Abstract
Brassinosteroids (BRs) are plant specific steroidal hormones that play diverse roles in regulating a broad spectrum of plant growth and developmental processes, as well as, in responding to various biotic and abiotic stresses. Extensive research over the years has established stress-impact-mitigating role of BRs and associated compounds in different plants exposed to various abiotic and biotic stresses, suggesting the idea that they may act as immunomodulators, thus opening new approaches for plant resistance against hazardous environmental conditions. In this research the characterization of the transcriptional response of 11 transcription factors (TFs) belonging to BRASSINAZOLE-RESISTANT 1 (BZR1) TF family of Zea mays L. was analyzed in seedlings subjected to different stress conditions. Being important regulators of the BR synthesis, BZR TFs might have stress resistance related activities. However, no stress resistance related functional study of BZR TFs has been reported in maize so far. In silico analyses of the selected 11 TFs validated the features of their protein domains, where the highest degree of similarity observed with recognized BZR TFs of rice and Sorghum bicolor. Additionally, we investigated the organ-specific expression of 11 ZmBZR in maize seedlings. Five of them did not show any transcript accumulation, suggesting that ZmBZR expression might be regulated in a manner dependent on plant developmental stage. For the remaining six ZmBZR, their ubiquitous expression in the whole plant indicates they could function as growth regulators during maize development. More importantly, in response to various stress conditions, the spatial transcript accumulation of all ZmBZR varies along the plant. All six ZmBZR showed up-regulation against N starvation, hypoxia and salt stress. On the contrary, heat stress clearly down-regulated gene expression of all ZmBZR analysed. Consistently with the expression results, the distribution of stress-related cis-acting elements in the promoter of these genes inferred that the maize BZR TFs might play some roles in regulating the expression of the corresponding genes in response to multifarious stresses. In conclusion, these data reveal that BZR TFs have stress signaling activity in maize, in addition to their confirmed role in regulating plant physiology and morphology.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Brassinosteroids (BRs) are plant specific steroidal hormones that play diverse roles in regulating a broad spectrum of plant growth and developmental processes. They regulate multiple physiological functions including seed germination, cell elongation and division, senescence, vascular-differentiation, reproduction, root development, photomorphogenesis and respond to various biotic and abiotic stresses (Saini et al. 2015; Singh and Savaldi-Goldstein 2015). Molecular studies evidenced cross-talk between BRs and other phytohormones and hypothesised the existence of synergistic effects between exogenous BR treatments and endogenous levels of other hormones (Gruszka 2013; Zhu et al. 2013). Extensive research over the years has led to the idea that BRs could act as stress-impact-mitigating compounds in different plants exposed to various abiotic stresses such as high temperature, low temperature in terms of chilling and freezing, salinity, light, drought, metals/metalloids and organic pollutants (Derevyanchuk et al. 2016; Vardhini and Anjum 2015 and references therein). Some studies also suggest that BR treatments could promote plant resistance against many pathogens, such as fungi, bacteria, and virus (He et al. 2007; Kemmerling et al. 2007; Chinchilla et al. 2009). Essentially, BRs seem to act as immunomodulators when applied at the appropriate concentration and at the correct stage of plant development, thus opening new approaches for the improvement of plant resistance against hazardous environmental conditions.
Most of the information about BR signalling has been obtained from Arabidopsis. Molecular studies have demonstrated that BRs are perceived at the cell membrane by the BRASSINOSTEROID INSENSITIVE 1 (BRI1) receptor kinase, which upon ligand binding heterodimerizes with BRI1-ASSOCIATED RECEPTOR KINASE (BAK1). The fully activated BRI1/BAK1 triggers a series of downstream phosphorylation events and subsequently inactivates the GSK3/Shaggy-like protein kinase BIN2, a pivotal negative regulator of BR signaling (Li et al. 2001), which lead to the regulation of the expression of a large set of genes involved in plant growth and development (Sun et al. 2010). Downstream, BZR1 (BRASSINAZOLE RESISTANT1) and BES1 (BRASSINOSTEROID INSENSITIVE 1-ETHYL METHANESULFONATE-SUPPRESSOR 1), two closely related TFs belonging to the BRASSINAZOLE-RESISTANT (BZR) TF family, are rapidly dephosphorylated by protein phosphatase 2A (PP2A) (Tang et al. 2011). The dephosphorylated BZR1 and BES1 accumulate in the nucleus and directly bind to cis elements, known as E-box (CANNTG) and BR-response element (CGTGT/CG) of their target, regulating plant growth and development (Yu et al. 2011, Wu et al. 2016). Although the interaction between stress and BRs has long been observed (Nawaz et al. 2017), the underlying molecular mechanisms were far to be completely elucidated.
The BZR TF family appears to be involved in the regulation of various processes in plants. In Arabidopsis, BZR family proteins were thought to be the primary transcription factors regulating huge numbers of genes involved in BR signal output (Sun et al. 2010; Yu et al. 2011). Rice BZR family has been suggested to play a conserved role as in Arabidopsis (Tong and Chu 2012). Recent findings reveal that AtBZR1 positively regulates plant stress tolerance (Sahni et al. 2016); in Brassica rapa, BrBZR TFs family is suggested to be involved in regulating stress-related activities (Saha et al. 2015). While major studies have revealed the positive roles of these TFs in BR signal transduction in many plants (Yin et al. 2005), no genome-wide-in-depth study of the BZR TF family in maize has previously been reported.
In this work a comprehensive genome-wide analysis was carried out to characterize the BZR TFs family in maize. Eleven BZR TFs of Zea mays L. (ZmBZR) were characterized from a genome-wide survey and their expression profiles were assessed in different tissues. Considering that crop plants are subjected to combinations of abiotic stresses during their lifespan that greatly reduce productivity and that recent research suggests plants can be primed by chemical compounds to better tolerate different abiotic stresses, we proposed to better elucidate the role of BRs in stress response to test their effects in plant chemical priming. Being important regulators of the BR synthesis, BZR TFs might have stress resistance related activities. However, no stress resistance related functional study of BZR TFs has been reported in the monocot model plant Zea mays L. so far. The expression analyses on the candidate ZmBZR were evaluated to investigate their responses to several abiotic stresses such as low nitrate availability, hypoxia, salinity and heat. The obtained results provide a new start for the future studies of the BR signalling pathway in monocotyledons.
Materials and methods
Genome-wide identification of ZmBZR genes
To identify BZR TFs family members in Zea mays L., the Arabidopsis BZR1 amino acid sequence was used as query to search the maize Database (Phytozome). The conserved domains of the BZR were confirmed by Pfam (http://pfam.xfam.org). The list of genes analysed is reported in Supplemental Table S1, together with the primers utilized for reverse-transcription quantitative real-time PCR (RT-qPCR) expression analysis. Primers were designed with Primer3 web tool (version 0.4.0; http://frodo.wi.mit.edu/primer3/) and further verified with the PRATO web tool (http://prato.daapv.unipd.it). GRASSIUS database (http://grassius.org/) was used for gene nomenclature.
Phylogenetic analysis and classification of ZmBZR genes
The full amino acid sequences of BZR TFs members from maize, rice, sorghum, Nicotiana and Arabidopsis were aligned by CLUSTALW program. The gene IDs of BZR members in maize, rice, sorghum, Nicotiana and Arabidopsis are shown in Supplemental Table S2. Maize BZR genes were placed on 10 maize chromosomes according to their positions given in the GRAMENE maize database (available online: http://www.gramene.org). The distribution of ZmBZR genes on the maize chromosomes was drawn by MapInspect (available online: http://mapinspect.software.informer.com) and modified manually with annotation.
Cis-elements in the promoter regions of ZmBZR genes
To predict cis-acting regulatory DNA elements (cis-elements) in promoter regions of maize BZR genes, 2000 bp genomic DNA sequences upstream of the initiation codon (ATG) was analyzed by the PLACE website (available online: http://www.dna.affrc.go.jp/PLACE/signalscan.html).
Plant materials and growth conditions
Seeds of maize (Zea mays L.), inbred line B73, were washed in distilled water and germinated on wet filter paper at 25 °C in the dark. After 3 days, maize seedlings were transferred in a controlled environmental chamber in 500 ml tanks containing a Hoagland-modified nutrient solution (changed every 2 days), according to the following composition (µM): KNO3 (1000), CaCl2 (200), MgSO4 (200), KH2PO4 (40), FeNaEDTA (10), H3BO3 (4.6), MnCl2 (0.9), ZnCl2 (0.09), CuCl2 (0.036), NaMoO4 (0.01). This nutrient solution and a day/night cycle of 14 h/10 h at 25/20 °C air temperature, 70/90% relative humidity, and 280 µmol m− 2 s− 1 photon flux density were utilized as standard conditions to grow control plants for each treatment. Four different stress treatments were imposed on maize seedlings: (i) nutritional, (ii) hypoxic, (iii) salt and (iv) heat stress. For nutritional stress, seedlings were grown in a nitrogen-depleted nutrient solution (KNO3 derived from the nutrient solution supplied to the control plants was replaced by 1 mM KCl). Hypoxic stress conditions were achieved by not bubbling air through the liquid solution for the entire experiment. For salt stress, a 100 mM NaCl concentration, which corresponds to severe salt stress in maize (Farooq et al. 2015; Henry et al. 2015; Zörb et al. 2015), was employed. Finally, an intense heat stress, generally greater than 4 °C above optimum that in the case of maize is 25 °C (Hatfield and Prueger 2015), was performed by growing seedlings in a day/night cycle at 35/30 °C air temperature. After 5 days, control and treated plants were harvested by cutting the seedlings in four different parts (as illustrated in Fig. 5a), immediately frozen in liquid nitrogen and kept at − 80 °C for subsequent RNA extraction. An average of 20 randomly selected seedlings were used per sample in each experiment. Each experiment was repeated in triplicate.
RNA extraction and cDNA synthesis
Total RNA was extracted from 250 mg frozen tissue using the TRIzol method (Invitrogen, San Giuliano Milanese, Italy). Subsequently, an aliquot of total RNA was treated with RQ1 RNAse-free DNAse (Promega, Milano, Italy). Total RNA (1 µl) was quantified using a Nanodrop 1000 (Thermo Scientific, Nanodrop Products, Wilmington, DE, USA). Finally, cDNA was synthesized from 500 ng of total RNA mixed with 1 µl of 10 µM oligo-dT, as described by Trevisan et al. (2011).
Reverse-transcription quantitative real-time PCR (RT-qPCR)
Relative quantification of transcripts by RT-qPCR was performed in a StepOne Real-Time PCR System (Applied Biosystems, Monza, Italy). Reactions were performed using SYBR Green chemistry (Applied Biosystems), following the manufacturer’s instructions. Reverse-transcribed RNA (2.5 ng) was used as template in each reaction as indicated by Manoli et al. (2014). Melting-curve analysis confirmed the absence of multiple products and primer dimers. Data were exported and analysed according to the method of Livak and Schmittgen (2001) and MIQE guidelines (Bustin et al. 2009), using MEP (membrane proteinPB1A10.07c, primers: forward 5′-TGTACTCGGCAATGCTCTTG-3′ and reverse 5′-TTTGATGCTCCAGGCTTACC-3′), as the reference gene (Manoli et al. 2012). Only transcripts showing amplification with quantification cycle (Cq) < 35 were selected for subsequent gene expression analysis.
Results
Sequence analysis and phylogenetic classification
In order to identify BZR genes in Zea mays L. genome (B 73 RefGen_V3), the Arabidopsis BZR protein sequences were used as query to perform a genome-wide search. As a result, a total of 11 BZR genes were identified in the phytozome database (Table 1).
The predicted sizes of the 11 ZmBZR TFs ranged from 139 to 651 amino acids, and the predicted isoelectric points varied from 5.11 to 10.7. The major domains of the 11 ZmBZR proteins were identified by Pfam (Punta et al. 2012). Results showed that all BZR proteins possessed BZR signature that is essential for their activity as transcription factors. Two proteins exhibited also a glycoside hydrolase catalytic domain.
Chromosomal location, gene structure, and motif analysis of CPPs in maize
A physical map was drawn to show the distribution of ZmBZR on different chromosomes of maize (Fig. 1). The 11 putative ZmBZR gene candidates were distributed across 7 of the 10 chromosomes in the maize genome. Among them, chromosome 3 had three ZmBZR genes. Two ZmBZR genes were located on each of chromosomes 4 and 7. One ZmBZR gene was situated on chromosomes 1, 2 and 9. Chromosomes 6, 8 and 10 do not include any ZmBZR genes.
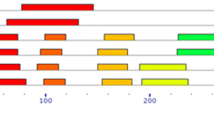
Additionally, the DNA sequences of the 11 ZmBZR TFs were determined based on the Z. mays L. whole-genome sequence. Analysis of the intron and exon distribution showed that most of the genes exhibited similar splicing patterns (Fig. 2). A BLAST search in NCBI database to compare the 11 ZmBZRs with BZRs of other species revealed that the deduced amino acid sequences of ZmBZRs shared the highest similarity levels with other monocots, as rice and Sorghum bicolor BZR TFs. The sequence similarity ranged from 50 to 94%, more specifically, 3 ZmBZRs shared greater than 80% similarity with rice BZR TFs. The similarity among the ZmBZR TFs sequences ranged from 57 to 96%, and 11 BrBZRs shared greater than 80% similarity within the species, indicating their probable duplication (Fig. 3).
Schematic representation of motifs and intro-exon distribution identified in Z. mays L. BZR proteins. The intron–exon organization patterns of 11 ZmBZR TFs are shown in panel a. Different aminoacidic motifs are indicated by different colours in panel b, and the names of all members are shown on the left side of the figure, along with their phylogenetic relatedness. The amino acidic composition of each motif is reported in panel c. (Color figure online)
The aminoacidic sequences of Arabidopsis, rice, sorghum BZR TFs from NCBI were retrieved to construct a phylogenetic tree with 11 deduced amino acid sequences of ZmBZR using the NJ method (Fig. 3). In this analysis, the close homologs of Arabidopsis BZR and BZR-homolog (BEH) were included, but they resulted distant from all the maize accessions studied. Four ZmBZR TFs formed a tight group with rice BZR1. The remaining five BZR TFs were closely grouped in a separate part of the phylogenetic tree and exhibited distant relationships with both Arabidopsis BZR TFs and the other ZmBZR. This result suggests that expansion of BZR1 and BZR2/BES1 took place after the divergence of dicots and monocots. The presence of BZR TFs specific for monocots could also be assumed.
The subcellular in silico localization of the 11 ZmBZR TFs were carried out by Protcomp 9.0 from Softberry. As transcription factor family proteins, 10 ZmBZRs were identified to have nuclear localization (Table 1). ZmBZR4 was predicted to be located in plastid of maize cells. Three ZmBZR (ZmBZR1, 5 and 7) were predicted to be located in both nucleus and plastid. Interestingly, the elements having a role in transcript localization were the CAT-box (cis-acting regulatory element related to meristem expression), Motif I (cis-acting regulatory element root specific) and Skn-1_motif (cis-acting regulatory element required for endosperm expression).
Cis-acting elements analysis
Phytohormones such as auxin, ethylene, abscisic acid (ABA), gibberellins (GAs) and jasmonic acid (JA), are involved in various processes throughout plants to overcome stress conditions. To identify the putative cis-acting regulatory elements in ZmBZR TFs, about 2000-bp of the gene CDS, from the protein start codons (ATG) were analyzed by PLACE database. The results showed that the ZmBZR genes contain various resistance- and hormone-related cis-acting elements (Fig. 4). Many key cis-elements that were related to environmental stress signal responsiveness were identified, such as MBS (MYB binding site, involved in drought-inducibility), TC-rich repeats (defence and stress-responsive element), HSE (heat shock element), LTR (low temperature-responsive element) and several light responsive elements such as G-box, Sp1, GAG-motif, and ACE (Fig. 4). Furthermore, cis-elements involved in phytohormone signaling, such as ABRE (abscisic acid-responsive element), ERE (ethylene-responsive element), TCA-element (salicylic acid-responsive element), CGTCA-motif (MeJA-responsive element), TGACG-motif (MeJA-responsive element), and P-box (gibberellin-responsive element) were also identified. ZmBZR genes also contained elements contributing to tissue-specific expression, including meristem specific elements (CCGTCC-box, CAT-box, CCGTCC-box, OCT) seed elements (RY-element), endosperm specific elements (GCN4_motif, Skn-1-like motif), trichome differentiation elements (MBSI) and vascular expression elements (AC-I, AC-II). Moreover, ZmBZR genes contained other functional elements, such as light-responsive elements and circadian control elements.
a Number of each cis-acting element in the promoter region (1 kb upstream of ATG site) of ZmBZR genes. b The statistics of total number of ZmBZR genes including corresponding cis-acting elements (red dot) and total number of cis-acting elements in ZmBZR1 gene family (blue box). Based on the functional annotation, the cis-acting elements were classified into three major classes: stress-, hormone-, development- and light responsiveness- related cis-acting elements. The regulatory elements and their descriptions are included in Supplementary Table S3. (Color figure online)
ZmBZR gene expression analyses
Expression analysis under unstressed conditions
Five of the 11 selected ZmBZR1 genes (ZmBRZ2, ZmBRZ3, ZmBRZ6, ZmBRZ7 and ZmBRZ8) were discarded since they evidenced very low amounts of transcripts in maize seedlings (data not shown). Therefore, the subsequent expression analyses were carried out on the remaining ZmBZR genes (ZmBRZ1, ZmBRZ4, ZmBRZ5, ZmBRZ9, ZmBZR10 and ZmBRZ11), as showed in Fig. 5b. ZmBZR10 displayed the highest mRNA abundance in all tissues analysed. In the apical region of root it displayed values of expression 2/4-fold higher than those measured for ZmBZR1, ZmBZR4, ZmBZR5, ZmBZR9 and ZmBZR11 (section A), while in the maturation zone (section B), mRNA levels of ZmBZR10 were 2/3-fold higher than the other. In stem region (section C) its expression was 6/10-fold higher than the others, while in leaves (section D) its transcripts were 2.5-fold more abundant than those of ZmBZR9 and 3/4-fold higher than those detected for the other ZmBZRs. Except for ZmBZR5, which showed no significant differences in terms of spatial distribution of transcripts within plant, all the remaining genes displayed the highest amount of transcripts in stem.
a Schematic picture showing the division of maize seedling for sampling material for expression analyses. A and B represent two maize root zones. The section A is enriched in meristem, transition and the elongation zone. The mature zone of the root is named B. In C and D samples are included aerial parts (stem and leaves, respectively). b RT-qPCR validation of six ZmBZR genes (ZmBZR1, ZmBZR4, ZmBZR5, ZmBZR9, ZmBZR10, ZmBZR11) in four different plant portions. Seedlings were grown in a Hoagland-modified nutrient solution for 5 days. The levels of ZmBZR gene expression were measured in total mRNAs from: meristem-, transition- and elongation-enriched root zone (A), root maturation zone (B), stem (C) and leaves (D). Data were expressed as a.u., arbitrary units
Expression analysis under stress conditions
Figure 6 describes the change in transcript level measured in each of the four plant seedlings sections after stress treatments for the six ZmBZR genes selected, independently from their relative abundance. Under N starvation conditions, an increase in transcript accumulation of these genes was observed in the root tissues. In particular in the section A (root meristem enriched in transition and elongation zone) the amount of transcripts of ZmBZR1, ZmBZR5 and ZmBZR11 was 35/50% higher than those observed in control plants, ZmBZR4 and ZmBZR9 showed an increase of transcript level of 70/80% and ZmBZR10 expression level was twofold higher respect to the control. In the stem and leaves (section C and D, respectively) the increase of transcript level was very low or insignificant for of all six ZmBZR genes. When seedlings were subjected to hypoxic stress, ZmBZR11 was found to be the most responsive gene, with a twofold transcript increase in sections A and D, and a 2.5-fold increase in the section C compared with the control plants. A twofold transcript increase, limited to sections C and D, was also detected for ZmBZR4 and, only in leaves, for ZmBZR1. An increase in the transcript level was registered also for the ZmBZR5, ZmBZR9 and ZmBZR10, even though to a lower extent.
Heat map representation of RT-qPCR of differential relative expression of six ZmBZR genes in four plant sections (A, meristem-, transition- and elongation-enriched root zone; B, root maturation zone; C, stem; D, leaves). Analysis was conducted using two independent biological repetitions. The expression levels were normalized against the maize MEP gene. Data for each region are reported as stressed/non-stressed RT-qPCR relative expression values. The colour bar indicates high to low expression respect to the control. (Color figure online)
As also observed in the case of N deficiency, when plants were subjected to NaCl treatment roots were the most responsive tissues. In particular, ZmBZR11 expression increased of threefold in section A and of twofold in section B, ZmBZR9 expression increased of twofold both in section A and B. Furthermore, in this case the gene transcript accumulation was significantly induced also in leaves tissues for ZmBZR1, ZmBZR4, ZmBZR5 and ZmBZR10, differently from what observed in the case of N starvation that affected only the level of transcript in roots.
Finally, heat stress induced an opposite effect on gene expression by down-regulating ZmBZR transcript amount in nearly all tissues. In particular in root apex (section A), a transcript reduction of between 40/50% and of 60/80% was observed for ZmBZR4, ZmBZ10, ZmBZR11 and for ZmBZR1, ZmBZR5, ZmBZR9, respectively. The same trend, even if less marked, was also observed in the root maturation zone, with a decrease of ZmBZR10 and ZmBZR9 transcription of 40 and 70%, respectively. Furthermore, ZmBZR1 showed a strong down regulation (− 60%) of its expression also in shoot.
Figure 7 describes the transcript accumulation of all ZmBZR genes along the plant in response to various stress conditions. As mentioned before, in non-stressed maize seedlings the most abundant mRNA levels were generally detected in the stem region for all ZmBZR genes. However, in response to nitrogen starvation, the highest transcript amount was detected in the root apex for ZmBZR1, ZmBZR4, ZmBZR9 and ZmBZR10, while ZmBZR11 shows the same mRNA level in both root apex and stem. ZmBZR5 did not evidence significant differences in the transcript spatial distribution within plant. A similar pattern of expression was observed in salt-stressed seedlings, although with higher variability among genes. Indeed, ZmBZR9 and ZmBZR11 showed a 1.5/2-fold increase of the mRNA abundance in the root apex while, as far as the remaining ZmBZR genes are concerned, this pattern of induction was less pronounced. Regarding hypoxic stress, no evident re-localization of ZmBZR1, ZmBZR5, ZmBZR9, and ZmBZR10 transcripts was showed. Conversely, O2-deprivation induced an increase of the transcript level both of ZmBZR4 and ZmBZR11 in the stem. Finally, heat stress triggered a re-localization of ZmBZR genes in the shoot, except for ZmBZR11, for which no differences in terms of tissue distribution was observed.
Spatial distribution of six ZmBZR genes differentially expressed after stress treatments in different plant portions: meristem-, transition- and elongation-enriched root zone (A), root maturation zone (B), stem (C) and leaves (D). Transcript abundance (%) is recorded in non-stressed maize seedlings (column C) and in response to 5 days of nutritional (N), hypoxic (O), salt and heat (T°) stress. Percentages are expressed as the ratio between the mRNA abundance measured in each specific plant zone and the global amount of transcript in the overall seedling
Discussion
The identification of a new class of plant endogenous steroidal hormones, named brassinosteroids (BRs), is the result of decades of research. Nowadays, the role of BRs both in regulating multiple physiological functions and in responding to various biotic and abiotic stresses is well established (Nawaz et al. 2017). BR perception and signal transduction involve a signaling cascade that transduces the BR signal from the cell surface to transcriptional activation in the nucleus (Kir et al. 2015). BZR1 transcription factor plays a key role in the downstream BR signaling pathway, by activating thousands of genes and repressing similar number of genes including BR biosynthetic genes via a feedback loop (Zhu et al. 2013). Considering that BRs are unable to be transported long distance, it has been proposed that BZR1 transcription factors may also serve as major connecting points among other signaling pathways (Saini et al. 2015). To understand how BRs regulate plant growth and development, as well as, they act in responding to stress conditions, a wide characterization of the transcriptional networks through which BRs regulate gene expression is necessary. To this aim the identification of BZR1 family members would be essential to elucidate the BR transcriptional networks. However, most of the information about BR signaling has been obtained from the model dicot species Arabidopsis thaliana. Although many authors suggest that Arabidopsis BZR1 TFs might play a conserved role also in rice (Tong and Chu 2012), specific components of this signaling pathway are far to be fully validated in maize.
In this work a systematic analysis was carried out to investigate the presence of BZR transcription factors in maize genome. A comprehensive set of 11 BZR transcription factors were identified and described from the current version (B73 RefGen_V3) of the B73 maize genome. In former publications, 6 and 15 BZR were identified in Arabidopsis and Brassica, respectively. BZR1 and its homologs represent a small family of plant specific proteins unrelated to any gene outside the plant kingdom (Wang et al. 2002). The presence of several members that share a similarity of more than 80% may suggest that they have overlapping or redundant functions. Motif and domain scanning showed that all of the maize BZR have the conserved BZR motifs, indicating that these maize BZR have the typical structures of the BZR TFs. Dissection of the functional domains of BZR proteins has revealed highly conserved N-terminal domains that have DNA binding activity both in vitro and in vivo (Yin et al. 2005). The BZR1 DNA binding domain (encoded by the first exon) is the most conserved region of the BZR1 proteins, as reported by He and collaborators (He et al. 2005).
BZR1 and BES1/BZR2 transcription factors are unique to plants and share high similarity at the amino acid level (Wang et al. 2002). Although the overall amino acid sequence identity among ZmBZR1, AtBZR1 and OsBZR1 is low, higher sequence identity is found in domains of important function. However, the homology between the two Arabidopsis TFs BZR1 and BZR2 (88%) is much higher than that observed between each of them and ZmBZRs. These date confirm the hypothesis that BZRs resulted from gene duplication from BZR1 only after the separation of dicots and monocots during evolution (Bai et al. 2007).
Additionally, we investigated the organ-specific expression of 11 ZmBZR genes in 5-days maize seedlings. Five of them did not show any transcript accumulation at this stage of development. The remaining genes (ZmBZR1, ZmBZR4, ZmBZR5, ZmBZR9, ZmBZR10 and ZmBZR11) were ubiquitously expressed in all the tissues examined, suggesting that they could function as growth regulators during maize development. In fact, recent studies have demonstrated that BR signaling pathway is required to regulate hypocotyl cell expansion (Gallego-Bartolomé et al. 2012; Li et al. 2012; Oh et al. 2012), as well as, to promote the transition from meristematic cells to primordial cells in the shoot (Oh et al. 2011; Zhiponova et al. 2013). In the root apex, BRs are further involved in controlling root growth, both coordinating root meristem size and also root cell elongation (Fridman et al. 2014; Heyman et al. 2013; Vilarrasa-Blasi et al. 2014; Vragovic et al. 2015). In all of these physiological processes BES1/BZR1 complex plays a pivotal role, interacting with several TFs in order to connect other signalling pathways. Interestingly, these observations could fit with our results, considering that both root apex and stem region registered the highest transcript accumulation in comparison with the other plant regions (i.e. root maturation zone and leaves) for most of the ZmBZR genes analysed in this work.
More importantly, in response to various stress conditions, the spatial transcript accumulation of all ZmBZR genes varied along the plant. This is not surprising considering than many studies have suggested essential roles for BRs in responding to various stresses; however, most of these results have been obtained by exogenously applied BRs, while the molecular basis of BR-mediated stress tolerance, including the involvement of BZR TFs, remain still elusive. Here, we demonstrate that all stress conditions tested cause a spatial transcript redistribution of BZR TFs throughout the young plant with respect to non-stressed conditions.
In response to N starvation, all six ZmBZR genes show an induction of their expression in the root system. The involvement of a BR signaling component in the regulation of the response to nutrients is to be expected, as, for example, phosphate deprivation reduces the expression of BR biosynthetic genes and shifts the intracellular localization of BZR1/BES1 (Singh et al. 2014); however, to date, it remains unclear how BR signaling is involved in N-stress responses. The application of exogenous brassinolide up-regulates a large number of NRT genes in Arabidopsis seedling roots grown on both high and low nitrate plates (Kiba et al. 2011). On the contrary, Trevisan et al. (2011) reported that the BR receptor-like kinase BRI1 expression was down-regulated after 5 days of nitrate depletion in maize. Similarly, the BRI1 kinase inhibitor 1 gene BKI1, a negative regulator involved in the BR signaling pathway, was up-regulated under N deficiency in cucumber (Zhao et al. 2015). These apparently conflicting data might be explained considering that BRs perform diverse functions by sharing signaling pathways with other phytohormones. For example, it has been demonstrated that ABA inhibits plant growth by suppressing BR signaling downstream of BR receptor (Zhang et al. 2009). An antagonistic interaction has been also evidenced between BRs and gibberellins, since the GA repressor DELLA directly interacts with BZR1 to inhibit its DNA binding and thus transcription activity in controlling photomorphogenesis (Sun et al. 2010; Li et al. 2012). In this scenario, given the apparent involvement of multiple phytohormones also in nitrogen signalling (Kiba et al. 2011), one future challenge will be to understand how BRs interact with other phytohormones to respond to N deficiency.
Regarding other abiotic stress conditions, such as hypoxia, salt and heat stress in plants, a large number of studies have demonstrated the ameliorating effect of exogenously applies BRs in promoting stress tolerance (Vardhini and Anjum 2015). This positive action is generally correlated with higher expression of stress marker genes, indicating that increased expression of stress responsive genes is responsible, at least in part, for the higher stress tolerance in BR-treated plants (Vardhini and Anjum 2015). In addition, it has been shown that application of BRs activates antioxidative pathways, including ROS-scavenging systems, as well as, non-enzymatic antioxidants, such as osmolytes like proline, glycine betaine, sorbitol, mannitol, and reduced glutathione, ascorbic acid that are needed for osmotic adjustment, stabilization of membranes, and ROS-scavenging (Fariduddin et al. 2014). However, it is still unclear whether BRs, directly or indirectly, modulate the responses of plants to oxidative stress. Interestingly, we found differential pattern of expression of all the ZmBZR genes in response to stress conditions. Most of these genes are highly up-regulated under both hypoxia and salt stress, suggesting that they might play a role in abiotic stress resistance in maize. Specifically ZmBZR4 and ZmBZR11 were found to be the most responsive gene under hypoxic conditions while ZmBZR9 and, again, ZmBZR11 were the most responsive to salt stress. An increase in the transcript level was also registered for the remaining genes, although, less pronounced. These data suggest that every single ZmBZR TFs may play a specific role in transducing different stress signals. Finally, it is worthy of attention the fact that heat stress clearly down-regulated gene expression of all ZmBZRs analysed. We speculate that this apparently contrasting result might be explained by considering the antagonistic interaction between BRs and ABA in regulating, for example, seed germination and dormancy during embryo maturation (Hu and Yu 2014). More consistently with our results, it has been demonstrated that high endogenous levels of ABA suppress BR-mediated responses in plant (Divi et al. 2010). In ABA deficient mutant aba1-1 in fact, pronounced effects of exogenously BRs applied were observed under heat stress conditions due to higher accumulation of heat shock protein 90 (Divi et al. 2010). In this scenario, ABA conceals the effects of BRs in heat stress plant response and this interaction might involve the expression of BZR genes.
Consistently with the expression results, the analysis of the promoter regions of ZmBZR genes revealed the presence of a variety of cis-acting elements, regulating gene time and space expression levels. In addition to the hormone response elements, several stress and development-related elements were identified. The analysis revealed both a common and specific distribution of elements involved in different processes. These findings support the hypothesis that ZmBZR TFs play key roles in resistance to stress, defence against pathogen invasion, and the vegetative and reproductive growth of the plants.
In conclusion, these data reveal that BZR TFs have stress signaling activity in maize, in addition to their confirmed role in regulating plant physiology and morphology.
Abbreviations
- BRs:
-
Brassinosteroids
- TFs:
-
Transcription factors
- BES:
-
BRASSINOSTEROID-INSENSITIVE 1-EMS-SUPPRESSOR
- BZR:
-
BRASSINAZOLE-RESISTANT
- Zm:
-
Zea mays L.
- RT-qPCR:
-
Reverse transcription quantitative real-time PCR
References
Bai MY, Zhang LY, Gampala SS, Zhu SW, Song WY, Chong K, Wang ZY (2007) Functions of OsBZR1 and 14–3-3 proteins in brassinosteroids signaling in rice. Proc Natl Acad Sci USA 104:13839–13844. https://doi.org/10.1073/pnas.0706386104
Bustin SA, Benes V, Garson JA, Hellemans J, Huggett J, Kubista M, Mueller R, Nolan T, Pfaffl MW, Shipley GL, Vandesompele J, Wittwer CT (2009) The MIQE guidelines: minimum information for publication of quantitative real-time PCR experiments. Clin Chem 55:611–622. https://doi.org/10.1373/clinchem.2008.112797
Chinchilla D, Shan L, He P, de Vries SC, Kemmerling B (2009) One for all: the receptor associated kinase BAK1. Trends Plant Sci 14:535–541. https://doi.org/10.1016/j.tplants.2009.08.002
Derevyanchuk M, Litvinovskaya MR, Khripach V, Kravets V (2016) Brassinosteroid-induced de novo protein synthesis in Zea mays under salinity and bioinformatic approach for identification of heat shock proteins. Plant Growth Regul 78:297–305
Divi UK, Rahman T, Krishna P (2010) Brassinosteroid-mediated stress tolerance in Arabidopsis shows interactions with abscisic acid, ethylene and salicylic acid pathways. BMC Plant Biol 10:151. https://doi.org/10.1186/1471-2229-10-151
Fariduddin Q, Yusuf M, Ahmad I, Ahmad A (2014) Brassinosteroids and their role in response of plants to abiotic stresses. Biol Plant 58:9–17. https://doi.org/10.1007/s10535-013-0374-5
Farooq M, Hussain M, Wakeel A, Siddique KHM (2015) Salt stress in maize: effects, resistance mechanisms, and management—a review. Agron Sustain Dev 35:461–481. https://doi.org/10.1007/s13593-015-0287-0
Fridman Y, Elkouby L, Holland N, Vragovic K, Elbaum R, Savaldi-Goldstein S (2014) Root growth is modulated by differential hormonal sensitivity in neighboring cells. Genes Dev 28:912–920. https://doi.org/10.1101/gad.239335.114
Gallego-Bartolomé J, Minguet EG, Grau-Enguix F, Abbas M, Locascio A, Thomas SG, Alabadí D, Blázquez MA (2012) Molecular mechanism for the interaction between gibberellin and brassinosteroid signaling pathways in Arabidopsis. Proc Natl Acad Sci USA 109:13446–13451. https://doi.org/10.1073/pnas.1119992109
Gruszka D (2013) The brassinosteroid signaling pathway–new key players and interconnections with other signaling networks crucial for plant development and stress tolerance. Int J Mol Sci 14:8740–8774. https://doi.org/10.3390/ijms14058740
Hatfield JL, Prueger JH (2015) Temperature extremes: effect on plant growth and development. Weather Clim Extremes 10:4–10. https://doi.org/10.1016/j.wace.2015.08.001
He JX, Gendron JM, Sun Y, Gampala SS, Gendron N, Sun CQ, Wang ZY (2005) BZR1 is a transcriptional repressor with dual roles in brassinosteroid homeostasis and growth responses. Science 307:1634–1638. https://doi.org/10.1126/science.1107580
He K, Gou X, Yuan T, Lin H, Asami T, Yoshida S, Russell SD, Li J (2007) BAK1 and BKK1 regulate brassinosteroid-dependent growth and brassinosteroid-independent cell-death pathways. Curr Biol 17:1109–1115. https://doi.org/10.1016/j.cub.2007.05.036
Henry C, Bledsoe SW, Griffiths CA, Kollman A, Paul MJ, Sakr S, Lagrimini LM (2015) Differential role for trehalose metabolism in salt-stressed maize. Plant Physiol 169:1072–1089. https://doi.org/10.1104/pp.15.00729
Heyman J, Cools T, Vandenbussche F, Heyndrickx KS, Van Leene J, Vercauteren I, Vanderauwera S, Vandepoele K, De Jaeger G, Van Der Straeten D, De Veylder L (2013) ERF115 controls root quiescent center cell division and stem cell replenishment. Science 342:860–863. https://doi.org/10.1126/science.1240667
Hu Y, Yu D (2014) BRASSINOSTEROID INSENSITIVE2 interacts with ABSCISIC ACID INSENSITIVE5 to mediate the antagonism of brassinosteroids to abscisic acid during seed germination in Arabidopsis. Plant Cell 26:4394–4408. https://doi.org/10.1105/tpc.114.130849
Kemmerling B, Schwedt A, Rodriguez P, Mazzota S, Frank M, Qamar SA, Mengiste T, Betsuyaku S, Parker JE, Müssig C et al (2007) The BRI1-associated kinase 1, BAK1, has a brassinolide-independent role in plant cell-death control. Curr Biol 17:1116–1122. https://doi.org/10.1016/j.cub.2007.05.046
Kiba T, Kudo T, Kojima M, Sakakibara H (2011) Hormonal control of nitrogen acquisition: roles of auxin, abscisic acid, and cytokinin. J Exp Bot 62:1399–1409. https://doi.org/10.1093/jxb/erq410
Kir G, Ye H, Nelissen H, Neelakandan AK, Kusnandar AS, Luo A, Inzé D, Sylvester AW, Yin Y, Becraft PW (2015) RNA interference knockdown of BRASSINOSTEROID INSENSITIVE1 in maize reveals novel functions for brassinosteroid signaling in controlling plant architecture. Plant Physiol 169:826–839. https://doi.org/10.1104/pp.15.00367
Li J, Nam KH, Vafeados D, Chory J (2001) BIN2, a new brassinosteroid-insensitive locus in Arabidopsis. Plant Physiol 127:14–22. https://doi.org/10.1104/pp.127.1.14
Li QF, Wang C, Jiang L, Li S, Sun SS, He JX (2012) An interaction between BZR1 and DELLAs mediates direct signaling crosstalk between brassinosteroids and gibberellins in Arabidopsis. Sci Signal 5:ra72. https://doi.org/10.1126/scisignal.2002908
Livak KJ, Schmittgen TD (2001) Analysis of relative gene expression data using real-time quantitative PCR and the 2(–Delta Delta C(T)) method. Methods 25:402–408. https://doi.org/10.1006/meth.2001.1262
Manoli A, Sturaro A, Trevisan S, Quaggiotti S, Nonis A (2012) Evaluation of candidate reference genes for qPCR in maize. J Plant Physiol 169:807–815. https://doi.org/10.1016/j.jplph.2012.01.019
Manoli A, Begheldo M, Genre A, Lanfranco L, Trevisan S, Quaggiotti S (2014) NO homeostasis is a key regulator of early nitrate perception and root elongation in maize. J Exp Bot 65:185–200. https://doi.org/10.1093/jxb/ert358
Nawaz F, Naeem M, Zulfiqar B, Akram A, Ashraf MY, Raheel M, Shabbir RN, Hussain RA, Anwar I, Aurangzaib M (2017) Understanding brassinosteroid-regulated mechanisms to improve stress tolerance in plants: a critical review. Environ Sci Pollut Res Int 24:15959–15975. https://doi.org/10.1007/s11356-017-9163-6
Oh MH, Sun J, Oh DH, Zielinski RE, Clouse SD, Huber SC (2011) Enhancing Arabidopsis leaf growth by engineering the BRASSINOSTEROID INSENSITIVE1 receptor kinase. Plant Physiol 157:120–131. https://doi.org/10.1104/pp.111.182741
Oh E, Zhu JY, Wang ZY (2012) Interaction between BZR1 and PIF4 integrates brassinosteroid and environmental responses. Nat Cell Biol 14:802–809. https://doi.org/10.1038/ncb2545
Peng W, MingSong X, Wang Z, Rong W, Hu W, Wang W, Li Y, Hou X (2016) Genome-wide analysis of the BES1 transcription factor family in Chinese cabbage (Brassica rapa ssp. pekinensis). Plant Growth Regul 80:291–301
Punta M, Coggill PC, Eberhardt RY, Mistry J, Tate J, Boursnell C, Pang N, Forslund K, Ceric G, Clements J et al (2012) The Pfam protein families database. Nucleic Acids Res 40:D290-D301. https://doi.org/10.1093/nar/gkr1065
Saha G, Park JI, Jung HJ, Ahmed NU, Kayum MA, Kang JG, Nou IS (2015) Molecular characterization of BZR transcription factor family and abiotic stress induced expression profiling in Brassica rapa. Plant Physiol Biochem 92:92–104. https://doi.org/10.1016/j.plaphy.2015.04.013
Sahni S, Prasad BD, Liu Q, Grbic V, Sharpe A, Singh SP, Krishna P (2016) Overexpression of the brassinosteroid biosynthetic gene DWF4 in Brassica napus simultaneously increases seed yield and stress tolerance. Sci Rep 6:28298. https://doi.org/10.1038/srep28298
Saini S, Sharma I, Pati PK (2015) Versatile roles of brassinosteroid in plants in the context of its homoeostasis, signaling and crosstalks. Front Plant Sci 6:950. https://doi.org/10.3389/fpls.2015.00950
Singh AP, Savaldi-Goldstein S (2015) Growth control: brassinosteroid activity gets context. J Exp Bot 66:1123–1132. https://doi.org/10.1093/jxb/erv026
Singh AP, Fridman Y, Friedlander-Shani L, Tarkowska D, Strnad M, Savaldi-Goldstein S (2014) Activity of the brassinosteroid transcription factors BRASSINAZOLE RESISTANT1 and BRASSINOSTEROID INSENSITIVE1-ETHYL METHANESULFONATE-SUPPRESSOR1/BRASSINAZOLE RESISTANT2 blocks developmental reprogramming in response to low phosphate availability. Plant Physiol 166:678–688. https://doi.org/10.1104/pp.114.245019
Sun Y, Fan XY, Cao DM, Tang W, He K, Zhu JY, He JX, Bai MY, Zhu S, Oh E et al (2010) Integration of brassinosteroid signal transduction with the transcription network for plant growth regulation in Arabidopsis. Dev Cell 19:765–777. https://doi.org/10.1016/j.devcel.2010.10.010
Tang W, Yuan M, Wang R, Yang Y, Wang C, Oses-Prieto JA, Kim TW, Zhou HW, Deng Z, Gampala SS et al (2011) PP2A activates brassinosteroid-responsive gene expression and plant growth by dephosphorylating BZR1. Nat Cell Biol 13:124–131. https://doi.org/10.1038/ncb2151
Tong H, Chu C (2012) Brassinosteroid signaling and application in rice. J Genet Genomics 39:3–9. https://doi.org/10.1016/j.jgg.2011.12.001
Trevisan S, Manoli A, Begheldo M, Nonis A, Enna M, Vaccaro S, Caporale G, Ruperti B, Quaggiotti S (2011) Transcriptome analysis reveals coordinated spatiotemporal regulation of haemoglobin and nitrate reductase in response to nitrate in maize roots. New Phytol 192:338–352. https://doi.org/10.1111/j.1469-8137.2011.03822.x
Vardhini BV, Anjum NA (2015) Brassinosteroids make plant life easier under abiotic stresses mainly by modulating major components of antioxidant defense system. Front Environ Sci 2:67. https://doi.org/10.3389/fenvs.2014.00067
Vilarrasa-Blasi J, González-García MP, Frigola D, Fàbregas N, Alexiou KG, López-Bigas N, Rivas S, Jauneau A, Lohmann JU et al (2014) Regulation of plant stem cell quiescence by a brassinosteroid signaling module. Dev Cell 30:36–47. https://doi.org/10.1016/j.devcel.2014.05.020
Vragovic K, Sela A, Friedlander-Shani L, Fridman Y, Hacham Y, Holland N, Bartom E, Mockler TC, Savaldi-Goldstein S (2015) Translatome analyses capture of opposing tissue-specific brassinosteroid signals orchestrating root meristem differentiation. Proc Natl Acad Sci USA 112:923–928. https://doi.org/10.1073/pnas.1417947112
Wang ZY, Nakano T, Gendron J, He J, Chen M, Vafeados D, Yang Y, Fujioka S, Yoshida S, Asami T, Chory J (2002) Nuclear-localized BZR1 mediates brassinosteroid-induced growth and feedback suppression of brassinosteroid biosynthesis. Dev Cell 2:505–513. (PMID: 11970900)
Wu XX, Chen JL, Xu S, Zhu ZW, Zha DS (2016) Exogenous 24-epibrassinolide alleviates zinc-induced toxicity in eggplant (Solanum melongena L.) seedlings by regulating the glutathione-ascorbate-dependent detoxification pathway. J Hortic Sci Biotech 91(4):412–420
Yin Y, Vafeados D, Tao Y, Yoshida S, Asami T, Chory J (2005) A new class of transcription factors mediates brassinosteroid-regulated gene expression in Arabidopsis. Cell 120:249–259. https://doi.org/10.1016/j.cell.2004.11.044
Yu X, Li L, Zola J, Aluru M, Ye H, Foudree A, Guo H, Anderson S, Aluru S, Liu P et al (2011) A brassinosteroid transcriptional network revealed by genome-wide identification of BES1 target genes in Arabidopsis thaliana. Plant J 65:634–646. https://doi.org/10.1111/j.1365-313X.2010.04449.x
Zhang S, Cai Z, Wang X (2009) The primary signaling outputs of brassinosteroids are regulated by abscisic acid signaling. Proc Natl Acad Sci USA 106:4543–4548. https://doi.org/10.1073/pnas.0900349106
Zhao W, Yang X, Yu H, Jiang W, Sun N, Liu X, Liu X, Zhang X, Wang Y, Gu X (2015) RNA-Seq-based transcriptome profiling of early nitrogen deficiency response in cucumber seedlings provides new insight into the putative nitrogen regulatory network. Plant Cell Physiol 56:455–467. https://doi.org/10.1093/pcp/pcu172
Zhiponova MK, Vanhoutte I, Boudolf V, Betti C, Dhondt S, Coppens F, Mylle E, Maes S, González-García MP et al (2013) Brassinosteroid production and signaling differentially control cell division and expansion in the leaf. New Phytol 197:490–502. https://doi.org/10.1111/nph.12036
Zhu JY, Sae-Seaw J, Wang ZY (2013) Brassinosteroid signalling. Development 140:1615–1620. https://doi.org/10.1242/dev.060590
Zörb C, Mühling KH, Kutschera U, Geilfus CM (2015) Salinity stiffens the epidermal cell walls of salt-stressed maize leaves: is the epidermis growth-restricting? PLos ONE 10:e0118406. https://doi.org/10.1371/journal.pone.0118406
Acknowledgements
This project and AM fellowship were supported by the grant “The role of Brassinosteroids in plant stress response and adaptation to environment”, funded by the Italian Ministry of Foreign Affairs and International Cooperation (Scientific and Technological Project of Great Relevance 2016 Italy-South Korea, No. PGR00214).
Author information
Authors and Affiliations
Contributions
The work presented here was carried out in collaboration among all authors. AM and ST performed the experiments and wrote the manuscript. SQ conceived and designed the project, analysed data, wrote the manuscript and obtained funds to support the project. SV contributed to concept the idea, helped in manuscript writing and obtained funds to support the project. All authors have read and approved the final manuscript.
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Manoli, A., Trevisan, S., Quaggiotti, S. et al. Identification and characterization of the BZR transcription factor family and its expression in response to abiotic stresses in Zea mays L.. Plant Growth Regul 84, 423–436 (2018). https://doi.org/10.1007/s10725-017-0350-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10725-017-0350-8