. Marker duplications were then related to their registered accession names. When the duplication analyses revealed discrepancies between the marker data and accession names, the involved accessions were checked for potential documentation errors and for potential synonymy. In case discrepancies remained unresolved, the sampled trees were re-examined phenotypically, and/or photo-documentation was studied in order to check for potential errors in variety identification.
Results
Microsatellite screening
The microsatellite variation detected in the total sample is presented in Table 3. Because the profiles of microsatellite U50187-SSR could not be scored consistently among the samples, variation for this marker was not recorded and was disregarded in all data analyses. For microsatellite CH01h02 two ranges of allele sizes were observed that were denoted as CH01h02 (a) and CH01h02 (b) and were considered as two separate loci throughout the analyses. Due to the absence of an amplification product, insufficient fluorescent signals or unreliable microsatellite profiles for individual samples, 7% missing values on average were recorded per locus. Three alleles per locus for one or more loci were observed in 190 DNA samples, suggesting a triploid genome for these accessions. For 10 samples even four alleles per locus were recorded for one or more loci. The average frequency of three and four alleles per locus was 5.7 and 0.2%, respectively.
Reference varieties were included in all molecular analyses. As a consequence, the microsatellite genotypes of each of the references were independently determined eight times. Apart from minor differences in absolute allele size, microsatellite genotypes observed for the reference varieties ‘Discovery’ and ‘Prima’ were generally in line with those published on the HiDras project website (Table 4). Discrepancies for ‘Discovery’ were found for microsatellite CH02c02b that showed only weak and inconsistent signals at 84 and 134 base pairs and for microsatellite CH05e04 for which no signal whatsoever was observed at 230 base pairs. For the latter microsatellite ‘Prima’ did not show any amplification product at the 230 and 234 base pairs either. In addition, discrepancies for ‘Prima’ included microsatellite Hi07f01, for which an additional, weak and inconsistent signal was observed at 219 base pairs, and microsatellite SdSSR. However, the latter case may be due to incorrect publishing of allele sizes on the HiDRAS website since the largest allele is presented first.
The number of observed alleles per locus ranged from 3 for Mald4-5ssr to 37 for CH05e03, and a total number of 296 alleles were scored for the total set of 16 loci (Table 3). Relatively high levels of diversity were detected by individual microsatellites, as PIC values per locus ranged from 0.507 for CH01h02 (a) to 0.912 for CH02c02a. The resolving power of the entire data set was estimated to be 6.4 × 10−20, meaning that the probability of observing identical genotypes purely by chance could be neglected.
Diversity within and among collections
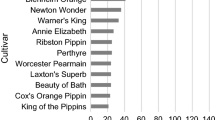
Among the 695 investigated trees, 475 different multilocus microsatellite genotypes were observed, suggesting a potential redundancy of 220 trees (32%). A UPGMA cluster analysis showed that in case of molecularly different samples, the degree of similarity was generally small, as similarity values between 0.9 and 1.0 were observed in only nine sample comparisons (results not shown). The number of different genotypes per collection ranged from 11 for PV3 to 205 for NPV (Fig. 2). Potential redundancies per individual collection could be derived from the difference between the sample size and the observed number of genotypes, and ranged from 0 for PV1 and PV2 to 36 (15%) for the NPV collection. The sum of the number of potential redundancies per collection was 73, which is 33% of the total number of 220 observed among the entire sample, indicating that the majority of the potential redundancies were between collections, rather than within collections. Apart from PV3, about half of each collection consisted of unique genotypes, i.e. microsatellite genotypes not observed in the other investigated collections, indicating that no single collection covered the total genotypic variation extensively. The number of microsatellite alleles recorded within collections ranged from 97 (33% of the total observed number) for PV3 to 237 (80%) for CGN. Thus, in contrast to genotypic variation, a large proportion of the total allelic diversity could be found in single collections. Accordingly, collections were found to harbour only low numbers of unique alleles, which was highest for the CGN collection with 37 unique alleles. These findings seem to indicate that the observed genotypic variation is for the larger extent due to genetic recombination rather than to a wide variety of distinct alleles, the high figure of the CGN collection reflecting the inclusion of the wild germplasm from Kazakhstan.
Relationships between molecular profiles and variety names
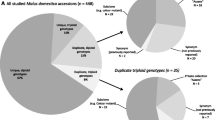
Comparison of passport data revealed 136 potential duplication groups involving a total of 349 accessions (Table 5A). The duplicate status was confirmed by identical microsatellite profiles in 144 cases (41%), while 6 cases (2%) showed a difference at only a single locus. The remaining 199 passport duplicates had to be rejected according to the molecular data. However, a posteriori variety information revealed that in 49 rejected cases the presumed similarity based on accession names was unjustified. For the trees that were phenotypically re-examined, the documented variety name appeared to be incorrect in 35 cases. In 8 cases it could not be determined which variety was misnamed, although in these cases in addition to distinct marker profiles also clear morphological differences were observed. Phenotypic re-examination of the remaining 107 rejected passport duplicates could so far not be carried out, but verification of these accessions is still ongoing. Thus, none of the rejected passport duplicates so far had to be ascribed to incorrect inferences of the microsatellite data, but appeared to be rather due to documentation and determination errors.
Comparison of molecular profiles revealed that the total sample, supplemented with the three reference varieties, contained 121 groups of marker duplicates comprising 344 trees (Table 5B). Out of these 344 accessions, 139 (40%) were confirmed by the passport data, whereas the remaining 205 accessions (60%) appeared not in line with the passport data. However, discrepancies between the marker profiles and the passport data could be resolved after verification, either by synonymy, varietal mutations, sampling of the interstock instead of the grafted variety, incorrect documented variety names or observed phenotypic similarity. Only in 11 cases, trees appeared phenotypically distinct despite a similar microsatellite genotype.
Based on the microsatellite profiles and subsequent verification process of discrepancies, the varietal names of 45 accessions were corrected. Thus, the combined results of our duplication analyses indicated the high potential of the application of microsatellite markers to trace documentation and determination errors and to assist in variety identification.
Discussion
Microsatellite characterization
Characterization and evaluation of plant genetic resources is one of the main priorities defined by the European Cooperative Programme for Plant Genetic Resources (ECPGR website: http://www.ecpgr.cgiar.org/). For this purpose, molecular marker techniques have increasingly been used to complement traditional approaches (Bretting and Widrlechner 1995; Spooner et al. 2005). Here we presented the results of a study using a highly informative set of 16 microsatellite markers to characterize apple genetic resources obtained from eight Dutch collections.
Based on the level of variation observed in the present study, the estimated probability of observing identical multilocus microsatellite genotypes by chance was 6.4 × 10−20, indicating the high resolution potential of the marker set used. Whereas this probability may have been overestimated because the underlying assumptions of random mating and Hardy–Weinberg equilibrium may not be valid for apple collections and because a few markers may not segregate independently due to their location on the same chromosome, the resolving power may have been underestimated because of the existence of triploid tree specimens, since diploidy was assumed in the estimation procedure. The high estimated resolving power indicates that samples showing identical microsatellite profiles may be considered identical or to represent closely related genotypes. The latter may include varieties that differ in only a single or few mutations but that nevertheless show distinct phenotypical characteristics. A few examples were observed in the present study, such as the pair of varieties ‘Gronsvelder Klumpke’ and ‘Eijsdener Klumpke’ that are known to represent color mutants that could not be distinguished based on the investigated microsatellites.
When compared with other accessions, the majority of investigated accessions showed either identical multilocus microsatellite profiles, suggesting identity, or differences at more than one marker locus, indicating that a lower resolving power would have sufficed to distinguish unrelated accessions. A standard set of 12 microsatellites and 8 reference accessions has been agreed upon by the ECPGR Malus/Pyrus working group following a workshop in East Malling, United Kingdom in 2006 (Kate Evans, personal communication). The standards include the microsatellites CH01h01, CH01h10 and CH02d08 and the variety ‘Prima’ that were also used in the present study. This will enable linking of the present data with ongoing and future initiatives, albeit with a reduced resolving power of 1.2 × 10−4.
Microsatellites are generally considered one of the most reliable marker technologies in collaborative projects (Röder et al. 2002). However, a low frequency of discrepancies in marker scores are usually observed when identical samples are analyzed at different laboratories (Jones et al. 1997; Bredemeijer et al. 2002). Similarly, in the present study discrepancies were observed when microsatellite scores of the reference varieties ‘Discovery’ and ‘Prima’ were compared with those published on the HiDRAS project website. Assuming that genetically identical samples of the reference varieties were used in both projects, discrepancies may have been due to the use of different selection thresholds for allelic intensities or to incorrect allele size identification if stutter bands resulting from PCR artifacts occurred. Consistent differences in absolute allele size may either have resulted from experimental variation or from variation in primer design, such as the use of primers with or without pigtail extension (Brownstein et al. 1996). However, relative allele size differences are more relevant and reliable than absolute allele sizes (Liebhard et al. 2002). Therefore, methodological variation necessitates the use of reference varieties and exchange of marker profiles and scoring protocols in order to streamline calibration of marker scores between different laboratories (Bredemeijer et al. 2002).
To increase the efficiency of the analysis, some of the investigated microsatellites that were labeled with a similar fluorescent dye were combined into a single multiplex. These combinations were chosen carefully based on pre-existing knowledge about the size range of identified alleles. In the present study, scoring difficulties due to overlapping size ranges in multiplexed samples using similar fluorescent dyes were encountered for the marker combinations CH02c02b/CH05e04 and CH02d08/U50187-SSR. In case the microsatellite profiles could not be scored unambiguously, missing values were recorded, thereby reducing the efficiency of the analysis. Therefore, for diversity studies it is recommended to avoid the use of similarly labeled markers in multiplexes, as long as the full size range of the alleles is not completely known.
Collection management
Based on the microsatellite data, a redundancy of 32% was observed in the total sample of 695 accessions, of which the major part could be ascribed to duplication between collections rather than within collections. It should be noted that this estimated level of redundancy may be biased because only the CGN collection was investigated to the full extent, whereas the study materials from the other collections were not selected randomly. The inclusion of accessions that were presumed to occur in more than one collection may have caused over-estimation, while the avoidance of large groups of accessions with identical documented name may have caused under-estimation of the level of redundancy. Despite the observed overlap between collections, the investigated collections were each 50% unique. These findings reflect NGO policies. NGOs mostly aim to collect the commonly known Dutch varieties widely used throughout the country, such as ‘Glorie van Holland’ and ‘Notarisappel’, as well as the varieties of which the use was traditionally largely limited to the region in which the NGO is located. Although a certain level of duplication, both within and between collections provides for a safety backup system, high levels of duplication should be avoided on the basis of efficiency grounds (Van Treuren and van Hintum 2003). Maintenance costs are particularly high for crops that are not conserved in the form of seeds but instead are maintained by vegetative propagation in field collections of trees, such as apple (Hokanson et al. 1998). The microsatellite data were used by the collection holders to critically re-examine the composition of their collections, which resulted in the removal of certain accessions in case of unintended internal redundancy and the replacement of accessions in case of misidentifications. All unintended internal duplicates were removed from the CGN collection in order to improve efficiency. Redundancies among collections are much more difficult to avoid because NGOs generally have their individual incentives and largely operate independently from each other with respect to collection composition. Therefore, curators do not tend to remove accessions from their collection because of the presence of the same variety in other collections.
Obviously, duplication among collections reduces the risk of an absolute loss of accessions. In that respect, the commonly known varieties that can generally be found in various collections have a low probability of becoming lost, in contrast to the varieties that were observed only in single collections in the present study. None of the investigated collections appeared to cover the total genetic diversity extensively because of the presence of unique diversity within each collection. Collecting the total diversity in a single collection would be an elusive goal because of the large number of apple varieties involved and the consequent high maintenance costs. The establishment of a network of co-operating collections may be considered as an efficient alternative option. Increased co-operation was initiated through the joint documentation of collections by developing a common database with variety descriptors that currently applies to 2,418 accessions from 16 collections. This database, that is so far available only in the Dutch language, is accessible via the internet at www.appelcollecties.nl.
Continuity of collections is a key issue as a long as a safety back-up system for unique accessions is lacking. However, during the annual meeting of collection holders, developments at individual collections can be discussed and emergency plans organized when necessary. Nevertheless, a future objective should be the representation of varieties in a minimum of two collections in order to minimize the risk of loosing diversity. In 2008, the apple collections of the ‘POMologische Vereniging Noord-Holland’, the ‘Noordelijke Pomologische Vereniging’ and the Centre for Genetic Resources, the Netherlands were included in the Multi-Lateral System (MLS) of the International Treaty on Plant Genetic Resources for Food and Agriculture (Fowler et al. 2003).
Variety identification
The combined analysis of passport and molecular duplicates showed that for many of the studied accessions the molecular data profiles did not fit the variety names registered for those accessions. Many discrepancies could be resolved following a renewed verification process by crop experts through detailed examination of the documented variety names and/or through investigation of fruit specimens of the trees involved. Investigation of fruits appeared impossible for about 50% of the observed discrepancies because of the absence of fruits in consecutive years.
For a few small duplication groups involving a total of six accessions, trees with identical variety names differed at only a single microsatellite locus, which may be ascribed to spontaneous mutation at marker loci during vegetative propagation. For only 11 accessions the discrepancy between molecular and passport data remained after the renewed verification process, that is, trees registered under different variety names indeed showed phenotypic differences despite identical molecular profiles. Considering the high estimated resolving power of the investigated microsatellites, it seems highly unlikely that unrelated accessions would show identical marker profiles. It remains to be verified whether these findings were possibly due to sampling error or experimental error. In addition, it cannot be ruled out that identical genotypes show phenotypic differences under different environmental conditions or that in fact related varieties were involved that differ in only a single or few mutations.
In most cases, initial discrepancies between molecular and passport data could be explained by documentation errors or determination errors. Also in other studies, misidentified apple accessions were uncovered through genotype matching using highly informative microsatellite markers (Hokanson et al. 1998). Discrepancies in our study were revealed by the duplication analysis, but it can be expected that documentation and determination errors also apply to accessions that appeared to be unique in the present study. Documentation data may be further improved by ongoing verification of variety names by collection holders and by the development of a marker database that can be used for reference purposes.
Based on the results of the present study, a database was established with confirmed duplicates and verified varieties, currently holding the microsatellite data of 121 apple varieties represented in 227 accessions (see supplementary table). Twenty-six of the included varieties (21%) are genotypes with 3 alleles at a single locus or multiple loci. Twelve of these varieties are known to be triploids, whereas the ploidy level of the rest of this group of 26 was unknown.
In our study, ten samples with even four alleles per locus were encountered, which could indicate a tetraploid genome or a duplication event of chromosomal segments. Both triploid and tetraploid varieties are known to exist for apple (Brown 1998).
Concluding remarks
For apple, microsatellite genotyping appeared a very efficient tool for enhancing collection management of genetic resources and for variety identification. The latter may be further facilitated by cross-referencing with molecular datasets of other ongoing projects, such as the microsatellite fingerprinting initiative of the entire apple collection at Brogdale, United Kingdom. The microsatellites used in the present study were selected, amongst others, for their linkage to known trait loci. Thus far unexplored variation could be of interest to plant breeders in their search for desired characters in the development of new apple varieties.
References
Blommers LHM (1982) Collecting and preserving apple and pear cultivars in the Netherlands. Acta Hortic 142:15–22
Botstein D, White RL, Skolnick M, Davis RW (1980) Construction of a genetic linkage map in man using restriction fragment length polymorphisms. Am J Hum Genet 32:314–331
Bredemeijer GMM, Cooke RJ, Ganal MW, Peeters R, Isaac P, Noordijk Y, Rendell S, Jackson J, Röder MS, Wendehake K, Dijcks M, Amelaine M, Wickaert V, Bertrand L, Vosman B (2002) Construction and testing of a microsatellite database containing more than 500 tomato varieties. Theor Appl Genet 105:1019–1026
Bretting PK, Widrlechner MP (1995) Genetic markers and plant genetic resource management. Plant Breed Rev 13:11–86
Brown SK (1998) Genetics of apple. In: Janick J (ed) Plant breeding reviews Vol 9. Wiley-Interscience, pp 333–366
Brownstein MJ, Carpten JD, Smith JR (1996) Modulation of non-templated nucleotide addition by Taq DNA polymerase: primer modifications that facilitate genotyping. Biotechniques 20:1004–1010
Cevik V, King GJ (2002) High-resolution genetic analysis of the Sd-1 aphid resistance locus in Malus spp. Theor Appl Genet 105:346–354
Coart E, Vekemans X, Smulders MJM, Wagner I, Van Huylenbroeck J, Van Bockstaele E, Roldán-Ruiz I (2003) Genetic variation in the endangered wild apple (Malus sylvestris (L.) Mill.) in Belgium as revealed by amplified fragment length polymorphism and microsatellite markers. Mol Ecol 12:845–857
Fowler C, Moore G, Hawtin G (2003) The international treaty on plant genetic resources for food and agriculture: a primer for the future of CGIAR. IPGRI, Rome
Fulton TM, Chunwongse J, Tanksley SD (1995) Microprep protocol for extraction of DNA from tomato and other herbaceous plants. Plant Mol Biol Rep 13:207–209
Gao Z (2005) Localization of candidate allergen genes on the apple (Malus domestica) genome and their putative allergenicity. Ph.D. thesis, Wageningen University, Wageningen
Gharghani A, Zamani Z, Talaie A, Oraguzie NC, Fatahi R, Hajnajari H, Wiedow C, Gardiner SE (2009) Genetic identity and relationships of Iranian apple (Malus × domestica Borkh.) cultivars and landraces, wild Malus species and representative old apple cultivars based on simple sequence repeat (SSR) marker analysis. Genet Res Crop Evol. doi:10.1007/s10722-008-9404-0
Gianfranceschi L, Soglio V (2004) The European project HiDRAS: innovative multidisciplinary approaches to breeding high quality disease resistant apples. Acta Hortic 663:327–330
Gianfranceschi L, Seglias N, Tarchini R, Komjanc M, Gessler C (1998) Simple sequence repeats for the genetic analysis of apple. Theor Appl Genet 96:1069–1076
Goulão L, Oliveira CM (2001) Molecular characterisation of cultivars of apple (Malus × domestica Borkh.) using microsatellite (SSR and ISSR) markers. Euphytica 122:81–89
Guilford P, Prakash S, Zhu JM, Rikkerink E, Gardiner S, Bassett H, Forster R (1997) Microsatellites in Malus × domestica (apple): abundance, polymorphism and cultivar identification. Theor Appl Genet 94:249–254
Hokanson SC, Szewc-McFadden AK, Lamboy WF, McFerson JR (1998) Microsatellite (SSR) markers reveal genetic identities, genetic diversity and relationships in a Malus × domestica Borkh. core subset collection. Theor Appl Genet 97:671–683
Hokanson SC, Lamboy WF, Szewc-McFadden AK, McFerson JR (2001) Microsatellite (SSR) variation in a collection of Malus (apple) species and hybrids. Euphytica 118:281–294
Jaccard P (1908) Nouvelles recherches sur la distribution florale. Bull Société Vaudoise Sci Naturelles 44:223–270
Janick J, Cummins JN, Brown SK, Hemmat M (1996) Apples. In: Janick J, Moore JN (eds) Fruit breeding, tree and tropical fruits, vol II. John Wiley, New York, pp 1–77
Jarne P, Lagoda PJL (1996) Microsatellites, from molecules to populations and back. Trends Ecol Evol 11:424–429
Jones CJ, Edwards KJ, Castaglione S, Winfield MO, Sala F, van de Wiel C, Bredemeijer G, Vosman B, Matthes M, Daly A, Brettschneider R, Bettini P, Buiatti M, Maestri E, Malcevschi A, Marmiroli N, Aert R, Volckaert G, Rueda J, Linacero R, Vazquez A, Karp A (1997) Reproducibility testing of RAPD, AFLP and SSR markers in plants by a network of European laboratories. Mol Breed 3:381–390
Knoop JH (1758) Pomologia, dat is beschrijvingen en afbeeldingen van de beste zoorten van appels en peeren, welke in Neder- en Hoog-Duitschland, Frankryk, Engeland en elders geagt zyn, en tot dien einde gecultiveert worden. Ferwerda & Tresling, Leeuwarden, the Netherlands
Koopman WJM, Li Y, Coart E, Van de Weg WE, Vosman B, Roldán-Ruiz I, Smulders MJM (2007) Linked vs. unlinked markers: multilocus microsatellite haplotype-sharing as a tool to estimate gene flow and introgression. Mol Ecol 16:243–256
Liebhard R, Gianfranceschi L, Koller B, Ryder CD, Tarchini R, Van de Weg WE, Gessler C (2002) Development and characterization of 140 new microsatellites in apple (Malus × domestica Borkh.). Mol Breed 10:217–241
Queller DC, Strassmann JE, Hughes CR (1993) Microsatellites and kinship. Trends Ecol Evol 8:285–288
Ragan WH (1926) Nomenclature of the apple: a catalogue index of the known varieties referred to in American publications from 1804 to 1904. USDA Bur Plant Ind Bull 56
Röder MS, Wendehake K, Korzun V, Bredemeijer G, Laborie D, Bertrand L, Isaac P, Rendell S, Jackson J, Cooke RJ, Vosman B, Ganal MW (2002) Construction and analysis of a microsatellite-based database of European wheat varieties. Theor Appl Genet 106:67–73
Rohlf FJ (1993) NTSYSpc numerical taxonomy and multivariate analysis system, version 2.1. Exeter Software, New York
Silfverberg-Dilworth E, Matasci CL, Van de Weg WE, Van Kaauwen MPW, Walser M, Kodde LP, Soglio V, Gianfranceschi L, Durel CE, Costa F, Yamamoto T, Koller B, Gessler C, Patocchi A (2006) Microsatellite markers spanning the apple (Malus × domestica Borkh.) genome. Tree Genet Genomics 2:202–224
Spooner D, van Treuren R, de Vicente MC (2005) Molecular Markers for Genebank Management. IPGRI Technical Bulletin No. 10. International Plant Genetic Resources Institute, Rome, Italy
Van Treuren R, van Hintum ThJL (2003) Marker-assisted reduction of redundancy in germplasm collections: genetic and economic aspects. Acta Hortic (ISHS) 623:139–149
Van Treuren R, van Hintum ThJL, van de Wiel CCM (2008) Marker-assisted optimization of an expert-based strategy for the acquisition of modern lettuce varieties to improve a genebank collection. Genet Res Crop Evol 55:319–330
Vinatzer BA, Patocchi A, Tartarini S, Gianfranceschi L, Sansavini S, Gessler C (2004) Isolation of two microsatellite markers from BAC clones of the Vf scab resistance region and molecular characterization of scab-resistant accessions in Malus germplasm. Plant Breed 123:321–326
Acknowledgments
The authors would like to thank Hennie Rossel, Coen Ballintijn, Rein Baars, Rene Smulders, Reinder Elema, ‘Boomkwekerijen Jos Frijns & Zonen B.V’, the ‘Noordelijke Pomologische Vereniging’ and the ‘Stichting Fruithof’ for their involvement in the project. We are grateful to Hennie Rossel for his major role in the selection of the study material, to Peter Goossens and Wendy van’t Westende for carrying out the molecular analyses, to Eric van de Weg for selecting the microsatellites and designing the multiplexes used in the project, and to two anonymous reviewers for their useful comments on an earlier version of the manuscript. This project was financed by the Dutch Ministry of Agriculture, Nature and Food Quality.
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
van Treuren, R., Kemp, H., Ernsting, G. et al. Microsatellite genotyping of apple (Malus × domestica Borkh.) genetic resources in the Netherlands: application in collection management and variety identification. Genet Resour Crop Evol 57, 853–865 (2010). https://doi.org/10.1007/s10722-009-9525-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10722-009-9525-0