Abstract
CAS (Cell Annotation Software) is a novel tool for analysis of microscopic images and selection of the cell soma or nucleus, depending on the research objectives in medicine, biology, bioinformatics, etc. It replaces time-consuming and tiresome manual analysis of single images not only with automatic methods for object segmentation based on the Statistical Dominance Algorithm, but also semi-automatic tools for object selection within a marked region of interest. For each image, a broad set of object parameters is computed, including shape features and optical and topographic characteristics, thus giving additional insight into data. Our solution for cell detection and analysis has been verified by microscopic data and its application in the annotation of the lateral geniculate nucleus has been examined in a case study.
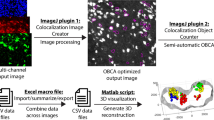
Similar content being viewed by others
Avoid common mistakes on your manuscript.
Introduction
Studying the principles of the structure of neuronal nets can bring us closer to unraveling the functioning of heterogenic populations of neurons within the central nervous system. The investigation of these nets involves consideration of the characteristics of neurons and their distribution within a particular nervous structure. In some cases, the distribution of neurons is enough to predict their characteristics. Often neurons form repetitive, regularly organized groups within which they share common physiological and neurochemical characteristics. For example, this phenomenon is pronounced within the visual cortex (Dyck & Cynader 1993; Kaas 2012; Ohki et al. 2005) and the spinal cord (Merkulyeva et al. 2016). However, in many other cases the structure of the object of interest is not so highly ordered; however, this does not mean that neurons are randomly distributed. A good example is a laminated visual thalamic structure, the lateral geniculate nucleus, LGN, which contains neurons differentiated by multiple features, such as input from ipsilateral or contralateral eye, the visuotopic position of retinal projections, or the type of projecting ganglion cell (Sanderson 1971; Payne & Peters 2002; Schiller 2010). In such cases, we expect these neuronal nets to have a complex structure; a full description of them can be obtained only by means of thorough analysis of cell parameters measured within the entire structure. Obviously, this leads to a significant increase in the effort needed to detect and analyze neuronal data, but this can be significantly facilitated with the use of specialized software.
The software applications designed for this purpose offer a wealth of opportunities for manual cell detection; however, in many cases only restricted opportunities for automatic cell detection are provided, usually based on brightness and color filters. Sometimes these techniques are suitable, but there are many cases in which biological objects have staining that is too difficult to be entrusted to automatic detection modes. Examples include a high level of noise, such as stained fibers and dendritic arbors, or high variability of the color intensity of objects of interest. Moreover, subsequent analysis is performed in statistical data processing software such as Statistica (StatSoft) and SPSS Statistics (IBM). These applications have a specific way of organizing data, but image processing software cannot export data to these programs; therefore, a lot of additional work is necessary for data preparation.
Cell Annotation Software, CASFootnote 1, was created in order to meet the needs arising from investigations of objects in histological specimens. ’Object’ is a general term we use throughout whole paper, but it should be understood as cell soma or cell nucleus, since either can be selected with CAS depending on the chosen parameters. The selection procedure ensures that any artifacts that are a result of tissue processing are considered. Comparative analysis of neuronal populations is also possible. Moreover, easy navigation between various levels of analysis and final data storage in files whose format is easily accessible to other software makes CAS a unique and fully functional program.
The program is designed to work with histologically and immunohistologically prepared specimens of neuronal tissue for which topological information about neural object placement is crucial. It allows different parts of stratified structures (e.g. laminae of neocortex, thalamic nuclei, spinal cord) to be compared and their intrinsic inhomogeneity to be examined. Definition of a region of interest of any shape is very practical and easy to apply. The semi-automatic detection of objects depends on only a few parameters; the rules of how to set these parameters are given in this work. Data unification in the X-axis allows for better comparison between several specimens; this is important in ontogenetic research, in which the absolute size of brain structures changes progressively.
Brightness normalization is important for quantitative comparison of the degree of histochemical or immunohistochemical staining of tissues. This makes it possible to analyze the amount of stained substance in objects indirectly, which in turn allows examination of the accumulation (or degradation) of a substance during maturation of the nervous system or under experimental conditions
Combining the aforementioned functionality of this tool makes CAS a powerful solution which facilitates histological structure analysis and removes the manual work which is normally necessary with other open-source software.
The basic discussion of selecting the most accurate image processing algorithms for object detection is presented in “Object Segmentation”. This draws attention to many problems which must be considered in order to choose a suitable method for data segmentation. Detecting objects requires automatic description of shape, size, and distribution. The implemented measures and their interpretations are given in “Shape Parameters”. Finally, Section “Case Study” presents an example of the application of CAS in research on the LGN. The “Conclusions” summarize the article.
Object Segmentation
Although it has already been addressed, the problem of correct segmentation between cell soma/nucleus and backgrounds in extensive groups of microscopic images does not yet seem to have been universally solved. This is interesting because the problem of object detection in biplanar images (in which there are only objects and a well-defined background) is widely discussed in the literature and there are many methods with various degrees of accuracy for solving it (Irshad et al. 2014).
The standard approach is based on a family of binarization methods and other more advanced algorithms dedicated to this problem. In most cases, researchers conduct object segmentation manually, exploiting the tool-set in ImageJ/Fiji (Papadopulos et al. 2007; Collins & et al. 2007; Schneider et al. 2012; Schindelin et al. 2012; Hartig 2001). Researchers, who specialize in image processing, have also created solutions dedicated to specific applications. Examples include the ImageJ plug-in (Forero et al. 2010; Pool et al. 2008), an ImageJ framework expansion (Gulyás et al. 2016), software for the Matlab environment (La Torre et al. 2013), and stand-alone applications such as CellProfiler (Kamentsky et al. 2011; Carpenter et al. 2006). Commercial solutions also exist, but their license usually precludes them from comparison. In many of the aforementioned solutions, extended H-minima is exploited for segmentation, while in our work the Statistical Dominance Algorithm SDA (Piorkowski 2016) is applied; this is novel in this field.
This section presents the common problems which should be considered when object segmentation is performed on microscopic images. The discussion starts with the choice of color space and its influence on the accuracy of the location of the border of the final object. The difficulties which must be overcome by binarization procedures are then described. Finally, a comparison of the standard procedures exploiting the extended H-minima method and the novel approach based on the SDA algorithm applied in the presented software is given.
Adjusting Color space
Data gathered by microscopic examination is stored as color images. Generally, the correct approach to further analysis of color data assumes the necessity of color-space normalization (Ing et al. 2016). However, in the case of microscopic nerve tissue data, which is the main objective of CAS development, normalization is not necessary due to the monochromatic nature of the input data, which holds most information in one or two channels of the three used for acquisition (usually in RGB format). Moreover, the differences between channels are not substantial. Previous attempts at selecting the most representative color channel of various tissues have shown that the best solution is to choose the channel with the highest variance (in RGB) for processing (Kolodziejczyk et al. 2014).
Standard approach to image binarization
The standard approach to object segmentation has already been established and is based on microscopic preparations, a commonly used solution that was introduced in neuro-science (Selinummi et al. 2005; Baecker 2010; Hartig 2001). This technique consists of two stages of image processing: background correction and application of one of a range of binarization methods.
Background correction aims to remove the very common phenomenon of uneven illumination. This can be achieved in various ways, for instance by high-pass filtering (which removes low frequencies). However, the rolling ball algorithm, which is the most widely applied, is based on background subtraction, as suggested by Sternberg (1983). This procedure subtracts average illuminance in a usually disc-shaped given neighborhood and prompts the operator to specify its dimensions. It is assumed that this size (the radius) should be a few times larger than the object’s area in order to avoid distorting objects while maintaining the effective removal of uneven illumination. A discussion on the influence of changing radius on the performance of the rolling ball algorithm was presented by Ekstrom et al. (2016).
Binarization is the second stage; it assigns 1s for pixels belonging to the object and 0s for the background. The easiest approach uses global thresholding, which defines one threshold value for the whole image. This is an efficient solution for a group of selected images; however, when larger variations in image content are considered or noise and distortions are present, more sophisticated approaches are necessary. In such cases, adaptive methods are useful, such as the essential method introduced by Otsu (1975). ImageJ/Fiji software (Hartig 2001) also implements other methods which can be exploited (e.g. Huang, Li, Max Entropy, Renyi Entropy, Shanbhag, Yen, etc.). The common advantage of all adaptive methods is the lack of necessity for the operator to set parameters, hence the human influence on data processing is removed. Still, these techniques may misinterpret areas of images that have different object density and background type than the rest of the image, or where images containing tissues have varying structures.
Extended H-minima Technique
Two of the basic methods of object segmentation in biplanar images are the H-minima technique and the extended regional version (extended H-minima) described by Soille (1999). The main idea assumes that objects are sought that differ in intensity from the background by least the value of H. This algorithm is often applied for cellular nuclei segmentation or detection of whole cells (Cheng et al. 2009; Jung & Kim 2010; Koyuncu et al. 2016; Gertych et al. 2016). In the case of LGN images, this method gives unstable results. Figure 1a presents an exemplary part of an image and the segmentation results achieved for consecutive H parameter values: Fig. 1b – d. One can see that different cells are present with increasing H. In the next steps, some cells disappear, while others appear or reappear; therefore, image segmentation is not consistent, which is not acceptable.
Statistical Dominance Algorithm
We suggest a novel approach for object segmentation based on the recently introduced Statistical Dominance Algorithm (SDA) Piorkowski (2016), which exploits statistics to specify image content. For each pixel in a monochromatic image, a number of neighboring pixels are defined such that the currently analyzed pixel is dominant over them (or inversely, neighboring pixels are dominant over the current pixel). As with H-minima, it is possible to define an additional threshold that must be reached to assume dominance. Thus, the output image contains statistics of point distribution in the input data; however, it is interesting that this method makes it possible to keep the original shapes. Because relations between points instead of illumination differences are computed, sensitivity to uneven illumination and structure variation diminishes. Moreover, this technique can derive indirect information about an object’s area.
The concept of the SDA algorithm is based on a statistical calculation of how many points dominate the central point of a neighborhood. The algorithm seems simple, especially if it is presented in the form of the code given in Listing 1, where:
-
imgin – input image,
-
imgout – output image,
-
SX, SY – width and height of input/output image,
-
R – radius of neighborhood,
-
N – size of neighborhood mask (also size of mirror margin used here), N = ⌈R⌉
-
T - the threshold to be checked (especially for noisy images); its value usually is positive.
The SDA algorithm for each pixel statistically determines its reference in the neighborhood. If a point dominates most of the surrounding points, it is qualified as a fragment of the sought object; otherwise, it is treated as part of the background. Adjusting the minimum level of brightness accordingly can eliminate background noise. Figure 2 shows an illustrative picture of a sample bitmap that assumes dark colors for objects (levels 0, 10, ...) and bright for background (levels 90, 80, ...). The fragment of the input image (see Fig. 2a) contains four different objects:
-
in the upper left: fragment of larger, uniform tissue (or aggregation of objects);
-
in the lower left corner: regular cell, with lighter luminance;
-
in the upper right corner: cell with dendrites, with a darker luminance;
-
in the bottom right corner: background luminance unevenness (levels 60–70–80).
Using the SDA algorithm, statistics were calculated for each point of the input image to produce an output image (see Fig. 2b). The following parameters were adopted: t h r e s h o l d = 50 and a disc-shaped mask with radius R = 3.75 (see Fig. 2c) that covers an area of 45 pixels (output range: 0–45). By analyzing the received statistics the following can be concluded:
-
objects of a small, round shape, regardless of real brightness, are clearly promoted if only the minimum threshold condition is met;
-
both cells have comparable statistical values despite the difference in brightness in the input image; the normalization of these values is shown in the real microscopic image (Fig. 3b);
-
dendrites have the highest statistical values;
-
uniform larger objects are suppressed and can be removed by simple binarization (Fig. 2d); this corresponds to the real case presented in Fig. 5.
The aforementioned features are quite well suited to biplanar images. In the case of additional interference, spots with a different average brightness than the background (Fig. 4) can be removed using adaptive binarization, e.g. Otsu.
The above considerations show that the characteristics of the SDA algorithm justify its use in the analysis of nerve cells. Preliminary studies have shown the utility of segmentation algorithms in other tissues and histopathological images for which classical methods such as Otsu or adaptive thresholding do not provide accurate solutions (Kowal et al. 2013).
As described, it is necessary to set two parameters: the size of the neighborhood, specified by radius R (which defines a circular neighborhood), and minimal difference threshold T, which is analogous to the binarization threshold H exploited in H-minima. In the case of the neighborhood radius, the procedure is similar to the one used in background subtraction: usually a radius two or three times longer than the object is sufficient and testing for larger values does not change the outcome (this is presented more precisely in “Comparison of Object Detection Methods”). The selection of thresholding value, as in H-minima, also demands operator interaction. However, some datasets containing large groups of images give similar, stable results for the same parameter settings (a group of images accessible at BrainMaps (2005) was used for verification). The novel approach implemented in CAS assumes exploitation of the SDA algorithm in the background subtraction stage, followed by manual or automatic application of global or adaptive binarization. The image obtained from SDA should also be binarized, which can be done automatically (in the case of CAS, the Otsu method is suggested as it can remove some types of disturbance; refer to Fig. 4), or the global threshold can be set manually. The presented research focuses on object detection with the application of SDA, whose compatibility with other techniques has been proved.
The main advantage of SDA compared to other techniques is the representation of objects not as a binary map, but as an estimation of the probability that each pixel belongs to the object. This is well visualized in Fig. 5.
Example of SDA (c,d) over performance on standard segmentation methods (b) for cell detection. Image is taken from work (Mikula et al. 2007) and presents the sagittal section of the molecular layer, Nissl stained in cerebellum brain of the cat characterised by uneven nuclei distribution
This image is too difficult for the standard object segmentation method, but was successfully segmented by applying SDA. The complex input data (see Fig. 5a) shows sagittal sections of cerebellum cut on a Leica microtome and stained with cresyl violet. The greatest challenge here is the presence of different tissues in the image. The standard approach based on binarization does not correctly segment the cells from the (darker) tissue – the granular layer – regardless of whether the manual or automatic global thresholding method implemented in ImageJ/Fiji is used. An example of the outcome is depicted in Fig. 5b. Since the SDA’s output gives greyscale values (see light-grey and dark-grey objects in Fig. 5c), it was possible to continue with further segmentation and select an appropriate threshold in order to obtain accurate cell detection, as annotated on the original image in Fig. 5c. This feature proves the attractiveness of the SDA method for semi-automatic segmentation of cells in nerve tissues (Kopacz & Piorkowski 2017). The presented solution does not perform automatic segmentation of clustered nuclei; although this problem is still unresolved, some approaches are addressed by (Koyuncu et al. 2016; Jung & Kim 2010; Cheng et al. 2009; Gertych et al. 2012; Skobel et al. 2017).
Comparison of Object Detection Methods
In order to choose a method for further processing, the agreement in object detection in biplanar microscopic images was evaluated. Good quality resolution of objects and backgrounds was assured in all examples. For the standard approach, the global threshold was selected manually to assure correct segmentation. The radius applied in background subtraction methods and SDA was set to 50 pixels.
The goal of the first experiment was to evaluate the similarity of object segmentation masks achieved for various threshold values applied for standard and SDA methods. A similar problem was presented by (Ekstrom et al. 2016), but with lower precision. Here, the tests were performed on images presenting the LGN and perigeniculate nucleus PGN of 123-day-old cats acquired with a Leica MDI6000 inverted fluorescence microscope (Fig. 6a shows an example.). The coincidence of object detection was evaluated using the Dice coefficient (Dice 1945), which is expressed by following formula:
where TP stands for true positives (count of pixels belonging to objects), FP describes false positives (count of pixels wrongly assigned to the object), and FN is a false negative (count of object pixels which were wrongly assigned to the background). In medical research, coefficient values above 0.9 are assumed to have good correspondence. Figure 6b presents a 3D plot in which varying threshold values are given on the X and Y axes, while the Dice coefficient score change is presented by the surface. It can be observed that for several threshold combinations, the coefficient value reaches and exceeds 0.8–0.9; the maximum value of 0.99 clearly shows that both methods can be applied interchangeably.
We then concentrated on the relation between SDA parameters applied for images of objects of varying sizes. The influence of varying radius on detected object area is presented in Fig. 6d for the image presented in Fig. 6c, which was taken from a public dataset prepared for the Nucleus Counting Challenge competition that took place at the 2015 BioImage Informatics Conference. From this plot, one can see that the outcome stabilizes for radii above 20 pixels; the bigger the detected threshold value, the smaller the influence of data size. Of course, when the image is resized, the radius should be scaled as well. Moreover, the radius is always related to the object size, but the relation to the threshold value is nevertheless maintained.
The suggested methodology for object segmentation has been also tested on various tissues taken from an interactive multi-resolution brain atlas (BrainMaps 2005; Mikula et al. 2007). This database provided us with tissues from various species prepared using different types of staining; this gave a good overview of the possible problems that should be addressed in the developed tool. Using CAS, it was possible to perform segmentation quickly (about 2 minutes per slice), as presented in Fig. 7, while the parameters set in the program are gathered in Table 1. As one can see, the program enables accurate segmentation and labeling of objects regardless of their shape and the input data type.
Examples of object segmentaton with CAS on images presenting various tissues with diverse stainig. The exemplary data was gathered from BrainMaps (2005). The description contains – spieces: stain, plane, methodx
Discussion
The presented experiments and considerations address all the problems encountered in achieving accurate object segmentation in microscopic images of various tissues. From the elaborated results, it was decided that the CAS tool should interpret input images using only one data channel. The SDA was suggested as a segmentation method; however, to maintain compatibility with other research tools, the extended H-minima is also accessible. Finally, setting method parameters demands knowledge about object size, which for method radius is straightforward, while the SDA threshold should be adjusted and the final binarisation threshold can be chosen automatically using the Otsu approach.
Shape Parameters
It is not sufficient to develop techniques for automatic detection of objects in images: in most tasks, especially when general insight into object population is sought, this is the starting point for further analysis which demands thorough inspection of object features. Depending on the requirements, the desired characteristics vary, yet in general they concentrate on basic shape features which allow analysis of not only circularity or elongation, but also spatial and size distribution within a sample. This area of research is very tiresome, therefore automation is of great importance. Below, several shape parameters implemented in CAS are detailed and an interpretation is given with some examples.
General Features
The area and perimeter of an object are computed in order to obtain general information about the detected cell soma or cell nucleus (for complex shapes a fractal area could be considered Mazurek and Oszutowska-Mazurek 2014). Initially, the number of pixels belonging to the object is treated as the area value; similarly, pixels on the object boundary form the perimeter. However, for convenience it is also possible to compute these features in real units if this information was stored during data collection. Figure 8 presents an exemplary cell with the perimeter denoted by a white line, while Table 2 contains general features. The next feature – a centroid – holds information about object placement in the entire image. However, when necessary, it is possible in CAS to define start (x = 0%) and end (x = 100%) reference points and recalculate the x centroid coefficient for all objects.
Other global information about an object is provided by its intensity. The average brightness is defined for image I as the average intensity of all pixels p that belong to the object:
Since the intensities may vary depending on the data sampling procedure, this measure is very subjective when comparison between samples is considered. Therefore, a reference brightness was defined for which a reference intensity RI denoted by the user is exploited in order to obtain objective object brightness information:
Circularity Measures
An object’s shape is described by topological attributes (Russ 1998; Gonzalez & Woods 2001; Jahne 2002), of which only those not affected by translation and rotation were chosen in this research. These have found applications in research, where shape plays an important role (Nurzynska et al. 2013; Nurzynska & Piorkowski 2016; Piorkowski et al. 2017). The formulation of many attributes exploits information about area A and perimeter P to define a circularity measure. Here are some examples implemented in the described software:
where A M A X is a maximal area calculated as a multiplication of the maximal width and height of the object, and M A X I S is the length of the major axis of an object.
Table 3 presents examples of various shapes and their calculated features. The objects are ordered by ascending area size to better check the influence of scale on the result. It can be observed that in case of the C 1 parameter, a value equal to 1 corresponds to a circular shape, while a smaller value corresponds to a more elongated shape. In contrast, smaller values of C 2 and C 3 better describe round shapes, while bigger values are obtained for elongated ones.
It is difficult to find a pattern in which C 4, C 5, and Roughness are considered. The results gathered for C 4 seem completely unrelated to the shape, as one can see that using only the circumference for description is insufficient. A similar observation is also seen for the Roughness parameter, for which only area was exploited in the calculation. However, for C 5, the feature value is still strongly related to the object size, yet smaller values represent more circular shapes when objects with comparable area are considered (for instance, check the outcomes in rows 4–6, 7 with 8, etc.).
Next, when the Malinowska parameter is taken into account, values close to 0 describe circular objects, while higher values are related to longer shapes; this is also true of the Shapeless parameter. Moreover, we can additionally state that this works inversely for C 1 when the formulas are compared. Finally, Roundness is equal to 1 for circular objects and gets smaller for longer shapes.
Other
The last group of parameters that describe objects should help to organize this data and obtain more spatial information about it. First of all, it is possible to mark a region (bigger patch) on the image and work only with the objects belonging to it. Such a region can be automatically divided into three perpendicular horizontal layers. Next, for each object the major and minor axes are found. Here, two algorithms are exploited:
- Ellipse Fit:
-
– chooses the best fitted ellipse that can be inscribed in the object. The major and minor axes correspond to those of the ellipse. Moreover, the centroid can be recalculated to overlap the ellipse center point.
- Feret:
-
– a shape feature parameter that computes the relation between the two longest and perpendicular dimensions of an object. Principal component analysis (PCA) is exploited to gather the results. The major axis corresponds to the direction of the first eigenvector; the minor axis is related to the second one.
In both cases, the reference angle is computed as the angle between the object’s major axis and the image’s x-axis. This is used to position objects better in the input data.
Case Study
Data Processing Example
Comprehensive investigation of cell populations is especially needed in sophisticated structures in which characteristics of neurons are dependent upon their location. A good example is the LGN of cats; this is the visual thalamic nucleus, which provides the main transmission of information from the retina to the primary visual cortex. The LGN has a 6-layered structure in which the principal ones, A, A1, and C M receive input from different eyes: the ipsilateral side (A1) and the contralateral side (A, C M) (Payne and Peters, 2002, pp. 5–9). Moreover, the retinal afferents from different parts of the visual space project to the LGN, thus forming a highly ordered map—the so-called visuotopic map (Sanderson 1971)—which is depicted in Fig. 9. There is a lot of information in the literature about the non-uniform distribution of neurons in LGN layers according to their functional features (Bowling & Wieniawa-Narkiewicz 1986; LeVay 1977; Mitzdorf & Singer 1977). For example, particular types of LGN relay cells—so-called Y cells (Burnat et al. 2002; Kaplan 2014; Schiller 2010)—tend to be located close to the interlaminar borders of layers A and A1 (Bickford et al. 1998; Bowling & Wieniawa-Narkiewicz 1986; Mitzdorf & Singer 1977); in general, they tend to be located in the representation of the visual field periphery (Hoffmann et al. 1972; LeVay 1977). Herein, we used LGN slices labeled with a specific Y cell marker for demonstration of CAS: SMI-32 antibodies (labeling nonphosphorylated heavy-chain neurofilament proteins) (Bickford et al. 1998; Carden et al. 2000). These antibodies visualize the cell soma and proximal dendrites of immunopositive cells. They are complex to detect because the cell soma is often labeled unevenly and the unvisualized nucleus disturbs the outline of the soma.
The scheme of the frontal slice of the LGN with aligned visuotopic coordinates (Sanderson 1971). A, A1, C M layers of the LGN. The visuotopic coordinates assigned in degrees are on the top of the figure
Histology and Image Acquisition
All experimental procedures were approved by the Ethics Commission of the Pavlov Institute of Physiology. Experiments were performed in strong accordance with the requirements of Council Directive (2010/63/EU) of the European Parliament on the protection of animals used for experimental and other scientific purposes.
All presented research exploits images of the LGN of 123-day-old cats. The animals were perfused transcardially with 0.9% NaCl (0.6L) and 4% paraformaldehyde (1.0L, 0.1M PBS at pH = 7.4) under deep anesthesia (zoletil and xylazine mixture (1:3)). After perfusion, the brain was stored in 20 and 30% sucrose until it sank. The visual thalamus was cut on a freezing microtome into 50 μ m frontal sections. Sections were collected in 0.1M PBS, pH = 7.4. Immunohistochemical staining was performed with Vectastain reagents (Vector Laboratories, Peterborough, United Kingdom) in accordance with the manufacturer’s recommendations. One of three primary antibodies was applied for antigen labeling:
-
monoclonal mouse primary antibodies to NeuN (Millipore, MAB377, in 1:5000 dilution);
-
monoclonal mouse primary antibodies to non-phosphorylated Neurofilament H (NF-H) (Covance, SMI-32, in 1:3000 dilution);
-
polyclonal rabbit primary antibodies to the parvalbumin (Abcam, ab11427, in 1:10000 dilution).
The primary antibodies were labeled with:
-
Biotinylated secondary antibodies (goat anti-rabbit IgG, BA-1000, or horse anti-mouse IgG, BA-9200; Vector Laboratories, Peterborough, United Kingdom) for further DAB/Ni staining protocol for light microscopy.
-
Fluorochrome-conjugated secondary antibodies (Alexa Fluor488 goat anti-mouse IgG or Alexa Fluor568 goat anti-rabbit IgG, Thermo Fisher Scientific, Waltham, MA, USA) for cell detection with a fluorescence microscope.
DAB-reacted slices were analyzed with an Olympus microscope (Olympus Corporation, Tokyo, Japan, a 10 × magnification lens) using a Nikon photo camera (Nikon Corporation, Tokyo, Japan). Fluorescent slices were analyzed with a Leica DMI6000 inverted fluorescence microscope (at the Center for Molecular and Cell Technologies, Research Park, St. Petersburg State University).
The image data preparation procedure:
-
1.
Mark the layer borders.
-
2.
Mark the sub-layer parts of layers.
-
3.
Outline all labelled cells with appropriate parameters one-by-one.
-
4.
Measure data and copy the results to an external file.
-
5.
Complete the measurement file with additional information about the layer, sublayer, cell location, etc.
-
6.
Perform cell distribution normalization on the X axis.
-
7.
Perform cell brightness normalization using a reference region.
Table 4 compares how the analyses of input data were performed before and after the dedicated software was implemented. Actually, in CAS the order of steps is slightly different due to its automatic character. Firstly, parameters necessary for correct cell soma segmentation are sought, followed by manual annotation of layers A, A1 and C M (see Fig. 10b). Next, the automatic procedures compute the three sub-layers and detect cells (see Fig. 10c). Manual correction is sometimes necessary when several cells are detected as one. In such cases, the software supports division of the object and deletion of unsuitably selected objects and artifacts. It is worth noting that at the end of the selection, the cutting of dendrites can be performed in CAS; therefore, the soma is selected in all cells using uniform parameters. Although, there are solutions which may need to preserve dendrites for visualization of the LGN network (Morgan et al. 2016). In the case of manual outlining, the cell soma was detected by an operator for each cell in the delineation process. The automatic cell detection procedure, manual correction, and dendrite cutting are shown in Fig. 11. Next, the reference brightness region and points for spatial normalization are selected. Finally, all prepared data can be saved for further processing. The summary table contains the number and location marks of each cell soma, chosen measurement parameters, and additional normalized data. In addition, the current status of the processed image (with region borders and chosen objects) can be saved in a CSV file for subsequent correction of measurements. Also, the processed image can be saved in two additional forms. The first form has labeled cells with their serial numbers (similar to those in the text file) (Fig. 10a) and is used for an experimental protocol. The second, depicted in Fig. 10d, has bordered regions of interest and color dots in the place of labeled cells, which significantly facilitates the primary assessment of cell distribution and subsequent data presentation.
SMI-32 antibodies are just one of many markers used to study the LGN and other neuronal structures. To demonstrate the abilities of the software, we would like to additionally show the detection of small inhibitory interneurons labeled with antibodies against parvalbumin (Stichel et al. 1988) and all neurons within the LGN labeled with NeuN antibodies (Mullen et al. 1992). We examined cell detection in images obtained with light and fluorescent microscopes. The original images are shown in Figs. 12a,c and 13a,c; the results of detection are shown in Figs. 12b,d and 13b,d. The same quality of detection could be achieved in slices prepared for light as for fluorescent microscopy. The parvalbumin positive cells (see Fig. 12) were easy to detect and manual correction was almost not needed. In the case of NeuN-positive cells (Fig. 13), the difficulties of cell detection can be seen. The cell borders are detected correctly, but their density does not allow each cell to be recognized separately. This was due to the high neuronal density in the examined nucleus and the use of 50 μ m thick slices containing several layers of cells that overlap each other in the resulting image. This was overcome with manual correction tools and the program can still be used to measure restricted sample areas in structures of interest.
The examples of cell detection with CAS. The samples of tissue with NeuN positive cells acquired under light (a, c, e) and fluorescent (b, d, f) microscopes. The original images are on the top and the processed ones on the bottom. The middle shows the operation applied for object detection: red – automatic cell detection; cyan – cell division line; yellow – removed objects (when placed next to red line, the division line (cyan) was omitted for clarity); green – manually drawn elements
The application of CAS in research on the distribution of neuronal populations in the LGN considerably reduced the workload needed in data analysis. Two of the most time-consuming procedures— outlining cell borders and primary data preparation (adding of slice/region marks in the result table, spatial and brightness normalization)—are done automatically, thus shortening the analysis. In order to obtain some measurable data, 10 samples of spinal cord, LGN, and cortex were prepared. In the experimental group, various types of immunohistochemical staining were applied: parvalbumin, calretinin, NeuN, and SMI-32. The number of cells varied from 84 to 210 (with average equal to 142.9). Manual outlining (depending on complexity) took from 540 to 1380 seconds (average 873), while CAS needed from 10 to 30 seconds (average 19 seconds) for automatic cell segmentation. The computation was performed on a personal computer running Windows 10 with a 3.2GHz Intel CPU and 16GB of RAM. Additional manual cell improvement was applied to some slices: the operator checked the correctness of automatic selection and removal or cutting of some cells, which took from 180 to 600 seconds (average 378). In the results, we can see that 55 times speed-up was achieved for automatic cell selection, which is the most tiresome work. Of course, to improve the result, some additional semi-automatic action was necessary, which resulted in achieving a final speed-up of 2.29 times. Figure 14 presents an example of manual and automatic location of cell borders while Table 5 summarises the performance.
In addition, the software has convenient instruments for data presentation and re-examination. It is also worth pointing out that if changes are made (e.g. removing or adding cells, etc.), the results are recomputed automatically without operator intervention, which makes corrections and changes easy to apply. As a result, we have a convenient tool to detect specific changes of the LGN under experimental conditions or as a consequence of age-related development.
Conclusions
This work addresses the problem of automatic cell soma or nucleus segmentation and annotation in microscopic images. A detailed comparison of the standard image processing method against a novel approach exploiting SDA was presented. The research shows that the method used in CAS enables object border detection with comparable quality to standard methods. In difficult cases in which there are more types of object to be recognized, it outperforms the current leading techniques. In addition, shape features were introduced for description of object characteristics and specific analysis and interpretation were performed. Additional features implemented in CAS which enable data scale changes, data division into layers, and other functionality were presented with a case study of semi-automatic annotation of the LGN of cats.
Cell Annotation Software was designed to meet the need for a system which combines the various functionalities necessary for segmentation and analysis of objects in neural specimens. The presented approach proved to fulfill all demands and allows: saving topological information about cell distribution by region definition; comparative analysis of different slices and investigation of changes in the neuronal population due to x-axis normalization; and quantitative comparison of the degree of histological or immunohistological staining of tissues in slices when a change in the production quantity of a particular substance due to the influence of experimental factors is investigated. Moreover, simple recalculation of results and storage in open-format files make this software easy to incorporate in research pipelines.
The practical application of CAS in research conducted at the Pavlov Institute of Physiology proved to be a great culmination of this research, but a new development frontier has also arisen. In future work, we plan to address the problem of multiple labeling, working with tissues with various cell types and 3D reconstruction.
Information Sharing Statement
In the article we worked with data from the following data sources:
-
1.
Dataset provided for the Nucleus Counting Challenge (Fig. 3, 6c) (Maric 2015).
-
2.
BrainMaps: An Interactive Multiresolution Brain Atlas (BrainMaps 2005) decribed by Mikula et al. (2007) (Figs. 5 and 7)
-
3.
Data acquired by A. Mikhalkin (other figures).
The presented Cell Annotation Software can be downloaded from http://home.agh.edu.pl/pioro/cas/.
References
Baecker, V. (2010). Workshop: Image processing and analysis with ImageJ and MRI Cell Image Analyzer.
Bickford, M. E., Guido, W., & Godwin, D. W. (1998). Neurofilament proteins in Y-cells of the cat lateral geniculate nucleus: Normal expression and alteration with visual deprivation. The Journal of neuroscience, 18(16), 6549–57.
Bowling, D. B., & Wieniawa-Narkiewicz, E. (1986). The distribution of on- and off-centre X- and Y-like cells in the A layers of the cat’s lateral geniculate nucleus. The Journal of Physiology, 375, 561–72.
BrainMaps (2005). BrainMaps: An Interactive Multiresolution Brain Atlas. http://brainmaps.org.
Burnat, K., Vandenbussche, E., & Ernicki, B. (2002). Global motion detection is impaired in cats deprived early of pattern vision. Behavioural Brain Research, 134(1-2), 59–65.
Carden, W.B., Guido, W., Ziburkus, J., Datskovskaia, A., Godwin, D.W., & Bickford, M.E. (2000). A novel means of Y cell identification in the developing lateral geniculate nucleus of the cat. Neuroscience Letters, 295, 5–8.
Carpenter, A. E., Jones, T. R., Lamprecht, M. R., Clarke, C., Kang, I. H., Friman, O., Guertin, D. A., Chang, J. H., Lindquist, R. A., Moffat, J., et al. (2006). Cellprofiler: image analysis software for identifying and quantifying cell phenotypes. Genome Biology, 7(10), R100.
Cheng, J., Rajapakse, J. C., & et al. (2009). Segmentation of clustered nuclei with shape markers and marking function. IEEE Transactions on Biomedical Engineering, 56(3), 741–748.
Collins, T. J., & et al. (2007). Imagej for microscopy. Biotechniques, 43(1 Suppl), 25–30.
Dice, L. R. (1945). Measures of the amount of ecologic association between species. Ecology, 26(3), 297–302.
Dyck, R. H., & Cynader, M. S. (1993). An interdigitated columnar mosaic of cytochrome oxidase, zinc, and neurotransmitter-related molecules in cat and monkey visual cortex. Proceedings of the National Academy of Sciences of the United States of America, 90(19), 9066–9069.
Ekstrom, A., Suvanto, R. W., Yang, T., Ye, B., & Zhou, J. (2016). Robust neuron counting based on fusion of shape map and multi-cue learning. In International conference on brain and health informatics (pp. 313). Springer.
Forero, M. G., Pennack, J. A., & Hidalgo, A. (2010). Deadeasy neurons: automatic counting of hb9 neuronal nuclei in drosophila. Cytometry Part A, 77(4), 371–378.
Gertych, A., Joseph, A. O., Walts, A. E., & Bose, S. (2012). Automated detection of dual p16/ki67 nuclear immunoreactivity in liquid-based pap tests for improved cervical cancer risk stratification. Annals of Biomedical Engineering, 40(5), 1192–1204.
Gertych, A., Ma, Z., Tajbakhsh, J., Velásquez-Vacca, A, & Knudsen, B. S. (2016). Rapid 3-d delineation of cell nuclei for high-content screening platforms. Computers in Biology and Medicine, 69, 328–338.
Gonzalez, R. C., & Woods, R. E. (2001). Digital image processing, 2nd Edn. MA, USA: Addison-Wesley Longman Publishing Co., Inc.
Gulyás, M., Bencsik, N., Pusztai, S., Liliom, H., & Schlett, K. (2016). Animaltracker: An imagej-based tracking api to create a customized behaviour analyser program. Neuroinformatics, 14(4), 479–481.
Hartig, S. M. (2001). Basic image analysis and manipulation in ImageJ. In Current protocols in molecular biology. Wiley. ISBN 9780471142720. https://doi.org/10.1002/0471142727.mb1415s102.
Hoffmann, K., Stone, J., & Sherman, S. M. (1972). Relay of receptive-field properties in dorsal lateral geniculate nucleus of the cat. Journal Neurophysiology, 35(5), 518–531.
Ing, N., Salman, S., Ma, Z., Walts, A., Knudsen, B., & Gertych, A. (2016). Machine learning can reliably distinguish histological patterns of micropapillary and solid lung adenocarcinomas. In Information technologies in medicine, advances in intelligent systems and computing (Vol. 471, pp. 193–206). Springer.
Irshad, H., Veillard, A., Roux, L., & Racoceanu, D. (2014). Methods for nuclei detection, segmentation, and classification in digital histopathology: A review—current status and future potential. IEEE Reviews in Biomedical Engineering, 7, 97–114.
Jahne, B. (2002). Digital image processing: Concepts, algorithms and scientific applications, 5th Edn. NJ, USA: Springer-Verlag New York, Inc., Secaucus.
Jung, C., & Kim, C. (2010). Segmenting clustered nuclei using H-minima transform-based marker extraction and contour parameterization. IEEE Transactions on Biomedical Engineering, 57(10), 2600–2604.
Kaas, J. H. (2012). Evolution of columns, modules, and domains in the neocortex of primates. Proceedings of the National Academy of Sciences of the United States of America, 109(Suppl), 10,655–10,660.
Kamentsky, L., Jones, T. R., Fraser, A., Bray, M. A., Logan, D. J., Madden, K. L., Ljosa, V., Rueden, C., Eliceiri, K. W., & Carpenter, A. E. (2011). Improved structure, function and compatibility for cellprofiler: Modular high-throughput image analysis software. Bioinformatics, 27(8), 1179–1180.
Kaplan, E. (2014). The new visual neurosciences. In Werner, J.S., & Chalupa, L.M. (Eds.) The M, P, and K pathways of the primate visual system revisited (pp. 215–227). Cambridge: MIT Press.
Kolodziejczyk, A., Ladniak, M., & Piorkowski, A. (2014). Constructing software for analysis of neuron, glial and endothelial cell numbers and density in histological nissl-stained rodent brain tissue. Journal of Medical Informatics and Technologies, 23, 77–86.
Kopacz, B., & Piorkowski, A. (2017). A Review of the Efficient Algorithm Implementation for Image Processing in the ImageJ and Matlab Environments (Przeglad moŻliwości wydajnej implementacji algorytmów przetwarzania obrazów w środowisku Matlab). Studia Informatica, 38(132), 177–190.
Kowal, M., Filipczuk, P., Obuchowicz, A., Korbicz, J., & Monczak, R. (2013). Computer-aided diagnosis of breast cancer based on fine needle biopsy microscopic images. Computers in Biology and Medicine, 43 (10), 1563–1572.
Koyuncu, C. F., Akhan, E., Ersahin, T., Cetin-Atalay, R., & Gunduz-Demir, C. (2016). Iterative H-minima-based marker-controlled watershed for cell nucleus segmentation. Cytometry Part A, 89, 338–349.
La Torre, A., Alonso Nanclares, L., Muelas Pascual, S., Peña Sanchez, J. M., Oroquieta, F., & et al. (2013). 3d segmentations of neuronal nuclei from confocal microscope image stacks. Frontiers in neuroanatomy, 7, 1–10.
LeVay, S., Ferster, D (1977). Relay cell classes in the lateral geniculate nucleus of the cat and the effects of visual deprivation. The Journal of Comparative Neurology, 172(4), 563–84.
Maric, D. (2015). 2015 BioImage Informatics Conference: Nucleus Counting Challenge. https://isg.nist.gov/BII_2015/webPages/pages/nucleusCounting/NucleusCounting.html.
Mazurek, P., & Oszutowska-Mazurek, D. (2014). From the slit-island method to the ising model: Analysis of irregular grayscale objects. International Journal of Applied Mathematics and Computer Science, 24(1), 49–63.
Merkulyeva, N., Veshchitskii, A., Makarov, F., Gerasimenko, Y., & Musienko, P. (2016). Distribution of 28 kDa calbindin-immunopositive neurons in the cat spinal cord. Frontiers in Neuroanatomy, 9, 166.
Mikula, S., Trotts, I., Stone, J. M., & Jones, E. G. (2007). Internet-enabled high-resolution brain mapping and virtual microscopy. Neuroimage, 35(1), 9–15.
Mitzdorf, U., & Singer, W. (1977). Laminar segregation of afferents to lateral geniculate nucleus of the cat: an analysis of current source density. Journal of Neurophysiology, 40(6), 1227–44.
Morgan, J. L., Berger, D. R., Wetzel, A. W., & Lichtman, J. W. (2016). The fuzzy logic of network connectivity in mouse visual thalamus. Cell, 165(1), 192–206.
Mullen, R. J., Buck, C. R., & Smith, A. M. (1992). NeuN, a neuronal specific nuclear protein in vertebrates. Development, 116, 201–211.
Nurzynska, K., & Piorkowski, A. (2016). The correlation analysis of the shape parameters for endothelial image characterisation. Image Analysis and Stereology, 35(3), 149–158.
Nurzynska, K., Kubo, M., & Muramoto, K. (2013). Shape parameters for automatic classification of snow particles into snowflake and graupel. Meteorological Applications, 20(3), 257–265.
Ohki, K., Chung, S., Ch’ng, Y. H., Kara, P., & Reid, R. C. (2005). Functional imaging with cellular resolution reveals precise micro-architecture in visual cortex. Nature, 433(7026), 597–603.
Otsu, N. (1975). A threshold selection method from gray-level histograms. Automatica, 11(285–296), 23–27.
Papadopulos, F., Spinelli, M., Valente, S., Foroni, L., Orrico, C., Alviano, F., & Pasquinelli, G. (2007). Common tasks in microscopic and ultrastructural image analysis using imagej. Ultrastructural Pathology, 31(6), 401–407.
Payne, B. R., & Peters, A. (2002). The cat primary visual cortex. In Payne, B.R. & Peters, A. (Eds.) The concept of cat primary visual cortex (pp. 1–129). San Diego, London, Boston, New York, Sydney, Tokyo, Toronto: Academic Press.
Piorkowski, A. (2016). A statistical dominance algorithm for edge detection and segmentation of medical images. In Information technologies in medicine, advances in intelligent systems and computing (Vol. 471, pp. 3–14). Springer.
Piorkowski, A., Nurzynska, K., Gronkowska-Serafin, J., Selig, B., Boldak, C., & Reska, D. (2017). Influence of applied corneal endothelium image segmentation techniques on the clinical parameters. Computerized Medical Imaging and Graphics, 55, 13–27.
Pool, M., Thiemann, J., Bar-Or, A., & Fournier, A. E. (2008). Neuritetracer: a novel imagej plugin for automated quantification of neurite outgrowth. Journal of Neuroscience Methods, 168(1), 134–139.
Russ, J. (1998). The image processing handbook, 3rd Edn. Springer, and IEEE Press: CRC Press.
Sanderson, K. J. (1971). The projection of the visual field to the lateral geniculate and medial interlaminar nuclei in the cat. The Journal of Comparative Neurology, 143(1), 101–108.
Schiller, P. H. (2010). Parallel information processing channels created in the retina. Proceedings of the National Academy of Sciences of the United States of America, 107(40), 17,087–17,094.
Schindelin, J., Arganda-Carreras, I., Frise, E., Kaynig, V., Longair, M., Pietzsch, T., Preibisch, S., Rueden, C., Saalfeld, S., & Schmid, B. et al (2012). Fiji: an open-source platform for biological-image analysis. Nature Methods, 9(7), 676–682.
Schneider, C. A., Rasband, W. S., Eliceiri, K. W., & et al. (2012). NIH Image to ImageJ: 25 years of image analysis. Nature Methods, 9(7), 671–675.
Selinummi, J., Seppala, J., Yli-Harja, O., & Puhakka, J. A. (2005). Software for quantification of labeled bacteria from digital microscope images by automated image analysis. Biotechniques, 39(6), 859–862.
Skobel, M., Kowal, M., & Korbicz, J. (2017). Nuclei recognition using iterated conditional modes approach. In International conference on computer recognition systems (pp. 326–335). Springer.
Soille, P. (1999). Morphological Image Analysis: Principles and Applications. NJ, USA: Springer-Verlag New York, Inc. Secaucus.
Sternberg, S. R. (1983). Biomedical image processing. Computer, 16(1), 22–34.
Stichel, C. C., Singer, W., & Heizmann, C. W. (1988). Light and electron microscopic immunocytochemical localization of parvalbumin in the dorsal lateral geniculate nucleus of the cat: evidence for coexistence with GABA. The Journal of Comparative Neurology, 268(1), 29–37.
Acknowledgements
The authors thank Shkorbatova P. for NeuN primary antibody, prof. Musienko P. for microscopic equipment, and the Center for Molecular and Cell Technologies, Research Park, St. Petersburg State University.
K. Nurzynska was supported by statutory funds for young researchers (BKM/507/RAU2/2016) from the Institute of Informatics, Silesian University of Technology, Poland.
A. Piorkowski was supported by AGH University of Science and Technology, Faculty of Geology, Geophysics and Environmental Protection as a part of a statutory project.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
Open Access This article is distributed under the terms of the Creative Commons Attribution 4.0 International License (https://creativecommons.org/licenses/by/4.0), which permits use, duplication, adaptation, distribution, and reproduction in any medium or format, as long as you give appropriate credit to the original author(s) and the source, provide a link to the Creative Commons license, and indicate if changes were made.
About this article
Cite this article
Nurzynska, K., Mikhalkin, A. & Piorkowski, A. CAS: Cell Annotation Software – Research on Neuronal Tissue Has Never Been so Transparent. Neuroinform 15, 365–382 (2017). https://doi.org/10.1007/s12021-017-9340-2
Published:
Issue Date:
DOI: https://doi.org/10.1007/s12021-017-9340-2