Abstract
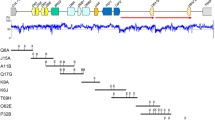
Artificial breeding is an important project to protect, recover and reintroduce endangered species. Knowledge of the population’s genetic diversity at functional loci is important for the establishment of effective captive breeding programs. The major histocompatibility complex (MHC) genes are ideal candidate genetic markers to inform planned breeding, due to their high levels of polymorphism and importance in the main immune coding region of the vertebrate genome. In this study, we constructed BAC-based contigs and isolated six functional MHC class I genes from the giant panda (Ailuropoda melanoleuca), which we designated Aime-C, Aime-F, Aime-I, Aime-K, Aime-L and Aime-1906. Analyses of the tissue expression patterns and full-length cDNA sequences of these class I genes revealed that Aime-C, -F, -I and -L could be considered classical class I loci, due to their extensive expression patterns and normal exonic structures. In contrast, Aime-K and -1906 appeared to be nonclassical genes based on their tissue-specific expression patterns and the presence of an abnormal exon 7 in both genes. We established techniques for genotyping exons 2 and 3 of the classical loci using locus-specific single strand conformation polymorphism (SSCP) and sequence analysis. In the Chengdu captive population, we identified one monomorphic locus (Aime-F) and three polymorphic loci with different numbers of alleles (4/4/4 exon 2 alleles at Aime-C/I/L and 6/5/5 exon 3 alleles at Aime-C/I/L). The distributions of the Aime-C, -I and -L alleles among members of different families were in good agreement with the known pedigree relationships, suggesting that the genotyping results are reliable. Therefore, the MHC-I genotyping techniques established in this study may provide a powerful tool for the future design of scientific breeding or release/reintroduction programs.
Article PDF
Similar content being viewed by others
Avoid common mistakes on your manuscript.
References
Ebenhard T. Conservation breeding as a tool for saving animal species from extinction. Trends Ecol Evol, 1995, 10:438–443
Seddon P J, Armstrong D P, Maloney R F. Developing the science of reintroduction biology. Conserv Biol, 2007, 21:303–312
Frankham R. Genetic adaptation to captivity in species conservation programs. Mol Ecol, 2008, 17:325–333
Robert A. Captive breeding genetics and reintroduction success. Biol Conserv, 2009, 142:2915–2922
Chong A Y Y. Genetic variation in the MHC of the Collared peccary: A potential model for the effects of captive breeding on the MHC. Univ Sydney Undergr Res J, 2009, 1:98–116
Sommer S. The importance of immune gene variability (MHC) in evolutionary ecology and conservation. Front Zool, 2005, 2:1–18
Frankham R, Ballou J D, Briscoe D A. Introduction to Conservation Genetics. Cambridge: Cambridge University Press, 2002
Klein J. The Natural History of the Major Histocompatibility Complex. New York: Wiley & Sons, 1986
Xu T J, Sun Y N, Chen S L. Allelic variation, balancing selection and positive selected sites detected from MHC class I a gene of olive flounder. Genetica, 2010, 138:1251–1259
Bos D H, Waldman B. Evolution by recombination and transspecies polymorphism in the MHC class I gene of Xenopus laevis. Mol Biol Evol, 2006, 23:137–143
Glaberman S, Du Pasquier L, Caccone A, et al. Characterization of a nonclassical class I MHC gene in a reptile, the galápagos marine iguana (Amblyrhynchus cristatus). PLoS One, 2008, 3:1–11
Birch J, Codner G, Guzman E, et al. Genomic location and characterisation of nonclassical MHC class I genes in cattle. Immunogenetics, 2008, 60:267–273
Piertney S, Oliver M. The evolutionary ecology of the major histocompatibility complex. Heredity, 2006, 96:7–21
Kassiotis G, Garcia S, Simpson E, et al. Impairment of immunological memory in the absence of MHC despite survival of memory T cells. Nat Immunol, 2002, 3:244–250
Bernatchez L, Landry C. MHC studies in nonmodel vertebrates: What have we learned about natural selection in 15 years? J Evol Biol, 2003, 16: 363–377
Radwan J, Kawalko A, Wojcik J M, et al. MHC-DRB3 variation in a free-living population of the European bison, Bison bonasus. Mol Ecol, 2006, 16:531–540
State Forestry Administration of China. The Third National Survey Report on Giant Panda in China (in Chinese). Beijing: Science Press, 2006
Xie Z, Gipps J. The 2012 International Studbook for Giant Panda (Ailuropoda melanoleuca). Beijing: Chinese Association of Zoological Garden, 2012
Wan Q H, Zeng C J, Ni X W, et al. Giant panda genomic data provide insight into the birth-and-death process of mammalian major histocompatibility complex class II genes. PLoS One, 2009, 4:e4147
Wan Q H, Zhu L, Wu H, et al. Major histocompatibility complex class II variation in the giant panda (Ailuropoda melanoleuca). Mol Ecol, 2006, 15:2441–2450
Chen Y Y, Zhang Y Y, Zhang H M, et al. Natural selection coupled with intragenic recombination shapes diversity patterns in the major histocompatibility complex class II genes of the giant panda. J Exp Zool B (Mol Dev Evol), 2010, 314B:208–223
Wan Q H, Zhang P, Ni X W, et al. A novel HURRAH protocol reveals high numbers of monomorphic MHC class II loci and two asymmetric multi-locus haplotypes in the Père David’s Deer. PLoS One, 2011, 6:e14518
Pan H J, Wan Q H, Fang S G. Molecular characterization of major histocompatibility complex class I genes from the giant panda (Ailuropoda melanoleuca). Immunogenetics, 2008, 60:185–193
Shen F J, Zhang Z H, He W, et al. Microsatellite variability reveals the necessity for genetic input from wild giant pandas (Ailuropoda melanoleuca) into the captive population. Mol Ecol, 2009, 18:1061–1070
Sambrook J, Russell D W. iMolecular Cloning: A Laboratory Manual, 3rd ed. Cold Spring Harbor. New York: Cold Spring Harbor Laboratory Press, 2001
Zeng C J, Pan H J, Gong S B, et al. Giant panda BAC library construction and assembly of a 650-kb contig spanning major histocompatibility complex class II region. BMC Genomics, 2007, 8:1–10
Sunnucks P, Wilson A C C, Beheregaray L B, et al. SSCP is not so difficult: The application and utility of single-stranded conformation polymorphism in evolutionary biology and molecular ecology. Mol Ecol, 2000, 9:1699–1710
Bjorkman P J, Saper M A, Samraoui B, et al. The foreigh antigen bing site and T cell recognition regions of class I histocompatibility antigens. Nat Immunol, 1987, 329:512–518
Rousset F. GENEPOP’007: A complete re-implementation of the GENEPOP software for windows and linux. Mol Ecol Resour, 2008, 8:103–106
Belkhir K, Borsa P, Chikhi L, et al. Genetix version 4.05, Logiciel sous windows TM pour la Génétique des populations, Montpellier: Université de Montpellier II, 2004
Kelley J, Walter L, Trowsdale J. Comparative genomics of major histocompatibility complexes. Immunogenetics, 2005, 56:683–695
Kurtz J, Kalbe M, Aeschlimann P B, et al. Major histocompatibility complex diversity influences parasite resistance and innate immunity in sticklebacks. Proc R Soc B, 2004, 271:197–204
Westerdahl H, Waldenstrom J, Hansson B, et al. Associations between malaria and MHC genes in a migratory songbird. Proc R Soc B, 2005, 272:1511–1518
Spurgin L G, Richardson D S. How pathogens drive genetic diversity: MHC, mechanisms and misunderstandings. Proc R Soc B, 2010, 277:979–988
Author information
Authors and Affiliations
Corresponding author
Additional information
This article is published with open access at Springerlink.com
Rights and permissions
This article is published under an open access license. Please check the 'Copyright Information' section either on this page or in the PDF for details of this license and what re-use is permitted. If your intended use exceeds what is permitted by the license or if you are unable to locate the licence and re-use information, please contact the Rights and Permissions team.
About this article
Cite this article
Zhu, Y., Sun, D., Ge, Y. et al. Isolation and characterization of class I MHC genes in the giant panda (Ailuropoda melanoleuca). Chin. Sci. Bull. 58, 2140–2147 (2013). https://doi.org/10.1007/s11434-012-5582-4
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11434-012-5582-4