Abstract
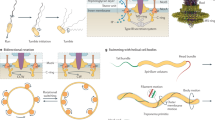
The motion of small bacteria consists of two phases: relatively long runs alternate with intermittent stops, back-ups, or tumbles, depending on the species. In polar monotrichous bacteria, the flagellum is anchored at the cell pole inherited from the parent generation (old pole) and is surrounded by a chemoreceptor cluster. During forward swimming, the leading pole is always the pole recently formed in cell division (new pole). The flagella of the peritrichous bacterium Escherichia coli often form a bundle behind the old pole. Its cell orientation and receptor positioning during runs generally mimic that of monotrichous bacteria. When encountering a solid surface, peritrichous bacteria exhibit a circular motion with the leading pole dipping downward. Some polar monotrichous bacteria also perform circular motion near solid boundaries, but during back-ups. In this case, the leading pole points upward. Very little is known about behavior near milieu-air interfaces. Biophysical simulations have revealed some of the mechanisms underlying these phenomena, but leave many questions unanswered. Combining biophysics with molecular techniques will certainly advance our understanding of bacterial locomotion.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Berg H C. E. coli in motion. In: Biological and Medical Physics Biomedical Engineering. New York: Springer-Verlag, 2003
Wadhams G H, Armitage J P. Making sense of it all: bacterial chemotaxis. Nat Rev Mol Cell Biol, 2004, 5: 1024–1037
Blair D F. How bacteria sense and swim. Annu Rev Microbiol, 1995, 49: 489–522
Sourjik V, Armitage J P. Spatial organization in bacterial chemotaxis. EMBO J, 2010, 29: 2724–2733
Wei Y, Wang X, Liu J, et al. The population dynamics of bacteria in physically structured habitats and the adaptive virtue of random motility. Proc Natl Acad Sci USA, 2011, 108: 4047–4052
Ramia M, Tullock D L, Phanthien N. The role of hydrodynamic interaction in the locomotion of microorganisms. Biophys J, 1993, 65: 755–778
Berg H C. Random Walks in Biology. Princeton: Princeton University Press, 1993
Dusenbery D B. Minimum size limit for useful locomotion by freeswimming microbes. Proc Natl Acad Sci USA, 1997, 94: 10949–10954
Berg H C, Brown D A. Chemotaxis in Escherichia coli analysed by Three-dimensional Tracking. Nature, 1972, 239: 500–504
Montesinos E. Change in size of Chromatium minus cells in relation to growth rate, sulfur content, and photosynthetic activity: A comparison of pure cultures and field populations. Appl Environ Microbiol, 1987, 53: 864–871
Ahmed T, Shimizu T S, Stocker R. Microfluidics for bacterial chemotaxis. Integr Biol, 2010, 2: 604–29
Mitchell J G, Martinez-Alonso M, Lalucat J, et al. Velocity changes, long runs, and reversals in the Chromatium minus swimming response. J Bacteriol, 1991, 173: 997–1003
Darnton N C, Turner L, Rojevsky S, et al. On torque and tumbling in swimming Escherichia coli. J Bacteriol, 2007, 189: 1756–1764
Macnab R M, Ornston M K. Normal-to-curly flagellar transitions and their role in bacterial tumbling. Stabilization of an alternative quaternary structure by mechanical force. J Mol Biol, 1977, 112: 1–30
Goto T, Nakata K, Baba K, et al. A fluid-dynamic interpretation of the asymmetric motion of singly flagellated bacteria swimming close to a boundary. Biophys J, 2005, 89: 3771–3779
Taylor B L, Koshland Jr. D E. Reversal of flagellar rotation in monotrichous and peritrichous bacteria: generation of changes in direction. J Bacteriol, 1974, 119: 640–642
Xie L, Altindal T, Chattopadhyay S, et al. Bacterial flagellum as a propeller and as a rudder for efficient chemotaxis. Proc Natl Acad Sci USA, 2011, 108: 2246–2251
Stocker R. Reverse and flick: Hybrid locomotion in bacteria. Proc Natl Acad Sci USA, 2011, 108: 2635–2636
Armitage J P, Macnab R M. Unidirectional, intermittent rotation of the flagellum of Rhodobacter sphaeroides. J Bacteriol, 1987, 169: 514–518
Armitage J P, Schmitt R. Bacterial chemotaxis: Rhodobacter sphaeroide and Sinorhizobium meliloti—variations on a theme? Microbiology, 1997, 143: 3671–3682
Vladimirov N, Lebiedz D, Sourjik V. Predicted auxiliary navigation mechanism of peritrichously flagellated chemotactic bacteria. PLoS Comput Biol, 2010, 6: e1000717
Berg H C, Purcell E M. Physics of chemoreception. Biophys J, 1977, 20: 193–219
Homma M, Oota H, Kojima S, et al. Chemotactic responses to an attractant and a repellent by the polar and lateral flagellar systems of Vibrio alginolyticus. Microbiology, 1996, 142: 2777–2783
Kudo S, Imai N, Nishitoba M, et al. Asymmetric swimming pattern of Vibrio alginolyticus cells with single polar flagella. FEMS Microbiol Lett, 2005, 242: 221–225
Magariyama Y, Masuda S, Takano Y, et al. Difference between forward and backward swimming speeds of the single polarflagellated bacterium, Vibrio alginolyticus. FEMS Microbiol Lett, 2001, 205: 343–347
Cabeen M T, Charbon G, Vollmer W, et al. Bacterial cell curvature through mechanical control of cell growth. EMBO J, 2009, 28: 1208–1219
Koyasu S, Shirakihara Y. Caulobacter crescentus flagellar filament has a right-handed helical form. J Mol Biol, 1984, 173: 125–130
Li G, Tang J X. Low flagellar motor torque and high swimming efficiency of Caulobacter crescentus swarmer cells. Biophys J, 2006, 91: 2726–2734
Li G L, Tam L K, Tang J X. Amplified effect of Brownian motion in bacterial near-surface swimming. Proc Natl Acad Sci USA, 2008, 105: 18355–18359
Vaituzis Z, Doetsch R N. Motility tracks: technique for quantitative study of bacterial movement. Appl Microbiol, 1969, 17: 584–588
Conrad J C, Gibiansky M L, Jin F, et al. Flagella and pili-mediated near-surface single-cell motility mechanisms in P. aeruginosa. Biophys J, 2011, 100: 1608–1616
Alley M, Maddock J R, Shapiro L. Requirement of the carboxyl terminus of a bacterial chemoreceptor for its targeted proteolysis. Science, 1993, 259: 1754–1757
Maddock J R, Shapiro L. Polar location of the chemoreceptor complex in the Escherichia coli cell. Science, 1993, 259: 1717–1723
Ping L, Weiner B, Kleckner N. Tsr-GFP accumulates linearly with time at cell poles, and can be used to differentiate ‘old’ versus ≠w’ poles, in Escherichia coli. Mol Microbiol, 2008, 69: 1427–1438
Thiem S, Kentner D, Sourjik V. Positioning of chemosensory clusters in E. coli and its relation to cell division. EMBO J, 2007, 26: 1615–1623
Greenfield D, McEvoy A L, Shroff H, et al. Self-organization of the Escherichia coli chemotaxis network imaged with super-resolution light microscopy. PLoS Biol, 2009, 7: e1000137
Sourjik V. Receptor clustering and signal processing in E. coli chemotaxis. Trends Microbiol, 2004, 12: 569–576
Boldog T, Grimme S, Li M, et al. Nanodiscs separate chemoreceptor oligomeric states and reveal their signaling properties. Proc Natl Acad Sci USA, 2006, 103: 11509–11514
Segall J E, Block S M, Berg H C. Temporal comparisons in bacterial chemotaxis. Proc Natl Acad Sci USA, 1986, 83: 8987–8991
Duke T A J, Bray D. Heightened sensitivity of a lattice of membrane receptors. Proc Natl Acad Sci USA, 1999, 96: 10104–10108
Sourjik V, Berg H C. Functional interactions between receptors in bacterial chemotaxis. Nature, 2004, 428: 437–441
Khursigara C M, Lan G, Neumann S, et al. Lateral density of receptor arrays in the membrane plane influences sensitivity of the E. coli chemotaxis response. EMBO J, 2011, 30: 1719–1729
Hansen C H, Sourjik V, Wingreen N S. A dynamic-signaling-team model for chemotaxis receptors in Escherichia coli. Proc Natl Acad Sci USA, 2010, 107: 17170–17175
Aquino G, Clausznitzer D, Tollis S, et al. Optimal receptor-cluster size determined by intrinsic and extrinsic noise. Phys Rev E, 2011, 83: 021914
Berg H C, Turner L. Cells of Escherichia coli swim either end forward. Proc Natl Acad Sci USA, 1995, 92: 477–479
Ping L. The asymmetric flagellar distribution and motility of Escherichia coli. J Mol Biol, 2010, 397: 906–916
Wadhams G H, Martin A C, Armitage J P. Identification and localization of a methyl-accepting chemotaxis protein in Rhodobacter sphaeroides. Mol Microbiol, 2000, 36: 1222–1233
Thar R, Kühl M. Bacteria are not too small for spatial sensing of chemical gradients: An experimental evidence. Proc Natl Acad Sci USA, 2003, 100: 5748–5753
Hill J, Kalkanci O, McMurry J L, et al. Hydrodynamic surface interactions enable Escherichia coli to seek efficient routes to swim upstream. Phys Rev Lett, 2007, 98: 068101
Vigeant M A S, Ford R M, Wagner M, et al. Reversible and irreversible adhesion of motile Escherichia coli cells analyzed by total internal reflection aqueous fluorescence microscopy. Appl Environ Microbiol, 2002, 68: 2794–2801
Hsu R, Ganatos P. The motion of a rigid body in viscous-fluid bounded by a plane Wall. J Fluid Mech, 1989, 207: 29–72
Shum H, Gaffney E A, Smith D J. Modelling bacterial behaviour close to a no-slip plane boundary: the influence of bacterial geometry. Philos Trans R Soc Lond A, 2010, 466: 1725–1748
Cabeen M T, Jacobs-Wagner C. The bacterial cytoskeleton. Annu Rev Genet, 2010, 44: 365–392
van den Ent F, Johnson C M, Persons L, et al. Bacterial actin MreB assembles in complex with cell shape protein RodZ. EMBO J, 2010, 29: 1081–1090
Frankel R B, Bazylinski D A, Johnson M S, et al. Magneto-aerotaxis in marine coccoid bacteria. Biophys J, 1997, 73: 994–1000
Aswad D, Koshland Jr. D E. Isolation, characterization and complementation of Salmonella typhimurium chemotaxis mutants. J Mol Biol, 1975, 97: 225–235
Lauga E, DiLuzio W R, Whitesides G M, et al. Swimming in circles: motion of bacteria near solid boundaries. Biophys J, 2006, 90: 400–412
Frymier P D, Ford R M, Berg H C, et al. Three-dimensional tracking of motile bacteria near a solid planar surface. Proc Natl Acad Sci USA, 1995, 92: 6195–9
DiLuzio W R, Turner L, Mayer M, et al. Escherichia coli swim on the right-hand side. Nature, 2005, 435: 1271–1274
Maeda K, Imae Y, Shioi J I, et al. Effect of temperature on motility and chemotaxis of Escherichia coli. J Bacteriol, 1976, 127: 1039–1046
Oleksiuk O, Jakovljevic V, Vladimirov N, et al. Thermal robustness of signaling in bacterial chemotaxis. Cell, 2011, 145: 312–321
Lowe G, Meister M, Berg H C. Rapid rotation of flagellar bundles in swimming bacteria. Nature, 1987, 325: 637–640
Boehm A, Kaiser M, Li H, et al. Second messenger-mediated adjustment of bacterial swimming velocity. Cell, 2010, 141: 107–116
Paul K, Nieto V, Carlquist W C, et al. The c-di-GMP binding protein YcgR controls flagellar motor direction and speed to affect chemotaxis by a backstop brake mechanism. Mol Cell, 2010, 38: 128–139
Berg H C, Turner L. Chemotaxis of bacteria in glass-capillary arrays—Escherichia coli, motility, microchannel plate, and light-scattering. Biophys J, 1990, 58: 919–930
Magariyama Y, Ichiba M, Nakata K, et al. Difference in bacterial motion between forward and backward swimming caused by the wall effect. Biophys J, 2005, 88: 3648–3658
Loosdrecht M C M, Norde W, Lyklema J, et al. Hydrophobic and electrostatic parameters in bacterial adhesion. Aquat Sci, 1990, 52: 103–114
Magariyama Y, Sugiyama S, Muramoto K, et al. Simultaneous measurement of bacterial flagellar rotation rate and swimming speed. Biophys J, 1995, 69: 2154–2162
Nakai T, Kikuda M, Kuroda Y, et al. Speed, trajectory and increment in the number of cells of singly flagellated bacteria swimming close to boundaries. J Biomech Sci Eng, 2009, 4: 2–10
Vigeant M, Ford R. Interactions between motile Escherichia coli and glass in media with various ionic strengths, as observed with a three-dimensional-tracking microscope. Appl Environ Microbiol, 1997, 63: 3474–3479
McCarter L L. Polar flagellar motility of the Vibrionaceae. Microbiol Mol Biol Rev, 2001, 65: 445–462
Di Leonardo R, Dell’Arciprete D, Angelani L, et al. Swimming with an image. Phys Rev Lett, 2011, 106: 038101
Lemelle L, Palierne J F, Chatre E, et al. Counter-clockwise circular motion of bacteria swimming at air/liquid interface. J Bacteriol, 2010, 192: 6307–6308
Zhang H F, Maslov K, Stoica G, et al. Functional photoacoustic microscopy for high-resolution and noninvasive in vivo imaging. Nat Biotech, 2006, 24: 848–851
Garrity G M, Brenner D J, Krieg N R, et al. The Proteobacteria. In: Garrity G M, Brenner D J, Krieg N R, et al., eds. Bergey’s Manual of Systematic Bacteriology. Vol. 2. New York: Springer-Verlag, 2005
Lefèvre C T, Song T, Yonnet J P, et al. Characterization of bacterial magnetotactic behaviors by using a magnetospectrophotometry Assay. Appl Environ Microbiol, 2009, 75: 3835–3841
Matsunaga T, Sakaguchi T, Tadakoro F. Magnetite formation by a magnetic bacterium capable of growing aerobically. Appl Microbiol Biotechnol, 1991, 35: 651–655
Harshey R M. Bacterial motility on a surface: Many ways to a common goal. Annu Rev Microbiol, 2003, 57: 249–273
Luchsinger R H, Bergersen B, Mitchell J G. Bacterial swimming strategies and turbulence. Biophys J, 1999, 77: 2377–2386
Young K D. The selective value of bacterial shape. Microbiol Mol Biol Rev, 2006, 70: 660–703
Weibel D B, DiLuzio W R, Whitesides G M. Microfabrication meets microbiology. Nat Rev Microbiol, 2007, 5: 209–218
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Ping, L. Cell orientation of swimming bacteria: From theoretical simulation to experimental evaluation. Sci. China Life Sci. 55, 202–209 (2012). https://doi.org/10.1007/s11427-012-4298-7
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-012-4298-7