Abstract
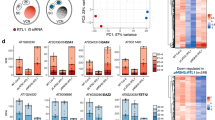
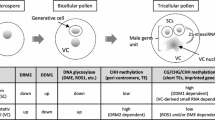
In plants, each pollen mother cell undergoes two rounds of cell divisions to form a mature pollen grain, which contains a vegetative cell (VC) and two sperm cells (SC). As a companion cell, the VC carries the SCs to an ovule by germinating a pollen tube. In-depth sequencing analyses of mature pollen showed that microRNAs (miRNAs) and short interfering RNAs (siRNAs) are present in both the VC and SCs. Additionally, epigenetically-regulated transposable elements (TEs) are reactivated in the VC and these TE mRNAs are further processed into 21-nt epigenetically reactivated siRNA (easiRNA) in SCs, which prevent 24-nt siRNA accumulation and sequester miRNA loading. Small RNAs are thought to move from the VC to SCs, where they regulate gene expression and reinforce TE silencing. Here, we summarize current knowledge of the biogenesis and function of miRNAs, siRNAs, and easiRNAs in pollen, emphasizing how these different small RNAs coordinately contribute to sperm cell formation and TE silencing.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
McCormick S. Control of male gametophyte development. Plant Cell, 2004, 16(Suppl): S142–153
Eady C, Lindsey K, Twell D. The significance of microspore division and division symmetry for vegetative cell-specific transcription and generative cell differentiation. Plant Cell, 1995, 7: 65–74
Borg M, Twell D. Life after meiosis: patterning the angiosperm male gametophyte. Biochem Soc Trans, 2010, 38: 577–582
Rotman N, Durbarry A, Wardle A, Yang WC, Chaboud A, Faure JE, Berger F, Twell D. A novel class of MYB factors controls sperm-cell formation in plants. Curr Biol, 2005, 15: 244–248
Brownfield L, Hafidh S, Borg M, Sidorova A, Mori T, Twell D. A plant germline-specific integrator of sperm specification and cell cycle progression. PLoS Genet, 2009, 5: e1000430
Borg M, Brownfield L, Khatab H, Sidorova A, Lingaya M, Twell D. The R2R3 MYB transcription factor DUO1 activates a male germline-specific regulon essential for sperm cell differentiation in Arabidopsis. Plant Cell, 2011, 23: 534–549
Zheng B, He H, Zheng Y, Wu W, McCormick S. An ARID domain-containing protein within nuclear bodies is required for sperm cell formation in Arabidopsis thaliana. PLoS Genet, 2014, 10: e1004421
Rogers K, Chen X. Biogenesis, turnover, and mode of action of plant microRNAs. Plant Cell, 2013, 25: 2383–2399
Allen E, Xie Z, Gustafson AM, Carrington JC. microRNA-directed phasing during trans-acting siRNA biogenesis in plants. Cell, 2005, 121: 207–221
Jin H, Vacic V, Girke T, Lonardi S, Zhu JK. Small RNAs and the regulation of cis-natural antisense transcripts in Arabidopsis. BMC Mol Biol, 2008, 9: 6
Matzke MA, Mosher RA. RNA-directed DNA methylation: an epigenetic pathway of increasing complexity. Nat Rev Genet, 2014, 15: 394–408
Grant-Downton R, Hafidh S, Twell D, Dickinson H. Small RNA pathways are present and functional in the angiosperm male gametophyte. Mol Plant, 2009 2:500–512
Chambers C, Shuai B. Profiling microRNA expression in Arabidopsis pollen using microRNA array and real-time PCR. BMC Plant Biol, 2009, 9: 87
Grant-Downton R, Le Trionnaire G, Schmid R, Rodriguez-Enriquez J, Hafidh S, Mehdi S, Twell D, Dickinson H. microRNA and tasiRNA diversity in mature pollen of Arabidopsis thaliana. BMC Genomics, 2009, 10: 643
Slotkin RK, Vaughn M, Borges F, Tanurdzic M, Becker JD, Feijo JA, Martienssen RA. Epigenetic reprogramming and small RNA silencing of transposable elements in pollen. Cell, 2009, 136: 461–472
Borges F, Pereira PA, Slotkin RK, Martienssen RA, Becker JD. microRNA activity in the Arabidopsis male germline. J Exp Bot, 2011, 62: 1611–1620
Calarco JP, Borges F, Donoghue MT, Van Ex F, Jullien PE, Lopes T, Gardner R, Berger F, Feijo JA, Becker JD, Martienssen RA. Reprogramming of DNA methylation in pollen guides epigenetic inheritance via small RNA. Cell, 2012, 151: 194–205
Creasey KM, Zhai J, Borges F, Van Ex F, Regulski M, Meyers BC, Martienssen RA. miRNAs trigger widespread epigenetically activated siRNAs from transposons in Arabidopsis. Nature, 2014, 508: 411–415
Li J, Wu Y, Qi Y. MicroRNAs in a multicellular green alga Volvox carteri. Sci China Life Sci, 2014, 1: 36–45
Borges F, Gomes G, Gardner R, Moreno N, McCormick S, Feijo JA, Becker JD. Comparative transcriptomics of Arabidopsis sperm cells. Plant Physiol, 2008, 148: 1168–1181
Mi S, Cai T, Hu Y, Chen Y, Hodges E, Ni F, Wu L, Li S, Zhou H, Long C, Chen S, Hannon GJ, Qi Y. Sorting of small RNAs into Arabidopsis argonaute complexes is directed by the 5′ terminal nucleotide. Cell, 2008, 133: 116–127
Grant-Downton R, Kourmpetli S, Hafidh S, Khatab H, Le Trionnaire G, Dickinson H, Twell D. Artificial microRNAs reveal cell-specific differences in small RNA activity in pollen. Curr Biol, 2013, 23: R599–601
Brosnan CA, Voinnet O. Cell-to-cell and long-distance siRNA movement in plants: mechanisms and biological implications. Curr Opin Plant Biol, 2011, 14: 580–587
Marin-Gonzalez E, Suarez-Lopez P. “And yet it moves”: cell-to-cell and long-distance signaling by plant microRNAs. Plant Sci, 2012, 196: 18–30
Ron M, Alandete Saez M, Eshed Williams L, Fletcher JC, McCormick S. Proper regulation of a sperm-specific cis-nat-siRNA is essential for double fertilization in Arabidopsis. Genes Dev, 2010, 24: 1010–1021
Fu Q, Wang PJ. Mammalian piRNAs: Biogenesis, function, and mysteries. Spermatogenesis, 2014, 4: e27889
Lippman Z, May B, Yordan C, Singer T, Martienssen R. Distinct mechanisms determine transposon inheritance and methylation via small interfering RNA and histone modification. PLoS Biol, 2003, 1: e67
McCue AD, Nuthikattu S, Slotkin RK. Genome-wide identification of genes regulated in trans by transposable element small interfering RNAs. RNA Biol, 2013, 10: 1379–1395
Nuthikattu S, McCue AD, Panda K, Fultz D, DeFraia C, Thomas EN, Slotkin RK. The initiation of epigenetic silencing of active transposable elements is triggered by RDR6 and 21–22 nucleotide small interfering RNAs. Plant Physiol, 2013, 162: 116–131
Song X, Li P, Zhai J, Zhou M, Ma L, Liu B, Jeong DH, Nakano M, Cao S, Liu C, Chu C, Wang XJ, Green PJ, Meyers BC, Cao X. Roles of DCL4 and DCL3b in rice phased small RNA biogenesis. Plant J, 2012, 69: 462–474
Fei Q, Xia R, Meyers BC. Phased, secondary, small interfering RNAs in posttranscriptional regulatory networks. Plant Cell, 2013, 25: 2400–2415
Saze H, Kakutani T. Heritable epigenetic mutation of a transposon-flanked Arabidopsis gene due to lack of the chromatin-remodeling factor DDM1. EMBO J, 2007, 26: 3641–3652
Sarazin A, Voinnet O. Exploring new models of easiRNA biogenesis. Nat Genet, 2014, 46: 530–531
McCue AD, Nuthikattu S, Reeder SH, Slotkin RK. Gene expression and stress response mediated by the epigenetic regulation of a transposable element small RNA. PLoS Genet, 2012, 8: e1002474
Scarpin R, Sigaut L, Pietrasanta L, McCormick S, Zheng B, Muschietti J. Cajal Bodies are developmentally regulated during pollen development and pollen tube growth in Arabidopsis thaliana. Mol Plant, 2013, 6: 1355–1357
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at springerlink.fh-diploma.de
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
He, H., Yang, T., Wu, W. et al. Small RNAs in pollen. Sci. China Life Sci. 58, 246–252 (2015). https://doi.org/10.1007/s11427-015-4800-0
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11427-015-4800-0