Abstract
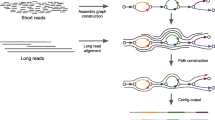
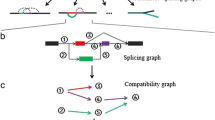
The fast development of next-generation sequencing technology presents a major computational challenge for data processing and analysis. A fast algorithm, de Bruijn graph has been successfully used for genome DNA de novo assembly; nevertheless, its performance for transcriptome assembly is unclear. In this study, we used both simulated and real RNA-Seq data, from either artificial RNA templates or human transcripts, to evaluate five de novo assemblers, ABySS, Mira, Trinity, Velvet and Oases. Of these assemblers, ABySS, Trinity, Velvet and Oases are all based on de Bruijn graph, and Mira uses an overlap graph algorithm. Various numbers of RNA short reads were selected from the External RNA Control Consortium (ERCC) data and human chromosome 22. A number of statistics were then calculated for the resulting contigs from each assembler. Each experiment was repeated multiple times to obtain the mean statistics and standard error estimate. Trinity had relative good performance for both ERCC and human data, but it may not consistently generate full length transcripts. ABySS was the fastest method but its assembly quality was low. Mira gave a good rate for mapping its contigs onto human chromosome 22, but its computational speed is not satisfactory. Our results suggest that transcript assembly remains a challenge problem for bioinformatics society. Therefore, a novel assembler is in need for assembling transcriptome data generated by next generation sequencing technique.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Martin J A, Wang Z. Next-generation transcriptome assembly. Nat Rev Genet, 2011, 12: 671–682
Schliesky S, Gowik U, Weber A P, et al. RNA-Seq assembly-are we there yet? Front Plant Sci, 2012, 3: 220
Li Z, et al. Comparison of the two major classes of assembly algorithms: overlap-layout-consensus and de-bruijn-graph. Brief Funct Genom, 2012, 11: 25–37
Myers E W. Toward simplifying and accurately formulating fragment assembly. J Comput Biol, 1995, 2: 275–290
Chevreux B, Pfisterer T, Drescher B, et al. Using the miraEST assembler for reliable and automated mRNA transcript assembly and SNP detection in sequenced ESTs. Genome Res, 2004, 14: 1147–1159
Mullikin J C, Ning Z. The phusion assembler. Genome Res, 2003, 13: 81–90
Margulies M, Egholm M, Altman W E, et al. Genome sequencing in microfabricated high-density picolitre reactors. Nature, 2005, 437: 376–380
Idury R M, Waterman M S. A new algorithm for DNA sequence assembly. J Comput Biol, 1995, 2: 291–306
Simpson J T, Wong K, Jackman S D, et al. ABySS: a parallel assembler for short read sequence data. Genome Res, 2009, 19: 1117–1123
Zerbino D R, Birney E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res, 2008, 18: 821–829
Grabherr M G, Haas B J, Yassour M, et al. Full-length transcriptome assembly from RNA-Seq data without a reference genome. Nat Biotechnol, 2011, 29: 644–652
Schulz M H, Zerbino D R, Vingron M, et al. Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics, 2012, 28: 1086–1092
Hsu F, Kent W J, Clawson H, et al. The UCSC known genes. Bioinformatics, 2006, 22: 1036–1046
Jiang L, Schlesinger F, Davis C A, et al. Synthetic spike-in standards for RNA-seq experiments. Genome Res, 2011, 21: 1543–1551
Langmead B, Trapnell C, Pop M, et al. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol, 2009, 10: R25
Au K F, Jiang H, Lin L, et al. Detection of splice junctions from paired-end RNA-seq data by SpliceMap. Nucleic Acids Res, 2010, 38: 4570–4578
Trapnell C, Pachter L, Salzberg S L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics, 2009, 25: 1105–1111
Trapnell C, Roberts A, Goff L, et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat Prot, 2012, 7: 562–578
Lander E S, Linton L M, Birren B, et al. Initial sequencing and analysis of the human genome. Nature, 2001, 409: 860–921
External RNA Controls Consortium. Proposed methods for testing and selecting the ERCC external RNA controls. BMC genomics, 2005, 6: 150
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 2.0 International License (https://creativecommons.org/licenses/by/2.0), which permits unrestricted use, distribution, and reproduction in any medium, provided the original work is properly cited.
About this article
Cite this article
Clarke, K., Yang, Y., Marsh, R. et al. Comparative analysis of de novo transcriptome assembly. Sci. China Life Sci. 56, 156–162 (2013). https://doi.org/10.1007/s11427-013-4444-x
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-013-4444-x