Abstract
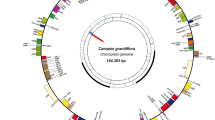
Magnolia grandiflora is an important medicinal, ornamental and horticultural plant species. The chloroplast (cp) genome of M. grandiflora was sequenced using a 454 sequencing platform and the genome structure was compared with other related species. The complete cp genome of M. grandiflora was 159623 bp in length and contained a pair of inverted repeats (IR) of 26563 bp separated by large and small single copy (LSC, SSC) regions of 87757 and 18740 bp, respectively. A total of 129 genes were successfully annotated, 18 of which included introns. The identity, number and GC content of M. grandiflora cp genes were similar to those of other Magnoliaceae species genomes. Analysis revealed 218 simple sequence repeat (SSR) loci, most composed of A or T, contributing to a bias in base composition. The types and abundances of repeat units in Magnoliaceae species were relatively conserved and these loci will be useful for developing M. grandiflora cp genome vectors. In addition, results indicated that the cp genome size in Magnoliaceae species and the position of the IR border were closely related to the length of the ycf1 gene. Phylogenetic analyses based on 66 shared genes from 30 species using maximum parsimony (MP) and maximum likelihood (ML) methods provided strong support for the phylogenetic position of Magnolia. The availability of the complete cp genome sequence of M. grandiflora provides valuable information for breeding of desirable varieties, cp genetic engineering, developing useful molecular markers and phylogenetic analyses in Magnoliaceae.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Verma D, Daniell H. Chloroplast vector systems for biotechnology applications. Plant Physiol, 2007, 145: 1129–1143
Clegg M T, Gaut B S, Learn G H, et al. Rates and patterns of chloroplast DNA evolution. Proc Natl Acad Sci USA, 1994, 91: 6795–6801
Shinozaki K, Ohme M, Tanaka M, et al. The complete nucleotide sequence of the tobacco chloroplast genome: its gene organization and expression. EMBO J, 1986, 5: 2043–2049
Ohyama K, Fukuzawa H, Kohchi T, et al. Chloroplast gene organization deduced from complete sequence of Liverwort Marchantia polymorpha chloroplast DNA. Nature, 1986, 322: 572–574
Terada R, Urawa H, Inagaki Y, et al. Efficient gene targeting by homologous recombination in rice. Nat Biotechnol, 2002, 20: 1030–1034
Kane N, Sveinsson S, Dempewolf H, et al. Ultra-barcoding in cacao (Theobroma spp.; Malvaceae) using whole chloroplast genomes and nuclear ribosomal DNA. Am J Bot, 2012, 99: 320–329
Zhang Y J, Ma P F, Li D Z. High-Throughput Sequencing of Six Bamboo Chloroplast Genomes: Phylogenetic Implications for Temperate Woody Bamboos (Poaceae: Bambusoideae). PLoS One, 2011, 6: e20596
Kanevski I, Maliga P, Rhoades D F, et al. Plastome engineering of ribulose-1, 5-bisphosphate carboxylase/oxygenase in tobacco to form a sunflower large subunit and tobacco small subunit hybrid. Plant Physiol, 1999, 119: 133–142
Craig W, Lenzi P, Scotti N, et al. Transplastomic tobacco plants expressing a fatty acid desaturase gene exhibit altered fatty acid profiles and improved cold tolerance. Transgenic Res, 2008, 17: 769–782
Jansen K J, Zhengqiu C, Linda A R, et al. Analysis of 81 genes from 64 plastid genomes resolves relationships in angiosperms and identifies genome-scale evolutionary patterns. Proc Natl Acad Sci USA, 2007, 104: 19369–19374
Xu F, Rudall P J. Comparative floral anatomy and ontogeny in Magnoliaceae. Plant Syst Evol, 2006, 258: 1–15
Yamada T, Imaichi R, et al. The outer integument and funicular outgrowth complex in the ovule of Magnolia grandiflora (Magnoliaceae). J Plant Res, 2003, 116: 189–198
Kim S, Park C W, Kim Y D, et al. Phylogenetic relationships in family Magnoliaceae inferred from ndhF sequences. Am J Bot, 2001, 88: 717–728
Sauquet H, Doyle J A, Scharaschkin T, et al. Phylogenetic analysis of Magnoliales and Myristicaceae based on multiple data sets: implications for character evolution. Bot J Linn Soc, 2003, 142: 125–186
Fu D L. Notes on Yulania Spach. J Wuhan Bot Res, 2001, 19: 191–198
Wang Q, Wang Z Z, Li Y L. Study on tissue culture of Magnolia grandiflora L.. Northwest Pharm J, 2001, 16: 11–13
Lee S, Chappell J. Biochemical and genomic characterization of terpene synthases in Magnolia grandiflora. Plant Physiol, 2008, 147: 1017–1033
Wang Z G, Wu C Q, Wang C Y, et al. Effect of Magnolia grandifore oil on lipid metabolism in hyperlipoidemic rat. Chin Trad Patent Med, 2010, 32: 1679–1682
Li X W, Hu Z G, Lin X H, et al. High-throughput pyrosequencing of the complete chloroplast genome of Magnolia officinalis and its application in species identification. Acta Pharm Sin, 2012, 47: 124–130
Cai Z Q, Penaflor C, Kuehl J V, et al. Complete plastid genome sequence of Drimys, Liriodendron, and Piper: implications for the phylogenetic relationships of magnoliids. BMC Evol Biol, 2006, 6: 77
Xu J W, Feng D J, Song G S, et al. The first introns in rice EPSP synthase enhance exogenous gene expression. Science China Ser C-Life Sci, 2003, 33: 224–230
Fukuzawa H, Kohchi T, Shirai H, et al. Coding sequences for chloroplast ribosomal protein S12 from the liverwort Marchantia polymorpha, are separated far apart on the different DNA Strand. FEBS Lett, 1986, 198: 11–15
Kim K J, Lee H L. Complete chloroplast genome sequences from Korean Ginseng (Panax schinseng Nees) and comparative analysis of sequence evolution among 17 vascular plants. DNA Res, 2004, 11: 247–261
Kuang D Y, Wu H, Wang Y L, et al. Complete chloroplast genome sequence of Magnolia kwangsiensis (Magnoliaceae): implication for DNA barcoding and population genetics. Genome, 2011, 54: 663–673
Xu G X, Guo C C, Shan H Y, et al. Divergence of duplicate genes in exon-intron structure. Proc Natl Acad Sci USA, 2012, 109: 1187–1192
Kaundun S S, Matsumoto S. Heterologous nuclear and chloroplast microsatellite amplification and variation in tea Camellia sinensis. Genome, 2002, 45: 1041–1048
Jiao Y, Jia H M, Li X W, et al. Development of simple sequence repeat (SSR) markers from a genome survey of Chinese Bayberry (Myrica rubra). BMC Genomics, 2012, 13: 201
Ravi V, Khurana J P, Tyagi A K, et al. An update on chloroplast genomes. Plant Syst Evol, 2008, 271: 101–122
Michael J M, Soltis, P S, Bell C D, et al. Phylogenetic analysis of 83 plastid genes further resolves the early diversification of eudicots. Proc Natl Acad Sci USA, 2010, 107: 4623–4628
Azuma H, Thien L B, Kawano S. Molecular phylogeny of Magnolia (Magnoliaceae) inferred from cpDNA sequences and evolutionary divergence of the floral scents. J Plant Res, 1999, 112: 291–306
Azuma H, Thien L B, Kawano S. Molecular phylogeny of Magnolia based on chloroplast DNA sequence data (trnK intron, psbA-trnH and atpB-rbcL intergenic spacer regions) and floral scent chemistry. In: Liu Y H, Fan H M, eds. Proceedings of the International Symposium on the Family Magnoliaceae. Beijing: Science Press, 2000. 219–227
Azuma H, Garcia-Franco J G, Rico-Gray V, et al. Molecular phylogeny of the Magnoliaceae: the biogeography of tropical and temperate disjunctions. Am J Bot, 2001, 88: 2275–2285
Ueda K, Yamashita J, Tamura M N. Molecular phylogeny of the Magnoliaceae. In: Liu Y H, Fan H M, eds. Proceedings of the International Symposium on the Family Magnoliaceae. Beijing: Science Press, 2000. 205–209
Wang Y L, Zhang S Z, Cui T C. The utility of trnL intron and trnL-trnF IGS in phylogenetic analysis of Magnoliaceae. Acta Bot Boreali-Occident Sin, 2003, 23: 247–252
Wang Y L, Li Y, Zhang S Z, et al. The utility of matk gene in the phylogenetic analysis of the genus Magnolia. Acta Phytotaxon Sin, 2006: 135–147
Author information
Authors and Affiliations
Corresponding author
Additional information
Contributed equally to this work
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Li, X., Gao, H., Wang, Y. et al. Complete chloroplast genome sequence of Magnolia grandiflora and comparative analysis with related species. Sci. China Life Sci. 56, 189–198 (2013). https://doi.org/10.1007/s11427-012-4430-8
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-012-4430-8