Abstract
A quantitative analysis of naturally-occurring regulatory networks, especially those present in mammalian cells, is difficult due to their high complexity. Much simpler gene networks can be engineered in model organisms and analyzed as isolated regulatory modules. Recently, several synthetic networks have been constructed in mammalian systems. However, most of these engineered mammalian networks have been characterized using steady-state population level measurements. Here, we use an integrated experimental-computational approach to analyze the dynamical response of a synthetic positive feedback network in individual mammalian cells. We observe a switch-like activation of the network with variable delay times in individual cells. In agreement with a stochastic model of the network, we find that increasing the strength of the positive feedback results in a decrease in the mean delay time and a more coherent activation of individual cells. Our results are important for gaining insight into biological processes which rely on positive feedback regulation.
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Positive feedback regulation plays an important role in many cellular signaling systems. Biological processes such as cell cycle regulation (Yao et al. 2008; Skotheim et al. 2008), apoptosis (Legewie et al. 2006), and vertebrate oocyte maturation (Ferrell and Machleder 1998) rely on positive feedback to generate a bistable switch in which cells transition from one distinct phenotype to another while residing in intermediate states only transiently. In these systems, cell-to-cell variability within a population of cells can result in a non-uniform response with individual cells switching between states at different times following induction (Rehm et al. 2002; Lai et al. 2004). A quantitative analysis of the dynamic behavior of positive feedback modules at the single-cell level may provide insight into how information flows through such motifs and can lead to a greater understanding of the switch-like responses that occur in many regulatory systems.
Characterizing the behavior of individual positive feedback modules in naturally occurring signaling pathways is difficult because these positive feedback loops are often embedded in extremely complicated networks. In contrast, simple engineered gene networks consisting of a single positive feedback loop can be analyzed as isolated regulatory modules. Several simple synthetic gene networks have been engineered and analyzed in model organisms such as E. coli and S. cerevisiae (Elowitz et al. 2002; Raser and O’Shea 2004; Volfson et al. 2006). The recent development of inducible mammalian transgene control systems has allowed for the construction of synthetic gene circuits in mammalian cells and organisms (Greber and Fussenegger 2007). Mammalian transgene control elements have been used to build synthetic mammalian networks such as toggle switches (Kramer et al. 2004), hysteretic switches (Kramer and Fussenegger 2005), and time-delay circuits (Weber et al. 2007). Most of these engineered mammalian networks have been characterized using steady-state population level measurements hindering an examination of the dynamic network behavior in individual cells (Longo and Hasty 2006). Obtaining quantitative single-cell measurements using fluorescence-activated cell sorting (FACS) is a valuable approach for analyzing gene expression levels in individual cells. In a recent study, the hysteretic response of a synthetic mammalian positive feedback network was monitored with single-cell FACS measurements allowing a bimodal response profile to be observed (May et al. 2008).
Some of the most widely used mammalian gene control systems are those which are regulated with tetracycline or the tetracycline analogue doxycycline (Gossen and Bujard 1992; Gossen et al. 1995; Rennel and Gerwins 2002). Here, we construct a synthetic mammalian positive feedback network using tetracycline-responsive control elements, and we utilize an integrated experimental-computational approach to analyze the dynamic response of the network at the single-cell level. Using a stochastic model of the network, we demonstrate that increasing the strength of the positive feedback results in a shorter mean delay time prior to activation and less variability in the activation time in individual cells. We confirm our theoretical predictions with quantitative single-cell measurements from a clonal population of mammalian cells harboring the synthetic circuit. Our findings may help us to predict the dynamic behavior of more complex cellular networks and may improve our ability to construct artificial gene networks that could be useful for gene therapy.
Materials and methods
Network construction
The GFP (green fluorescent protein) reporter vector, Hermes-HRSpuro-gfp (Rossi et al. 1998), was a gift from Helen Blau, Blau Laboratory, Stanford University, Stanford, CA. The autoregulatory vector was constructed from Hermes-HRSpuro-gfp by replacing the coding sequence for GFP with the coding sequence for the tet-responsive transactivator (rtTA). Oligonucleotide primers were used to PCR amplify the rtTA gene from the RTAb(+) vector (Rossi et al. 1998; a gift from Helen Blau, Blau Laboratory, Stanford University, Stanford, CA) and restriction endonucleases and T4 DNA ligase were used to insert the rtTA gene downstream of the O7-CMVm promoter. Plasmids were propagated in E. coli cells grown in LB and the antibiotic ampicillin. Cloning was confirmed by restriction digests visualized by gel electrophoresis, and the constructed vector was verified by utilizing the sequencing service provided by Eton Bioscience, Inc. The autoregulatory vector and the reporter vector were purified using Qiagen’s Plasmid Midi Kits.
Cell culture
NIH 3T3 cells were maintained in DMEM supplemented with 10% bovine calf serum, penicillin (100 units/ml), streptomycin (100 μg/ml), and l-glutamine (1%). Cells were grown in a 5% CO2, 37°C incubator.
Retroviral transductions
Retroviral constructs were cotransfected with pCL.Eco into 293T cells, and 42 h post-transfection filtered supernatant was used to infect NIH 3T3 cells. Low efficiency infections (infection rates less than 2%) were used to ensure single copy integration per cell. To generate cells harboring both vectors, cells were first infected with the autoregulatory vector and selected with puromycin hydrochloride (Sigma). Selected cells were subsequently superinfected with the reporter vector and GFP-positive cells were sorted by fluorescence-activated cell sorting (FACS).
Flow cytometry
Cells were washed in phosphate-buffered saline (PBS) and resuspended in PBS. Cells were sorted using a Becton–Dickinson FACSVantage SE flow cytometry system. Cells were analyzed with a Becton–Dickinson FACSCalibur flow cytometer with a 488-nm argon excitation laser and a 515–545-nm emission filter (FL1) at a high flow rate. Forward scatter values and fluorescence values were collected for at least 10,000 cells per sample. Data analysis was performed using MATLAB (the MathWorks, Inc.). A forward-scatter and side-scatter gate was used to minimize fluorescence variation due to cell size.
Modeling
Stochastic simulations were performed by implementing the Gillespie algorithm (Gillespie 1977) in C++.
Results
We have constructed a mammalian synthetic gene network that utilizes tetracycline (tet) responsive control elements. The synthetic network (Fig. 1a) consists of two vectors: (i) an autoregulatory vector which contains the coding sequence for a tet-responsive transactivator (rtTA) downstream of an O7-CMVm tet-regulatable promoter and (ii) a reporter vector which contains the gfp gene downstream of the O7-CMVm promoter. In the presence of doxycycline (dox), rtTA binds to tet operator sites on each O7-CMVm promoter thereby inducing its own expression and expression of the GFP reporter. The binding affinity of rtTA to the tet operator sites is determined by the concentration of dox. Thus, the strength of the positive feedback can be tuned by altering the concentration of dox.
Tetracycline-regulatable autoregulatory network. a Diagram of the synthetic positive feedback network. The O7-CMVm inducible promoter consists of seven tet operator sites (O7) fused to a minimal CMV promoter (CMVm). Vector 1 contains the rtTA gene downstream of the inducible promoter. Vector 2 contains the gfp gene downstream of the inducible promoter. In the presence of tetracycline, the rtTA dimers bind to the tet operator sites on the regulatable promoters and induce expression of rtTA and GFP. b GFP expression from a mammalian cell line harboring the synthetic network in the absence of dox (black) and in the presence of 1 μg/ml dox (green) for 24 h. c Normalized fluorescence as a function of dox. Experimental GFP expression levels were determined by FACS after cells from a clonal population were cultured in each dox concentration for 3 days. Error bars represent standard deviation of triplicate samples. Theoretical mean expression levels were determined from 20 runs of the simulation following induction at each dox concentration for 3 days. Experimental and theoretical data were individually normalized to conditions of no induction (0 μg/ml dox) and full induction (1 μg/ml dox). d Steady-state GFP expression levels versus dox concentration. A deterministic model (described in supplementary material) derived from the network reactions listed in Table 1 was used to determine stable steady-state GFP levels
Stable mammalian cells harboring the synthetic circuit were generated by transducing mouse embryonic fibroblasts (NIH 3T3 cells) with the two vectors. Clonal populations were produced by single-cell sorting the transduced cells and expanding individual cells into stable clonal cell lines. Cell lines were screened for inducibility by using flow cytometry to measure GFP expression levels from each clone after being cultivated in the absence and presence of dox (1 μg/ml for 24 h). GFP expression levels from a representative clone are shown in Fig. 1b. In all of the clones assayed, mean GFP expression increased with the addition of dox. We selected one clone for further quantitative analysis. The response of the clone to several dox concentrations was examined by culturing cells in dox concentrations ranging from 0 to 1 μg/ml for 3 days and measuring fluorescence levels using flow cytometry (Fig. 1c). Because mean fluorescence levels do not reach steady state values within 3 days of induction, the dose–response curve in Fig. 1c represents a transient response of the system. Simulated steady-state GFP expression levels as a function of dox (Fig. 1d) reveal that the system is bistable for a small range of dox concentrations.
While regulatory systems with positive feedback have the potential to exhibit bistable behavior, systems without feedback are monostable (Ferrell 2002; Xiong and Ferrell 2003; Mitrophanov and Groisman 2008). Simulated steady-state GFP levels as a function of dox for both the positive feedback system and for an analogous system without positive feedback (with constitutive rtTA expression instead of autoregulated rtTA expression) are shown in Fig. S2 (see supplementary material for model equations). The simulations show that the positive feedback results in a steep induction curve with the transition from low GFP expression levels to high GFP expression levels occurring over a much more narrow range of dox concentrations for the autoregulated system than for the no-feedback system. In contrast to the positive feedback system which exhibits hysteresis, the steady-state dose response curve for the no-feedback system does not depend on initial conditions. The simulations also reveal that the positive feedback system has significantly higher maximum GFP expression levels than the no-feedback system. Several key features produced by the simulations have been observed experimentally in a recent study which demonstrated that mammalian cells with autoregulated rtTA expression have a steeper induction curve and higher maximum expression levels than cells with constitutive rtTA expression (Markusic et al. 2005).
Stochastic modeling of the mammalian synthetic network
Cell-to-cell variability is present in any population of cells including genetically identical cell populations that have been exposed to the same environment (Elowitz et al. 2002; Raser and O’Shea 2005). Gene expression noise arising from the inherent randomness of biochemical processes such as transcription and translation can lead to cell-to-cell variation in the expression of a gene (Raj and van Oudenaarden 2008). To facilitate our exploration of the synthetic network’s dynamic behavior at the single-cell level, we used the Gillespie algorithm (Gillespie 1977) to perform stochastic simulations of the biochemical reactions involved in the positive feedback network. The model reactions include transcription, translation, mRNA degradation, and protein degradation of both the activator (rtTA) and the GFP reporter (Table 1). rtTA dimerization and operator binding are reflected in the transcriptional rates which are described by Hill-type functions. The association constant of the activator, K a, is determined by K a = S/(Ks + S) where S is the dox concentration and Ks is the dox concentration that results in half-maximal K a. The binding affinity of rtTA to the tet operator sites, and thus the strength of the positive feedback, is highest at high dox concentrations. Model parameter values were estimated using values in the literature where available, and remaining parameter values were chosen such that the model generates a dose–response curve that is in good agreement with experimentally observed dose–response behavior (Fig. 1c; Table S1). All stochastic simulations were equilibrated for a period of 50 h to allow variability in expression levels to stabilize prior to induction with dox. We used the coefficient of variation (CV, standard deviation divided by the mean) to quantify variability in GFP levels and found that the CV plateaus at approximately 0.35 (Fig. S3). We verified that the experimental level of cell-to-cell variability in the uninduced state is similar to that seen in our model by analyzing GFP expression levels determined by flow cytometry for cells cultured in the absence of dox. We found that the experimentally determined CV has a value of 0.43 and thus agrees well with the CV for the simulations. This level of cell-to-cell variability falls within the range in CV (0.1–0.6) that was recently reported for nearly 1,000 protein levels in human cells (Cohen et al. 2008).
Coherent activation with strong positive feedback
Using our stochastic model, we examined how the strength of the positive feedback affects the response of the network in individual cells with variable expression levels. Stochastic simulations were run to observe the dynamic behavior of the network following induction with 0.05, 0.1, and 1 μg/ml dox. Because these dox concentrations are all above the bistable regime (Fig. 1d), all trajectories will eventually reach a steady state with high GFP expression levels. At the lowest dox concentration, 0.05 μg/ml, the association constant K a is below half-maximal (K a = 0.38) resulting in weak positive feedback. At 0.1 μg/ml dox, K a is near half-maximal (K a = 0.56) resulting in intermediate strength positive feedback, and at 1 μg/ml dox, the strength of the positive feedback is near maximal with K a = 0.93. Our simulated results reveal that, at each of the three concentrations, there is a switch-like activation of the network with individual trajectories flipping to the high GFP steady state at different times following induction with dox (Fig. 2). At the lowest dox concentration (0.05 μg/ml), there is a mean delay of approximately 50 h before switching occurs (Fig. 2a). This mean delay time decreases as the positive feedback strength increases, with cells beginning to flip to the high expressing state within 10 h following induction at the highest dox concentration (Fig. 2c). The simulations also reveal that the timing of activation is extremely variable with low feedback strengths. In response to induction with 0.05 μg/ml dox, some trajectories start to switch to the high expression state as early as 50 h following induction while a significant fraction of the population is still expressing GFP at low basal levels at 200 h following induction (Fig. 2a). At 0.1 μg/ml, there is a smaller time window during which activation occurs ranging from approximately 20 to 100 h following induction (Fig. 2b). With strong positive feedback (1 μg/ml dox), there is a more coherent activation, with all trajectories beginning to switch to the high state within approximately 40 h (Fig. 2c).
To test our theoretical predictions, we employed flow cytometry to examine the temporal response of the network following induction with 0.05, 0.1, and 1 μg/ml dox. We measured the GFP expression levels of cells that were cultured in each dox concentration for 0, 10, 20, 30, 40, and 50 h. Following induction, fluorescence histograms show bimodality (Fig. 3a) thus confirming that there is a switch-like activation of the network. At each dox concentration, the percentage of cells with high expression levels increased over time. However, in agreement with our theoretical results, the dynamics of activation differed significantly for the three dox concentrations. Our measurements confirm our theoretical prediction that low feedback strengths result in a longer mean delay time prior to activation. The fluorescence distribution does not start to appear bimodal until 40 h following induction at the lowest feedback strength (0.05 μg/ml dox). In contrast, at 1 μg/ml dox, activation is much more rapid with a bimodal distribution appearing within 10 h following induction. Our experimental results also confirm that there is indeed less variability in the timing of activation in individual cells with strong positive feedback. This can be seen more clearly by examining the percentage of activated cells at each time point (Fig. 3b). At 1 μg/ml dox, about 90% of the cells are activated within 30 h. In contrast, at 0.05 and 0.1 μg/ml dox, there is a more gradual increase in the percentage of activated cells at each time point. The percentage of high expressing cells over time as determined by the model is also shown in Fig. 3b. We find that there is a good agreement between our simulations and our experimental data.
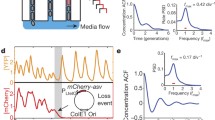
Time course of GFP expression following induction with different dox concentrations. a Fluorescence distributions from cells induced with 0.05, 0.1, or 1 μg/ml dox for 0, 10, 20, 30, 40, and 50 h. The relative frequency was obtained by normalizing the actual frequency by the frequency at the mode. b The percentage of cells in the activated state was determined from flow cytometry measurements. Error bars represent standard deviation of triplicate samples. Theoretical values were determined from 200 runs of the simulation at each concentration. c Five simulated trajectories are shown to demonstrate the lower and upper thresholds used for calculating τ1 and τ2. The length of τ1 and τ2 are shown for the blue trajectory. d Mean and standard deviation for τ1 and τ2 versus dox for 50 simulations. The inset graph shows the coefficient of variation versus dox concentration
Naturally-occurring regulatory networks with positive feedback
Our theoretical and experimental analysis of a simple mammalian positive feedback module has revealed that there is a variable delay time prior to the switch-like activation of the network, with the longest delay times and the least coherent activation occurring when the positive feedback strength is low. Similar signaling response characteristics have been observed in several naturally-occurring regulatory networks that contain positive feedback regulation. In S. cerevisiae, positive feedback in the G1/S regulatory network allows for a switch-like activation into cell cycle entry with a coordinated activation of individual cells, while a more discordant response is observed when positive feedback is completely removed from the network (Skotheim et al. 2008). Single-cell studies have shown that cell cycle entry in mammalian cells is also governed by a network which generates an all-or-none response via positive feedback regulation (Yao et al. 2008). However, quantifying the timing of cell cycle entry in single cells will be necessary to determine if there is also a coherent activation into cell cycle entry in mammalian cells.
In the signaling pathway that governs apoptosis, positive feedback is utilized to generate an all-or-none activation of the effector caspases which provoke cell death (Legewie et al. 2006), and the timing of effector caspase activation has been shown to vary considerably in individual mammalian cells (Rehm et al. 2002; Albeck et al. 2008). Although the delay time is variable and relatively long (several hours), once activated, effector caspases cleave their substrates and provoke cell death within minutes (Goldstein et al. 2000; Rehm et al. 2002). Thus, the response of effector caspases can be divided into two temporal components: a long and variable delay followed by a short and relatively robust period of activity. We next asked whether a complex regulatory architecture was necessary for producing this type of response, or if a simple positive feedback module could generate a similar response profile.
To address this question, we utilized our experimentally-validated stochastic model to examine the mean and standard deviation in the first and second portion of the response of our synthetic autoregulatory network. We define the first portion of the response (τ1) as the time period required for GFP expression levels to surpass basal expression levels (and thus exceed a lower threshold value), and we define the second portion of the response (τ2) as the time period required for GFP levels to increase from the lower threshold value to the maximum expression level (and thus exceed an upper threshold value) (Fig. 3c). The mean and standard deviation for τ1 and τ2 were determined from 50 simulations at several dox concentrations (Fig. 3d), and the coefficient of variation (CV) was calculated by dividing the standard deviation by the mean (Fig. 3d inset). We find that τ1 is highly dependent on the dox concentration (τ1 approaches infinity as the dox concentration approaches the bistable regime where some trajectories can remain in the low state indefinitely [Fig. 1d]). In contrast, (τ2) is rather insensitive to changes in dox. At dox concentrations below 0.1 μg/ml, τ1 has a higher mean and standard deviation than τ2. Thus, in this regime, there is a relatively long and variable delay period followed by a shorter, less variable period of activation. Therefore, the simple synthetic positive feedback network is in fact capable of generating a response profile that is similar to the apoptotic response.
We next investigated how the timing of activation (τ1) is affected by changes in the parameter values which determine the levels of the transactivator by performing a parameter sensitivity analysis. In our model, each of the parameter values involved in rtTA synthesis and degradation were increased and decreased by 5% (for a dox concentration of 1 μg/ml) and the mean value of τ1 was determined (Fig. 4a). We find that the timing of activation is most sensitive to changes in the basal transcription rate of rtTA (α A) and the fold-induction of transcription (f). With a 5% decrease in α A, the mean time of activation is increased by over 12 h (an increase of over 16%). Variability in the timing of activation (determined by the standard deviation in τ1) also depends strongly on the values of the transcriptional parameters α A and f (Fig. 4b). These results demonstrate that the timing of activation depends not only on the strength of the feedback but also depends strongly on the basal and induced transcription rates. In naturally-occurring regulatory networks such as the network governing apoptosis, precise regulation of these transcriptional rates may provide a mechanism for tuning the timing of activation.
Parameter sensitivity analysis. Each of the parameters involved in the production and degradation of rtTA mRNA and protein levels were increased and decreased by 5% (with S = 0.1 μg/ml). The mean (a) and standard deviation (b) in the time of activation (τ1) for 50 individual runs of the stochastic simulation is plotted
In many biological networks, it may be advantageous for cells to limit basal production of an autoregulatory protein to minimize energy expenditures. However, depending on the relevant timescale of the signaling program, it may be critical for all cells to be activated within a certain time period. Therefore, there may be a tradeoff between minimizing basal transcription levels and achieving a threshold level of coherency among the cells. In other signaling systems, such as those governing cell-fate determination (Wagers et al. 2002), the fraction of the cell population that switches states is highly regulated. Our sensitivity analysis of the synthetic positive feedback loop system reveals that the proportion of cells in the activated state can be regulated by manipulating the transcription rates. For example, stochastic simulations with several different basal rtTA transcription rates are shown in Fig. S4. At relatively high basal transcription rates (Fig. S4A, B), all trajectories are activated in a coherent manner. At lower transcription rates (Fig. S4C), a fraction of the trajectories remain inactivated during the 400 h time course. With the parameter values used to generate Fig. S4C, there is a single stable high GFP state, but the convergence of all trajectories to the steady state value is extremely slow. The low GFP state can be considered quasi-stable, because the trajectories may not switch out of this state within a biologically relevant time period (Mitrophanov and Groisman 2008).
Discussion
In this work we have utilized an integrated computational-experimental approach to analyze the dynamic response of a synthetic mammalian positive feedback network at the single-cell level. We have demonstrated that the switching-time characteristics of the synthetic network are highly dependent on the strength of the feedback with the shortest mean delay time and the most coherent response being produced by strong positive feedback.
The signaling properties of the simple synthetic positive feedback network are similar to signaling response characteristics that have been observed in several naturally-occurring, complex regulatory networks that contain positive feedback regulation. Using our experimentally-validated stochastic model we investigated whether or not our simple circuit could recapitulate the two temporal phases involved in caspase activation, and we found that the simple positive feedback module could in fact produce a similar temporal response profile. Using a parameter sensitivity analysis, we found that the timing of activation is highly sensitive to changes in the transcriptional rates of the transactivator. In naturally-occurring regulatory networks, this dependence on transcriptional levels may allow for precise regulation of the timing of activation.
In many biological signaling systems, including those regulating yeast polarization (Wedlich-Soldner et al. 2004), Xenopus oocyte maturation (Abrieu et al. 2001), and calcium signaling (Lewis 2001), multiple interconnected positive feedback loops are present. Building and characterizing an isolated synthetic positive feedback loop can facilitate the development of more complex synthetic regulatory networks, such as dual-positive-feedback networks, that closely resemble naturally-occurring signaling networks. Recent computational studies have suggested that interlinked slow and fast positive feedback loops can generate switches that are both rapidly inducible and resistant to noise (Brandman et al. 2005; Zhang et al. 2007). Analyzing the behavior of synthetic gene networks with interlinked positive feedback loops would be a valuable method for verifying the predicted performance advantages of interlinked dual-time feedback loops.
The approach used here to characterize the dynamic response of a simple engineered mammalian regulatory network can be utilized to provide greater insight into the signaling properties of the more complicated regulatory networks found in living cells and may also aid in the synthesis and analysis of more complex mammalian control systems.
References
Abrieu A, Dorée M, Fisher D (2001) The interplay between cyclin-B-Cdc2 kinase (MPF) and MAP kinase during maturation of oocytes. J Cell Sci 114:257–267
Albeck JG, Burke JM, Aldridge BB, Zhang M, Lauffenburger DA, Sorger PK (2008) Quantitative analysis of pathways controlling extrinsic apoptosis in single cells. Mol Cell 30:11–25. doi:10.1016/j.molcel.2008.02.012
Brandman O, Ferrell JE Jr, Li R, Meyer T (2005) Interlinked fast and slow positive feedback loops drive reliable cell decisions. Science 310:496–498. doi:10.1126/science.1113834
Cohen AA, Geva-Zatorsky N, Eden E, Frenkel-Morgenstern M, Issaeva I, Sigal A, Milo R, Cohen-Saidon C, Liron Y, Kam Z, Cohen L, Danon T, Perzov N, Alon U (2008) Dynamic proteomics of individual cancer cells in response to a drug. Science 332:1511–1516. doi:10.1126/science.1160165
Elowitz MB, Levine AJ, Siggia ED, Swain PS (2002) Stochastic gene expression in a single cell. Science 297:1183–1186. doi:10.1126/science.1070919
Ferrell JE Jr (2002) Self-perpetuating states in signal transduction: positive feedback, double-negative feedback and bistability. Curr Opin Cell Biol 14:140–148. doi:10.1016/S09550674(02)00314-9
Ferrell JE Jr, Machleder EM (1998) The biochemical basis of an all-or-none cell fate switch in Xenopus oocytes. Science 280:895–898. doi:10.1126/science.280.5365.895
Gillespie DT (1977) Exact stochastic simulation of coupled chemical reactions. J Phys Chem 81:2340–2361. doi:10.1021/j100540a008
Goldstein JC, Waterhouse NJ, Juin P, Evan GI, Green DR (2000) The coordinate release of cytochrome c during apoptosis is rapid, complete and kinetically invariant. Nat Cell Biol 2:156–162. doi:10.1038/35004029
Gossen M, Bujard H (1992) Tight control of gene expression in mammalian cells by tetracycline-responsive promoters. Proc Natl Acad Sci USA 89:5547–5551. doi:10.1073/pnas.89.12.5547
Gossen M, Freundlieb S, Bender G, Müller G, Hillen W, Bujard H (1995) Transcriptional activation by tetracyclines in mammalian cells. Science 268:1766–1769. doi:10.1126/science.7792603
Greber D, Fussenegger M (2007) Mammalian synthetic biology: engineering of sophisticated gene networks. J Biotechnol 130:329–345. doi:10.1016/j.jbiotec.2007.05.014
Kramer BP, Fussenegger M (2005) Hysteresis in a synthetic mammalian gene network. Proc Natl Acad Sci USA 102:9517–9522. doi:10.1073/pnas.0500345102
Kramer BP, Viretta AU, Daoud-El-Baba M, Aubel D, Weber W, Fussenegger M (2004) An engineered epigenetic transgene switch in mammalian cells. Nat Biotechnol 22:867–870. doi:10.1038/nbt980
Lai K, Robertson MJ, Schaffer DV (2004) The sonic hedgehog signaling system as a bistable genetic switch. Biophys J 86:2748–2757. doi:10.1016/S0006-3495(04)74328-3
Legewie S, Bluthgen N, Herzel H (2006) Mathematical modeling identifies inhibitors of apoptosis as mediators of positive feedback and bistability. PLoS Comput Biol 2:e120. doi:10.1371/journal.pcbi.0020120
Lewis RS (2001) Calcium signaling mechanisms in T lymphocytes. Annu Rev Immunol 19:497–521. doi:10.1146/annurev.immunol.19.1.497
Longo D, Hasty J (2006) Dynamics of single-cell gene expression. Mol Syst Biol 2:64. doi:10.1038/msb4100110
Markusic D, Oude-Elferink R, Das AT, Berkhout B, Seppen J (2005) Comparison of single regulated lentiviral vectors with rtTA expression driven by an autoregulatory loop or a constitutive promoter. Nucleic Acids Res 33:e63. doi:10.1093/nar/gni062
May T, Eccleston L, Herrmann S, Hauser H, Goncalves J, Wirth D (2008) Bimodal and hysteretic expression in mammalian cells from a synthetic gene circuit. PLoS ONE 3:e2372. doi:10.1371/journal.pone.0002372
Mitrophanov AY, Groisman EA (2008) Positive feedback in cellular control systems. Bioessays 30:542–555. doi:10.1002/bies.20769
Raj A, van Oudenaarden A (2008) Nature, nurture, or chance: stochastic gene expression and its consequences. Cell 135:216–226. doi:10.1016/j.cell.2008.09.050
Raser JM, O’Shea EK (2004) Control of stochasticity in eukaryotic gene expression. Science 304:1811–1814. doi:10.1126/science.1098641
Raser JM, O’Shea EK (2005) Noise in gene expression: origins, consequences, and control. Science 309:2010–2013. doi:10.1126/science.1105891
Rehm M, Dussmann H, Janicke RU, Tavare JM, Kogel D, Prehn JH (2002) Single-cell fluorescence resonance energy transfer analysis demonstrates that caspase activation during apoptosis is a rapid process. J Biol Chem 277:24506–24514. doi:10.1074/jbc.M110789200
Rennel E, Gerwins P (2002) How to make tetracycline-regulated transgene expression go on and off. Anal Biochem 309:79–84. doi:10.1016/S0003-2697(02)00250-6
Rossi FM, Guicherit OM, Spicher A, Kringstein AM, Fatyol K, Blakely BT, Blau HM (1998) Tetracycline-regulatable factors with distinct dimerization domains allow reversible growth inhibition by p16. Nat Genet 20:389–393. doi:10.1038/3871
Skotheim JM, Di Talia S, Siggia ED, Cross FR (2008) Positive feedback of G1 cyclins ensures coherent cell cycle entry. Nature 454:291–296. doi:10.1038/nature07118
Volfson D, Marciniak J, Blake WJ, Ostroff N, Tsimring LS, Hasty J (2006) Origins of extrinsic variability in eukaryotic gene expression. Nature 439:861–864. doi:10.1038/nature04281
Wagers AJ, Christensen JL, Weissman IL (2002) Cell fate determination from stem cells. Gene Ther 9:606–612. doi:10.1038/sj/gt/3301717
Weber W, Kramer BP, Fussenegger M (2007) A genetic time-delay circuitry in mammalian cells. Biotechnol Bioeng 98:894–902. doi:10.1002/bit.21463
Wedlich-Soldner R, Wai SC, Schmidt T, Li R (2004) Robust cell polarity is a dynamic state established by coupling transport and GTPase signaling. J Cell Biol 166:889–900. doi:10.1083/jcb.200405061
Xiong W, Ferrell JE Jr (2003) A positive-feedback-based bistable ‘memory module’ that governs a cell fate decision. Nature 426:460–465. doi:10.1038/nature02089
Yao G, Lee TJ, Mori S, Nevins JR, You L (2008) A bistable Rb-E2F switch underlies the restriction point. Nat Cell Biol 10:476–482. doi:10.1038/ncb1711
Zhang XP, Cheng Z, Liu F, Wang W (2007) Linking fast and slow positive feedback loops creates an optimal bistable switch in cell signaling. Phys Rev E 76:031924. doi:10.1103/PhysRevE.76.031924
Acknowledgements
We thank Dr. Matt Bennett and Dr. Will Mather for valuable discussions. This work was supported by a grant from the National Institutes of Health (GM69811-01).
Open Access
This article is distributed under the terms of the Creative Commons Attribution Noncommercial License which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
Author information
Authors and Affiliations
Corresponding author
Electronic supplementary material
Below is the link to the electronic supplementary material.
Rights and permissions
Open Access This is an open access article distributed under the terms of the Creative Commons Attribution Noncommercial License (https://creativecommons.org/licenses/by-nc/2.0), which permits any noncommercial use, distribution, and reproduction in any medium, provided the original author(s) and source are credited.
About this article
Cite this article
Longo, D.M., Hoffmann, A., Tsimring, L.S. et al. Coherent activation of a synthetic mammalian gene network. Syst Synth Biol 4, 15–23 (2010). https://doi.org/10.1007/s11693-009-9044-5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1007/s11693-009-9044-5