Abstract
Experimental studies on protein dynamics at atomic resolution by NMR-spectroscopy in solution require isolated 1H-X spin pairs. This is the default scenario in standard 1H-15N backbone experiments. Side chain dynamic experiments, which allow to study specific local processes like proton-transfer, or tautomerization, require isolated 1H-13C sites which must be produced by site-selective 13C labeling. In the most general way this is achieved by using site-selectively 13C-enriched glucose as the carbon source in bacterial expression systems. Here we systematically investigate the use of site-selectively 13C-enriched ribose as a suitable precursor for 13C labeled histidines and tryptophans. The 13C incorporation in nearly all sites of all 20 amino acids was quantified and compared to glucose based labeling. In general the ribose approach results in more selective labeling. 1-13C ribose exclusively labels His δ2 and Trp δ1 in aromatic side chains and helps to resolve possible overlap problems. The incorporation yield is however only 37% in total and 72% compared to yields of 2-13C glucose. A combined approach of 1-13C ribose and 2-13C glucose maximizes 13C incorporation to 75% in total and 150% compared to 2-13C glucose only. Further histidine positions β, α and CO become significantly labeled at around 50% in total by 3-, 4- or 5-13C ribose. Interestingly backbone CO of Gly, Ala, Cys, Ser, Val, Phe and Tyr are labeled at 40–50% in total with 3-13C ribose, compared to 5% and below for 1-13C and 2-13C glucose. Using ribose instead of glucose as a source for site-selective 13C labeling enables a very selective labeling of certain positions and thereby expanding the toolbox for customized isotope labeling of amino-acids.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
NMR spectroscopy enables high resolution studies of protein structures (Wuthrich 2001), dynamics (Palmer 2004) and interactions (Zuiderweg 2002). A key requirement for studies of protein dynamics, that are often directly linked to function (Mittermaier and Kay 2006), are isolated 1H-X spin pairs that are not affected by coupling with their neighbours. While being the default for dynamic studies of backbone amides (Akke and Palmer 1996; Ishima and Torchia 2003; Jarymowycz and Stone 2006; Loria et al. 1999), dynamics studies of amino acid side chains (Hansen and Kay 2011; Hansen et al. 2012; Lundstrom et al. 2009a; Millet et al. 2002; Muhandiram et al. 1995; Mulder et al. 2002; Paquin et al. 2008; Weininger et al. 2012a, c) often requires site selective 13C and/or 2H labeling (Lundstrom et al. 2012b). Studies of side chain dynamics not only complement existing backbone studies, but widen the view on certain processes and enable unique additional information of structure (Korzhnev et al. 2010; Neudecker et al. 2012), ring-flips (Weininger et al. 2014b; Yang et al. 2015), histidine tautomers (Weininger et al. 2017) and proton occupancy and transfer reactions (Hansen and Kay 2014; Wallerstein et al. 2015). For studies of structure, interaction and function site selective labeling is not strictly required but often advantageous, especially for large systems (Lundstrom et al. 2012a; Ruschak and Kay 2010; Tugarinov and Kay 2005) or in solid -state (Eddy et al. 2013).
In the most general way site-selective 13C labeling is achieved using glucose (Lundstrom et al. 2007; Teilum et al. 2006), glycerol (Ahlner et al. 2015), or pyruvate (Milbradt et al. 2015). These labeling schemes with precursors at the beginning of the biological pathways in bacteria, label many positions in all amino acids. Using precursors closer to the desired product result in a more exclusive labeling of certain positions. A well established case is the exclusive site selective labeling of methyl groups at high yields which results in superb NMR probes (Ruschak and Kay 2010; Tugarinov et al. 2006; Tugarinov and Kay 2005; Weininger et al. 2012b). Aromatic side chains can be targeted specifically by erythrose labeling (Kasinath et al. 2013; Weininger 2017) and more advanced chemically synthesized precursors for labeling of Trp (Schörghuber et al. 2015), Tyr and Phe (Lichtenecker et al. 2013) and most recently for His (Schörghuber et al. 2017). Also advanced in-vitro strategies using the SAIL approach have been developed for Trp (Miyanoiri et al. 2011), Tyr and Phe (Takeda et al. 2010).
Aromatic residues are an interesting target. They are bulky and form a substantial part of protein hydrophobic cores. They are also over-represented in binding sites (Lo Conte et al. 1999). Especially Tyr and Trp contribute significantly to the binding free energy (Bogan and Thorn 1998). They can be involved in specific aromatic–aromatic pair interactions (Burley and Petsko 1985, 1989), forming hydrogen bonds (Levitt and Perutz 1988), or interacting with cations (Mahadevi and Sastry 2013) or sulfur atoms (Valley et al. 2012). His and Tyr play important catalytic residues for enzyme activity (Bartlett et al. 2002). His has a pK a value close to physiological pH and can exist in three different states, one protonated and two different tautomeric neutral forms (Reynolds et al. 1973). It can act as a nucleophile, an acid/base catalyst (Fersht 1977), as a proton shuttle (Lindskog 1997), and a an hydrogen bond donor and acceptor (Krishna Deepak and Sankararamakrishnan 2016; Preimesberger et al. 2015).
Recently improved NMR methods 13C based aromatic side chain dynamics have been developed (Weininger et al. 2012a). The first studies of order parameters have been reported (Boyer and Lee 2008; Kasinath et al. 2013, 2015) and experiments to characterize dynamics on the ms (Weininger et al. 2012c) and µs (Weininger et al. 2014a) time-scales have been developed. Also site selective labeling has improved their use as structural probes (Milbradt et al. 2015) and residual dipolar couplings in aromatic side chains have been measured (Sathyamoorthy et al. 2013).
Here we present an easy and robust alternative approach using selectively labeled ribose in combination with unlabeled glucose. This approach is very close to standard 13C labeling using glucose. The only modification is the additional presence of ribose. Further, we quantify the 13C incorporation in all positions of the 20 amino acids. 1-13C ribose labeling leads to an exclusive labeling of Trp δ1 and His δ2 in aromatic side chains. His δ2 is an excellent probe for the tautomeric state of an histidine (Pelton et al. 1993; Vila et al. 2011; Weininger et al. 2017) Further these are the only positions in aromatic side chains that are per default immune against strong 1H-1H coupling artifacts in relaxation dispersion experiments (Weininger et al. 2013). The incorporation yield (37%) is however lower compared to 2-13C glucose (50%). Histidine positions β, α and CO become significantly labeled at around 50% in total by 3-, 4- or 5-13C ribose. His β does not become labeled at all using well established 1-13C or 2-13C glucose protocols and only 60% of this yield using 2-13C erythrose. Using ribose His Cβ becomes accessible for dynamics on the ms time-scale (Lundstrom et al. 2009b). Interestingly backbone CO of Gly, Ala, Cys, Ser, Val, Phe and Tyr are labeled at 40–50% in total with 3-13C ribose, compared to 5% and below for glucose. Also ribose seems to enter the chorismate pathway.
Finally, we show that the ribose-based approach for site-selective 13C labeling can be easily combined with the glucose approach, enabling a more custom labeling. A combined 1-13C ribose and 2-13C glucose labeling yields a isolated 13C incorporation in His δ2 of 75%.
Materials and methods
Expression and purification
Recombinant FKBP12 was expressed and purified as described (Weininger 2017). M9 minimal medium was subsidized at the beginning with 1 g/l 15N NH4Cl, 2 g/l unlabeled glucose 2 g/l selectively 13C enriched ribose, unless otherwise indicated. At the end the buffer was exchanged to NMR buffer and the protein was concentrated to ~12 mg/ml.
NMR spectroscopy
All spectra were run on 900 µM samples in 25 mM sodium phosphate, pH 7.0 and 10% (v/v) D2O at 25 °C and a static magnetic field strength of 14.1 T. For each sample, a 1H–15N plane of an HNCO, non-ct 1H–13C HSQCs for the aliphatic and aromatic regions, and a 1D spectrum on 13C were recorded for quantification of 13C incorporation. Intensities of different samples were referenced to intensities of a 1H–15N HSQC to account for small concentration deviations in the samples. Aromatic 13C relaxation studies were performed using L-optimized TROSY detected relaxation experiments (Weininger et al. 2012a). All spectra were processed using NMRPipe (Delaglio et al. 1995) and analysed using NMRView (Johnson 2004).
Data analysis
13C incorporation was resulting from ribose labeling was compared to glucose labeling (Weininger 2017). All positions of interest described in this article resulting from ribose labeling (and glucose labeling for comparison) were isolated and showed no signs of any 13C–13C 1J coupling. Intensities were normalized to the fully 13C enriched sample and expressed in %. By analysing multiple signals of the same kind, the relative error in the intensities of 13C covalently bound to 1H could be estimated to 1%. Errors for 13C not bound to 1H were estimated to 3%.
Results and discussion
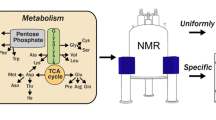
Ribose is a precursor that directly enters the pentose-5-phosphate way from which histidine and parts of tryptophan are built (Fig. 1 and SI Fig. 1 for more detail). This allows for a very distinct labeling of only the positions of interest. To make the labeling procedure as general and simple as possible and to avoid scrambling from ribose to other pathways, selective 13C labeled ribose is used in combination with unlabeled glucose. Further this allows for a possible combination of selective 13C ribose and glucose based labeling in a straightforward way. 13C incorporation was monitored for all side-chain positions, with exception of Tyr γ, His γ, and Trp δ2 and ε2. They all lack a directly attached proton which makes them harder to study and therefore less interesting. The resulting data provides information on background labeling, scrambling, and unexpected selective incorporations, as described below.
Site-selective 13C labeling of histidine and tryptophan
The above mentioned ribose labeling strategy leads to following isolated 13C labeling at the expected positions (Fig. 1) and the background labeling of other positions is much less than that obtained using glucose as the sole carbon source. The optimal amount of labeled ribose in the expression medium was tested using different amounts of 1-13C1-ribose (Fig. 2). A virtual maximum in 13C incorporation is at 2 g ribose per liter medium, whereas already at 1 g/l one is close to the maximum. 1 g/l seems to be the most economic concentration for close to optimal 13C incorporation per ribose needed. However one can still slightly increase the level of 13C incorporation by adding more ribose. In this study all (13C-site labeling) quantifications are done with 2 g/l ribose.
13C incorporation levels for the expected positions in His and Trp (see Fig. 1) are summarized in Table 1 (incorporation levels for all positions and amino acids using ribose labeling are listed in SI Table). For His δ2 and Trp δ1 the 13C incorporation using 1-13C ribose are 38 and 35%, respectively. This is a clear improvement compared to 1-13C glucose (26 and 26%), but doesn’t reach the yield of 2-13C glucose (52 and 49%). 2-13C glucose also results in isolated 13C positions which wasn’t clear from previous studies (Lundstrom et al. 2007). One potential problem of 2-13C glucose is, that it is effectively labeling Tyr ε* as well, which resonate in the same region as His δ2. 1-13C ribose however labels His δ2 exclusively (Fig. 3). Both His δ2 and Trp δ1 are not affected by 1H-1H strong coupling artifacts in relaxation dispersion experiments (Weininger et al. 2013) and His δ2 is a powerful probe for tracking the tautomeric state of histidines (Pelton et al. 1993; Vila et al. 2011; Weininger et al. 2017). Additionally 13C ribose enriched on positions 2–5 yields to very efficient and isolated labeling of Trp and His γ (though not directly shown for His), His β, His α and His CO. Especially His β is very useful since it doesn’t get isolated 13C labeled by 1-13C and 2-13C glucose and far less by 2-13C erythrose. Moreover His β is the only position that gives rise to signal in an aliphatic 1H13C HSQC that gets labeled above 3%, which means basically natural abundance. His CO seems to be labeled extremely efficient (71%) by 5-13C ribose while all other CO are below 15%. This might be a useful feature for selective HNCO experiments.
Tyr ε* His δ2 region of an aromatic 1H13C-HSQC of FKBP12. Signals arising from a 2-13C1-glucose labeled sample are shown in black, while signals from a 1-13C1-ribose labeled sample are shown in red. His δ2 signals are broadened because 15N was not decoupled. Asterisk represents an averaged signal of position 1 and 2 because of fast exchange of the aromatic rings on the NMR time-scale
13C relaxation of aromatic side chains
Both ribose and glucose labeling lead to site-selective 13C labeling in aromatic side-chains of Trp and His. By comparing R 1, R 2 and 13C NOE (Ferrage et al. 2008) for identical positions between ribose- and glucose-labeled samples, we observe an excellent agreement (Fig. 4). Thus, the two approaches give virtually the same result; potential long range 13C-13C couplings do not affect the results. While it is not clear if additional deuteration is needed for artifact free relaxation data (Kasinath et al. 2013) or not (Weininger et al. 2012a) in general, this will not affect aromatic positions that get labeled with ribose. Both His δ2 and Trp δ1 do only have one proton in 2J distance of the 13C of interest. This protons are nitrogen bound and exchange with the solvent. If they matter one has to change the solvent but not the labeling protocol. 13C relaxation dispersion experiments both for CPMG (Weininger et al. 2012c) and R 1ρ (Weininger et al. 2014a) were previously validated for glucose labeled samples. These experiments can be directly applied to samples resulting from ribose labeling, since the relaxation behaviour is identical.
Site-selective 13C labeling in non standard positions
Since ribose is a precursor closer to the end product then glucose the 13C background in other then the desired positions (Fig. 1) is much reduced (SI Table 1). However, a few positions are worth mentioning, which become efficiently labeled with 13C. In contrast to glucose all positions labeled with ribose appear to result in isolated 13C, no signs of 13C-13C couplings could be detected. 1-13C ribose only labels Tyr ζ above 10%. Since Phe ζ doesn’t show any significant 13C incorporation this might be a false positive resulting from a less reliable 13C direct detected 1D experiment. 2-13C ribose only labels Tyr ε and Phe ε to around 15%, indicating some cross over to the chorismate pathway. Indeed ribose 5-phosphate can be transformed to erythrose 4- phosphate via sedoheptulose 7-phosphate by transketolase transaldolase and transaldolase. (Schwender et al. 2003) 3-13C ribose leads to a significant 13C incorporation (30–50%) in the backbone carbonyl of Gly, Ala, Cys, Lys, Val, Trp, Phe and Tyr. 4-13C and 5-13C ribose show some weak incorporation pattern of 2-13C and 1-13C glucose, respectively. Despite the backbone carbonyl none of the positions show a higher or even close 13C incorporation compared to glucose. However they result in spectra with a reduced amount of signals and any 13C-13C couplings.
Combined labeling of ribose and glucose
Since the described labeling scheme is based on 13C labeled ribose and unlabeled glucose and the scrambling from ribose into other pathways is low, 13C labeling both from ribose and glucose can be easily combined. This was demonstrated in an approach where protein was expressed using 2 g/l 1-13C ribose and 2 g/l 2-13C glucose. Both precursors are labeling aromatic His δ2 and Trp δ1, while 2-13C glucose is additionally labeling Trp ζ3 and ζ2 and Phe and Tyr ε*. Theoretical considerations expect a labeling yield in His δ2 and Trp δ1 of about 70%: About 37% of histidine is produced from 1 to 13C ribose with 99% 13C incorporation in δ2 and about 63% is produced from 2 to 13C glucose with 51% 13C incorporation in δ2. By this approach one would maximizes the 13C labeling of His δ2. Of course this is just useful if signals from His δ2 are isolated from Tyr ε*. The experiment confirms this considerations. 75% of His δ2 and Trp δ1 get site selectively 13C labeled. This approach is generating samples with the highest sensitivity of isolated His δ2 and Trp δ1, outperforming the 2-13C glucose approach by 50% and thus nicely expanding the toolkit for a more customized site selective 13C labeling.
Different ways of site-selective 13C labeling of histidine and tryptophan
Up to date there are three different approaches of site-selective 13C labeling of histidine (CO, α, β, δ2) and tryptophan (δ1). The most general is 2-13C glucose (Lundstrom et al. 2007) which effectively (around 50%) labels His α and δ2, as well as Trp δ1. Additionally different aromatic sites (Phe and Tyr ε, and Trp ζ3 and ζ2) and α positions (all except Leu) get 13C labeled and accessible for NMR dynamic studies as well. The other two, using ribose (this work) or precursors closer to the products (Schörghuber et al. 2015, 2017) are more discriminating in the positions that get 13C labeled and can thereby solve potential overlap problems.
No precise values of 13C incorporation have been reported for the latter approaches (Schörghuber et al. 2015, 2017) nor have all positions been targeted (Trp δ1, and His α, β and δ2 are still missing). However this seems relatively straight forward to achieve and could be superior, because the starting compounds are closer to the products. The ribose approach (this work) has the disadvantage of a lower 13C incorporation in His δ2 and Trp δ1 (37%), is about the same for His α, and superior for His β and His CO, compared to the 2-13C glucose approach. If wanted 13C incorporation in His δ2 and Trp δ1 can be maximized to 75% at the cost of not selectively targeting these position anymore.
The ribose approach is about twice as expensive (for His δ2 and Trp δ1, and more for other positions) as the glucose approach, the compounds by Schörghuber require organic synthesis. Both effect the use as a standard method at the moment, but this should improve if they get more established. Even now they are very useful and superior for certain applications (overlap or sensitivity issues, new positions available). Since these compounds are just added to the regular expression medium, their use is as straight forward as any glucose labeling. They both label aromatic sites highly selective (Trp δ1 and His δ2 for ribose, Trp δ1 or His δ2 for Schörghubers compounds, after some adaptation), however the approach by Schörghuber is more discriminating for His CO.
Conclusions
We have shown that ribose as a source for site-selective 13C labeling of histidine and tryptophan yields more selective incorporation patterns than what is achieved using glucose. By this it is possible to study aromatic His δ2 signals, that are very diagnostic of the tautomeric states of histidine, without possible interference of Tyr ε* signals. If there is no interference one can maximize (75%) the 13C incorporation in His δ2 and Trp δ1 by a combination of 1-13C ribose and 2-13C glucose. Further ribose labeling leads to an improved site selective 13C incorporation in the aliphatic moiety of histidine compared to the glucose approach. Especially His β, which is not accessible by the standard 1-13C or 2-13C glucose approach, becomes significantly 13C labeled with 56% and available studies of dynamics.
References
Ahlner A, Andresen C, Khan SN, Kay LE, Lundstrom P (2015) Fractional enrichment of proteins using [2-C-13]-glycerol as the carbon source facilitates measurement of excited state C-13 alpha chemical shifts with improved sensitivity. J Biomol NMR 62:341–351
Akke M, Palmer AG (1996) Monitoring macromolecular motions on microsecond–millisecond time scales by R1r–R1 constant-relaxation-time NMR spectroscopy. J Am Chem Soc 118:911–912
Bartlett GJ, Porter CT, Borkakoti N, Thornton JM (2002) Analysis of catalytic residues in enzyme active sites. J Mol Biol 324:105–121
Bogan AA, Thorn KS (1998) Anatomy of hot spots in protein interfaces. J Mol Biol 280:1–9
Boyer JA, Lee AL (2008) Monitoring aromatic picosecond to nanosecond dynamics in proteins via C-13 relaxation: expanding perturbation mapping of the rigidifying core mutation, V54A, in eglin C. Biochemistry 47:4876–4886
Burley SK, Petsko GA (1985) Aromatic-aromatic interaction: a mechanism of protein structure stabilization. Science 229:23–28
Burley SK, Petsko GA (1989) Electrostatic interactions in aromatic oligopeptides contribute to protein. Stability Trends Biotech 7:354–359
Delaglio F, Grzesiek S, Vuister GW, Zhu G, Pfeifer J, Bax A (1995) Nmrpipe—a multidimensional spectral processing system based on unix pipes. J Biomol NMR 6:277–293
Eddy MT, Belenky M, Sivertsen AC, Griffin RG, Herzfeld J (2013) Selectively dispersed isotope labeling for protein structure determination by magic angle spinning NMR. J Biomol NMR 57:129–139. doi:10.1007/s10858-013-9773-3
Ferrage F, Piserchio A, Cowburn D, Ghose R (2008) On the measurement of 15N-{1H} nuclear Overhauser effects. J Magn Reson 192:302–313
Fersht A (1977) Enzyme structure and mechanism. W. H. Freeman, New York
Hansen AL, Kay LE (2011) Quantifying millisecond time-scale exchange in proteins by CPMG relaxation dispersion NMR spectroscopy of side-chain carbonyl groups. J Biomol NMR 50:347–355
Hansen AL, Kay LE (2014) Measurement of histidine pK(a) values and tautomer populations in invisible protein states. Proc Natl Acad Sci USA 111:E1705-E1712
Hansen AL, Lundstrom P, Velyvis A, Kay LE (2012) Quantifying millisecond exchange dynamics in proteins by CPMG relaxation dispersion NMR using side-chain H-1 probes. J Am Chem Soc 134:3178–3189
Ishima R, Torchia DA (2003) Extending the range of amide proton relaxation dispersion experiments in proteins using a constant-time relaxation-compensated CPMG approach. J Biomol NMR 25:243–248
Jarymowycz VA, Stone MJ (2006) Fast time scale dynamics of protein backbones: NMR relaxation methods, applications, and functional consequences. Chem Rev 106:1624–1671
Johnson BA (2004) Using NMRView to visualize and analyze the NMR spectra of macromolecules. Meth Mol Biol 278:313–352
Kasinath V, Valentine KG, Wand AJ (2013) A C-13 Labeling strategy reveals a range of aromatic side chain motion in calmodulin. J Am Chem Soc 135:9560–9563
Kasinath V, Fu YN, Sharp KA, Wand AJ (2015) A sharp thermal transition of fast aromatic-ring dynamics in ubiquitin. Angew Chem Int Edit 54:102-+
Korzhnev DM, Religa TL, Banachewicz W, Fersht AR, Kay LE (2010) A transient and low-populated protein-folding intermediate at atomic resolution. Science 329:1312–1316
Krishna Deepak RN, Sankararamakrishnan R (2016) N-H...N hydrogen bonds involving histidine imidazole nitrogen atoms: a new structural role for histidine residues in proteins. Biochemistry 55:3774–3783. doi:10.1021/acs.biochem.6b00253
Levitt M, Perutz MF (1988) Aromatic rings act as hydrogen-bond acceptors. J Mol Biol 201:751–754
Lichtenecker RJ, Weinhaupl K, Schmid W, Konrat R (2013) Alpha-Ketoacids as precursors for phenylalanine and tyrosine labelling in cell-based protein overexpression. J Biomol NMR 57:327–331
Lindskog S (1997) Structure and mechanism of carbonic anhydrase. Pharmacol Ther 74:1–20
Lo Conte L, Chothia C, Janin J (1999) The atomic structure of protein-protein recognition sites. J Mol Biol 285:2177–2198
Loria JP, Rance M, Palmer AG (1999) A relaxation-compensated Carr-Purcell-Meiboom-Gill sequence for characterizing chemical exchange by NMR spectroscopy. J Am Chem Soc 121:2331–2332
Lundstrom P et al (2007) Fractional C-13 enrichment of isolated carbons using [1-C-13]- or [2-C-13]-glucose facilitates the accurate measurement of dynamics at backbone C-alpha and side-chain methyl positions in proteins. J Biomol NMR 38:199–212
Lundstrom P, Hansen DF, Vallurupalli P, Kay LE (2009a) Accurate measurement of alpha proton chemical shifts of excited protein states by relaxation dispersion NMR spectroscopy. J Am Chem Soc 131:1915–1926
Lundstrom P, Lin H, Kay LE (2009b) Measuring (13)C(beta) chemical shifts of invisible excited states in proteins by relaxation dispersion NMR spectroscopy. J Biomol NMR 44:139–155
Lundstrom P, Ahlner A, Blissing AT (2012a) Isotope labeling methods for large systems isotope labeling. Biomol Nmr 992:3–15
Lundstrom P, Ahlner A, Blissing AT (2012b) Isotope labeling methods for relaxation measurements isotope. Labeling Biomol Nmr 992:63–82
Mahadevi AS, Sastry GN (2013) Cation-pi interaction: its role and relevance in chemistry, biology, and material science. Chem Rev 113:2100–2138
Milbradt AG, Arthanari H, Takeuchi K, Boeszoermenyi A, Hagn F, Wagner G (2015) Increased resolution of aromatic cross peaks using alternate C-13 labeling and TROSY. J Biomol NMR 62:291–301
Millet O, Muhandiram DR, Skrynnikov NR, Kay LE (2002) Deuterium spin probes of side-chain dynamics in proteins. 1. Measurement of five relaxation rates per deuteron in C-13-labeled and fractionally H-2-enriched proteins in solution. J Am Chem Soc 124:6439–6448
Mittermaier A, Kay LE (2006) Review - New tools provide new insights in NMR studies of protein dynamics. Science 312:224–228
Miyanoiri Y, Takeda M, Jee J, Ono AM, Okuma K, Terauchi T, Kainosho M (2011) Alternative SAIL-Trp for robust aromatic signal assignment and determination of the chi(2) conformation by intra-residue NOEs. J Biomol NMR 51:425–435
Muhandiram DR, Yamazaki T, Sykes BD, Kay LE (1995) Measurement of H-2 T-1 and T-1P relaxation-times in uniformly C-13-labeled and fractionally H-2-labeled proteins in solution. J Am Chem Soc 117:11536–11544
Mulder FAA, Hon B, Mittermaier A, Dahlquist FW, Kay LE (2002) Slow internal dynamics in proteins: application of NMR relaxation dispersion spectroscopy to methyl groups in a cavity mutant of T4 lysozyme. J Am Chem Soc 124:1443–1451
Neudecker P et al (2012) Structure of an intermediate state in protein folding and aggregation. Science 336:362–366
Palmer AG (2004) NMR characterization of the dynamics of biomacromolecules. Chem Rev 104:3623–3640
Paquin R, Ferrage F, Mulder FAA, Akke M, Bodenhausen G (2008) Multiple-timescale dynamics of side-chain carboxyl and carbonyl groups in proteins by C-13 nuclear spin relaxation. J Am Chem Soc 130:15805-+
Pelton JG, Torchia DA, Meadow ND, Roseman S (1993) Tautomeric states of the active-site histidines of phosphorylated and unphosphorylated Iii(Glc), a signal-transducing protein from Escherichia-coli, using 2-dimensional heteronuclear nmr. Techniq Prot Sci 2:543–558
Preimesberger MR, Majumdar A, Rice SL, Que L, Lecomte JT (2015) Helix-capping histidines: diversity of N-H...N hydrogen bond strength revealed by (2 h)JNN scalar couplings. Biochemistry 54:6896–6908. doi:10.1021/acs.biochem.5b01002
Reynolds WF, Peat IR, Freedman MH, Lyerla JR Jr (1973) Determination of the tautomeric form of the imidazole ring of L-histidine in basic solution by carbon-13 magnetic resonance spectroscopy. J Am Chem Soc 95:328–331
Ruschak AM, Kay LE (2010) Methyl groups as probes of supra-molecular structure, dynamics and function. J Biomol NMR 46:75–87. doi:10.1007/s10858-009-9376-1
Sathyamoorthy B, Singarapu KK, Garcia AE, Szyperski T (2013) Protein conformational space populated in solution probed with aromatic residual dipolar C-13-H-1 Couplings. Chembiochem 14:684–688
Schörghuber J, Sara T, Bisaccia M, Schmid W, Konrat R, Lichtenecker RJ (2015) Novel approaches in selective tryptophan isotope labeling by using Escherichia coli overexpression media. Chembiochem 16:746–751
Schörghuber J, Geist L, Platzer G, Konrat R, Lichtenecker RJ (2017) Highly selective stable isotope labeling of histidine residues by using a novel precursor in E. coli-Based overexpression systems. Chembiochem 18:1487–1491. doi:10.1002/cbic.201700192
Schwender J, Ohlrogge JB, Shachar-Hill Y (2003) A flux model of glycolysis and the oxidative pentosephosphate pathway in developing Brassica napus embryos. J Biol Chem 278:29442–29453. doi:10.1074/jbc.M303432200
Takeda M, Ono AM, Terauchi T, Kainosho M (2010) Application of SAIL phenylalanine and tyrosine with alternative isotope-labeling patterns for protein structure determination. J Biomol NMR 46:45–49
Teilum K, Brath U, Lundstrom P, Akke M (2006) Biosynthetic C-13 labeling of aromatic side chains in proteins for NMR relaxation measurements. J Am Chem Soc 128:2506–2507
Tugarinov V, Kay LE (2005) Methyl groups as probes of structure and dynamics in NMR studies of high-molecular-weight proteins. Chembiochem 6:1567
Tugarinov V, Kanelis V, Kay LE (2006) Isotope labeling strategies for the study of high-molecular-weight proteins by solution NMR spectroscopy. Nat Prot 1:749–754. doi:10.1038/nprot.2006.101
Valley CC, Cembran A, Perlmutter JD, Lewis AK, Labello NP, Gao J, Sachs JN (2012) The methionine-aromatic motif plays a unique role in stabilizing protein structure. J Biol Chem 287:34979–34991
Vila JA, Arnautova YA, Vorobjev Y, Scheraga HA (2011) Assessing the fractions of tautomeric forms of the imidazole ring of histidine in proteins as a function of pH. Proc Natl Acad Sci USA 108:5602–5607. doi:10.1073/pnas.1102373108
Wallerstein J, Weininger U, Khan MA, Linse S, Akke M (2015) Site-specific protonation kinetics of acidic side chains in proteins determined by pH-dependent carboxyl (13)C NMR relaxation. J Am Chem Soc 137:3093–3101
Weininger U (2017) Site-selective 13C labeling of proteins using erythrose. J Biomol NMR 67:191–200. doi:10.1007/s10858-017-0096-7
Weininger U, Diehl C, Akke M (2012a) C-13 relaxation experiments for aromatic side chains employing longitudinal- and transverse-relaxation optimized NMR spectroscopy. J Biomol NMR 53:181–190
Weininger U, Liu Z, McIntyre DD, Vogel HJ, Akke M (2012b) Specific 12CbD212CgD2S13CeHD2 isotopomer labeling of methionine to characterize protein dynamics by 1H and 13C NMR relaxation dispersion. J Am Chem Soc 134:18562–18565. doi:10.1021/ja309294u
Weininger U, Respondek M, Akke M (2012c) Conformational exchange of aromatic side chains characterized by L-optimized TROSY-selected C-13 CPMG relaxation dispersion. J Biomol NMR 54:9–14
Weininger U, Respondek M, Low C, Akke M (2013) Slow Aromatic ring flips detected despite near-degenerate NMR frequencies of the exchanging nuclei. J Phys Chem B 117:9241–9247
Weininger U, Brath U, Modig K, Teilum K, Akke M (2014a) Off-resonance rotating-frame relaxation dispersion experiment for C-13 in aromatic side chains using L-optimized TROSY-selection. J Biomol NMR 59:23–29
Weininger U, Modig K, Akke M (2014b) Ring flips revisited: C-13 relaxation dispersion measurements of aromatic side chain dynamics and activation barriers in basic pancreatic trypsin inhibitor. Biochemistry 53:4519–4525
Weininger U, Modig K, Geitner AJ, Schmidpeter PAM, Koch JR, Akke M (2017) Dynamics of aromatic side chains in the active site of FKBP12. BioChemistry 56:334–343. doi:10.1021/acs.biochem.6b01157
Wuthrich K (2001) The way to NMR structures of proteins. Nat Struct Biol 8:923–925. doi:10.1038/Nsb1101-923
Yang CJ, Takeda M, Terauchi T, Jee J, Kainosho M (2015) Differential large-amplitude breathing motions in the interface of FKBP12 drug complexes. Biochemistry 54:6983–6995. doi:10.1021/acs.biochem.5b00820
Zuiderweg ERP (2002) Mapping protein-protein interactions in solution by. NMR spectroscopy biochemistry 41:1–7. doi:10.1021/bi011870b
Acknowledgements
Protein expression and purification was performed by the Lund Protein Production Platform (LP3), Lund University, Sweden (http://www.lu.se/lp3). This research was supported by the Royal Physiographic Society of Lund and the Deutsche Forschungsgemeinschaft (WE 5587/1–1).
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Weininger, U. Site-selective 13C labeling of histidine and tryptophan using ribose. J Biomol NMR 69, 23–30 (2017). https://doi.org/10.1007/s10858-017-0130-9
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s10858-017-0130-9