Abstract
Here, we present the results of studies of Japanese Hebeloma collections. The four species described by Imai as Hebeloma (H. fimicola, H. helvolescens, H. humosum, and H. tomoeae) are not from the genus Hebeloma, but are members of Agrocybe, Homophron, or Pholiota. Recombinations are made. Hebeloma crustuliniforme f. microspermum, described by Hongo, is a synonym of H. nanum. Three species of Hebeloma are described as new to science, all currently known only from Japan. Two of these species, H. asperosporum and H. cinnamomeum, are members of H. sect. Denudata while the third species H. citrisporum belongs to H. sect. Velutipes. Japanese records of H. cavipes, H. eburneum, H. hygrophilum, H. subtortum, and H. velutipes are validated. In total, fifteen species of Hebeloma are confirmed from Japan; this is compared with previous checklists.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
A number of species of the genus Hebeloma appear to be indigenous, or possibly even endemic, to Japan. The latter include species such as Hebeloma luchuense and H. radicosoides (H. sect. Scabrispora), H. sagarae (H. sect. Myxocybe), for which neither observations nor sequence data were found from other countries (status 16 June 2021). Hebeloma lactariolens and H. vinosophyllum (H. sect. Porphyrospora) have been described from Japan, but have also been reported from other countries (e.g., Ho et al. 2014; Cho et al. 2016; Wu et al. 2019; Eberhardt et al. 2021a).
Katumoto (2010) produced a checklist where he listed 15 distinct species of Hebeloma recorded for Japan. The 15 species listed are H. crustuliniforme, H. crustuliniforme f. microspermum, H. fimicola, H. helvolescens, H. humosum, H. longicaudum, H. luchuense, H. mesophaeum, H. radicosoides, H. radicosum, H. sacchariolens, H. sinuosum, H. spoliatum, H. tomoeae, and H. vinosophyllum. Katumoto did not include in his list Hebeloma lactariolens; this was only recombined into Hebeloma in 2013 (Rees et al. 2013). Accessing the Global Biodiversity Information Facility (www.gbif.org accessed 10 June 2021) gives 202 occurrences of Hebeloma in Japan. Of these 202 occurrences, 166 reference a preserved specimen and 131 give a species name. Additional to the names given by Katumoto and Hebeloma lactariolens are also Hebeloma leucosarx (one record, no preserved specimen), Hebeloma microsporum (one record, but not this taxon, see below under H. cinnamomeum), Hebeloma pusillum (one record, no preserved specimen), and H. sordescens (one specimen recorded as H. testaceum from 1935). iNaturalist (www.iNaturalist.org accessed 10 June 2021) gives a single record of Hebeloma for Japan, H. crustuliniforme.
One reason for this rather small number of taxa and records is that while it is normally straightforward for a competent field mycologist to identify a mushroom as being a Hebeloma, determining the species has in the past been notoriously difficult. This has been partly because of the macroscopic similarity of species, but also because of the confused interpretation of species concepts by many authors (e.g., Moser 1983; Smith et al. 1983; Arora 1986; Hansen and Knudsen 1992; Breitenbach and Kränzlin 2000; Bon 2002; Vesterholt 2005). As a result of this confusion, many mycologists ignore Hebeloma in the field, which results in few records; within herbaria, many collections determined as Hebeloma remain undetermined at species level. For example, the TNS herbarium currently holds a total of 223 collections of Hebeloma, out of which a total of 210 were collected in Japan, but more than 70 collections remain unidentified to species.
In 2016, Beker et al. produced a monograph on Hebeloma in Europe, which was intended to provide a foundation for species names and concepts. Since that publication, several papers have appeared describing new species in Southeast Asia and Japan, as well as clarifying the infrageneric classification of species from these areas (for example: Eberhardt et al. 2020a, 2020b, 2021a).
We examined 106 collections of Hebeloma, including the holotypes of all species described, from Japan: H. fimicola, H. helvolescens, H. humosum, H. tomoeae (Imai 1938); H. vinosophyllum (Hongo 1965); H. crustuliniforme f. microspermum (Hongo 1966); H. lactariolens (Clémençon and Hongo 1994); H. luchuense (Fukiharu and Hongo 1995); H. radicosoides (Sagara et al. 2000); and H. sagarae (Eberhardt et al. 2020a).
With regard to the types studied, H. luchuense and H. radicosoides were confirmed within H. sect. Scabrispora, H. sagarae was confirmed within H. sect. Myxocybe, and both H. lactariolens and H. vinosophyllum were confirmed within H. sect. Porphyrospora (Eberhardt et al. 2020b). The Japanese collections of H. spoliatum, referred to by Eberhardt et al. (2020b), were similar to or conspecific with H. danicum, and it was noted that further research was required to decide on conspecificity. This is addressed here.
Within this paper, we also present and discuss our findings with regard to additional type studies of species of Hebeloma, described from Japan, and provide a list of Hebeloma species we have found during analysis of collections sent to us by citizen scientists published on Mushroom Observer (https://mushroomobserver.org/) and herbarium collections, particularly those from the National Museum of Nature and Science (TNS). Included are three species new to science: Hebeloma asperosporum is only known from the remote island of Amami Ōshima, while H. cinnamomeum and H. citrisporum are described from the island of Honshu.
Materials and methods
All material studied was dried material from, primarily, the National Museum of Nature and Science (TNS) and the Hokkaido University Museum (SAPA), but also included a few collections sent to us directly by U. Kawasaki. This was compared with material, sequenced and discussed in Beker et al. (2016), Cripps et al. (2019), and Eberhardt et al. (2020a, 2020b, 2021a).
Sequences were obtained from the dried basidiomes by direct sequencing. Internal transcribed spacer sequences were generated following methods detailed in Eberhardt (2012), and Cripps et al. (2019); RPB2 and TEF1a sequences following Eberhardt et al. (2021b); and sequences of a variable region (V6) of the mitochondrial SSU followed Gonzalez and Labarère (1998). Sequencing was carried out at LGC Genomics (Berlin). Sequences were edited using Sequencher vs. 4.9 (Gene Codes Corp., Ann Arbor, Michigan). Newly generated sequences were accessioned to GenBank (MT157290, MZ725546–MZ725550, MZ724681, MZ782100–MZ782148, MZ782867–MZ782889, and MZ782921–MZ782985). Tables 1 and 2 summarize all sequences used in the analyses.
Alignments were viewed and reformatted using aliview 1.27 (Larsson 2014). Sequence alignments were done online in MAFFT using the E-INS-i option (Katoh et al. 2005, 2019) or locally with the “Mafft-globalpair” setting of Mafft 7.471 (Katoh and Standley 2013). Maximum likelihood (ML) phylogenetic analyses were run in IQ-TREE (Nguyen et al. 2015) either locally (version 2.1.3) or online (Trifinopoulos et al. 2016). Branch support was obtained through 1000 replicates of ultrafast bootstrap (Minh et al. 2013; Hoang et al. 2018) and Shimodaira–Hasegawa (SH)-like approximate likelihood ratio tests (Guindon et al. 2010). Support values are given as SH-like approximate likelihood ratio test support [%] / ultrafast bootstrap support [%], for SH-like approximate likelihood ratio test support ≥ 80% and ultrafast bootstrap support ≥ 95%. These values were selected following the recommendations of Minh et al. (2021), who also recommend using SH-like approximate likelihood ratio test support alongside with (ultrafast) bootstrap. Model selection (Kalyaanamoorthy et al. 2017) was done using the BIC criterion, including FreeRate models and merging partitions if possible (protein coding loci were originally partitioned according to position (coding) and non-coding). The study was submitted to TREEBASE (acc. no. TB2:S28657). Trees were visualized using figtree 1.4.4 (Rambaut 2006–2018). Results of Beker et al. (2016), Grilli et al. (2016), Frings et al. (2020), and Tian and Matheny (2021) guided the selection of the roots for the topologies. The compatibility of different loci was assessed following the principle of Kauff and Lutzoni (2002), assuming a conflict to be significant if two different relationships for the same set of taxa, one being monophyletic and the other non-monophyletic, are supported by ≥ 85% SH-like approximate likelihood ratio test support or ≥ 95% ultrafast bootstrap support. Alignments of different loci were concatenated and analyzed, indicating branches with conflicting results from single locus analyses by dashed lines. Country codes in Figs. 2, 3, 4, 5 and 6 follow ISO 3166–1 alpha-2 (https://www.iso.org/iso-3166-country-codes.html, accessed 22 Jun 2021).
Maximum likelihood result (ITS) for placing Agrocybe humosa and A. imaii in a taxonomic context. Support values are from 1000 replicates of SH-like approximate likelihood ratio tests and ultrafast bootstrap. Support values ≥ 85% (SH-like approximate likelihood ratio tests) or 95% (ultrafast bootstrap) are shown. The last two letters indicate the country of origin
Maximum Likelihood result (ITS) for placing Homophron helvolescens in a taxonomic context. Support values are from 1000 replicates of SH-like approximate likelihood ratio tests and ultrafast bootstrap. Support values ≥ 85% (SH-like approximate likelihood ratio tests) or 95% (ultrafast bootstrap) are shown. Branches with at least one of the support values above the thresholds are thick lined. * Type collection included in clade. The last two letters indicate the country of origin
Maximum likelihood result (ITS) for placing Pholiota tomoeae in a taxonomic context. Support values are from 1000 replicates of SH-like approximate likelihood ratio tests and ultrafast bootstrap. Support values ≥ 85% (SH-like approximate likelihood ratio tests) or 95% (ultrafast bootstrap) are shown. Branches with at least one of the support values above the thresholds are thick lined. * Type collection included in clade. The last two letters indicate the country of origin
Maximum Likelihood result of concatenated ITS, RPB2, TEF1a, and mitSSU V6 sequences of members of Hebeloma sect. Denudata, subsects. Clepsydroida and Crustuliniformia, rooted with members of H. subsect. Echinospora. Relationships that include conflicting data are indicated by dashed lines. Support values ≥ 85% (SH-like approximate likelihood ratio tests) or 95% (ultrafast bootstrap) are shown. In some places support values 85–99% (SH-like approximate likelihood ratio tests) or 95–99% (ultrafast bootstrap) are represented by “*” and values of 100% by “#”. Branches with at least one of the support values above its threshold are thick lined. The last two letters indicate the country of origin
Details of morphological analyses were provided in Beker et al. (2016). The amount of macroscopic detail available to us varied hugely from collection to collection as it was dependent on the detail provided by the collector. Where one of the authors was the collector, each specimen was photographed and observed both in the field when characters were still fresh, and later in the laboratory. Fresh basidiomes of each specimen were dried using a food dehydrator (Snackmaster Express FD-60; Nesco/American Harvest, Milwaukee, WI, USA).
All microscopic analysis was carried out on dried material, using a Leica DMRZA2 microscope with a Leica DFC495 camera connected to a computer running Leica Application Suite (LAS) V4 software. A number of photographs were taken of the spores at × 500 and × 1600, which were then measured using the LAS software. Photographs were also taken of the cheilocystidia on the lamella edge at × 500 and of individual cystidia and basidia at × 1000. The material was then examined in 5% KOH. Again, photographs were taken of the spores at × 500 and × 1600 and of the cheilocystidia (and pleurocystidia if any were present) and basidia at × 500 and × 1000.
For each Hebeloma collection, wherever possible, at least 50 spores were measured in Melzer’s reagent, excluding the apiculus. The maximum length and width of each spore was measured, and its Q value (ratio of length to width) calculated. Average length, width, and Q value were calculated and recorded alongside the median, standard deviation, and 5% and 95% percentiles. The degree of dextrinoidity, ornamentation, and the degree of loosening of the perispore was observed and classified. For the assessment of the degrees of ornamentation (O0, O1, O2, O3, O4), of the loosening perispore (P0, P1, P2, P3), and for the dextrinoidity (D0, D1, D2, D3, D4), we used Beker et al. (2016) and Vesterholt (2005).
The average width of the widest part of the cheilocystidium in the vicinity of the apex appears to be an important character in the separation of species within Hebeloma (Vesterholt 2005). It is also important, when determining this average width near the apex, not to be selective with regard to the cystidia chosen for measurement. To determine the average width at the apex, about 100 cheilocystidia were measured on the lamella edge. For other measurements, some 20 cheilocystidia, separated from the lamella edge, were measured from each collection. Because of the complex shapes of the cheilocystidia four measurements were made: length, width at apex (A), width at narrowest point in central region (M), and maximum width in lower half (B), see Fig. 1. The measurements were given in this order, and an average value was calculated for each of these measurements. For each cheilocystidium, the ratios A/M, A/B, and B/M were calculated and averaged across all cheilocystidia measured. Measurements were made in 5% KOH and Melzer’s reagent.
For all other details with regard to our methodology, see Beker et al. (2016). Each collection studied has a database record number associated with that collection; we give these numbers as we intend to make the database publicly available.
Results
None of the four species described by Imai (1938), Hebeloma fimicola, H. helvolescens, H. humosum, and H. tomoeae belong to Hebeloma as recognized today. They belong to the genera Agrocybe, Homophron, and Pholiota and are recombined in the Taxonomy section with a short summary of their main characters. Hebeloma crustuliniforme f. microspermum is synonymized with H. nanum based on morphology (no sequence data obtained). Three new species of Hebeloma were discovered in the course of this study, two of these species, H. asperosporum and H. cinnamomeum, are members of H. sect. Denudata while the third species H. citrisporum belongs to H. sect. Velutipes.
The alignment for Agrocybe includes 24 ITS sequences and 717 positions. The result of the analysis is shown in Fig. 2. The topology is rooted with Agrocybe firma at a branch receiving 100/100 support (support not shown because branch used for rooting). The type species of Agrocybe is A. praecox, which is represented in the alignment by five sequences of different authors. The lectotype ITS of Agrocybe humosa (Hebeloma humosum) is placed in an unsupported clade together with sequences identified as A. smithii; the holotype ITS of A. imaii (H. fimicola) is placed in a clade with sequences identified as A. pediades.
The alignment for Homophron includes 35 ITS sequences and 708 positions. Homophron spadiceum is the type species of the genus. We have made an effort to sample all variation from UNITE SH1186588.08FU (which includes the epitype of Ho. spadiceum), because it was not possible to see the species limits of Ho. spadiceum. The result of the analysis is shown in Fig. 3. The topology is rooted with Lacrymaria spp. The clade indicated as “Homophron spadiceum s.l.” consists of a number of subclades, some appear to be geographically restricted, among which Ho. helvolescens is in a subclade with sequences from Japan and India.
The alignment for Pholiota (P. sect. Flammuloides in Tian and Matheny 2021) includes 26 ITS sequences and 771 positions. The result of the analysis is shown in Fig. 4. The sequences of the two syntypes (one designated as lectotype below) of P. tomoeae are within the P. brunnescens clade, which includes sequences from America, Asia, and Europe.
Hebeloma asperosporum is morphologically and molecularly distinct from all other described Hebeloma species. Morphologically, it is a member of H. subsect. Clepsydroida, distinguished by its small, strongly ornamented spores, on average less than 10.5 µm long, indextrinoid and with consistently loosening perispore.
Likewise, Hebeloma cinnamomeum is morphologically and molecularly distinct from all other described Hebeloma species. Morphologically, it is a member of H. subsect. Clepsydroida, distinguished by its cinnamon-colored pileus and the spores with average Q less than 1.85 and the cheilocystidia with average width near the apex of less than 6.5 µm.
Four loci (ITS, alignment 708 positions; mitSSU V6, alignment 476 positions; RPB2, alignment 779 positions; and TEF1a, alignment 717 positions) have been employed to infer the phylogenetic relationships of taxa in Hebeloma sect. Denudata (Fig. 5). The topology is rooted with members of H. subsect. Echinospora. All loci analyzed individually resolved the two available sequences of H. asperosporum as monophyletic with high support (min. 94% SH-like approximate likelihood ratio test and 97% ultrafast bootstrap support, in most loci higher), but it was on a long branch and the position of the species within the section was unresolved. The sequences obtained for H. cinnamomeum form an independent lineage only in the ITS (support 92%/99%) and V6 (84%/–). An ITS tree with all H. cinnamomeum sequences (not shown) is available via TreeBase. In the TEF1a result, H. cinnamomeum is paraphyletic in relation to H. ammophilum, H. cavipes, and H. sordidulum and in the RPB2 result also in relation to H. vaccinum.
Only ITS and V6 resolve H. asperosporum and H. cinnamomeum sequences as species clades in relation to other members of H. sect. Denudata, with SH-like approximate likelihood ratio test support above 90% and ultrafast bootstrap support above 95% in ITS, and with approximately 100% in both tests for H. asperosporum and with 87% SH-like approximate likelihood ratio test support, but without relevant ultrafast bootstrap support, for H. cinnamomeum in V6.
The topologies obtained for the different loci (see Treebase submission) were in conflict. This was largely due to H. asperosporum and H. matritense being placed with relevant support by one or both support criteria as sister to different species clades in the results for different loci. The result of the analysis of the concatenated alignment is shown in Fig. 5. The topology is rooted with H. echinosporum and H. populinum. Here, H. asperosporum receives 100% support by both criteria and H. cinnamomeum 94%/97% support. The inclusion of members of H. sect. Clepsydroida in two reciprocally monophyletic groups is weakly (86% in the lesser supported clade) supported by SH-like approximate likelihood ratio tests. However, conflicts between the evolutionary hypothesis of different loci concern this part of the tree.
Hebeloma citrisporum is morphologically and molecularly distinct from all other described Hebeloma species. The predominantly gently clavate cheilocystidia suggest that this species belongs in H. sect. Velutipes, which is supported by molecular results (Fig. 6).
Maximum Likelihood result of concatenated ITS, RPB2, and TEF1a sequences of members of Hebeloma sect. Velutipes, rooted with H. sacchariolens (H. sect. Sacchariolentia). Relationships that include conflicting data are indicated by dashed lines. Support values ≥ 80% (SH-like approximate likelihood ratio tests) or 95% (ultrafast bootstrap) are shown. In some places support values 85–99% ((SH-like approximate likelihood ratio tests) or 95–99% (ultrafast bootstrap) are represented by “*” and values of 100% by “#”. Branches with at least one of the support values above its threshold are thick lined. The last two letters indicate the country of origin
Three loci (ITS, alignment 715 positions; RPB2, alignment 779 positions; and TEF1a, alignment 712 positions) have been employed to infer the phylogenetic relationships of taxa in Hebeloma sect. Velutipes. Sequences of H. citrisporum are monophyletic in all single locus trees and the branch supported by 90–97%/95–100%. The only conflict among the single locus results (see Treebase submission) is between TEF1a and the RPB2 in the placement of H. erebium, paraphyletic as sister of H. celatum (TEF1a) or as sister to H. celatum, H. citrisporum, and H. quercetorum (RPB2). The ITS does not resolve H. celatum, H. erebium, and H. quercetorum in separate clusters. The result of the analysis of the concatenated data is shown in Fig. 6. Here, the lineage of H. citrisporum receives 100%/100% support and is supported as sister taxon of H. quercetorum as in the single locus results of RPB2 and TEF1a.
In addition to the species mentioned above (H. nanum, H. asperosporum, H. cinnamomeum, H. citrisporum), and H. danicum (often recorded as H. spoliatum), H. luchuense and H. radicosoides (H. sect. Scabrispora), H. sagarae (H. sect. Myxocybe) and H. lactariolens and H. vinosophyllum (H. sect. Porphyrospora), the following taxa were identified from the Japanese material examined: H. cavipes (e.g., TNS-F-44988; GenBank ITS = MZ782120), H. eburneum (TNS-F-38029; no DNA sequence data obtained), H. hygrophilum (TNS-F-61595; GenBank ITS = MZ782130), H. subtortum (TNS-F-61666; GenBank ITS = MZ782131), and H. velutipes (TNS-F-75075; GenBank ITS = MZ782137). This gives a total of 15 species that we can confirm are present in Japan.
Taxonomy
Agrocybe humosa (S. Imai) Beker & U. Eberh., comb. nov.
MB840285
Basionym: Hebeloma humosum S. Imai, Journal of the Faculty of Agriculture of the Hokkaido Imperial University 43(2): 230 (1938).
Typification: Japan, Hokkaido, Sapporo, Ishikari, on badly decayed wood or on humus in gardens, 25 Jun 1935, S. Imai, Syntypes (SAPA 10000033 and SAPA 10000034). Lectotype designated here (MBT10001751, SAPA 10000033), database record HJB1000376. GenBank ITS = MZ725546.
Description of the lectotype: Spores elliptical, smooth, pale brown, thick-walled, with a conspicuous germ pore, on av. 11.1 × 7.7 µm, with av. spore Q 1.44. The cheilocystidia are ventricose; the pleurocystidia are similar.
Japanese name: Ki-wakafusa-take (Imai 1938).
Remarks: There are two collections (SAPA 10000033 and SAPA 10000034) at SAPA, marked as syntypes. The material selected as lectotype fits the original description by Imai (1938) who gives the spore size as 10–1 l.5 × 7–8 µm. This matches most closely with our measurements for SAPA 10000033 (10.1–13.3 × 6.9–9.5 µm, av. 11.1 × 7.7 µm, av. Q 1.44) for which we also have an ITS sequence. The other collection, SAPA 10000034, also appears to be an Agrocybe, with elliptical, smooth, pale brown, thick-walled spores, with a conspicuous germ pore, on av. 13.3 × 8.9 µm (11.8–14.7 × 8.0–10.1 µm), with av. spore Q 1.50. The cheilocystidia are ventricose; the pleurocystidia are similar.
The spores of A. smithii (Watling and Bigelow 1983) and of the type of A. flexuosipes (Eberhardt et al., in press) are around the same size (11–13.5 × 6.5–8 µm respectively 12.1 × 7.7 µm) as the spores of A. humosa. Whereas the spores of the former two species are described as (indistinctly) ornamented, the spores observed in the lectotype of A. humosa are smooth. The spores of A. putaminum are described as 10–13 × 5–7 µm, thus somewhat narrower than in A. humosa, but also smooth (Maire 1913).
Both morphology (without annulus, yellowish tones in pileus, spore size, and prominent germ pore) and sequence data (Fig. 2) suggest that A. humosa is closely related with A. smithii, A. putaminum, or other related species (Nauta 2005). Another closely related species is A. flexuosipes (Eberhardt et al., in press). However, none of these species matches the type of A. humosa sufficiently closely for synonymization.
Agrocybe imaii Beker & U. Eberh., nom. nov.
MB840286
Basionym: Hebeloma fimicola [as “fimicolum”] S. Imai, Journal of the Faculty of Agriculture of the Hokkaido Imperial University 43(2): 227 (1938); non Agrocybe fimicola (Speg.) Singer, Lilloa 23: 209 (1952).
Etymology: In honor of S. Imai, the describer of this species.
Typification: Japan, Hokkaido, Sapporo, Ishikari, on horse dung under trees, Jun 1935, S. Imai, Holotype (SAPA 10000036), database record HJB1000377. GenBank ITS = MZ725547.
Description of the holotype: The spores are elliptical, smooth, pale brown, thick-walled, with a conspicuous germ pore, on av. 12.9 × 8.9 µm, with av. spore Q 1.44. The cheilocystidia are ventricose; pleurocystidia only near lamella edge. The pileipellis has a thin gelatinous layer, with small pieces of hypha embedded within it.
Japanese name: Baba-wakafusa-take (Imai 1938).
Remarks: Hebeloma fimicola is an Agrocybe. Our examination of the type material was in close agreement with the description of Imai (1938). It is possible, even likely, that A. imaii is a later synonym of A. pediades s.l., and, given its habitat on dung, may be A. pediades var. fimicola (Speg.) Nauta (Agrocybe fimicola (Speg) Sing.) (Nauta 2005), but we rather leave it to experts on the genus based on multilocus sequence data to come to a conclusion. Based on ITS data, Agrocybe pediades occurs in many countries including China, India, Iraq, Pakistan, and Sri Lanka (see Fig. 2 and UNITE SH2290413.08FU).
Hebeloma asperosporum Beker & U. Eberh., sp. nov., Figs. 7 and 8, Mycobank MB840287
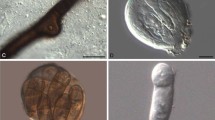
Hebeloma asperosporum holotype (TNS-F-75705) a spores and b spore ornamentation in Melzer’s reagent × 1600, c basidium in Melzer’s reagent × 1000, d spores and e spore ornamentation in KOH × 1600, f cheilocystidia in KOH × 1000, g cheilocystidia in KOH × 1000, h cheilocystidia in Melzer’s reagent × 1000, i caulocystidia in KOH × 500, j cutis in KOH × 125, k subcutis in KOH × 500, and l caulocystidia in KOH × 1000. Scale bars 10 µm, in j 100 µm. Photos H.J. Beker
Diagnosis: The small, strongly ornamented spores, on average less than 10.5 µm long, indextrinoid and with consistently loosening perispore distinguishes this species from other known members of H. section Denudata.
Etymology: From asper (Latin) meaning rough and spora (Latin) to emphasize the rough spores.
Typification: Japan, Kagoshima, Amami-Ŏshima Island, Yamato, Amami Forest Polis Park (approx. 28.316056N, 129.344778E, alt. approx. 214 m asl.) under Castanopsis sieboldii, Pinus luchuensis, and Quercus glauca, 17 Nov 2015, K. Hosaka, Holotype (TNS-F-75705), database record HJB16251). GenBank ITS = MZ782138.
Description: Pileus (21) 29–52 (58) mm diameter, hemispherical, convex often umbonate; margin often crenulate or eroded, often spotting, not hygrophanous; usually almost unicolored with color at center clay-buff, honey or dark pinkish buff, paler at margin. Lamellae emarginate, white, cream to brown, with strongly white fimbriate edge and droplets visible with the naked eye, number of full-length lamellae 70–72. Stipe (26) 32–56 (77) mm long, (6) 7–10 (13) mm diameter at median, widening towards a clavate base, surface cream, ivory to pale brown but discoloring from the base upwards, fibrillose, at apex pruinose. Context in pileus white to cream, firm, in stipe stuffed; taste and smell not recorded. Spore deposit color not recorded.
Basidiospores based on n = 45 spores of the holotype, 5% to 95% percentile range 9.0–10.8 × 5.4–6.6 µm, with median 9.5 × 5.9 µm and av. 9.6 × 5.9 µm with S. D. length 0.6 µm and width 0.35 µm; Q value 5% to 95% percentile range 1.53–1.77, with median 1.60 and av. 1.62 with S. D. 0.08; spore size based on two collections, medians 9.5–10.1 × 5.9–6.3 µm and av. 9.6–10.0 × 5.9–6.2 µm with av. S. D. length 0.565 µm and width 0.33 µm, av. Q 1.61–1.62, amygdaloid or limoniform, with small apiculus and rounded apically, with a distinct thinning of the apical wall and a strongly prominent papilla, guttulate with one or sometimes more oily drops, usually very distinctly ornamented (ornamentation visible at low magnification), with a strongly loosening perispore on almost every mature spore and indextrinoid hardly changing color in Melzer’s reagent (O3/4; P3; D0); spore color under the microscope brown. Basidia 22–30 × 6–8 µm, with av. Q 3.2–3.6 µm, cylindrical to clavate, without pigmentation, 4–spored. Cheilocystidium width near apex holotype 5% to 95% percentile range 5.7–9.2 µm, with median 7.7 µm and av. 7.6 µm with S. D. 1.1 µm; across two collections median 7.4–7.7 µm and av. 7.4–7.6 µm; examining approx. 20 selected cheilocystidia of each of the two collections yields a range for the avs. of 44–45 × 7.4–7.6 × 3.9–4.3 × 5.3–6.0 µm and 44 × 7.6 × 4.3 × 6.0 µm av. for holotype. Cheilocystidium av. ratios A/M: 1.79–1.91, A/B: 1.30–1.49, B/M: 1.36–1.41, mainly clavate-ventricose, often capitate stipitate or clavate stipitate, sometimes with one or two septa (occasionally clamped) often with thickening of the median wall. Pleurocystidia absent. Caulocystidia similar to cheilocystidia but larger, up to 110 μm long. Pileipellis is an ixocutis with an epicutis up to 150 µm thick, with gelatinized, often encrusted hyphae up to 6 µm wide. Subcutis pale yellow under the microscope and the trama below the cutis made up of cylindrical or thickly sausage-shaped cells up to 16 µm wide. Clamp connections present throughout the basidiome.
Ecology and distribution: In mixed woodlands with Pinus, Quercus and Castanopsis. The growth habit was caespitose to scattered. To date, Hebeloma asperosporum is recorded only from the island Amami-Ŏshima.
Additional collections examined: Japan, Kagoshima, Amami-Ŏshima Island, Yamato, Amami Forest Polis Park (approx. 28.316056N, 129.344778E, alt. approx. 214 m asl.) under Castanopsis sieboldii, Pinus luchuensis and Quercus glauca, 17 Nov 2015, K. Hosaka (TNS-F-75704, HJB16250).
Japanese name: Amami-wakafusa-take (newly proposed here).
Remarks: The mainly clavate-ventricose cheilocystidia together with distinctly to strongly ornamented spores support the placement of this taxon in H. subsect. Clepsydroida within H. sect. Denudata. The occasional presence of cystidia with thickening of the wall in the middle is further evidence of this placement. Molecularly, the position of the species within H. sect. Denudata is supported, but its assignment to subsection is not. Within H. sect. Denudata, the small spores, on average less than 10.5 µm long, indextrinoid but strongly ornamented and with consistently loosening perispore distinguishes this species from all other known members of H. section Denudata. While the description is based on just two collections, its morphological and molecular separation, from all known species of Hebeloma, leave us in no doubt that this should be regarded as a distinct species.
Of course, our description and knowledge of this taxon is limited to just these two collections from the same locality, hence our concept of this species is limited. A sequence published from China as H. alpinum (MW554385, Zouh, unpublished, submitted 26 Jan 2021; no UNITE SH to date) is, apart from obvious editing errors, identical to our H. asperosporum sequences. It is hoped that publishing this description will encourage other mycologists to search for this taxon.
Hebeloma cinnamomeum Beker & U. Eberh., sp. nov., Figs. 9 and 10, Mycobank MB840288
Hebeloma cinnamomeum holotype (TNS-F-82067) a spores and b spore ornamentation in Melzer’s reagent × 1600, c spores and d spore ornamentation in KOH × 1600, e cheilocysidia in Melzer’s reagent × 1000, f cheilocysidia in KOH × 1000, g basidia in KOH × 1000, h cheilocystidia in KOH × 1000, i caulocystidia in KOH × 1000, j caulocystidia in KOH × 500, k epicutis hyphae and l subcutis in KOH × 500, and m cutis in KOH × 125. Scale bars 10 µm, in m 100 µm. Photos H.J. Beker
Diagnosis: The cinnamon-colored pileus, the spores with average Q less than 1.85 and the cheilocystidia with average width near the apex of less than 6.5 µm, are characters that separate this species from others in H. sect. Denudata.
Etymology: From cinnamomeus (adj. Latin) describing the cinnamon-colored pileus.
Typification: Japan, Ishikawa, Wajima, Mii, Nakanagatani (approx. 37.39059N, 136.899196E, alt. approx. 229 m asl.), under Pinus densiflora, Quercus serrata and Q. variabilis, 16 Oct 2016, T. Kasuya TKB3261, Holotype (TNS-F-82067), database record HJB15837. Genbank ITS = MZ782111.
Description: Pileus (20) 21–46 (51) mm diameter, hemispherical, often convex, occasionally umbonate; margin involute, particularly when young, not hygrophanous; usually almost unicolored, usually cinnamon, rarely more brick colored, sometimes slightly paler towards margin. Lamellae emarginate, depth up to 4 mm, white, cream to brown, with white fimbriate edge but without droplets on the lamella edge, number of full-length lamellae 52–64. Stipe (24) 30–60 (64) mm long, 4–8 (11) mm diameter at median, often widening towards a clavate base, surface cream, ivory to pale brown rarely weakly discoloring from the base upwards, velute, at apex pruinose. Context in pileus white to cream, firm, in stipe stuffed; taste and smell not recorded. Spore deposit color not recorded.
Basidiospores based on n = 90 spores of the holotype, 5% to 95% percentile range 9.1–12.5 × 5.4–7.1 µm, with median 10.2 × 6.0 µm and av. 10.5 × 6.1 µm with S. D. length 1.1 µm and width 0.59 µm; Q value 5% to 95% percentile range 1.49–1.97, with median 1.70 and av. 1.72 with S. D. 0.16; spore size based on 23 collections, medians 9.5–11.2 × 5.7–6.3 µm and av. 9.6–11.2 × 5.8–6.3 µm with av. S. D. length 0.76 µm and width 0.45 µm, av. Q 1.60–1.83, amygdaloid, occasionally limoniform, with small apiculus and rounded apically, with a distinct thinning of the apical wall and sometimes a weak papilla, usually guttulate with one or sometimes more oily drops, weakly to distinctly ornamented (ornamentation not conspicuous at low magnification), with a perispore hardly loosening and weakly but sometimes distinctly dextrinoid, becoming at most pale brown or yellow brown in Melzer’s reagent, (O2/3; P0/1; D1/2); spore color under the microscope yellow brown. Basidia 22–33 × 6–8 µm, with av. Q 3.0–3.9 µm, cylindrical to clavate, without pigmentation, 4–spored. Cheilocystidium width near apex holotype 5% to 95% percentile range 5.9–7.5 µm, with median 6.7 µm and av. 6.6 µm with S.D. 0.54 µm; across 23 collections median 5.0–6.7 µm and av. 5.0–6.6 µm; examining approx. 20 selected cheilocystidia of each of the 23 collections yields a range for the avs. of 50–61 × 5.0–6.6 × 3.3–3.9 × 4.6–7.0 µm and 54 × 6.6 × 3.7 × 4.9 µm av. for holotype. Cheilocystidium av. ratios A/M: 1.42–1.81, A/B: 0.86–1.37, B/M: 1.35–1.86, mainly clavate-ventricose, rarely clavate stipitate, sometimes with one or two septa (occasionally clamped) often with thickening of the median wall, rarely with thick yellow content. Pleurocystidia absent. Caulocystidia similar to cheilocystidia but larger, up to 110 μm long. Pileipellis is an ixocutis with an epicutis up to 160 µm thick, with gelatinized, often encrusted hyphae up to 6 µm wide. Subcutis cinnamon colored and the trama below the cutis made up of cylindrical or ellipsoid, often thickly sausage-shaped cells up to 12 µm wide. Clamp connections present throughout the basidiome.
Ecology and distribution: In deciduous or mixed woodlands apparently associated with Quercus or Pinus, but occasionally Carpinus and Fagus were also present. The growth habit was mainly scattered, but occasionally caespitose and rarely solitary. To date, all collections of Hebeloma cinnamomeum have been recorded only from the Japanese island of Honshu.
Additional collections examined: Japan, Ibaraki, Tsukuba, Tsukuba Botanical Garden, (approx. 36.1015N, 140.1131E, alt. approx. 40 m asl.), 16 Oct 2000, D. Yoshioka (TNS-F-1387, HJB16173); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Evergreen Broad-leaved Forest Section,” (approx. 36.101518N, 140.11308E, alt. approx. 40 m asl.) on soil under Quercus myrsinifolia and other evergreen Quercus spp., 27 Oct 2011, Y. Muramatsu (TNS-F-42284, HJB16213); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Evergreen Broad-leaved Forest Section” (approx. 36.101518N, 140.11308E, alt. approx. 40 m asl.) on soil under Quercus myrsinifolia and other evergreen Quercus spp., 9 Nov 2011, Matsumoto (TNS-F-43723, HJB16214); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Evergreen Broad-leaved Forest Section” (approx. 36.101518N, 140.11308E, alt. approx. 40 m asl.) on soil under Quercus myrsinifolia and other evergreen Quercus spp., 4 Dec 2011, not recorded (TNS-F-44535, HJB16216); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Evergreen Broad-leaved Forest Section” (approx. 36.1015N, 140.1131E, alt. approx. 40 m asl.) on soil under Quercus myrsinifolia and other evergreen Quercus spp., 1 Nov 2012, K. Nishibori (TNS-F-49380, HJB16228); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Rockeries Section (High Altitudes)” (approx. 36.101513N, 140.113035E, alt. approx. 40 m asl.) on soil under evergreen Quercus spp., 1 Nov 2012, N.P. Thao (TNS-F-49387, HJB16229); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Evergreen Broad-leaved Forest Section” (approx. 36.1015N, 140.1131E, alt. approx. 40 m asl.) on soil under Quercus myrsinifolia and other evergreen Quercus spp., 1 Nov 2012, N.P. Thao (TNS-F-49389, HJB16230); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Fern Section” (approx. 36.1008N, 140.1126E, alt. approx. 38 m asl.), 10 Oct 2013, Dawa (TNS-F-58915, HJB16234); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Fern Section” (approx. 36.10078N, 140.112572E, alt. approx. 38 m asl.) on soil under Quercus sp., 13 Nov 2013, K. Hosaka (TNS-F-59097, HJB16235); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Fern Section” (approx. 36.10078N, 140.112572E, alt. approx. 38 m asl.) on soil under Quercus sp., 13 Nov 2013, K. Hosaka (TNS-F-59098, HJB16236); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Fern Section” (approx. 36.1008N, 140.1126E, alt. approx. 38 m asl.), 5 Nov 2014, K. Nishibori (TNS-F-72963, HJB16243); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Cool-temperate Deciduous Broad-leaved Forest Section” (approx. 36.103102N, 140.111201E, alt. approx. 43 m asl.) on soil in deciduous woodland under Carpinus sp. and Fagus sp., 13 Nov 2014, K. Hosaka (TNS-F-73028, HJB16244); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Warm-temperate Deciduous Broad-leaved Forest Section” (approx. 36.102944N, 140.11268E, alt. approx. 35 m asl.) on soil in deciduous woodland, 22 Oct 2015, K. Nishibori (TNS-F-73984, HJB16245); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Fern Section” (approx. 36.1008N, 140.1126E, alt. approx. 38 m asl.) on soil, 18 Nov 2015, J. Yamazaki (TNS-F-74039, HJB16246); Ibaraki, Tsukuba, Tsukuba Botanical Garden “Endangered Species Section” (approx. 36.1015N, 140.1131E, alt. approx. 40 m asl.) on soil, 25 Nov 2015, J. Yamazaki (TNS-F-74060, HJB16247); Ibaraki, Tsukuba, Usui, Tsukuba-Fureai-no-Sato (approx. 36.201127N, 140.108664E, alt. approx. 90 m asl.), 16 Oct 2005, collector not recorded (TNS-F-11893, HJB16177); Ishikawa, Nomi, Tatsunokuchi, Ishikawa Zoo (approx. 36.433759N, 136.545712E, alt. approx. 50 m asl.) under Quercus serrata, 4 Oct 2001, H. Mori (TNS-F-38024, HJB16199); Ishikawa, Wajima, Mii, Nakanagatani (approx. 37.39059N, 136.899196E, alt. approx. 40 m asl.), 16 Oct 2016, T. Kasuya (HJB15838); Tottori, Daisen, Mt. Daisen (approx. 36.314665N, 139.800149E, alt. approx. 35 m asl.), 12 Oct 2008, Y. Taneyama (TNS-F-44863, HJB16220); Tottori, Daisen, Mt. Daisen (approx. 36.314665N, 139.800149E, alt. approx. 35 m asl.), 12 Oct 2008, M. Nabe (TNS-F-44864, HJB16221); Yamanashi, Kofu, Atago, Mt. Atago, Kodomono-kuni (approx. 35.6702N, 138.5803E, alt. approx. 410 m asl.) under Pinus densiflora and Pinus resinosa, 11 Nov 2011, K. Hosaka (TNS-F-46098, HJB16226); Yamanashi, Fujiyoshida, Yamanashi Institute of Environmental Sciences (approx. 35.477904N, 138.795091E, alt. approx. 839 m asl.) under Pinus densiflora, 19 Sep 2013, K. Hosaka (TNS-F-60045, HJB16238).
Japanese name: Mizuho-wakafusa-take (newly proposed here).
Remarks: The mainly clavate-ventricose cheilocystidia together with distinctly ornamented spores support the placement of this taxon in H. subsect. Clepsydroida within H. sect. Denudata. The occasional presence of cystidia with thickening of the wall in the middle is further evidence of this placement. All loci apart from mitSSU V6 support a shared lineage of H. cinnamomeum with H. cavipes (the type species of H. subsect. Clepsydroida) rather than with H. crustuliniforme (the type of H. subsect. Crustuliniformia) or H. hiemale (the type of H. subsect. Hiemalia). Within H. sect. Denudata, the spores with average Q less than 1.85, the cheilocystidia with average width near the apex of less than 6.5 µm and the cinnamon-colored pileus are characters that separate this species from others within this subsection. The pileus color is an eye-catching feature that may well enable field identification, or at least a reasonable suspicion of determination, of this species.
While our morphological description is based on 23 collections, many (but not all) are from a relatively confined area within the Tsukuba Botanical Garden, which is planted with endogenous plants from Japan. The description provided may be too narrow, given the similar habitat for many of these collections. In addition, a sequence from a Quercus dentata root tip collected in Hokkaido (LC068995, Arai et al. unpublished, submitted 27-Jul-2015) clusters in ML analyses (not shown) with H. cinnamomeum sequences, supporting the assumptions on associates and adding evidence that it is an endogenous species in Japan. This sequence currently (9 June 2021) forms a singleton UNITE SH hypothesis (SH2292369.08FU) at the < 0.5% level, but owing to probable editing errors in this sequence this SH has to be treated with caution.
Note that cited collection TNS-F-44863 was originally identified as Hebeloma microsporum, the name under which it is recorded on GBIF, and which was mentioned in the introduction.
Hebeloma citrisporum Beker & U. Eberh., sp. nov. Figs. 11 and 12, Mycobank MB840289
Hebeloma citrisporum holotype (BR 5020214140236 V) a spores and b spore ornamentation in KOH × 1600, c basidium in KOH × 1000, d spores and e spore ornamentation in Melzer’s reagent × 1600, f cheilocystidium in KOH × 1000, g caulocystidia in KOH × 1000, h epicutis hyphae in KOH × 500, i cheilocystidium in KOH × 1000, j caulocystida in KOH × 500, k subcutis in KOH × 500, and l cutis in KOH × 125. Scale bars 10 µm, in l 100 µm. Photos H.J. Beker
Diagnosis: The citriform spores, distinctly ornamented and rather strongly dextrinoid but with the perispore only somewhat loosening in a few spores, with average width less than 7.5 µm, and the predominantly gently clavate cheilocystidia with average width near the apex greater than 6.5 µm and average ratio of basal width to median width (B/M) at most 1.35, distinguish this species from other Hebeloma.
Etymology: From citrus (noun, Latin) meaning lemon, lemon tree and spora (Latin) to emphasize the limoniform spores.
Typification: Japan, Ibaraki, Tsukuba, (approx. 36.0869N, 140.1197E, alt. approx. 30 m asl.) in urban pondside roadside under Quercus myrsinifolia and Q. serrata, 24 Oct 2016, U. Kawasaki UK270, MO258729, Holotype (BR 5020214140236 V), database record HJB15833. GenBank ITS = MZ782110.
Description: Pileus (21) 28–63 (68) mm diameter, hemispherical, often umbonate, occasionally convex; margin usually smooth, occasionally involute or wavy, not hygrophanous; usually bicolored, usually clay buff, distinctly paler towards margin. Lamellae emarginate, white, cream to brown, with white fimbriate edge but without droplets on the lamella edge, number of full-length lamellae 52–64. Stipe (22) 31–64 (84) mm long, (6) 7–9 (10) mm diameter at median, widening towards a bulbous base, surface cream, ivory to pale brown not discoloring on handling, velute, slightly fibrillose, at apex pruinose to floccose. Context in pileus and stipe white to cream, firm; taste and smell not recorded. Spore deposit color not recorded.
Basidiospores based on n = 90 spores of the holotype, 5% to 95% percentile range 10.4–13.1 × 6.2–7.4 µm, with median 11.6 × 6.9 µm and av. 11.6 × 6.8 µm with S. D. length 0.86 µm and width 0.40 µm; Q value 5% to 95% percentile range 1.53–1.90, with median 1.69 and av. 1.70 with S. D. 0.11; spore size based on 5 collections, medians 11.3–12.3 × 6.6–7.1 µm and av. 11.3–12.1 × 6.7–7.0 µm with av. S. D. length 0.82 µm and width 0.37 µm, av. Q 1.61–1.83, amygdaloid or limoniform, with small apiculus and rounded apically, with a distinct thinning of the apical wall and a very strong, very prominent papilla, usually guttulate with one or sometimes more oily drops, weakly to distinctly ornamented (ornamentation not conspicuous at low magnification), with a perispore somewhat to distinctly loosening in many spores and rather strongly dextrinoid, becoming medium brown in Melzer’s reagent, (O2/3; P1/2; D3); spore color under the microscope yellow to yellow brown. Basidia 26–36 × 7–9 µm, with av. Q 3.4–3.6 µm, cylindrical to clavate, without pigmentation, 4-spored. Cheilocystidium width near apex holotype 5% to 95% percentile range 4.8–8.4 µm, with median 6.6 µm and av. 6.6 µm with S. D. 0.54 µm; across 5 collections median 6.6–8.9 µm and av. 6.6–9.1 µm; examining approx. 20 selected cheilocystidia of each of the 5 collections yields a range for the avs. of 54–68 × 6.6–9.1 × 5.7–3.9 × 5.3–7.0 µm and 54 × 6.6 × 4.7 × 5.3 µm av. for holotype. Cheilocystidium av. ratios A/M: 1.31–1.58, A/B: 1.08–1.38, B/M: 1.12–1.35, mainly gently clavate, occasionally clavate-ventricose or ventricose, sometimes with one or two septa. Pleurocystidia absent. Caulocystidia similar to cheilocystidia but larger, up to 135 μm long. Pileipellis is an ixocutis with an epicutis up to 115 µm thick, with gelatinized, often encrusted hyphae up to 7 µm wide. Subcutis yellow brown and the trama below the cutis made up of cylindrical or thickly sausage-shaped cells up to 18 µm wide. Clamp connections present throughout the basidiome.
Ecology and distribution: All records to date suggest Quercus as the most likely ectomycorrhizal host. The growth habit was mainly scattered, but occasionally caespitose. To date, all collections of Hebeloma citrisporum have been recorded only from the Japanese island of Honshu, in the vicinity of Tokyo.
Additional collections examined: Japan, Tokyo, Shibuya, Yoyogi, Meiji Shrine (approx. 35.6781N, 139.6999E, alt. approx. 60 m asl.) under Quercus sp., 8 Nov 2011, K. Hosaka KH-JPN11-529 (TNS-F-44560, HJB16217); Ibaraki, Tsukuba (approx. 36.086858N, 140.119679E, alt. approx. 29 m asl.) in urban pondside roadside under Quercus myrsinifolia and Quercus serrata, 4 Oct 2016, U. Kawasaki UK267 (MO255034, HJB15832); Ibaraki, Tsukuba (approx. 36.085N, 140.12E, alt. approx. 25 m asl.) in mixed woodland, 20 Oct 2017, U. Kawasaki UK338 (MO295075, HJB17412); Ibaraki, Tsukuba (approx. 36.085N, 140.12E, alt. approx. 25 m asl.), 27 Oct 2017, U. Kawasaki UK350 (MO299321, HJB17414).
Japanese name: Tokiwa-wakafusa-take (newly proposed here).
Remarks: The predominantly gently clavate towards the apex cheilocystidia, with a few ventricose or clavate-ventricose, supports the placement of H. citrisporum in H. sect. Velutipes. Within this section, the citriform spores, distinctly ornamented and rather strongly dextrinoid but with the perispore only somewhat loosening in a few spores, with average width less than 7.5 µm, and the cheilocystidia with average width near the apex greater than 6.5 µm and average ratio of basal width to median width (B/M) at most 1.35, distinguish this species. While citriform spores are commonly found within species of H. sect. Velutipes, rarely are they as regular and eye-catching as for this species. The species and its placement in H. sect. Velutipes are supported by molecular data (Fig. 6).
The description is based on just five collections from a relatively small geographic area around Tokyo. We would suspect that it is far more widespread, at least on Honshu Island. A Quercus dentata root tip sequence from Hokkaido (AB979728, Arai et al. 2017) might have been formed by H. citrisporum. Currently (9 June 2021), the sequence forms a singleton UNITE SH (SH1235508.08FU) at the 3% level. It differs from our H. citrisporum sequences by three indels, two singleton deletions and one 15 bp insertion which we have not observed in our material.
Hebeloma danicum Gröger, Z. Mykol. 53(1): 53 (1987).
Japanese name: Ashinaga-numeri (Kawamura 1954).
Remarks: A detailed description and molecular analyses including this species are provided in Beker et al. (2016). In Eberhardt et al. (2020b) the authors commented that Japanese collections studied that were referred to H. spoliatum were similar to or conspecific with H. danicum, but that further research was necessary to decide upon any conspecificity. Following further morphological and molecular analysis using multiple loci, we cannot see any reason to differentiate the Japanese collections, albeit for some of the collections (but importantly not all), there are some minor morphological differences. Specifically, the range for the number of complete lamellae for our European collections is 58–64, while for the Japanese collections this is 60–72, and the range of spore lengths for our European collections is 9.5–10.3 µm while the range for our Japanese collections is 8.5–10.0 µm. Both these parameters affect the use of the dichotomous key provided in Beker et al. (2016), and might therefore lead to some confusion with other species of H. sect. Scabrispora, in particular H. laterinum, H. melleum, or H. pumilum. However, all three of these species can be straightforwardly distinguished: H. laterinum with its blackening stipe, H. melleum with its smaller basidiomes and H. pumilum, with its consistently loosening and very well visible perispore.
Hebeloma nanum Velen., Novitates mycologicae: 117 (1939).
= Hebeloma crustuliniforme f. microspermum Hongo, Journal of Japanese Botany 41: 169 (1966).
Japanese name: Kotsubu-o-wakafusa-take (Hongo 1966).
Remarks: A detailed description of H. nanum and molecular analyses of H. sect. Naviculospora are provided in Beker et al. (2016). The holotype of Hebeloma crustuliniforme f. microspermum has small (av. 7.9 × 4.8 µm) spores, prompting the choice of name. The spores, however, are weakly ornamented and strongly dextrinoid (unlike H. crustuliniforme). The cheilocystidia are very irregular, subclavate to subcylindrical, again very unlike H. crustuliniforme, but typical for H. sect. Naviculospora, as is Hongo’s description of the pileus: “viscid, pale tan, often tinged brownish-alutaceous, especially on the disc, sometimes irregularly cracked.” Within this section, the narrow spores and rarely ventricose cheilocystidia are typical for H. nanum, with which this taxon is certainly conspecific. We were not able to generate any molecular data from the type material. Hebeloma crustuliniforme f. microspermum was described by Hongo (1966) from pine forests in Akiba-yama, Niitsu, Niigata Prefecture. We have examined and sequenced another collection of H. nanum from Japan (TNS-F-55100; GenBank ITS = MZ782125).
Hebeloma nanum is widespread across the globe and as well as in Japan and northern Europe; we are aware of confirmed collections from India, Canada, China, and the USA (unpublished results). The species probably corresponds to UNITE species hypothesis SH1951515.08FU, including the type (from Czechia) and, in addition to the above, sequence data from Pakistan. It appears that H. nanum most commonly occurs on sandy soil, in post-fire regenerating boreal and temperate, coniferous forests (Beker et al. 2016; Hughes et al. 2020).
Homophron helvolescens (S. Imai) Beker & U. Eberh., comb. nov.
MB840290
Basionym: Hebeloma helvolescens S. Imai, Journal of the Faculty of Agriculture of the Hokkaido Imperial University 43(2): 229 (1938).
Typification: Japan, Hokkaido, Sapporo Botanic Garden, 4 Jul 1932, on the ground around the tree trunks in open woods, S. Imai, Holotype (SAPA 10000035), database record HJB1000374. GenBank ITS = MZ725549.
Description of type: The spores are elliptical to cylindrical, sometimes fabiform, smooth, pale brown, without a germ pore, on av. 9.9 × 5.3 µm, with av. spore Q 1.87. The cheilocystidia are metuloid; the pleurocystidia are similar.
Japanese name: Ko-wakafusa-take (Imai 1938).
Remarks: Our examination of the type material is in good agreement with the description given by Imai (1938). Imai also mentions the caespitose habit of this species, and the presence of crystals on the cystidia. Morphologically, it appears to be closely related or even conspecific with Ho. spadiceum.
Spore size of Ho. spadiceum is, according to Vašutová (2008), 8–9.5 × 4–5 μm, average 8.7 × 4.7 μm, Q 1.6–2.2 (−2.3) and according to Enderle (1989) who later collected the specimen that was selected as epitype by Örstadius (2001) (8.0)9–10.2(11) × 4.2–5.2(5.6) µm. Both authors stress the light spore colour of the species, under the microscope in water and in the spore deposit, according to Vašutová (2008) brick-beige (S40Y60M5, Küppers 1999). Vašutová (2008) suspected that Smith (1972) concept of Psathyrella spadicea might not coincide with what is called Ho. spadiceum in Europe, citing a difference in spore deposit colour as indication. Imai (1938) described the spores of Ho. helvolescens as cinnamon; S40Y60M5 (Küppers 1999) is lighter and more greenish than cinnamon; under the microscope the spores of Ho. helvolescens appear distinctly colored.
The ITS of the eptitype of Ho. spadiceum is sequenced, as is the holotype of Ho. helvolescens and both are included in the “Homophron spacideum s.l.” clade of Fig. 3. This clade includes a number of supported (and unsupported) subclades that appear to have restricted geographical distributions. However, the sequence variation underlying these clades is so small (see scale in Fig. 3) and the data is only from a single locus that we hesitate to recognize them as hypotheses of distinct species. Vašutová et al. (2008) stressed that ITS is not sufficient for delimiting species in the Psathyrellaceae. Thus, it is not clear whether Ho. helvolescens is indeed a later synonym of Ho. spadiceum.
Pholiota tomoeae (S. Imai) Beker & U. Eberh. comb. nov.
MB840291
Basionym: Hebeloma tomoeae S. Imai, Journal of the Faculty of Agriculture of the Hokkaido Imperial University 43(2): 226 (1938).
Typification: Japan, Hokkaido, Sapporo, 14 Oct 1937, on the humus ground or decayed wood under trees, in gardens or roadsides, S. Imai, Syntypes (SAPA 10000031 and SAPA 10000032). Lectotype designated here (MBT10001755, SAPA 10000031), database record HJB1000379. GenBank ITS = MZ724681.
Description of syntypes: The spores are smooth, ellipsoid to ovate, yellowish brown to brown and with a very small indistinct germ pore. For SAPA 1000031 the spores are on av. 7.6 × 4.9 µm, with av. spore Q 1.56, while for SAPA 1000032 they are on av. 7.9 × 4.8 µm, with av. spore Q 1.63. In both cases, the pleurocystidia are mainly ventricose, hyaline to yellowish in 5% KOH, while the cheilocystidia are more utriform in shape.
Japanese name: Tomoe-take (Imai 1938).
Remarks: Based on morphology and ITS sequence data (Fig. 4), the syntypes represent the same taxon. Although neither of the syntypes is in particularly good condition, of the two collections, SAPA 10000031 yielded better sequence data.
The description is in good agreement with the description given by Imai (1938). Although Imai does not mention burnt ground as the habitat, morphologically and molecularly both syntypes of H. tomoeae are in good agreement with Pholiota brunnescens, which is associated with burnt soil or wood (Matheny et al. 2018). It appears likely that P. tomoeae is a later synonym of P. brunnescens; we rather leave it to Pholiota experts to synonymize the two species, also considering that the phylograms based on several loci published by Matheny et al. (2018) and Tian and Matheny (2021) suggest that P. brunnescens sl may be rather variable and is not supported by bootstrap (Tian and Matheny 2021).
Discussion
For the newly described species as well as for the species described by Imai (1938), we had both morphological and molecular data available and the conclusions from both kinds of data supported each other. With regard to Imai’s species, it is clear that, at the time he described these four species, the concepts of brown-spored genera were rather different from today. We cannot be sure that all of the new combinations are indeed non-redundant species. It is beyond the scope of this study to explore the limits of species from genera other than Hebeloma. We only have ITS data available for the types which may not be sufficient to distinguish species in Agrocybe, around Homophron spadiceum or Pholiota brunnescens. This situation leads to a dilemma between different principles of best taxonomic practices formulated by Aime et al. (2021): On the one hand, the publication of superfluous names ought to be avoided, on the other we view the recombination into the genus we consider the correct one as the most efficient means to communicate our findings to the experts for the respective genera. The experts, in our view, should make the decision on the synonymy or otherwise of A. imaii with A. pediades, A. humosa with A. smithii, Ho. helvolescens with Ho. spadiceum, and P. tomae with P. brunnescens.
Based on the almost completed study of Hebeloma types worldwide, we are confident that the new Hebeloma species, H. asperosporum, H. cinnamomeum, and H. citrisporum, are good species and all of them can, based on our current knowledge, be identified by ITS alone.
The presence of conflicts between single locus results in the genus Hebeloma was previously known. Conflicts have been discussed earlier by Grilli et al. (2016) for H. sect. Velutipes and by Eberhardt et al. (2016) for H. sect. Clepsydroida. The recent recognition of additional taxa in H. sect. Clepsydroida, i.e., H. asperosporum, H. cinnamomeum, and H. sordidulum (Eberhardt et al., submitted) appears to have increased the conflict. While more advanced analysis methods might have given a better representation than concatenation, of what the available data suggest about species relationships, it would not have changed the fact that additional loci are needed to resolve infrageneric relationships. In the case of H. sect. Denudata, additional loci are also needed for the delimitation of subsections. Currently, we do not have molecular support that the morphological characters that are used for distinguishing and delimiting H. subsects. Clepsydroida, Crustuliniformia, and Hiemalia have evolved concertedly among the members of each subsection.
We are not aware of any records outside of Japan of the three new species described here (but note the remark made above with regard to a possible collection of H. asperosporum from China). The species may be endemic to these islands, but more collections, particularly of H. asperosporum and H. citrisporum, are needed before such a hypothesis could be either accepted or rejected.
With regard to the list of Hebeloma for Japan produced by Katumoto (2010) which listed 15 species, we can remove the four Imai species (H. fimicola, H. helvolescens, H. humosum, H. tomoeae), which are not Hebeloma. Additionally, it may be safer to reject H. longicaudum and H. sinuosum that should be regarded as dubious names and without material to examine, it is difficult to assign them. Hebeloma crustuliniforme f. microspermum is H. nanum, a species for which we have another collection, confirming its presence in Japan. Collections previously referred to H. spoliatum should likely be referred to H. danicum as discussed above. Collections previously referred to H. radicosum should likely be referred to H. sagarae as discussed in Eberhardt et al. (2020b).
We are unable to confirm the presence, in Japan, of H. crustuliniforme, H. mesophaeum, and H. sacchariolens. It is possible that those collections which were called H. crustuliniforme in the past are H. cavipes or H. velutipes, both of which have often been mistakenly determined as H. crustuliniforme (Vesterholt et al. 2014). It is likely that species previously determined as H. sacchariolens did indeed belong to H. sect. Sacchariolentia but during our studies, we have not identified any collections from this section. Four collections recorded as H. sacchariolens were in fact Pholiota sp., H. danicum and H. vinosophyllum (2). With regard to H. mesophaeum, again it is likely that species previously determined as H. mesophaeum did indeed belong to H. sect. Hebeloma but during our studies, the only species from this section that we have encountered is H. hygrophilum. The final three species Katumoto mentioned (H. luchuense, H. radicosoides, H. vinosophyllum) are certainly present in Japan and as yet the first two of these have not been recorded from anywhere outside Japan.
In summary, we can add, as verified for Japan, H. cavipes, H. eburneum, H. hygrophilum, H. subtortum, and H. velutipes, as well as the three new species described here, bringing the total number of Hebeloma species that we can confirm to be present in Japan to fifteen.
Searches in GenBank and UNITE (Kõljalg et al. 2005; Johnson et al. 2008; Nilsson et al. 2019) for sequence data from Japan strongly suggest that H. sordescens (AB848488, Miyamoto et al. 2014) and H. rostratum (MT596467, Faverol-Longo et al. unpublished, direct submission 08-Jun-2020) occur in the country, while sequences from H. sect. Hebeloma (e.g., AB211272 or AB327182, Nara 2006; Obase et al. 2007) could belong to various species of the H. mesophaeum complex; AB211274 (Nara 2006) might be H. helodes or H. auraniotumbrinum; AB848487 (Miyamoto et al. 2014) or some others could be H. incarnatulum rather than H. velutipes and EU711239 (Roy et al. 2009) or some others could be H. leucosarx rather than H. velutipes. LC009707 (Maeno and Sagara, unpublished, direct submission 6 Nov 2014) might be H. melleum or a closely related taxon.
Given the, relatively, small number of collections available for this study, it can be assumed that there are many more species of Hebeloma to be discovered from the islands of Japan. We hope that this publication will encourage the collection and determination of Hebeloma in the country. In lieu of a key for Hebeloma in Japan (which would be deficient, based on too few collections), we refer to an interactive identification tool for Hebeloma that is currently under development (Bartlett et al. 2021).
Data availability
The majority of the material was obtained through public collections or, in the case of the private collection of H.J.B., will be accessioned to a public collection when the project is finished. Sequence data were submitted to GenBank. Alignments and trees were submitted to TreeBASE.
Code availability
Not applicable.
References
Aime MC, Miller AN, Aoki T, Bensch K, Cai L, Crous PW, Hawksworth DL, Hyde KD, Kirk PM, Lücking R, May TW, Malosso E, Redhead SA, Rossman AY, Stadler M, Thines M, Yurkov AM, Zhang N, Schoch CL (2021) How to publish a new fungal species, or name, version 3.0. IMA Fungus 12:11. https://doi.org/10.1186/s43008-021-00063-1
Arai H, Tamai Y, Yajima T, Obase K, Miyamoto T (2017) Ectomycorrhizal fungal communities associated with Quercus dentata in a coastal broadleaf forest. Mycosphere 8:561–567. https://doi.org/10.5943/mycosphere/8/4/5
Arora D (1986) Mushrooms Demystified. Ten Speed Press, Berkeley, California
Bartlett P, Eberhardt U, Schütz N, Beker HJ (2021) Machine learning for species identification: the Hebeloma project from database to website. Biodivers Inf Sci Standards 5:e73972. https://doi.org/10.3897/biss.5.73972
Beker HJ, Eberhardt U, Vesterholt J (2016) Hebeloma (Fr.) P. Kumm. Fungi Europaei, vol 14. Edizioni Tecnografica, Lomazzo
Beker HJ, Eberhardt U, Vesterholt J (2010) Hebeloma hiemale Bres. in arctic/alpine habitats. N Am Fungi 5:51–65
Bon M (2002) Clé de détermination du genre Hebeloma (Fr.) Kumm. Documents Mycologiques 30(123):3–40
Breitenbach J, Kränzlin F (2000) Fungi of Switzerland. Part 3. Volume 5. Mykologia, Luzern, Switzerland
Cho HJ, Lee H, Park JY, Park MS, Kim NK, Eimes JA, Kim C, Han S-K, Lim YW (2016) Seven new recorded species in five genera of the Strophariaceae in Korea. Mycobiology 44(3):137–145. https://doi.org/10.5941/MYCO.2016.44.3.137
Clémençon H, Hongo T (1994) Notes on three Japanese Agaricales. Mycoscience 35:21–27
Cripps C, Eberhardt U, Schütz N, Beker HJ, Evenson VS, Horak E (2019) The genus Hebeloma in the Rocky Mountain alpine zone. MycoKeys 46:1–54. https://doi.org/10.3897/mycokeys.46.32823
Dentinger BTM, Didukh MY, Moncalvo J-M (2011) Comparing COI and ITS as DNA barcode markers for mushrooms and allies (Agaricomycotina). PLoS ONE 6:e25081. https://doi.org/10.1371/journal.pone.0025081
Eberhardt U (2012) Methods for DNA barcoding fungi. In: Kress JW, Erickson DL (eds) DNA Barcodes: methods and protocols. Humana Press Imprint (Springer), New York, pp 183–205. https://doi.org/10.1007/978-1-61779-591-6_9
Eberhardt U, Beker HJ, Vila J, Vesterholt J, Llimona X, Gadjieva R (2009) Hebeloma species associated with Cistus. Mycol Res 113:153–162. https://doi.org/10.1016/j.mycres.2008.09.007
Eberhardt U, Beker HJ, Vesterholt J, Dukik K, Walther G, Vila J, Fernández Brime S (2013) European species of Hebeloma section Theobromina. Fungal Divers 58:103–126. https://doi.org/10.1007/s13225-012-0188-3
Eberhardt U, Beker HJ, Vesterholt J (2015) Decrypting the Hebeloma crustuliniforme complex: European species of Hebeloma section Denudata subsection Denudata. Persoonia 35:101–147. https://doi.org/10.3767/003158515X687704
Eberhardt U, Beker HJ, Vesterholt J, Schütz N (2016) The taxonomy of the European species of Hebeloma section Denudata subsections Hiemalia, Echinospora subsect. nov. and Clepsydroida subsect. nov. and five new species. Fungal Biol 120:72–103. https://doi.org/10.1016/j.funbio.2015.09.014
Eberhardt U, Beker HJ, Schütz N, Mikami M, Kasuya T (2020a) Rooting Hebelomas: the Japanese ‘Hebeloma radicosum’ is a distinct species, Hebeloma sagarae sp. nov. (Hymenogastraceae, Agaricales). Phytotaxa 456 (2):125–144. https://doi.org/10.11646/phytotaxa.456.2.1
Eberhardt U, Beker HJ, Schütz N, Pedersen OS, Sysouphanthong P, Læssøe T (2020b) Adventurous cuisine in Laos: Hebeloma parvisporum, a new species in Hebeloma section Porphyrospora. Mycologia 112:172–184. https://doi.org/10.1080/00275514.2019.1680220
Eberhardt U, Schütz N, Beker HJ, Lee S, Horak E (2021a) Hebeloma in the Malay Peninsula: Masquerading within Psathyrella. MycoKeys 77:117–141. https://doi.org/10.3897/mycokeys.77.57394
Eberhardt U, Beker HJ, Borgen T, Knudsen H, Schütz N, Elborne SA (2021b) A survey of Hebeloma (Hymenogastraceae) in Greenland. MycoKeys 79:17–118. https://doi.org/10.3897/mycokeys.79.63363
Enderle M (1989) Bemerkenswerte Agaricales (Psathyrella)-Funde VIII. – Beitr Kenntn Pilze Mitteleurop 5:55–74
Frings RA, Maciá-Vicente JG, Buße S, Čmoková A, Kellner H, Hofrichter M, Hennicke F (2020) Multilocus phylogeny- and fruiting feature-assisted delimitation of European Cyclocybe aegerita from a new Asian species complex and related species. Mycol Prog 19:1001–1016. https://doi.org/10.1007/s11557-020-01599-z
Fukiharu T, Hongo T (1995) Ammonia fungi of Iriomote island in the southern Ryukyus, Japan, and a new ammonia fungus: Hebeloma luchuense. Mycoscience 36(4):425–430. https://doi.org/10.1007/BF02268627
GBIF.org (10 June 2021) GBIF Occurrence Download. https://doi.org/10.15468/dl.f75uja
Gonzalez P, Labarère J (1998) Sequence and secondary structure of the mitochondrial small-subunit rRNA V4, V6, and V9 domains reveal highly species-specific variations within the genus Agrocybe. Appl Environ Microbiol 64:4149–4160
Grilli E, Beker HJ, Eberhardt U, Schütz N, Leonardi M, Vizzini A (2016) Unexpected species diversity and contrasting evolutionary hypotheses in Hebeloma sections Sinapizantia and Velutipes in Europe. Mycol Prog 15(1):1–46. https://doi.org/10.1007/s11557-015-1148-6
Guindon S, Dufayard J-F, Lefort V, Anisimova M, Hordijk W, Gascuel O (2010) New algorithms and methods to estimate maximum-likelihood phylogenies: assessing the performance of PhyML 3.0. Syst Biol 59:307–321. https://doi.org/10.1093/sysbio/syq010
Hallen HE, Watling R, Adams GC (2003) Taxonomy and toxicity of Conocybe lactea and related species. Mycol Res 107:969–979
Hansen L, Knudsen H (1992) Nordic Macromycetes 2. Polyporales, Boletales, Agaricales, Russulales. Nordsvamp, Kopenhagen
Ho B-TQ, Pham N-DH, Shimizu K, Fukiharu T, Truong BN, Suzuki A (2014) The first record of Hebeloma vinosophyllum (Strophariaceae) in Southeast Asia. Mycotaxon 128:25–36. https://doi.org/10.5248/128.25
Hoang DT, Chernomor O, von Haeseler A, Minh BQ, Vinh LS (2018) UFBoot2: improving the ultrafast bootstrap approximation. Mol Biol Evol 35:518–522. https://doi.org/10.1093/molbev/msx281
Holec J, Kolařík M, Bizio E (2014) Pholiota chocenensis—a new European species of section Spumosae (Basidiomycota, Strophariaceae). Mycol Prog 13:399–406. https://doi.org/10.1007/s11557-013-0926-2
Holec J, Kolařík M, Borgarino D, Bidaud A, Moreau P-A (2016) Pholiota highlandensis var. citrinosquamulosa (Fungi, Agaricales) is conspecific with Pholiota gallica. Nova Hedwigia 103:251–263
Hongo T (1965) Notes on Japanese larger fungi (17). J Jap Bot 40(10):311–318
Hongo T (1966) Notes on Japanese larger fungi (18). J Jap Bot 41(6):165–172
Hughes KW, Matheny PB, Miller AN, Petersen RH, Iturriaga TM, Johnson KD, Methven AS, Raudabaugh DB, Bruns T (2020) Pyrophilous fungi detected after wildfires in the Great Smoky Mountains National Park expand known species ranges and biodiversity estimates. Mycologia 112:677–698. https://doi.org/10.1080/00275514.2020.1740381
Imai S (1938) Studies on the Agaricaceae of Hokkaido I. J Fac Agric Hokkaido Imperila Univ 43(2):1–378
Johnson M, Zaretskaya I, Raytselis Y, Merezhuk Y, McGinnis S, Madden TL (2008) NCBI BLAST: a better web interface. Nucleic Acids Res 36 (Web Server issue):W5-W9. https://doi.org/10.1093/nar/gkn201
Kalyaanamoorthy S, Minh BQ, Wong TKF, von Haeseler A, Jermiin LS (2017) Fast model selection for accurate phylogenetic estimates. Nat Meth 14:587–589. https://doi.org/10.1038/nmeth.4285
Katoh K, Standley DM (2013) MAFFT multiple sequence alignment software version 7: improvements in performance and usability. Mol Biol Evol 30:772–780. https://doi.org/10.1093/molbev/mst010
Katoh K, Kuma K, Toh H, Miyata T (2005) MAFFT version 5: improvement in accuracy of multiple sequence alignment. Nucleic Acids Res 33:511–518. https://doi.org/10.1093/nar/gki198
Katoh K, Rozewicki J, Yamada KD (2019) MAFFT online service: multiple sequence alignment, interactive sequence choice and visualization. Brief Bioinform 20(4):1160–1166. https://doi.org/10.1093/bib/bbx108
Katumoto K (2010) List of Fungi Reported in Japan. The Kanto Branch of the Mycological Society of Japan, Narashino
Kauff F, Lutzoni F (2002) Phylogeny of the Gyalectales and Ostropales (Ascomycota, Fungi): among and within order relationships based on nuclear ribosomal RNA small and large subunits. Mol Phylogenet Evol 25:138–156. https://doi.org/10.1016/S1055-7903(02)00214-2
Kawamura S (1954) Icones of Japanese Fungi, vol 5. Kazamashobo, Tokyo
Kõljalg U, Larsson K-H, Abarenkov K, Nilsson RH, Alexander IJ, Eberhardt U, Erland S, Høiland K, Kjøller R, Larsson E, Pennanen T, Sen R, Taylor AF, Tedersoo L, Vralstad T, Ursing BM (2005) UNITE: a database providing web-based methods for the molecular identification of ectomycorrhizal fungi. New Phytol 166:1063–1068
Küppers H (1999) Dumont’s Farbenatlas. Dumont, Köln
Larsson A (2014) AliView: a fast and lightweight alignment viewer and editor for large data sets. Bioinformatics 30:3276–3278. https://doi.org/10.1093/bioinformatics/btu531
Larsson E, Örstadius L (2008) Fourteen coprophilous species of Psathyrella identified in the Nordic countries using morphology and nuclear rDNA sequence data. Mycol Res 112:1165–1185. https://doi.org/10.1016/j.mycres.2008.04.003
Maire R (1913) Études mycologiques, Fascicule 1. Annales Mycologici 11:331–358
Malysheva EF, Kiyashko A (2011) Contribution to the study of Agrocybe pediades complex (Agaricales) in Russia based on nrITS sequences. Mycol Balc 8:115–124
Matheny PB, Bougher NL (2017) Fungi of Australia: Inocybaceae. Australian Biological Resources Study. CSIRO Publishing, Canberra
Matheny PB, Swenie RA, Miller AN, Petersen RH, Hughes KW (2018) Revision of pyrophilous taxa of Pholiota described from North America reveals four species—P. brunnescens, P. castanea, P. highlandensis, and P.molesta. Mycologia 110:997–1016. https://doi.org/10.1080/00275514.2018.1516960
Minh BQ, Nguyen MAT, von Haeseler A (2013) Ultrafast approximation for phylogenetic bootstrap. Mol Biol Evol 30:1188–1195. https://doi.org/10.1093/molbev/mst024
Minh BQ, Lanfear R, Trifinopoulos J, Schrempf D, Schmidt HA (2021) IQ-TREE version 2.1.2: Tutorials and Manual. Phylogenomic software by maximum likelihood. March 15, 2021. http://www.iqtree.org/doc/iqtree-doc.pdf
Miyamoto Y, Nakano T, Hattori M, Nara K (2014) The mid-domain effect in ectomycorrhizal fungi: range overlap along an elevation gradient on Mount Fuji, Japan. ISME J 8:1739–1746. https://doi.org/10.1038/ismej.2014.34
Moser M (1983) Die Röhrlinge und Blätterpilze. Kleine Kryptogamenflora II2/2. Springer, Stuttgart
Nara K (2006) Ectomycorrhizal networks and seedling establishment during early primary succession. New Phytol 169:169–178
Nauta MM (2005) Genus Agrocybe. In: Noordeloos ME, Kuyper TW, Vellinga EC (eds) Flora Agaricina Neerlandica, vol 6. Taylor and Francis, Boca Raton, pp 204–221
Nguyen L-T, Schmidt HA, von Haeseler A, Minh BQ (2015) IQ-TREE: a fast and effective stochastic algorithm for estimating maximum likelihood phylogenies. Mol Biol Evol 32:268–274. https://doi.org/10.1093/molbev/msu300
Nilsson RH, Larsson K-H, Taylor AFS, Bengtsson-Palme J, Jeppesen TS, Schigel D, Kennedy P, Picard K, Glöckner FO, Tedersoo L, Saar I, Kõljalg U, Abarenkov K (2019) The UNITE database for molecular identification of fungi: handling dark taxa and parallel taxonomic classifications. Nucleic Acids Res 47:D259–D264. https://doi.org/10.1093/nar/gky1022
Obase K, Tamai Y, Yajima T, Miyamoto T (2007) Mycorrhizal associations in woody plant species at the Mt. Usu volcano, Japan. Mycorrhiza 17:209–215. https://doi.org/10.1007/s00572-006-0097-y
Örstadius L (2001) Psathyrella spadicea – taxonomy and nomenclature. Windahlia 24:19–24
Örstadius L, Ryberg M, Larsson E (2015) Molecular phylogenetics and taxonomy in Psathyrellaceae (Agaricales) with focus on psathyrelloid species: introduction of three new genera and 18 new species. Mycol Prog 14(25):1–42. https://doi.org/10.1007/s11557-015-1047-x
Rambaut A (2006–2018) FigTree. Tree figure drawing tool version 14.4.4. Institute of Evolutionary Biology, University of Edinburgh. http://tree.bio.ed.ac.uk/software/figtree/. Accessed 20 Mar 2019
Rees B, Midgley D, Marchant A, Lerkins A, Orlovich D (2013) Morphological and molecular data for Australian Hebeloma species do not support the generic status of Anamika. Mycologia 105(4):1043–1058. https://doi.org/10.3852/12-404
Roy M, Yagame T, Yamato M, Iwase K, Heinz C, Faccio A, Bonfante P, Selosse M-A (2009) Ectomycorrhizal Inocybe species associate with the mycoheterotrophic orchid Epipogium aphyllum but not its asexual propagules. Ann Bot 104:595–610. https://doi.org/10.1093/aob/mcn269
Sagara N, Hongo T, Murakami Y, Hashimoto T, Nagamasu H, Fukiharu T, Asakawa Y (2000) Hebeloma radicosoides sp. nov., an agaric belonging to the chemoecological group ammonia fungi. Mycol Res 104:1017–1024. https://doi.org/10.1017/S0953756299002439
Schoch CL, Seifert KA, Huhndorf S, Robert V, Spouge JL, Levesque CA, Chen W, Fungal Barcoding Consortium (2012) Nuclear ribosomal internal transcribed spacer (ITS) region as a universal DNA barcode marker for Fungi. Proc Natl Acad Sci USA 109:6241–6246. https://doi.org/10.1073/pnas.1117018109
Slot JC, Hallstrom KN, Matheny PB, Hosaka K, Mueller G, Robertson DL, Hibbett DS (2010) Phylogenetic, structural and functional diversification of nitrate transporters in three ecologically diverse clades of mushroom-forming fungi. Fungal Ecol 3:160–177. https://doi.org/10.1016/j.funeco.2009.10.001
Smith AH (1972) The North American species of Psathyrella. Mem New York Bot Gard 24:1–633
Smith AH, Evenson VS, Mitchel DH (1983) The veiled species of Hebeloma in the Western United States. University of Michigan Press, Ann Arbor
Tedersoo L, Anslan S, Bahram M, Drenkhan R, Pritsch K, Buegger F, Padari A, Hagh-Doust N, Mikryukov V, Gohar D, Amiri R, Hiiesalu I, Lutter R, Rosenvald R, Rähn E, Adamson K, Drenkhan T, Tullus H, Jürimaa K, Sibul I, Otsing E, Põlme S, Metslaid M, Loit K, Agan A, Puusepp R, Varik I, Kõljalg U, Abarenkov K (2020) Regional-scale in-depth analysis of soil fungal diversity reveals strong pH and plant species effects in Northern Europe. Front Microbiol 11:1953. https://doi.org/10.3389/fmicb.2020.01953
Thorn RG, Reddy CA, Harris D, Paul EA (1996) Isolation of saprophytic basidiomycetes from soil. Appl Environ Microbiol 63:4288–4292
Tian E-J, Matheny PB (2021) A phylogenetic assessment of Pholiota and the new genus Pyrrhulomyces. Mycologia 113(1):146–167. https://doi.org/10.1080/00275514.2020.1816067
Trifinopoulos J, Nguyen LT, von Haeseler A, Minh B (2016) W-IQ-TREE: a fast online phylogenetic tool for maximum likelihood analysis. Nucleic Acids Res 44(W1):W232–W235. https://doi.org/10.1093/nar/gkw256
Vašutová M (2008) Taxonomic studies on Psathyrella sect. Spadiceae. Czech Mycol 60:137–171
Vašutová M, Antonín V, Urban A (2008) Phylogenetic studies in Psathyrella focusing on sections Pennatae and Spadiceae—new evidence for the paraphyly of the genus. Mycol Res 112:1153–1164. https://doi.org/10.1016/j.mycres.2008.04.005
Vesterholt J (2005) The Genus Hebeloma, vol 3. Fungi of Northern Europe. Svampetryk, Tilst
Vesterholt J, Eberhardt U, Beker HJ (2014) Epitypification of Hebeloma crustuliniforme. Myc Progr 13:553–562. https://doi.org/10.1007/s11557-013-0938-y
Vu D, Groenewald M, de Vries M, Gehrmann T, Stielow B, Eberhardt U, Al-Hatmi A, Groenewald JZ, Cardinali G, Houbraken J, Boekhout T, Crous PW, Robert V, Verkley GJM (2019) Large-scale generation and analysis of filamentous fungal DNA barcodes boosts coverage for kingdom fungi and reveals thresholds for fungal species and higher taxon delimitation. Stud Mycol 92:135–154. https://doi.org/10.1016/j.simyco.2018.05.001
Watling R, Bigelow HE (1983) Observations on the Bolbitaceae. Mycotaxon 17:377–397
Wu F, Zhou L-W, Yang Z-L, Bau T, Li T-H, Dai Y-C (2019) Resource diversity of Chinese macrofungi: edible, medicinal and poisonous species. Fungal Diversity 98:1–76. https://doi.org/10.1007/s13225-019-00432-7
Yangdol R, Sharma YP, Bhattacherjee S, Acharya K (2016) Molecular, physical and biochemical characterization of an edible mushroom, Psathyrella spadicea (P. Kumm.) Singer, from cold desert of Ladakh, India. Curr Res Environ Appl Mycol 6:334–343. https://doi.org/10.5943/cream/6/4/12
Acknowledgements
We are very much obliged to A. Bogaerts and P. Ballings of the Botanic Garden Meise (BR) for help with handling various loans from a variety of herbaria. We also thank the herbaria C, DUKE, MONT, TNS, and UWO for their help. We are indebted to E. Bizio, T. Borgen, E. Campo, C. Cripps, S.-A. Elborne, C. Hay, Mr or Ms Dawa, J. Horman, U. Kawasaki, H. Knudsen, D. Lewis, E. Grilli, E. Ludwig, T. Martin, Mr or Ms Matsumoto, H. Mori, Y. Muramatsu, M. Nabe, K. Nishibori, K. Osaku, S. Pizzardo, H. Shirayama, N. Siegel, S. Takehashi, Y. Taneyama, N.P. Thao, R. Vilgalys, J. Yamazaki, and D. Yoshioka for supplying us with interesting and exciting Hebeloma collections. This study was financed in part by JSPS KAKENHI Grants 24680085 and 20K06805 to KH and 15K16279 to TK. We thank the reviewers and the editor for their helpful comments and suggestions.
Funding
Open Access funding enabled and organized by Projekt DEAL. This research was funded in part by the last author and in part by JSPS KAKENHI Grants 24680085 and 20K06805 to KH and 15K16279 to TK.
Author information
Authors and Affiliations
Contributions
H.J.B., K.H., and T.K. obtained the material studied; all authors contributed to the identifications; U.E. and N.S. were responsible for the molecular analyses; H.J.B., K.H., and T.K. were responsible for the species descriptions. H.J.B. and U.E. prepared the first draft. All authors contributed to and approved of the manuscript.
Corresponding author
Ethics declarations
Conflict of interest
None.
Additional information
Publisher's note
Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Section editor: Zhu-Liang Yang
Rights and permissions
Open Access This article is licensed under a Creative Commons Attribution 4.0 International License, which permits use, sharing, adaptation, distribution and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons licence, and indicate if changes were made. The images or other third party material in this article are included in the article's Creative Commons licence, unless indicated otherwise in a credit line to the material. If material is not included in the article's Creative Commons licence and your intended use is not permitted by statutory regulation or exceeds the permitted use, you will need to obtain permission directly from the copyright holder. To view a copy of this licence, visit http://creativecommons.org/licenses/by/4.0/.
About this article
Cite this article
Eberhardt, U., Schütz, N., Bartlett, P. et al. Revisiting Hebeloma (Hymenogastraceae, Agaricales) in Japan: four species recombined into other genera but three new species discovered. Mycol Progress 21, 447–472 (2022). https://doi.org/10.1007/s11557-021-01757-x
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11557-021-01757-x