Abstract
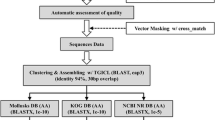
cDNA library of Myrmecia incisa H4301 was constructed using λ phage vectors. The library consisted of 1.5×106 clones with a recombination rate of 90%, and 1942 clones were randomly sequenced. All 1854 readable expressed sequence tags (ESTs) were clustered into 596 non-redundant sequences (NRSs), among which 126 NRSs were from 1384 ESTs, showing a high redundancy. Among the 596 NRSs, 30 were ribosomal RNA, and 152 significantly matched with those available in NCBI database and JGI Genome Portal, the latter were divided into nine subcategories. Overall, 59 NRSs were involved in photosynthesis, the respiratory electron transport chain, ATP synthesis, oxidation reduction, fatty acid biosynthesis, glucose metabolism, protein metabolism, and small molecular metabolism, suggesting that these genes were abundantly transcribed during energy and substance metabolism. Acyl-carrier protein, ferrodoxin and fatty acid elongase genes obtained from this cDNA library enabled presumption of a possible biosynthesis pathway of ArA in M. incisa. Codon usage analysis of 142 NRSs with 17798 codons in the predicted coding regions showed that the average G+C content level of the total codons was 55.39%, and that of the third position in base trimers was 66.42%, indicating a strong bias toward cytosine and/or guanosine in this algal genome. Among all synonymous codons, NAG was most favored, while NUA was most avoided. Phylogenic trees inferred from ribulose-1,5-bisphosphate carboxylase/oxygenase small subunit genes and the extra partial sequences of 18S rRNA obtained from this library demonstrated that M. incisa belonged to Trebouxiophyceae and was significantly clustered with M. incisa SAG 2007, Lobosphaera tirolensis, M. bisecta, and Parietochloris incisa, but was clearly distant from P. pseudoalveolaris and P. alveolar. Transmission electron microscopy revealed pyrenoids traversed by many parallel thylakoids membranes, while starch grains were only clearly observed when cells were grown under nitrogen stress. Based on combined investigation of the phylogeny and morphological characteristics, it is proposed that M. incisa be kept in the genus Myrmecia in which there might be two parallel groups, one living freely and another symbiotic species.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Reisigl H. Zur systematik und ökologie alpiner Bodenalgen (in German). Österr Bot Z, 1964, 111: 402–499
Cărăuş I. Algae of Romania-a distributional checklist of actual algae. Doctor Dissertation. Bacău: Universitatea din Bacău, 2002
Watanabe S, Hirabayashi S, Boussiba S, et al. Parietochloris incisa comb. nov. (Trebouxiophyceae, Chlorophyta). Phycol Res, 1996, 44: 107–108
Karsten U, Friedl T, Schumann R, et al. Mycosporine-like amino acids and phylogenies in green algae: Prasiola and its relatives from the Trebouxiophyceae (Chlorophyta). J Phycol, 2005, 41: 557–566
Bigogno C, Khozin-Goldberg I, Boussiba S, et al. Lipid and fatty acid composition of the green oleaginous alga Parietochloris incisa, the richest plant source of arachidonic acid. Phytochemistry, 2002, 60: 497–503
Bigogno C, Khozin-Goldberg I, Cohen Z. Accumulation of arachidonic acid-rich triacylglycerols in the microalga Parietochloris incisa (Trebouxiophyceae, Chlorophyta). Phytochemistry, 2002, 60: 135–143
Khozin-Goldberg I, Bigogno C, Shrestha P, et al. Nitrogen starvation induces the accumulation of arachidonic acid in the freshwater green alga Parietochloris incisa (Trebouxiophyceae). J Phycol, 2002, 38: 991–994
Merzlyak M N, Chivkunova O B, Gorelova O A, et al. Effect of nitrogen starvation on optical properties, pigments, and arachidonic acid content of the unicellular green alga Parietochloris incisa (Trebouxiophyceae, Chlorophyta). J Phycol, 2007, 43: 833–843
Li C Y, Du D H, Yu S Y, et al. Characterization of a ω3 fatty acid desaturase gene from Myrmecia incisa and its relative transcription during the stress course of nitrogen starvation (in Chinese). J Fish China, 2010, 34: 1343–1353
Tong M, Yu S Y, Ouyang L L, et al. Comparison of increased arachidonic acid content in Myrmecia incisa cultured during the course of nitrogen or phosphorus starvation (in Chinese). J Fish China, 2011, 35: 763–773
Yu S Y, Liu S C, Li C Y, et al. Submesoscale characteristics and transcription of a fatty acid elongase gene from a freshwater green microalgae, Myrmecia incisa Reisigl. Chin J Oceanol Limnol, 2011, 29: 87–95
Zhang C W, Cohen Z, Khozin-Goldberg I, et al. Characterization of growth and arachidonic acid production of Parietochloris incisa comb. nov. (Trebouxiophyceae, Chlorophyta). J Appl Phycol, 2002, 14: 453–460
Liu J G, Cohen Z, Richmond A. Fatty acids profile in a high cell density culture of arachidonic acid-rich Parietochloris incisa comb. nov. (Trebouxiophyceae, Chlorophyta) exposed to high PFD. Chin J Oceanol Limnol, 2002, 20:149–156
Wang C, Sun Y, Xing R, et al. Effect of liquid circulation velocity and cell density on the growth of Parietochloris incisa in flat plate photobioreactors. Biotechnol Bioproc Eng, 2005, 10: 103–108
Bigogno C, Khozin-Goldberg I, Adlerstein D, et al. Biosynthesis of arachidonic acid in the oleaginous microalga Parietochloris incisa (Chlorophyceae): Radiolabeling studies. Lipids, 2002, 37: 209–216
Iskandarov U, Khozin-Goldberg I, Cohen Z. Identification and characterization of Δ12, Δ6, and Δ5 desaturases from the green microalga Parietochloris incisa. Lipids, 2010, 45: 519–530
Iskandarov U, Khozin-Goldberg I, Ofir R, et al. Cloning and characterization of the Δ6 ployunsaturated fatty acid elongase from the green microalga Parietochloris incisa. Lipids, 2009, 44: 545–554
Asamizu E, Nakamura Y, Sato S, et al. A large scale structural analysis of cDNAs in a unicellular green alga, Chlamydomonas reinhardtii. I. Generation of 3433 non-redundant expressed sequence tags. DNA Res, 1999, 6: 369–373
Asamizu E, Miura K, Kucho K, et al. Generation of expressed sequence tags from low-CO2 and high-CO2 adapted cells of Chlamydomonas reinhardtii. DNA Res, 2000, 7: 305–307
Shrager J, Hauser C, Chang C W, et al. Chlamydomonas reinhardtii genome project. A guide to the generation and use of the cDNA information. Plant Physiol, 2003, 131: 401–408
Asamizu E, Nakamura Y, Miura K, et al. Establishment of publicly available cDNA material and information resource of Chlamydomonas reinhardtii (Chlorophyta) to facilitate gene function analysis. Phycologia, 2004, 43: 722–726
Jain M, Shrager J, Harris E H, et al. EST assembly supported by a draft genome sequence: an analysis of the Chlamydomonas reinhardtii transcriptome. Nucl Acids Res, 2007, 35: 2074–2083
Becker B, Feja N, Melkonian M. Analysis of expressed sequence tags (ESTs) from the scaly green flagellate Scherffelia dubia Pascher emend. Melkonian et Preisig. Protist, 2001, 152: 139–147
Simon A, Glöckner G, Felder M, et al. EST analysis of the scaly green flagellate Mesostigma viride (Streptophyta): Implications for the evolution of green plants (Viridiplantae). BMC Plant Biol, 2006, 6: 2
Lanier W, Moustafa A, Bhattacharya D, et al. EST analysis of Ostreococcus lucimarinus, the most compact eukaryotic genome, shows an excess of introns in highly expressed genes. PLoS ONE, 2008, 3: e2171
Shi J, Pan K, Yu J, et al. Analysis of expressed sequence tags from the marine microalga Nannochloropsis oculata (Eustigmatophyceae). J Phycol, 2008, 44: 99–102
Alkayal F, Albion R L, Tillett R L, et al. Expressed sequence tag (EST) profiling in hyper saline shocked Dunaliella salina reveals high expression of protein synthetic apparatus components. Plant Sci, 2010, 179: 437–449
Stanier R Y, Kunisawa M M, Cohen-Bazir G. Purification and properties of unicellular blue-green algae (order Chlorococcales). Bacteriol Rev, 1971, 35: 171–201
Barnes W M. PCR amplification of up to 35-kb DNA with high fidelity and high yield from λ bacteriophage templates. Proc Natl Acad Sci USA, 1994, 91: 2216–2220
Wong T K M, Ho C L, Lee W W, et al. Analyses of expressed sequence tags from Sargassum binderi (Phaeophyta). J Phycol, 2007, 43: 528–534
Scala S, Carels N, Galciatore A, et al. Genome properties of the diatom Phaeodactylum tricornutum. Plant Physiol, 2002, 129: 993–1002
Peden J F. Analysis of codon usage. Doctor dissertation. Sutton Bonington: University of Nottingham, 1999
Tamura K, Dudley J, Nei M, et al. MEGA4: Molecular evolutionary genetics analysis (MEGA) software version 4.0. Mol Biol Evol, 2007, 24: 1596–1599
Thompson J D, Gibson T J, Plewniak F, et al. The ClustalX windows interface flexible strategies for multiple sequence alignment aided by quality analysis tools. Nucleic Acids Res, 1997, 25: 4876–4882
Felsenstein J. Confidence limits on phylogenies: An approach using the bootstrap. Evolution, 1985, 39: 66–70
Campbell C R, Plank C O. Preparation of plant tissue for laboratory analysis. In: Kalra Y P, ed. Handbook of Reference Methods for Plant Analysis. Boca Raton: CRC Press. 1998. 37–49
Patanjali S R, Parimoo S, Weissman S M. Construction of a uniform-abundance (normalized) cDNA library (reassociation/hybridization/hydroxyapatite). Proc Natl Acad Sci USA, 1991, 88: 1943–1947
Lambert K N, Williamson V M. cDNA library construction from small amounts of RNA using paramagnetic beads and PCR. Nucleic Acids Res, 1993, 21: 775–776
Hackett J, Scheetz T, Yoon H, et al. Insights into a dinoflagellate genome through expressed sequence tag analysis. BMC Genomics, 2005, 6: 80
Gurr M I, Harwood J L, Frayn K N. Lipid Biochemistry: An Introduction, 5th ed. Osney Mead, Oxford: Blackwell Science Ltd, a Blackwell Publishing Company, 2002
Los D A, Murata N. Structure and expression of fatty acid desaturases. Biochim Biophys Acta, 1998, 1394: 3–15
Leonard A E, Pereira S L, Sprecher H, et al. Elongation of long-chain fatty acids. Prog Lipid Res, 2004, 43: 36–54
Yu S Y, Li H, Tong M, et al. Identification of a Δ6 fatty acid elongase gene for arachidonic acid biosynthesis localized to the endoplasmic reticulum in the green microalga Myrmecia incisa Reisigl. Gene, 2012, 493: 219–227
Shi X F, Huang J F, Liang C R, et al. Is there a close relationship between synonymous codon bias and codon-anticodon binding strength in human genes? Chin Sci Bull, 2001, 46: 1015–1019
Grantham R, Gautier C, Gouy M, et al. Codon catalog usage is a genome strategy modulated for gene expressivity. Nucleic Acids Res, 1981, 9: r43–r74
Shpaer E G. Constraints on codon context in Escherichia coli genes their possible role in modulating the efficiency of translation. J Mol Biol, 1986, 188: 555–564
Ohno S. Universal rule for coding sequence construction: TA/CG deficiency-TG/CT excess. Proc Natl Acad Sci USA, 1988, 85: 9630–9634
Andreyeva V M. On the taxonomy of the genus Myrmecia Printz (Chlorococcales, Chlorococcaceae) (in Russian). Bot Mag, 1978, 63: 442–460
Friedl T. Inferring taxonomic positions and testing genus level assignments in coccoid green lichen algae: A phylogenetic analysis of 18S ribosomal RNA sequences from Dictyochloropsis reticulata and from members of the genus Myrmecia (Chlorophyta, Trebouxiophyceae cl. nov.). J Phycol, 1995, 31: 632–639
Pröschold T, Leliaert F. Systematics of the green algae: Conflict of classic and modern approaches. In: Brodie J, Lewis J, eds. Unravelling the Algae: the Past, Present, and Future of Algal Systematics. Boca Raton: CRC Press, 2007. 123–153
Friedl T, Zeltner C. Assessing the relationships of some coccoid green lichen algae and the Microthamniales (Chlorophyta) with 18S ribosomal RNA gene sequence comparisons. J Phycol, 1994, 30: 500–506
Metting B. The systematics and ecology of soil algae. Bot Rev, 1981, 47: 195–312
Griffiths D J. The pyrenoid. Bot Rev, 1970, 36: 29–58
Tschermak-Woess E, Plessl A. Über zweierlei typen der sukzedanen teilung und ein auffallendes teilungsverhalten des chromatophors bei einer neuen Protococale, Myrmecia pyriformis (in German). Österr Bot Z, 1948, 95: 194–207
Watanabe S, Floyd G L. Comparative ultrastructure of the zoospores of nine species of Neochloris (Chlorophyta). Plant Syst Evol, 1989, 168: 195–219
Dayrat B. Towards integrative taxonomy. Biol J Linn Soc, 2005, 85: 407–415
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Electronic supplementary material
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Ouyang, L., Du, D., Yu, S. et al. Expressed sequence tags analysis revealing the taxonomic position and fatty acid biosynthesis in an oleaginous green microalga, Myrmecia incisa Reisigl (Trebouxiophyceae, Chlorophyta). Chin. Sci. Bull. 57, 3342–3352 (2012). https://doi.org/10.1007/s11434-012-5159-2
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5159-2